
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Youri Pavlov | + 1766 word(s) | 1766 | 2020-12-02 06:50:32 | | | |
| 2 | Lily Guo | -59 word(s) | 1707 | 2020-12-14 07:43:32 | | |
Video Upload Options
Recent studies on tumor genomes revealed that mutations in genes of replicative DNA polymerases cause a predisposition for cancer by increasing genome instability. The past 10 years have uncovered exciting details about the structure and function of replicative DNA polymerases and the replication fork organization. The principal idea of participation of different polymerases in specific transactions at the fork proposed by Morrison and coauthors 30 years ago and later named “division of labor,” remains standing, with an amendment of the broader role of polymerase δ in the replication of both the lagging and leading DNA strands. However, cancer-associated mutations predominantly affect the catalytic subunit of polymerase ε that participates in leading strand DNA synthesis.
1. Introduction
Research in the past decade has revealed the lofty role of alterations in replicative DNA polymerases (pols) in sporadic and hereditary cancer[1][2]. The predisposition to tumorigenesis is attributed to the low fidelity of DNA replication by inaccurate pol versions[3][4]. Among the replicative B-family enzymes, pol ε stands out. The alterations in the proofreading exonuclease domain caused by mutations in the POLE gene (see Table 1 for the nomenclature of DNA polymerase subunits in humans and in yeast and mouse models) are proven to be causative factors in the etiology of the malignant transformation (Figure 1), with predominant, but not exclusive prevalence, in colon and endometrial cancers. The review analyzes how the modern understanding of the replication fork based on the synthesis of information gained in model systems and genomics of tumors may explain the peculiarities of the connection of pols and cancer in humans.
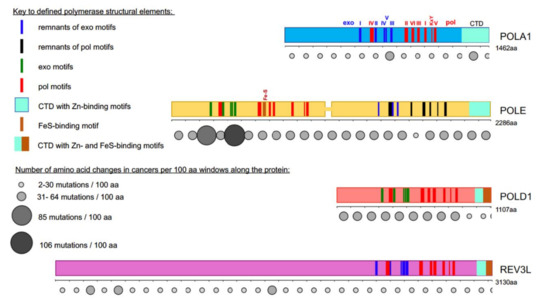
Figure 1. Most cancer-associated mutations affect the catalytically active half of POLE. Colored bars represent the main subunits of DNA pols, a catalytic subunit of pol α, POLA1, in light blue; of pol ε, POLE, in yellow; of pol δ, POLD1, in red; and pol ζ, REV3L, in purple. Note that POLE is a tandem of active pol (N-terminal half) and inactive pol (C-terminal half)[5][6]. Evolutionarily conserved motifs characteristic for all exonuclease (exo) domains, are labeled I-V in green and pol domains are labeled I-VI and KxY in red[5][6][7][8][9][10][11][12]. The order of the motifs along all four proteins is the same, but they occupy different parts of the whole protein. For example, REV3L has a very long N-terminal part not related to pols. In POL1, REV3L, and the C-terminal half of POLE, the exonuclease motifs are inactivated during evolution; they are shown in blue. Inactivated pol motifs in the C-terminal half of POLE are shown in black. The key for these and other elements of the pol primary structure is in the left upper quarter of the figure. Rows of circles of different sizes and shades of grey below the catalytic pol subunits represent the number of missense mutations found in tumors along the protein regions in 100 amino acid increments. Variants were collected from the cBioPortal database from a curated non-overlapping collection of tumor genomes (https://www.cbioportal.org/ (cbioportal.org)). A guide explaining the relation between size and intensity of grey to the number of mutations found in the database in the 100 amino acids interval is on the left lower quarter of the figure.
2. Progress on the Structure-Function of B-Family DNA Polymerases and Organization of the Replication Fork
The past ten years brought groundbreaking discoveries about B-family DNA pols. Currently, with the help of X-ray crystallography and the improvement of cryo-EM resolution, we understand the atomic details of the structures of catalytic cores and whole complexes (Table 1) of yeast and human (Table 1) primase-pol α [13][14][15], yeast and human pol δ[16][17][18], yeast pol ε alone or in complex with CMG[19][20][21][22], yeast pol ζ [23](Figure 3).
