Polyploidy has played an important evolutionary role in the genus Festuca (Poaceae), and several ploidy levels (ranging from 2n = 2x = 14 to 2n = 12x = 84) have been detected to date. Our understanding of ploidy diversity has greatly improved over the last two decades owing to the usage of flow cytometry (FCM). This valuable method makes it possible to not only estimate the genome size in a large number of individuals and populations but also infer ploidy levels. FCM has been used successfully in Festuca and its use is very useful in detecting a possible overlooked cytogenetic diversity that is generally manifested in the morphological variability exhibited in some of its taxa.
1. Introduction
Polyploidy, or whole-genome duplication (WGD), has long been recognised as a major force in angiosperm evolution
[1]. This phenomenon is known to have potential effects on certain life-history traits involving genome structure, gene expression, biochemistry, reproductive development, ecophysiology, competitive ability, growth rates and phenotype
[2][3]. The most visible and widespread immediate effect is undoubtedly the increase in cell size
[4]. In this respect, several studies have indicated that the polyploidy can induce detectable morphology variation in both microcharacters (e.g., stomata and pollen size) and the dimensions of vegetative (e.g., culm and leaf lengths) and reproductive organs (spikelet, lemma, anther and seed sizes, among others) or in generative features (e.g., flowers number per spikelet or inflorescences per plant)
[5][6][7].
Grasses are one of the best models with which to study polyploidy and its effects on genome evolution, as they are more than 80% polyploid or palepolyploid
[8].
Festuca L. is one of the largest genera of the Poaceae family, with more than 500 perennial species distributed mostly in the temperate regions of the world
[9][10]. About 70% of the fescue taxa have undergone at least a whole-genome duplication event during their evolutionary history
[11]. A wide range of ploidy levels ranging from diploid (2n = 2
x = 14) to dodecaploid (2n = 12
x = 84), with basic chromosome number
x = 7, have been reported to date
[11][12]. The taxonomic difficulties of the genus are well known, including many polyploidy groups with hardly distinguishable species.
Festuca yvesii Sennen & Pau
(Festuca sect.
Festuca) is a highly variable species comprising several cytotypes (Table 1). According to the most recent taxonomic treatment,
F. yvesii includes four subspecies widely distributed throughout the Iberian Peninsula
[13][14]. The subsp.
yvesii (2n = 42, 56) and the subsp.
altopyrenaica (Fuente & Ortúñez) Mart.-Sagarra & Devesa (2n = 28) have an eminently Pyrenean distribution, while the subsp.
summilusitana (Franco & Rocha Afonso) Mart.-Sagarra & Devesa and the subsp.
lagascae (Cebolla & Rivas Ponce) Mart.-Sagarra & Devesa, on which this study focuses, are found in the centre, west and north of the Iberian Peninsula. The subsp.
summilusitana (2n = 42, 70, 84) is highly polymorphic, and its taxonomy has changed in recent years
[15][16], with this being closely related to the detection of different chromosome numbers (Table 1), type of substrate and its geographical distribution
[12][17]. However, Devesa et al.
[13] showed that the diagnostic features employed in order to differentiate the segregated entities are highly variable and are blurred when the overall material is analysed, being practically impossible to assign specimens to an entity without knowing other nontaxonomic data. The subsp.
lagascae grows in the easternmost part of the Central System and in disjunct populations of northern Spain, and all of them are hexaploids (2n = 42)
[17][18]. The octoploid level (2n = 56) has been mentioned exceptionally on the basis of the chromosome counts obtained for the conflictive populations from the Cantabrian Mountains
[13][17] (Table 1). This taxon has undergone several changes in its nomenclature, and differs morphologically from the subsp.
summilusitana by having smaller sizes in certain characters (culm, lemma and panicle, among others). However, the distinction between both subspecies is sometimes difficult owing to the overlap in the range of diagnostic characters, with some transitional populations
[13].
