
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Sangdun Choi | -- | 3274 | 2022-09-29 13:47:13 | | | |
| 2 | Peter Tang | Meta information modification | 3274 | 2022-09-30 03:49:06 | | |
Video Upload Options
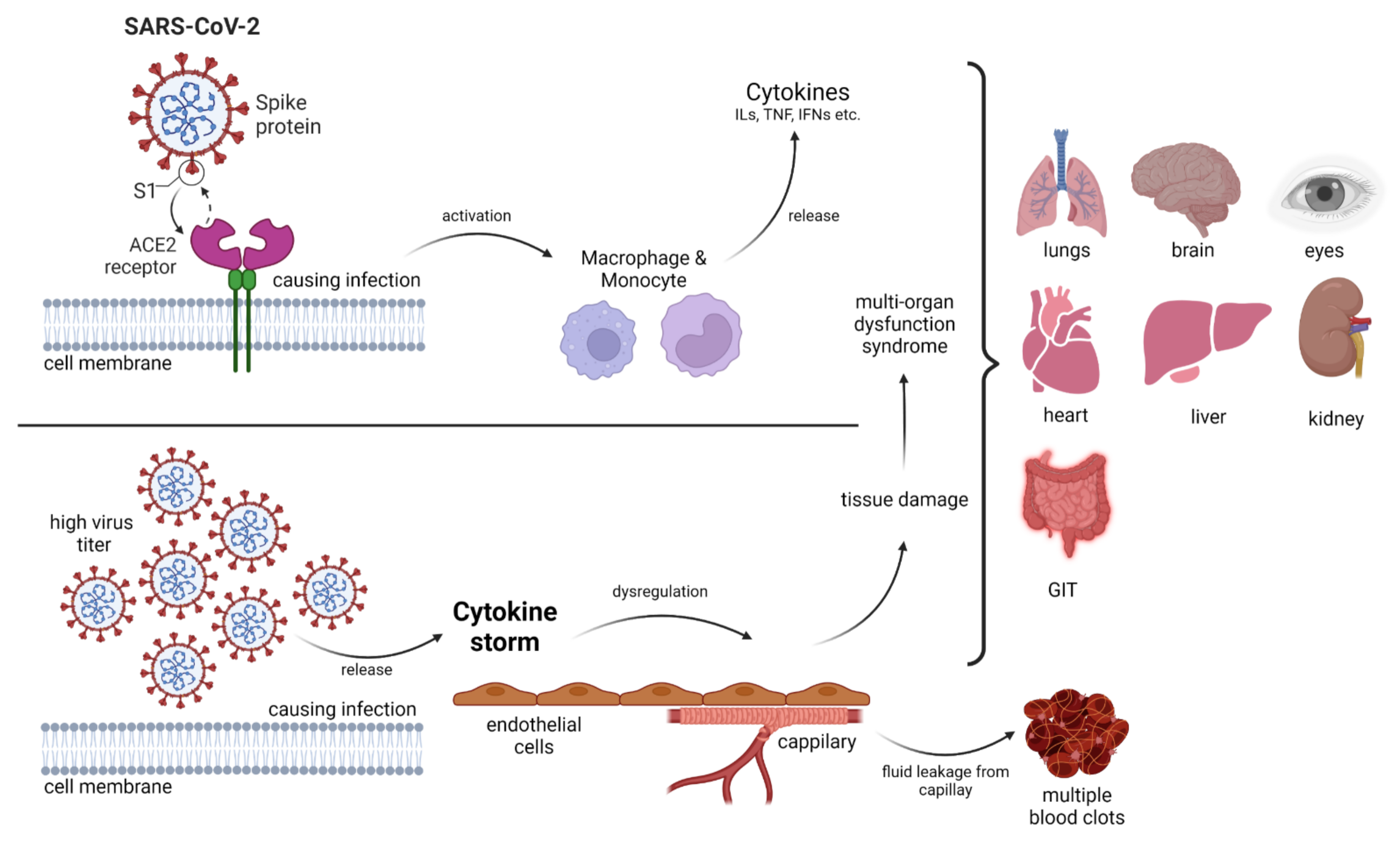
The innate immune system facilitates defense mechanisms against pathogen invasion and cell damage. Toll-like receptors (TLRs) assist in the activation of the innate immune system by binding to pathogenic ligands. This leads to the generation of intracellular signaling cascades including the biosynthesis of molecular mediators. TLRs on cell membranes are adept at recognizing viral components. Viruses can modulate the innate immune response with the help of proteins and RNAs that downregulate or upregulate the expression of various TLRs. In the case of COVID-19, molecular modulators such as type 1 interferons interfere with signaling pathways in the host cells, leading to an inflammatory response. Coronaviruses are responsible for an enhanced immune signature of inflammatory chemokines and cytokines. TLRs have been employed as therapeutic agents in viral infections as numerous antiviral Food and Drug Administration-approved drugs are TLR agonists.