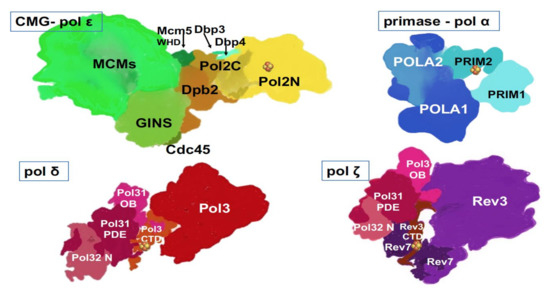
Figure 3. Multi-subunit replicative DNA pols. Artistic representations were made based on crystal and cryo-EM structures and models of human primase-pol α[14], yeast pol ε[22][24], yeast pol δ[17], yeast pol ζ [23][66]. Two latter structures were determined with a truncated third subunit (Table 1), Pol 32, without the C-terminal part, and thus, this part is missing from our drawings. Fe-S cluster (  ) is present in each of the four pol complexes.
) is present in each of the four pol complexes.
As we can see from the list, two human pols’ structures can only be modeled based on solved yeast counterparts, thus making the solution of structures of human pols a high priority. The structures of yeast and human pols appear to be similar in general features but differ in nuances. For example, human pol δ has an additional small subunit, p12 (Table 1) hypothesized to regulate pol δ activity during normal replication versus conditions of DNA damage or replicative stress[18][25]. The catalytic subunit of human pol ζ has extended the N-terminal part of unknown significance (Table 1, Figure 1 and Figure 3). Structural and functional studies helped understand transactions in the active site of polymerases and within the pol complexes. Examples of success are the basis of RNA primer synthesis by primase and transfer of RNA primer 3′-end into pol α active site to start DNA synthesis; the reasons for the high fidelity of pols δ and ε; and the ability of pol ζ to extend mismatches or unpaired DNA ends found opposite lesions.
One exciting finding is that all DNA pols coordinate Fe-S clusters, known regulatory/structural elements of various proteins[26], alluding to the connection of iron metabolism in mitochondria to replication and novel opportunities for regulation of pol reactions (Figure 1 and Figure 3)[27][28]. The cluster can accept or donate electrons and might be involved in sensing the redox potential of cells and DNA damage[29][30]. The first finding was the detection of the Fe-S cluster in the second subunit of archaeal and yeast primase[31], which was proven to play a seminal role in the primer synthesis by human enzyme[32][33][34]. Then, Fe-S clusters were found and verified in C-terminal regions of yeast and human pols δ and ζ (Figure 1 and Figure 3) and were shown to be necessary for pol function[27][35][36]. The Fe-S cluster in the catalytic subunit of pol ε was found in an unusual location: in the N-terminal half in the vicinity of pol II motif (Figure 1), structurally characterized, and shown to be necessary for pol but not exo activity in functional assays[37][38]. A recent study revealed the unique sensitivity of pol ε to suppression of Fe-S biosynthesis in basal-like breast cancer cell lines[39].
Another sensational discovery was the sharing of subunits between pols δ and ζ[35], Table 1, Figure 3. For quite an extended period, pol ζ was referred to as a two subunit enzyme consisting of catalytic Rev3 and accessory subunit Rev7[40], later found to be one REV3 to two REV7 subunit complex[41]. The two-subunit complex possessed quite low and variable activity[42][43][44]. The pol δ’s two accessory subunits appeared to be two additional subunits of pol ζ necessary for the full activity[35][45][46][47]. It appeared that the inconsistent activity of former “two-subunit” preps resulted from uncontrolled traces of a genuine four subunit enzyme [46]. The role of such subunit sharing between the main replicative pols and pol ζ is under debate. In the original paper describing the discovery, an elegant mechanism of switches of pol’s catalytic subunits on the already present core of PCNA/POLD2/POLD3 was proposed[35], and the role of Fe-S clusters in CTDs of both pols recognized, but possible details of the process have never been elaborated. The argument against the switch mechanism is the stability of multi-subunit complexes of pols δ and ζ[45][46]. Pols’ architecture with shared subunits might reflect evolutionary relationships and structural requirements[23].
New findings lead to a better understanding of replication fork in eukaryotes (Figure 4). The CMG complex bound to the C-terminal part of pol ε travels along on the leading strand. This tight association explains the participation of pol ε in the leading strand synthesis and exclusion of this pol from synthesis and proofreading on the lagging strand. In yeast pol, the accessory subunits of Pol2, Dpb3/Dpb4, may serve as “staples” rigidly connecting the C-terminal part with the active N-terminal part[22]. However, if this rigidity were stable, any transactions by other DNA pols on the leading strand (for example, when the switch to translesion pol is necessary for DNA damage bypass) would have been blocked by the N-terminal half of Pol2 stuck with the primer terminus, but this is not the case. Pol δ proofreads errors made by pol ε[48][49] and pol ζ, with other translesion DNA synthesis pols, operate on the leading strand to the same extent as on the lagging strand [49][50]. Therefore, there should be a mechanism of how the active part of the catalytic subunit of pol ε abandons the 3’-end of the nascent leading strand and yields to other pols.