Table 1. Ploidy level and chromosome number reported in the literature for subspecies of F. yvesii by chromosome counting or FCM analysis. The reference column provides the original source of publication to each chromosome count. Superscript indicates the taxon name it was published under. The asterisk (*) indicates chromosome counts in extra-Iberian material.
| Subspecies (According to Devesa et al., [13]) |
2n |
Method |
References |
| F. yvesii subsp. yvesii |
2n = 6x = 42 * |
Counting 1 |
[19] |
| |
2n = 8x = 56 |
Counting 1 |
[17] |
| F. yvesii subsp. altopyrenaica |
2n = 4x = 28 |
Counting 2 |
[20] |
| F. yvesii subsp. summilusitana |
2n = 6x = 42 |
Counting 3,4 |
[17][18] |
| |
2n = 10x = 70 |
Counting and FCM 4 |
[12][17] |
| |
2n = 12x = 84 |
Counting and FCM 4 |
[12] |
| F. yvesii subsp. lagascae |
2n = 6x = 42 |
Counting 5 |
[17] |
| |
2n = 8x = 56 |
Counting 1 |
[17] |
1 F. yvesii, 2 F. altopyrenaica, 3 F. gredensis, 4 F. summilusitana, 5 F. curvifolia.
2. Plant Material and Geographical Origin
Individuals of
F. yvesii subsp.
summilusitana and
F. yvesii subsp.
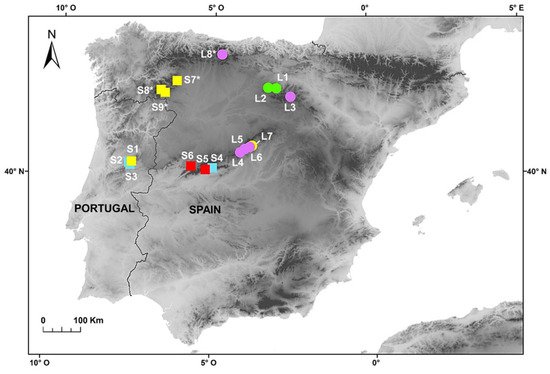
lagascae were collected throughout their range (Iberian Peninsula) in order to carry out karyological, genome size and morphological analyses (Figure 1). Field sampling was specifically carried out in several populations from the Sierra da Estrela (S1–S3), the Sierra de Gredos and Candelario (S4–S6), the Sierra de la Demanda and Cebollera (L1–L3), the Sierra de Guadarrama (L4–L7), the Cantabrian Mountains (L8) and in some mountains in the northwest (S7–S9) (Figure 1). The species were identified according to Devesa et al.
[13].
Figure 1. Populations of F. yvesii subsp. summilusitana (squares) and F. yvesii subsp. lagascae (circles) sampled in the Iberian Peninsula. Ploidy-level variation is shown using coloured symbols: purple, 6x; green, 8x; yellow, 10x; light blue, 12x; and red, 14x. The asterisks (*) indicate populations estimated from dried material.
Genome size (2C values) and DNA-ploidy levels were estimated by flow cytometry (FCM). A total of 94 individuals from 13 populations were analysed (Figure 1). Fresh leaves were selected from each of the individuals and stored in individual plastic bags at 4 °C for 3–5 days until use. Additionally, seven silica-gel-dried samples obtained from four populations were also analysed as a proxy for the validation of DNA-ploidy levels. With regard to the karyological study, we revisited those conflictive populations in which substantial differences were found between DNA-ploidy level and the number of chromosomes reported in the literature (i.e., populations S4–S6 from the Sierra de Gredos and Candelario and populations L4–L7 from the Sierra de Guadarrama; see Figure 1). The seeds of numerous individuals were collected from these seven selected populations during the fruiting period (late July–August) and stored in paper bags for their later use in chromosome counting and morphometric analyses. More than 200 root tips per population were analysed in order to obtain good metaphases plates. Finally, a total of 135 specimens from these populations were examined to explore the morphological variability between cytotypes. Voucher specimens of the populations studied were deposited in the Herbarium at the University of Cordoba (COFC), Spain.