1. Introduction
2. Structure of Coronavirus
|
Sr. No. |
SPs |
PDB ID |
Residues |
Physiological Significance |
Reference |
|---|---|---|---|---|---|
|
1 |
E |
7K3G |
76–109 |
Virus assembly, morphogenesis, viral–host interaction, membrane permeability |
[46] |
|
2 |
M |
8CTK |
220–260 |
Virus assembly, protein interactions (M–M, M–S, M–N) |
[47] |
|
3 |
N |
6VY0, 6YUN |
422 |
Abundant RNA-binding protein, virion genome packaging |
[48] |
|
4 |
S |
6VYB |
1273 |
Main antigen component, triggers the host immune response |
[49] |
|
Sr. No. |
NSPs |
PDB ID |
Residues |
Physiological Significance |
Reference |
|---|---|---|---|---|---|
|
1 |
NSP1 |
7K3N |
180 |
Protein synthesis, prevents antiviral activity of host cells, degrades host mRNA |
|
|
2 |
NSP2 |
7MSW |
638 |
Genome replication, disruption of intracellular host signaling |
|
|
3 |
NSP3 (Papain-like protease, PLpro) |
7KAG, 6WEY, 6WUU, 7LG0 |
1945 |
Integral to viral replication, post-translational processing of the two polyproteins, suppresses host protein synthesis |
|
|
4 |
NSP4 |
3GZF |
500 |
Protects new replicated virions, replication and assembly of viral structures in host cell |
|
|
5 |
NSP5 (3C-like protease, 3CLpro) |
6LU7 |
306 |
Protein cleavage capacity (conserved feature) |
|
|
6 |
NSP6 |
- |
290 |
Induction of autophagosomes, inhibition of viral components to reach host lysosomes |
|
|
7 |
NSP7 |
7JLT |
83 |
Primase complex (NSP7-NSP8), hetero-oligomeric complex (NSP7-NSP8-RdRp), viral replication |
|
|
8 |
NSP8 |
7JLT |
198 |
Primase complex (NSP7-NSP8), hetero-oligomeric complex (NSP7-NSP8-RdRp), viral replication |
|
|
9 |
NSP9 |
6WXD |
113 |
RNA synthesis, carries viral RNA to the host cell, responsible for proliferation |
|
|
10 |
NSP10 |
6ZPE |
139 |
Cofactor activation for replicative enzymes, complex NSP10-NSP14, viral RNA proofreading |
|
|
11 |
NSP11 |
- |
13 |
Cleavage product of PP1a by 3CLpro/MPro |