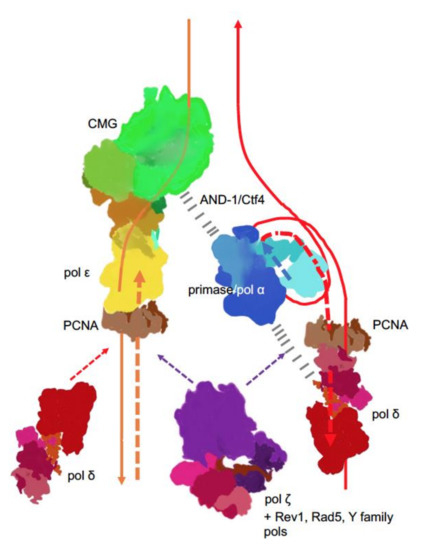
Figure 4. Main players at the eukaryotic replication fork. The CMG complex unwinds DNA, primase-Pol α synthesizes short RNA-DNA primers that are extended by pol δ. On the lagging strand, pol δ synthesis is halted when the pol reaches the previous Okazaki fragment. On the leading strand, pol ε takes over and contributes to around 80% of the bulk strand synthesis. Pol δ occasionally proofreads errors made by pol ε[48] and likely continues synthesis on the leading strand thereafter. The coordination of the whole process is likely achieved by interactions of primase-pol α with CMG via Ctf4 (yeast) or AND-1 (humans)[51][52][53][54] and some not precisely mapped pol α interactions with pol δ (dashed black lines)[55][56][57]. Replication stress caused by unusual DNA structures[58], DNA damage or defects in replisome[59][60][61][62] lead to recruitment and patches of synthesis of the fourth member of the B-family, pol ζ [63], along with translesion pols and accessory factors to mitigate replication problems, depending on the nature of replication problems.
The lagging DNA strand is synthesized in relatively short Okazaki fragments whose size coincides with the nucleosomal repeat (165 bp), as measured under conditions of constrained ligation [64][65]. The evidence from the distribution of inaccurate pol α-dependent mutations in yeast seems to support this estimate[66]]. The need for the ligation of short DNA fragments on the lagging strand led to a straightforward assumption that nicks and ssDNA regions are more prevalent in this strand. Such a property of the lagging strand would explain the more efficient operation of MMR on the lagging strand[67], or preferential damage of the lagging strand by DNA editing cytosine deaminases of the APOBEC family that act on ssDNA, shown in model systems[68][69] and tumors[70][71]. However, recent findings suggest that the leading DNA strand is discontinuous as well, and nicks in the yeast’s leading strand are even more frequent than in the lagging strand[65]. The effect is attributed to ribonucleotide excision, as seen in bacteria [72][73] and the preferential incorporation and repair of ribonucleotides in the leading DNA strand in yeast.
References
- Rayner, E.; van Gool, I.C.; Palles, C.; Kearsey, S.E.; Bosse, T.; Tomlinson, I.; Church, D.N. A panoply of errors: Polymerase proofreading domain mutations in cancer. Nat. Rev. Cancer 2016, 16, 71–81, doi:10.1038/nrc.2015.12.
- Barbari, S.R.; Shcherbakova, P.V. Replicative DNA polymerase defects in human cancers: Consequences, mechanisms, and implications for therapy. DNA Repair 2017, 56, 16–25, doi:10.1016/j.dnarep.2017.06.003.
- Loeb, L.A.; Springgate, C.F.; Battula, N. Errors in DNA replication as a basis of malignant changes. Cancer Res. 1974, 34, 2311–2321.
- Preston, B.D.; Albertson, T.M.; Herr, A.J. DNA replication fidelity and cancer. Semin. Cancer Biol. 2010, 20, 281–293, doi:10.1016/j.semcancer.2010.10.009.