3. Cytogenetic Variation
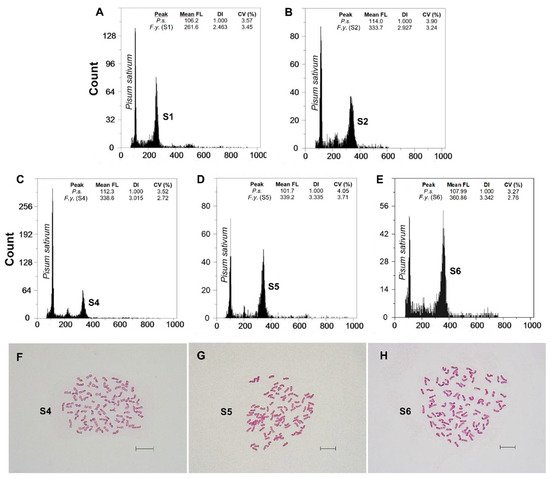
Fluorescence histograms of relative nuclear DNA content showed high-quality G1 peaks. The coefficients of variation (CVs) for fresh fescue samples were all within the normal range, below 5.0% (mean CVs = 3.48%), indicating reliable estimations (Figure 2A–E and Figure 3A–E). Genome size data for each subspecies and population are shown in Table 2.
Figure 2. (A–E) Flow cytometric histograms of relative PI fluorescence intensities and (F–H) mitotic metaphase chromosome preparations (for the S4, S5 and S6 populations) of F. yvesii subsp. summilusitana. Abbreviations: S1, ManTable S2. Sabugueiro; S4, Puerto del Pico (2n = 12x = 84); S5, Plataforma de Gredos (2n = 14x = 98); and S6, La Covatilla (2n = 14x = 98). Mean FL, mean relative fluorescence in picograms; DI, DNA index; CV (%), coefficient of variation of the peak in percent. Scale bar: 10 μm.
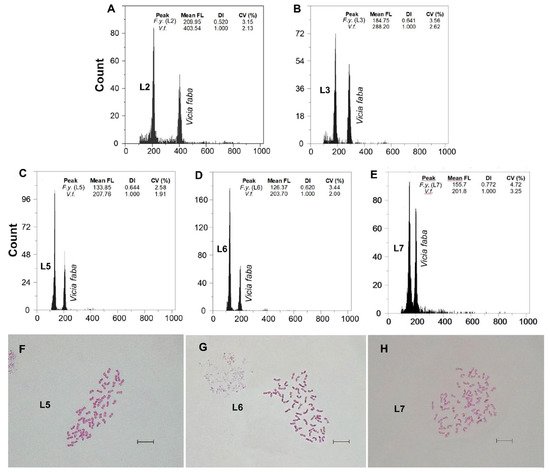
Figure 3. (A–E) Flow cytometric histograms of relative PI fluorescence intensities and (F–H) mitotic metaphase chromosome preparations (for the L5, L6 and L7 populations) of F. yvesii subsp. lagascae. Abbreviations: L2, Trigaza; L3, Puerto de Piqueras; L5, Puerto de Navacerrada (2n = 8x = 56), L6, Puerto de la Morcuera (2n = 8x = 56); and L7, Puerto de Canencia (2n = 10x = 70). Mean FL, mean relative fluorescence in picograms; DI, DNA index; CV (%), coefficient of variation of the peak in percent. Scale bar: 10 μm.