|
|
12 |
NSP12 (RNA polymerase, RdRp) |
6YYT |
932 |
RNA polymerase activity |
|
|
13 |
NSP13 |
6JYT |
601 |
Helicase activity |
|
|
14 |
NSP14 |
7R2V |
527 |
Viral RNA methylation, viral RNA proofreading, methyltransferase activity |
|
|
15 |
NSP15 |
6WXC |
346 |
Endoribonuclease activity |
|
|
16 |
NSP16 |
6WVN |
298 |
Viral replication, immune response evasion Viral RNA methylation, methyltransferase activity |
3. Overview of TLR Signaling
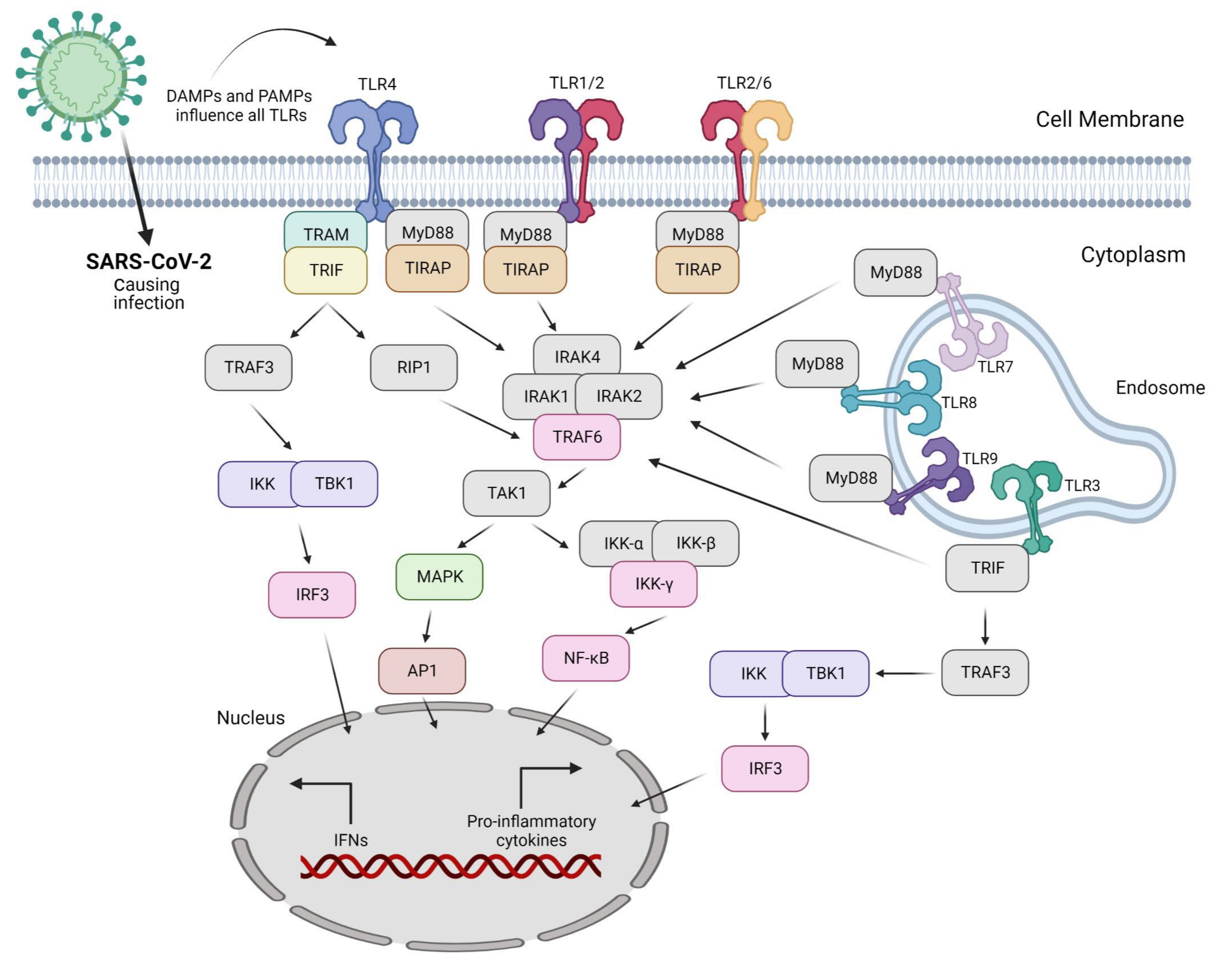
|
TLRs |
Ligand Recognition |
Form |
Localization |
Adaptor Molecules |
Negative Adaptors |
Response |
Reference |
|---|---|---|---|---|---|---|---|
|
TLR1 |
Triacyl lipopeptides, soluble factors |
Heterodimer |
Cell surface |
MyD88, Mal |
- |
NF-κB activation and proinflammatory cytokines |
|
|
TLR2 |
Hsp70, lipopeptide, HCV, Nonstructural protein 3 |
Heterodimer |
Cell surface |
MyD88, Mal |
- |
NF-κB activation and proinflammatory cytokines |
|
|
TLR3 |
dsRNA |
Homodimers |
Endosomal membrane |
TRIF |
SARM negatively regulates TRIF |
IRF activation, production of type 1 IFNs and proinflammatory cytokines |
|
|
TLR4 |
Lipopolysaccharide, Taxol, S protein of SARS-CoV-2 |
Homodimers |
Cell surface |
MyD88, Mal, TRIF, TRAM |
SARM negatively regulates TRIF and TRAM to consequently reduce inflammation |
Activation of NF-κB, pro-inflammatory cytokines, and IFN-inducible genes |
|
|
TLR5 |
Flagellin |
Homodimers |
Cell surface |
MyD88 |
- |
Activation of NF-κB and proinflammatory cytokines |
|
|
TLR6 |
Diacyl lipopeptides, lipoteichoic acid, fungal zymosan |
Heterodimer |
Cell surface |
MyD88, Mal/TIRAP |
- |
Activation of NF-κB and proinflammatory cytokines |
|
|
TLR7 |
SARS-CoV-2 ssRNA, imadozoquinoline |
Homodimers |
Endosomal membrane |
MyD88 |
- |
IRF activation, production of Type 1 IFNs and proinflammatory cytokines |
|
|
TLR8 |
SARS-CoV-2 ssRNA |
Endosomal membrane |
MyD88 |
- |