- Tahirov, T.H.; Makarova, K.S.; Rogozin, I.B.; Pavlov, Y.I.; Koonin, E.V. Evolution of DNA polymerases: An inactivated polymerase-exonuclease module in Pol epsilon and a chimeric origin of eukaryotic polymerases from two classes of archaeal ancestors. Biol. Direct 2009, 4, 11, doi:10.1186/1745-6150-4-11.
- Kazlauskas, D.; Krupovic, M.; Guglielmini, J.; Forterre, P.; Venclovas, Č. Diversity and evolution of B-family DNA polymerases. Nucleic Acids Res. 2020, 48, 10142–10156, doi:10.1093/nar/gkaa760.
- Delarue, M.; Poch, O.; Tordo, N.; Moras, D.; Argos, P. An attempt to unify the structure of polymerases. Protein Eng. 1990, 3, 461–467.
- Blanco, L.; Bernad, A.; Blasco, M.A.; Salas, M. A general structure for DNA-dependent DNA polymerases. Gene 1991, 100, 27–38, doi:10.1016/0378-1119(91)90346-d.
- Brautigam, C.A.; Steitz, T.A. Structural and functional insights provided by crystal structures of DNA polymerases and their substrate complexes. Curr. Opin. Struct. Biol. 1998, 8, 54–63, doi:10.1016/s0959-440x(98)80010-9.
- Hopfner, K.P.; Eichinger, A.; Engh, R.A.; Laue, F.; Ankenbauer, W.; Huber, R.; Angerer, B. Crystal structure of a thermostable type B DNA polymerase from Thermococcus gorgonarius. Proc. Natl. Acad. Sci. USA 1999, 96, 3600–3605, doi:10.1073/pnas.96.7.3600.
- Kähler, M.; Antranikian, G. Cloning and characterization of a family B DNA polymerase from the hyperthermophilic crenarchaeon Pyrobaculum islandicum. J. Bacteriol. 2000, 182, 655–663, doi:10.1128/jb.182.3.655-663.2000.
- Shevelev, I.V.; Hübscher, U. The 3′ 50′ exonucleases. Nat. Rev. Mol. Cell. Biol. 2002, 3, 364–376, doi:10.1038/nrm804.
- Pellegrini, L. The Pol alpha-Primase Complex. Subcell. Biochem. 2012, 62, 157–169, doi:10.1007/978-94-007-4572-8_9.
- Agarkar, V.B.; Babayeva, N.D.; Pavlov, Y.I.; Tahirov, T.H. Crystal structure of the C-terminal domain of human DNA primase large subunit: Implications for the mechanism of the primase-polymerase α switch. Cell Cycle 2011, 10, 926–931, doi:10.4161/cc.10.6.15010.
- Baranovskiy, A.G.; Babayeva, N.D.; Zhang, Y.; Gu, J.; Suwa, Y.; Pavlov, Y.I.; Tahirov, T.H. Mechanism of concerted rna-dna primer synthesis by the human primosome. J. Biol. Chem. 2016, 291, 10006–10020, doi:10.1074/jbc.M116.717405.
- Swan, M.K.; Johnson, R.E.; Prakash, L.; Prakash, S.; Aggarwal, A.K. Structural basis of high-fidelity DNA synthesis by yeast DNA polymerase delta. Nat. Struct. Mol. Biol. 2009, 16, 979–986, doi:910.1038/nsmb.1663.
- Jain, R.; Rice, W.J.; Malik, R.; Johnson, R.E.; Prakash, L.; Prakash, S.; Ubarretxena-Belandia, I.; Aggarwal, A.K. Cryo-EM structure and dynamics of eukaryotic DNA polymerase δ holoenzyme. Nat. Struct. Mol. Biol. 2019, 26, 955–962, doi:10.1038/s41594-019-0305-z.
- Lancey, C.; Tehseen, M.; Raducanu, V.S.; Rashid, F.; Merino, N.; Ragan, T.J.; Savva, C.G.; Zaher, M.S.; Shirbini, A.; Blanco, F.J.; et al. Structure of the processive human Pol δ holoenzyme. Nat. Commun. 2020, 11, 1109, doi:10.1038/s41467-020-14898-6.
- Hogg, M.; Osterman, P.; Bylund, G.O.; Ganai, R.A.; Lundstrom, E.B.; Sauer-Eriksson, A.E.; Johansson, E. Structural basis for processive DNA synthesis by yeast DNA polymerase varepsilon. Nat. Struct. Mol. Biol. 2014, 21, 49–55.