Table 2. Holoploid genome size estimations (mean, standard deviation of the mean and maximum and minimum values) for the sampled populations of each subspecies. Data regarding the population and number of individuals sampled (n) are provided. The last column shows the chromosome counts carried out in this study. Type of material analysed: F, fresh; D, dry.
| Taxon |
Mountain Range |
Locality |
n |
2C Mean ± SD (pg) |
2C Range (pg) |
Material |
DNA-Ploidy Level |
2n |
| F. yvesii subsp. summilusitana |
W range: Serra da Estrela (Portugal) |
S1: Castelo Branco, Manteigas |
9 |
22.89 ± 0.96 |
21.44–24.17 |
F |
10x |
- |
| |
S2: Castelo Branco, Sabugueiro |
8 |
26.60 ± 0.35 |
26.18–27.15 |
F |
12x |
- |
| |
S3: Castelo Branco, Torre |
6 |
26.58 ± 0.87 |
25.20–27.71 |
F |
12x |
- |
| |
C-W range: Sierra de Gredos and Candelario (Spain) |
S4: Ávila, Puerto del Pico |
5 |
26.89 ± 0.55 |
25.98–27.48 |
F |
12x |
84 |
| |
S5: Ávila, Plataforma de Gredos |
7 |
30.31 ± 0.49 |
29.68–31.16 |
F |
14x |
98 |
| |
S6: Salamanca, La Covatilla |
6 |
30.48 ± 0.73 |
29.85–31.91 |
F |
14x |
98 |
| |
NW mountains (Spain) |
S7: León, Villalibre de Somoza |
3 |
24.67 ± 1.47 |
23.01–25.84 |
D |
10x |
- |
| |
S8: Zamora, Sierra Segundera |
1 |
24.11 |
|
D |
10x |
- |
| |
S9: Zamora, Alto de San Juan |
1 |
24.82 |
|
D |
10x |
- |
| F. yvesii subsp. lagascae |
n range: Sierra de la Demanda and Cebollera (Spain) |
L1: Logroño, Valdezcaray |
6 |
13.79 ± 0.15 |
13.62–13.98 |
F |
6x |
- |
| |
L2: Burgos, Trigaza |
10 |
14.01 ± 0.32 |
13.60–14.46 |
F |
6x |
- |
| |
L3: Logroño/Soria, Puerto de Piqueras |
10 |
17.42 ± 0.29 |
17.06–18.00 |
F |
8x |
- |
| |
C range: Sierra de Guadarrama (Spain) |
L4: Madrid, Puerto de Guadarrama |
7 |
16.76 ± 0.22 |
16.57–17.21 |
F |
8x |
56 |
| |
L5: Madrid, Puerto de Navacerrada |
7 |
16.88 ± 0.24 |
16.63–17.36 |
F |
8x |
56 |
| |
L6: Madrid, Puerto de la Morcuera |
4 |
17.39 ± 0.65 |
16.71–18.23 |
F |
8x |
56 |
| |
L7: Madrid, Puerto de Canencia |
9 |
21.18 ± 0.62 |
20.51–22.31 |
F |
10x |
70 |
| |
North range: Cantabrian mountains (Spain) |
L8: Puerto de San Glorio |
2 |
19.31 ± 0.41 |
19.02–19.59 |
D |
8x |
- |
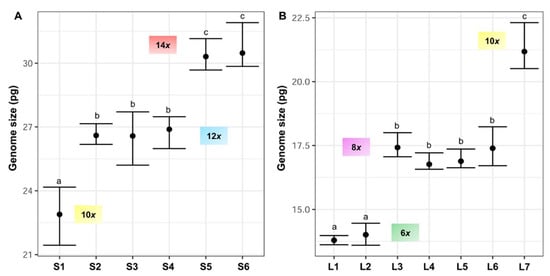
Nuclear 2C DNA content has been estimated for the first time in F. yvesii subsp. lagascae. The holoploid genome size estimates obtained were significantly different (up to 2.35-fold) for the two subspecies studied. The lowest 2C values were observed for Festuca yvesii subsp. lagascae, ranging from 13.60 to 22.31 pg, while the highest 2C values were found in the F. yvesii subsp. summilusitana individuals, ranging from 21.44 to 31.91 pg (Table 2, Figure 4). The analysis of variance (ANOVA) revealed significant differences between the 2C values (F12,81 = 837.8, p < 0.0001) at the subspecies level. Variability in genome size was found among the populations studied, with a maximum variation of 1.33-fold in the subsp. summilusitana (S1 vs. S6) and 1.54-fold in the subsp. lagascae (L1 vs. L7) (Table 2). Nevertheless, genome size estimates were homogeneous among individuals, with an average variation of 5.9%.