IRF activation, production of type 1 IFNs and proinflammatory cytokines |
||
|
TLR9 |
Unmethylated CPG-containing ssDNA, hemozoin from the malaria parasite |
Homodimers |
Endosomal membrane |
MyD88 |
- |
IRF activation, production of type 1 IFNs and proinflammatory cytokines |
4. Role of Antiviral Drugs Employing TLRs
|
Drugs |
TLRs |
Viruses |
Significance |
References |
|---|---|---|---|---|
|
Pam2CSK4 |
TLR2 |
Parainfluenza |
Reduced virus replication |
[138] |
|
INNA-051 |
TLR2 |
SARS-CoV-2 |
Reduces viral RNA load |
[139] |
|
PIKA |
TLR3 |
Influenza A |
Reduces virus load |
[140] |
|
Poly ICLC |
TLR3 |
HIV |
Release of IFN-α/β/γ |
[141] |
|
NA6 |
TLR4 |
Norovirus |
Induction of IFN-β |
[142] |
|
MPL |
TLR4 |
VZV |
Stimulate cytokines |
[143] |
|
Flagellin |
TLR5 |
Influenza A |
Reduces virus replication |
[144] |
|
CBLB502 |
TLR5 |
ConA |
Activation of NF-κB |
[145] |
|
Pam2CSK4 |
TLR6 |
Parainfluenza |
Reduces virus replication |
[138] |
|
INNA-051 |
TLR6 |
SARS-CoV-2 |
Reduces viral RNA load |
[139] |
|
GS-9620 |
TLR7 |
HIV |
Reactivates latency |
[109] |
|
Vesatolimod |
TLR7 |
HIV |
Modest delay in viral rebound |
[146] |
|
R848 |
TLR7/8 |
Zika |
Activation of NF-κB |
[147] |
|
GS-9688 |
TLR8 |
HBV |
Activation of dendritic and natural killer cells |
[148] |
|
ODN2395 |
TLR9 |
Parainfluenza |
Reduces viral replication |
[138] |
CBLB502—Entolimod; ConA—Concanavalin A; GS-9688—Selgantolimod; R848—Resiquimod; NA6—neoagarohexaose; VZV—Varicella-Zoster virus.
References
- Iwasaki, A.; Medzhitov, R. Control of Adaptive Immunity by the Innate Immune System. Nat. Immunol. 2015, 16, 343–353.
- Patra, M.C.; Choi, S. Recent Progress in the Development of Toll-like Receptor (TLR) Antagonists. Expert Opin. Ther. Pat. 2016, 26, 719–730.
- Nie, L.; Cai, S.-Y.; Shao, J.-Z.; Chen, J. Toll-Like Receptors, Associated Biological Roles, and Signaling Networks in Non-Mammals. Front. Immunol. 2018, 9, 1523.