- Jain, R.; Rajashankar, K.R.; Buku, A.; Johnson, R.E.; Prakash, L.; Prakash, S.; Aggarwal, A.K. Crystal structure of yeast DNA polymerase epsilon catalytic domain. PLoS ONE 2014, 9, e94835.
- Georgescu, R.; Yuan, Z.; Bai, L.; de Luna Almeida Santos, R.; Sun, J.; Zhang, D.; Yurieva, O.; Li, H.; O’Donnell, M.E. Structure of eukaryotic CMG helicase at a replication fork and implications to replisome architecture and origin initiation. Proc. Natl. Acad. Sci. USA 2017, 114, E697–E706, doi:10.1073/pnas.1620500114.
- Yuan, Z.; Georgescu, R.; Schauer, G.D.; O’Donnell, M.E.; Li, H. Structure of the polymerase ε holoenzyme and atomic model of the leading strand replisome. Nat. Commun. 2020, 11, 3156, doi:10.1038/s41467-020-16910-5.
- Malik, R.; Kopylov, M.; Gomez-Llorente, Y.; Jain, R.; Johnson, R.E.; Prakash, L.; Prakash, S.; Ubarretxena-Belandia, I.; Aggarwal, A.K. Structure and mechanism of B-family DNA polymerase ζ specialized for translesion DNA synthesis. Nat. Struct. Mol. Biol. 2020, doi:10.1038/s41594-020-0476-7.
- Goswami, P.; Abid Ali, F.; Douglas, M.E.; Locke, J.; Purkiss, A.; Janska, A.; Eickhoff, P.; Early, A.; Nans, A.; Cheung, A.M.C.; et al. Structure of DNA-CMG-Pol epsilon elucidates the roles of the non-catalytic polymerase modules in the eukaryotic replisome. Nat. Commun. 2018, 9, 5061, doi:10.1038/s41467-018-07417-1.
- Lee, M.; Zhang, S.; Wang, X.; Chao, H.H.; Zhao, H.; Darzynkiewicz, Z.; Zhang, Z.; Lee, E.Y.C. Two forms of human DNA polymerase δ: Who does what and why? DNA Repair 2019, 81, 102656, doi:10.1016/j.dnarep.2019.102656.
- Lill, R. Function and biogenesis of iron-sulphur proteins. Nature 2009, 460, 831–838.
- Netz, D.J.; Stith, C.M.; Stumpfig, M.; Kopf, G.; Vogel, D.; Genau, H.M.; Stodola, J.L.; Lill, R.; Burgers, P.M.; Pierik, A.J. Eukaryotic DNA polymerases require an iron-sulfur cluster for the formation of active complexes. Nat. Chem. Biol. 2012, 8, 125–132, doi:10.1038/nchembio.721.
- Baranovskiy, A.G.; Siebler, H.M.; Pavlov, Y.I.; Tahirov, T.H. Iron-Sulfur clusters in DNA polymerases and primases of eukaryotes. Methods Enzymol. 2018, 599, 1–20, doi:10.1016/bs.mie.2017.09.003.
- Rouault, T.A. The indispensable role of mammalian iron sulfur proteins in function and regulation of multiple diverse metabolic pathways. Biometals 2019, 32, 343–353, doi:10.1007/s10534-019-00191-7.
- Merino, E.J.; Boal, A.K.; Barton, J.K. Biological contexts for DNA charge transport chemistry. Curr. Opin. Chem. Biol. 2008, 12, 229–237, doi:10.1016/j.cbpa.2008.01.046.
- Klinge, S.; Hirst, J.; Maman, J.D.; Krude, T.; Pellegrini, L. An iron-sulfur domain of the eukaryotic primase is essential for RNA primer synthesis. Nat. Struct. Mol. Biol. 2007, 14, 875–877, doi:10.1038/nsmb1288.
- Baranovskiy, A.G.; Zhang, Y.; Suwa, Y.; Gu, J.; Babayeva, N.D.; Pavlov, Y.I.; Tahirov, T.H. Insight into the Human DNA Primase Interaction with Template-Primer. J. Biol. Chem. 2016, 291, 4793–4802, doi:10.1074/jbc.M115.704064.
- Holt, M.E.; Salay, L.E.; Chazin, W.J. A polymerase with potential: The Fe-S cluster in human DNA primase. Methods Enzymol. 2017, 595, 361–390, doi:10.1016/bs.mie.2017.07.002.