Figure 4. Holoploid genome size range and mean (2C values, in picograms) in the sampled populations of (A) F. yvesii subsp. summilusitana (S1–S6) and (B) F. yvesii subsp. lagascae (L1–L7). DNA-ploidy levels are marked (6x, 8x, 10x, 12x and 14x). Only estimates from fresh material are represented. Abbreviations (S1–S6 and L1–L7) correspond to the populations indicated in Table 2. Significant differences among populations at p < 0.05 are denoted by different letters.
Variation in DNA content was not continuous and five nonoverlapping DNA-ploidy levels were identified (Figure 4): hexaploids (2C = 13.60–14.46 pg), octoploids (2C = 16.57–18.23 pg), decaploids (2C = 20.51–24.17 pg), dodecaploids (2C = 25.20–27.71 pg) and tetradecaploids (2C = 29.68–31.91 pg).
Specifically, F. yvesii subsp. summilusitana individuals fell within ploidy levels of 10x (S1), 12x (S2, S3 and S4) and 14x (S5 and S6) (Figure 1 and Figure 4A), while individuals of F. yvesii subsp. lagascae had nuclear DNA contents that fell within ploidy levels of 6x (L1 and L2), 8x (L3, L4, L5 and L6) and 10x (L7) (Figure 1 and Figure 4B). Noticeably, the DNA content of dodecaploids was slightly less than twice that of hexaploids (Table 2).
The silica-gel-dried leaves yielded peaks of sufficient quality to assign DNA-ploidy level, although CVs were often higher (mean CVs = 4.9%). The preliminary estimates of dry individuals from northwestern populations of F. yvesii subsp. summilusitana (2C = 23.01–25.84 pg; S7, S8 and S9) fell within the same DNA content group as populations inferred as 10x, and those of F. yvesii subsp. lagascae (L8) were within the 8x range (Figure 1, Table 2). It should be noted that the dry samples always showed genome size estimates larger than the fresh samples with the same ploidy level, with a difference of up to 1.93 pg for the subsp. summilusitana (S1 vs. S9) and up to 2.55 pg for the subsp. lagascae (L4 vs. L8) (Table 2).
The karyological screenings carried out confirmed the ploidy level detected by FCM (Table 2, Figures 2F–H and 3F–H). The Spearman rank correlation showed a strong positive correlation between 2C nuclear DNA content and the chromosome number observed. New cytotypes were detected on the basic chromosome number. Unexpectedly high ploidy levels were detected for the F. yvesii subsp. summilusitana populations from the centre-west of the Iberian Peninsula (Table 2): Puerto del Pico population (S4) was dodecaploid (2n = 12x = 84; Figure 2F), while Plataforma de Gredos (S5) and La Covatilla (S6) were tetradecaploid (2n = 14x = 98; Figure 2G,H), the highest chromosome number reported in the Festuca genus to date. Two new somatic numbers were found for F. yvesii subsp. lagascae: 2n = 56 in the individuals from Puerto de Guadarrama (L4), Puerto de Navacerrada (L5; Figure 3F) and Puerto de Morcuera (L6; Figure 3G), and 2n = 70 in those from Puerto de Canencia (L7; Figure 3H), i.e., octoploid and decaploid, respectively (Table 2). In line with the FCM analyses, non-mixed-ploidy populations were detected.
4. Morphological Variation
In general, the largest sizes for all the characters were found in the subsp. summilusitana, with the exception of the auricle and awn lengths. In both subspecies, most of the morphological traits increased in size with increasing ploidy; however, no significant differences were observed with respect to all ploidy levels. The lemma, upper glume and seed lengths in particular responded significantly to changes in ploidy in both subspecies. The differences observed in the arithmetic mean obtained for most of the characters are significant only among the most extreme cytotypes of each subspecies, and even in these cases, the ranges of variation overlap.