- Liu, G.; Zhang, H.; Zhao, C.; Zhang, H. Evolutionary History of the Toll-Like Receptor Gene Family across Vertebrates. Genome Biol. Evol. 2020, 12, 3615–3634.
- Carty, M.; Bowie, A.G. Recent Insights into the Role of Toll-like Receptors in Viral Infection. Clin. Exp. Immunol. 2010, 161, 397–406.
- Abe, T.; Kaname, Y.; Hamamoto, I.; Tsuda, Y.; Wen, X.; Taguwa, S.; Moriishi, K.; Takeuchi, O.; Kawai, T.; Kanto, T.; et al. Hepatitis C Virus Nonstructural Protein 5A Modulates the Toll-like Receptor-MyD88-Dependent Signaling Pathway in Macrophage Cell Lines. J. Virol. 2007, 81, 8953–8966.
- Li, S.Y.; Chen, C.; Zhang, H.Q.; Guo, H.Y.; Wang, H.; Wang, L.; Zhang, X.; Hua, S.N.; Yu, J.; Xiao, P.G.; et al. Identification of Natural Compounds with Antiviral Activities against SARS-Associated Coronavirus. Antivir. Res. 2005, 67, 18.
- Maloney, G.; Schröder, M.; Bowie, A.G. Vaccinia Virus Protein A52R Activates P38 Mitogen-Activated Protein Kinase and Potentiates Lipopolysaccharide-Induced Interleukin-10. J. Biol. Chem. 2005, 280, 30838–30844.
- Stack, J.; Haga, I.R.; Schröder, M.; Bartlett, N.W.; Maloney, G.; Reading, P.C.; Fitzgerald, K.A.; Smith, G.L.; Bowie, A.G. Vaccinia Virus Protein A46R Targets Multiple Toll-like-Interleukin-1 Receptor Adaptors and Contributes to Virulence. J. Exp. Med. 2005, 201, 1007–1018.
- Blasius, A.L.; Beutler, B. Intracellular Toll-like Receptors. Immunity 2010, 32, 305–315.
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733.
- Onofrio, L.; Caraglia, M.; Facchini, G.; Margherita, V.; de Placido, S.; Buonerba, C. Toll-like Receptors and COVID-19: A Two-Faced Story with an Exciting Ending. Future Sci. OA 2020, 6, FSO605.
- Pearce, L.; Davidson, S.M.; Yellon, D.M. The Cytokine Storm of COVID-19: A Spotlight on Prevention and Protection. Expert Opin. Ther. Targets 2020, 24, 723–730.
- Kaushik, D.; Bhandari, R.; Kuhad, A. TLR4 as a Therapeutic Target for Respiratory and Neurological Complications of SARS-CoV-2. Expert Opin. Ther. Targets 2021, 25, 491–508.
- Kawasaki, T.; Kawai, T. Toll-like Receptor Signaling Pathways. Front. Immunol. 2014, 5, 461.
- O’Neill, L.A.J.; Bowie, A.G. The Family of Five: TIR-Domain-Containing Adaptors in Toll-like Receptor Signalling. Nat. Rev. Immunol. 2007, 7, 353–364.
- Takeuchi, O.; Akira, S. Innate Immunity to Virus Infection. Immunol. Rev. 2009, 227, 75–86.
- Chan, J.F.W.; Kok, K.H.; Zhu, Z.; Chu, H.; To, K.K.W.; Yuan, S.; Yuen, K.Y. Genomic Characterization of the 2019 Novel Human-Pathogenic Coronavirus Isolated from a Patient with Atypical Pneumonia after Visiting Wuhan. Emerg. Microbes Infect. 2020, 9, 221–236.
- Snijder, E.J.; Decroly, E.; Ziebuhr, J. The Nonstructural Proteins Directing Coronavirus RNA Synthesis and Processing. Adv. Virus Res. 2016, 96, 59.
- Shi, S.T.; Lai, M.M.C. Viral and Cellular Proteins Involved in Coronavirus Replication. In Coronavirus Replication and Reverse Genetics; Springer: Berlin/Heidelberg, Germany, 2005; Volume 287, p. 95.