- Baranovskiy, A.G.; Babayeva, N.D.; Zhang, Y.; Blanco, L.; Pavlov, Y.I.; Tahirov, T.H. Comment on The [4Fe4S] cluster of human DNA primase functions as a redox switch using DNA charge transport. Science 2017, 357, doi:10.1126/science.aan2396.
- Baranovskiy, A.G.; Lada, A.G.; Siebler, H.M.; Zhang, Y.; Pavlov, Y.I.; Tahirov, T.H. DNA Polymerase delta and zeta switch by sharing accessory subunits of DNA polymerase delta. J. Biol. Chem. 2012, 287, 17281–17287, doi:M112.351122 [pii].
- Bartels, P.L.; Stodola, J.L.; Burgers, P.M.J.; Barton, J.K. A Redox role for the [4Fe4S] cluster of yeast DNA polymerase δ. J. Am. Chem. Soc. 2017, 139, 18339–18348, doi:10.1021/jacs.7b10284.
- Jain, R.; Vanamee, E.S.; Dzikovski, B.G.; Buku, A.; Johnson, R.E.; Prakash, L.; Prakash, S.; Aggarwal, A.K. An iron-sulfur cluster in the polymerase domain of yeast DNA polymerase epsilon. J. Mol. Biol. 2014, 426, 301–308.
- Ter Beek, J.; Parkash, V.; Bylund, G.O.; Osterman, P.; Sauer-Eriksson, A.E.; Johansson, E. Structural evidence for an essential Fe-S cluster in the catalytic core domain of DNA polymerase ε. Nucleic Acids Res. 2019, 47, 5712–5722, doi:10.1093/nar/gkz248.
- Sviderskiy, V.O.; Blumenberg, L.; Gorodetsky, E.; Karakousi, T.R.; Hirsh, N.; Alvarez, S.W.; Terzi, E.M.; Kaparos, E.; Whiten, G.C.; Ssebyala, S.; et al. Hyperactive CDK2 activity in basal-like breast cancer imposes a genome integrity liability that can be exploited by targeting DNA polymerase ε. Mol. Cell 2020, in press, doi:10.1016/j.molcel.2020.10.016.
- Pavlov, Y.I.; Shcherbakova, P.V.; Rogozin, I.B. Roles of DNA polymerases in replication, repair, and recombination in eukaryotes. Int. Rev. Cytol. 2006, 255, 41–132, doi:S0074-7696(06)55002-8 [pii].
- Rizzo, A.A.; Vassel, F.M.; Chatterjee, N.; D'Souza, S.; Li, Y.; Hao, B.; Hemann, M.T.; Walker, G.C.; Korzhnev, D.M. Rev7 dimerization is important for assembly and function of the Rev1/Polζ translesion synthesis complex. Proc. Natl. Acad. Sci. USA 2018, 115, E8191–E8200, doi:10.1073/pnas.1801149115.
- Nelson, J.R.; Lawrence, C.W.; Hinkle, D.C. Thymine-thymine dimer bypass by yeast DNA polymerase zeta. Science 1996, 272, 1646–1649, doi:10.1126/science.272.5268.1646.
- Haracska, L.; Unk, I.; Johnson, R.E.; Johansson, E.; Burgers, P.M.; Prakash, S.; Prakash, L. Roles of yeast DNA polymerases delta and zeta and of Rev1 in the bypass of abasic sites. Genes Dev. 2001, 15, 945–954, doi:10.1101/gad.882301.
- Garg, P.; Stith, C.M.; Majka, J.; Burgers, P.M. Proliferating cell nuclear antigen promotes translesion synthesis by DNA polymerase zeta. J. Biol. Chem. 2005, 280, 23446–23450, doi:10.1074/jbc.C500173200.
- Johnson, R.E.; Prakash, L.; Prakash, S. Pol31 and Pol32 subunits of yeast DNA polymerase delta are also essential subunits of DNA polymerase zeta. Proc. Natl. Acad. Sci. USA 2012, 109, 12455–12460, doi:10.1073/pnas.1206052109.
- Makarova, A.V.; Stodola, J.L.; Burgers, P.M. A four-subunit DNA polymerase zeta complex containing Pol delta accessory subunits is essential for PCNA-mediated mutagenesis. Nucleic Acids Res. 2012, 40, 11618–11626, doi:11610.11093/nar/gks11948. Epub 12012 Oct 11612.
- Lee, Y.S.; Gregory, M.T.; Yang, W. Human Pol zeta purified with accessory subunits is active in translesion DNA synthesis and complements Pol eta in cisplatin bypass. Proc. Natl. Acad. Sci. USA. 2014, 111, 2954–2959, doi:2910.1073/pnas.1324001111.
- Bulock, C.R.; Xing, X.; Shcherbakova, P.V. DNA polymerase delta proofreads errors made by DNA polymerase epsilon. Proc. Natl. Acad. Sci. USA 2020, 117, 6035–6041, doi:10.1073/pnas.1917624117.
- Pavlov, Y.I.; Shcherbakova, P.V. DNA polymerases at the eukaryotic fork-20 years later. Mutat. Res. 2010, 685, 45–53, doi:10.1016/j.mrfmmm.2009.08.002.
- Pagès, V.; Johnson, R.E.; Prakash, L.; Prakash, S. Mutational specificity and genetic control of replicative bypass of an abasic site in yeast. Proc. Natl. Acad. Sci. USA 2008, 105, 1170–1175, doi:10.1073/pnas.0711227105.
- Simon, A.C.; Zhou, J.C.; Perera, R.L.; van Deursen, F.; Evrin, C.; Ivanova, M.E.; Kilkenny, M.L.; Renault, L.; Kjaer, S.; Matak-Vinković, D.; et al. A Ctf4 trimer couples the CMG helicase to DNA polymerase α in the eukaryotic replisome. Nature 2014, 510, 293–297, doi:10.1038/nature13234.
- Kilkenny, M.L.; Simon, A.C.; Mainwaring, J.; Wirthensohn, D.; Holzer, S.; Pellegrini, L. The human CTF4-orthologue AND-1 interacts with DNA polymerase alpha/primase via its unique C-terminal HMG box. Open Biol. 2017, 7, doi:10.1098/rsob.170217.
- Yuan, Z.; Georgescu, R.; Santos, R.L.A.; Zhang, D.; Bai, L.; Yao, N.Y.; Zhao, G.; O’Donnell, M.E.; Li, H. Ctf4 organizes sister replisomes and Pol α into a replication factory. Elife 2019, 8, doi:10.7554/eLife.47405.
- Rzechorzek, N.J.; Hardwick, S.W.; Jatikusumo, V.A.; Chirgadze, D.Y.; Pellegrini, L. CryoEM structures of human CMG-ATPγS-DNA and CMG-AND-1 complexes. Nucleic Acids Res. 2020, 48, 6980–6995, doi:10.1093/nar/gkaa429.
- Huang, M.E.; Le Douarin, B.; Henry, C.; Galibert, F. The Saccharomyces cerevisiae protein YJR043C (Pol32) interacts with the catalytic subunit of DNA polymerase alpha and is required for cell cycle progression in G2/M. Mol. Gen. Genet. 1999, 260, 541–550, doi:10.1007/s004380050927.
- Johansson, E.; Garg, P.; Burgers, P.M. The Pol32 subunit of DNA polymerase delta contains separable domains for processive replication and proliferating cell nuclear antigen (PCNA) binding. J. Biol. Chem. 2004, 279, 1907–1915, doi:10.1074/jbc.M310362200.
- Lewis, J.S.; Spenkelink, L.M.; Schauer, G.D.; Yurieva, O.; Mueller, S.H.; Natarajan, V.; Kaur, G.; Maher, C.; Kay, C.; O’Donnell, M.E.; et al. Tunability of DNA polymerase stability during eukaryotic DNA replication. Mol. Cell 2020, 77, 17–25, doi:10.1016/j.molcel.2019.10.005.
- Northam, M.R.; Moore, E.A.; Mertz, T.M.; Binz, S.K.; Stith, C.M.; Stepchenkova, E.I.; Wendt, K.L.; Burgers, P.M.; Shcherbakova, P.V. DNA polymerases ζ and Rev1 mediate error-prone bypass of non-B DNA structures. Nucleic Acids Res. 2014, 42, 290–306, doi:10.1093/nar/gkt830.
- Pavlov, Y.I.; Shcherbakova, P.V.; Kunkel, T.A. In vivo consequences of putative active site missense mutations in yeast replicative DNA polymerases α, ε, δ and ζ. Genetics 2001, 159, 47–64.
- Garbacz, M.; Araki, H.; Flis, K.; Bebenek, A.; Zawada, A.E.; Jonczyk, P.; Makiela-Dzbenska, K.; Fijalkowska, I.J. Fidelity consequences of the impaired interaction between DNA polymerase epsilon and the GINS complex. DNA Repair 2015, 29, 23–35, doi:10.1016/j.dnarep.2015.02.007.