- Guo, Y.R.; Cao, Q.D.; Hong, Z.S.; Tan, Y.Y.; Chen, S.D.; Jin, H.J.; Tan, K.S.; Wang, D.Y.; Yan, Y. The Origin, Transmission and Clinical Therapies on Coronavirus Disease 2019 (COVID-19) Outbreak-an Update on the Status. Mil. Med. Res. 2020, 7, 11.
- Han, Y.; Du, J.; Su, H.; Zhang, J.; Zhu, G.; Zhang, S.; Wu, Z.; Jin, Q. Identification of Diverse Bat Alphacoronaviruses and Betacoronaviruses in China Provides New Insights into the Evolution and Origin of Coronavirus-Related Diseases. Front. Microbiol. 2019, 10, 1900.
- Park, S.E. Epidemiology, Virology, and Clinical Features of Severe Acute Respiratory Syndrome -Coronavirus-2 (SARS-CoV-2; Coronavirus Disease-19). Clin. Exp. Pediatr. 2020, 63, 119–124.
- Gasmalbari, E.; Abbadi, O.S. Non-Structural Proteins of SARS-CoV-2 as Potential Sources for Vaccine Synthesis. Infect. Dis. Trop. Med. 2020, 6, e667.
- Gorla, U.S.; Rao, G.S.N.K. SARS-CoV-2: The Prominent Role of Non-Structural Proteins (NSPS) in COVID-19. Indian J. Pharm. Educ. Res. 2020, 54, S381–S389.
- Yoshimoto, F.K. The Proteins of Severe Acute Respiratory Syndrome Coronavirus-2 (SARS CoV-2 or n-COV19), the Cause of COVID-19. Protein J. 2020, 39, 198–216.
- Mousavizadeh, L.; Ghasemi, S. Genotype and Phenotype of COVID-19: Their Roles in Pathogenesis. J. Microbiol. Immunol. Infect. 2021, 54, 159–163.
- Mohammed, M.E.A. The Percentages of SARS-CoV-2 Protein Similarity and Identity with SARS-CoV and BatCoV RaTG13 Proteins Can Be Used as Indicators of Virus Origin. J. Proteins Proteom. 2021, 12, 81.
- Du, L.; He, Y.; Zhou, Y.; Liu, S.; Zheng, B.J.; Jiang, S. The Spike Protein of SARS-CoV--a Target for Vaccine and Therapeutic Development. Nat. Rev. Microbiol. 2009, 7, 226–236.
- Yuan, Y.; Cao, D.; Zhang, Y.; Ma, J.; Qi, J.; Wang, Q.; Lu, G.; Wu, Y.; Yan, J.; Shi, Y.; et al. Cryo-EM Structures of MERS-CoV and SARS-CoV Spike Glycoproteins Reveal the Dynamic Receptor Binding Domains. Nat. Commun. 2017, 8, 15092.
- Bojadzic, D.; Alcazar, O.; Chen, J.; Chuang, S.T.; Condor Capcha, J.M.; Shehadeh, L.A.; Buchwald, P. Small-Molecule Inhibitors of the Coronavirus Spike: ACE2 Protein-Protein Interaction as Blockers of Viral Attachment and Entry for SARS-CoV-2. ACS Infect. Dis. 2021, 7, 1519–1534.
- Satarker, S.; Nampoothiri, M. Structural Proteins in Severe Acute Respiratory Syndrome Coronavirus-2. Arch. Med. Res. 2020, 51, 482–491.
- Yan, X.; Hao, Q.; Mu, Y.; Timani, K.A.; Ye, L.; Zhu, Y.; Wu, J. Nucleocapsid Protein of SARS-CoV Activates the Expression of Cyclooxygenase-2 by Binding Directly to Regulatory Elements for Nuclear Factor-Kappa B and CCAAT/Enhancer Binding Protein. Int. J. Biochem. Cell Biol. 2006, 38, 1417–1428.
- Wang, Q.; Li, C.; Zhang, Q.; Wang, T.; Li, J.; Guan, W.; Yu, J.; Liang, M.; Li, D. Interactions of SARS Coronavirus Nucleocapsid Protein with the Host Cell Proteasome Subunit P42. Virol. J. 2010, 7, 99.