- Northam, M.R.; Garg, P.; Baitin, D.M.; Burgers, P.M.; Shcherbakova, P.V. A novel function of DNA polymerase zeta regulated by PCNA. EMBO J. 2006, 25, 4316–4325.
- Becker, J.R.; Nguyen, H.D.; Wang, X.; Bielinsky, A.K. Mcm10 deficiency causes defective-replisome-induced mutagenesis and a dependency on error-free postreplicative repair. Cell Cycle 2014, 13, 1737–1748, doi:10.4161/cc.28652.
- Kochenova, O.V.; Daee, D.L.; Mertz, T.M.; Shcherbakova, P.V. DNA polymerase ζ-dependent lesion bypass in Saccharomyces cerevisiae is accompanied by error-prone copying of long stretches of adjacent DNA. PLoS Genet. 2015, 11, e1005110, doi:10.1371/journal.pgen.1005110.
- Smith, D.J.; Whitehouse, I. Intrinsic coupling of lagging-strand synthesis to chromatin assembly. Nature 2012, 483, 434–438, doi:410.1038/nature10895.
- Sriramachandran, A.M.; Petrosino, G.; Méndez-Lago, M.; Schäfer, A.J.; Batista-Nascimento, L.S.; Zilio, N.; Ulrich, H.D. Genome-wide Nucleotide-Resolution Mapping of DNA Replication Patterns, Single-Strand Breaks, and Lesions by GLOE-Seq. Mol. Cell 2020, 78, 975–985, doi:10.1016/j.molcel.2020.03.027.
- Waisertreiger, I.S.; Liston, V.G.; Menezes, M.R.; Kim, H.M.; Lobachev, K.S.; Stepchenkova, E.I.; Tahirov, T.H.; Rogozin, I.B.; Pavlov, Y.I. Modulation of mutagenesis in eukaryotes by DNA replication fork dynamics and quality of nucleotide pools. Environ. Mol. Mutagen. 2012, 53, 699–724, doi:10.1002/em.21735.
- Pavlov, Y.I.; Mian, I.M.; Kunkel, T.A. Evidence for preferential mismatch repair of lagging strand DNA replication errors in yeast. Curr. Biol. 2003, 13, 744–748.
- Hoopes, J.I.; Cortez, L.M.; Mertz, T.M.; Malc, E.P.; Mieczkowski, P.A.; Roberts, S.A. APOBEC3A and APOBEC3B preferentially deaminate the lagging strand template during DNA replication. Cell Rep. 2016, 14, 1273–1282, doi:10.1016/j.celrep.2016.01.021.
- Bhagwat, A.S.; Hao, W.; Townes, J.P.; Lee, H.; Tang, H.; Foster, P.L. Strand-biased cytosine deamination at the replication fork causes cytosine to thymine mutations in Escherichia coli. Proc. Natl. Acad. Sci. USA 2016, 113, 2176–2181, doi:10.1073/pnas.1522325113.
- Seplyarskiy, V.B.; Soldatov, R.A.; Popadin, K.Y.; Antonarakis, S.E.; Bazykin, G.A.; Nikolaev, S.I. APOBEC-induced mutations in human cancers are strongly enriched on the lagging DNA strand during replication. Genome Res. 2016, 26, 174–182, doi:10.1101/gr.197046.115.
- Haradhvala, N.J.; Polak, P.; Stojanov, P.; Covington, K.R.; Shinbrot, E.; Hess, J.M.; Rheinbay, E.; Kim, J.; Maruvka, Y.E.; Braunstein, L.Z.; et al. Mutational strand asymmetries in cancer genomes reveal mechanisms of DNA damage and repair. Cell 2016, 164, 538–549, doi:10.1016/j.cell.2015.12.050.
- Cronan, G.E.; Kouzminova, E.A.; Kuzminov, A. Near-continuously synthesized leading strands in Escherichia coli are broken by ribonucleotide excision. Proc. Natl. Acad. Sci. USA 2019, 116, 1251–1260, doi:10.1073/pnas.1814512116.
- Burgers, P.M. Solution to the 50-year-old Okazaki-fragment problem. Proc. Natl. Acad. Sci. USA 2019, 116, 3358–3360, doi:10.1073/pnas.1900372116.