- Lu, X.; Pan, J.; Tao, J.; Guo, D. SARS-CoV Nucleocapsid Protein Antagonizes IFN-β Response by Targeting Initial Step of IFN-β Induction Pathway, and Its C-Terminal Region Is Critical for the Antagonism. Virus Genes 2011, 42, 37.
- Luo, H.; Chen, Q.; Chen, J.; Chen, K.; Shen, X.; Jiang, H. The Nucleocapsid Protein of SARS Coronavirus Has a High Binding Affinity to the Human Cellular Heterogeneous Nuclear Ribonucleoprotein A1. FEBS Lett. 2005, 579, 2623.
- Ruch, T.R.; Machamer, C.E. The Hydrophobic Domain of Infectious Bronchitis Virus E Protein Alters the Host Secretory Pathway and Is Important for Release of Infectious Virus. J. Virol. 2011, 85, 675–685.
- Kuo, L.; Hurst, K.R.; Masters, P.S. Exceptional Flexibility in the Sequence Requirements for Coronavirus Small Envelope Protein Function. J. Virol. 2007, 81, 2249–2262.
- Verdiá-Báguena, C.; Nieto-Torres, J.L.; Alcaraz, A.; Dediego, M.L.; Enjuanes, L.; Aguilella, V.M. Analysis of SARS-CoV E Protein Ion Channel Activity by Tuning the Protein and Lipid Charge. Biochim. Biophys. Acta 2013, 1828, 2026–2031.
- Venkatagopalan, P.; Daskalova, S.M.; Lopez, L.A.; Dolezal, K.A.; Hogue, B.G. Coronavirus Envelope (E) Protein Remains at the Site of Assembly. Virology 2015, 478, 75–85.
- Schoeman, D.; Fielding, B.C. Coronavirus Envelope Protein: Current Knowledge. Virol. J. 2019, 16, 69.
- Corse, E.; Machamer, C.E. Infectious Bronchitis Virus E Protein Is Targeted to the Golgi Complex and Directs Release of Virus-like Particles. J. Virol. 2000, 74, 4319–4326.
- Mortola, E.; Roy, P. Efficient Assembly and Release of SARS Coronavirus-like Particles by a Heterologous Expression System. FEBS Lett 2004, 576, 174–178.
- Fehr, A.R.; Perlman, S. Coronaviruses: An Overview of Their Replication and Pathogenesis. Methods Mol. Biol. 2015, 1282, 1–23.
- Narayanan, K.; Maeda, A.; Maeda, J.; Makino, S. Characterization of the Coronavirus M Protein and Nucleocapsid Interaction in Infected Cells. J. Virol. 2000, 74, 8127–8134.
- Liu, D.X.; Yuan, Q.; Liao, Y. Coronavirus Envelope Protein: A Small Membrane Protein with Multiple Functions. Cell Mol. Life Sci 2007, 64, 2043–2048.
- Arndt, A.L.; Larson, B.J.; Hogue, B.G. A Conserved Domain in the Coronavirus Membrane Protein Tail Is Important for Virus Assembly. J. Virol. 2010, 84, 11418–11428.
- Cubuk, J.; Alston, J.J.; Incicco, J.J.; Singh, S.; Stuchell-Brereton, M.D.; Ward, M.D.; Zimmerman, M.I.; Vithani, N.; Griffith, D.; Wagoner, J.A.; et al. The SARS-CoV-2 Nucleocapsid Protein Is Dynamic, Disordered, and Phase Separates with RNA. Nat. Commun. 2021, 12, 1936.
- Huang, Y.; Yang, C.; Xu, X.F.; Xu, W.; Liu, S. wen Structural and Functional Properties of SARS-CoV-2 Spike Protein: Potential Antivirus Drug Development for COVID-19. Acta Pharmacol. Sin. 2020, 41, 1141–1149.
- Züst, R.; Cervantes-Barragán, L.; Kuri, T.; Blakqori, G.; Weber, F.; Ludewig, B.; Thiel, V. Coronavirus Non-Structural Protein 1 Is a Major Pathogenicity Factor: Implications for the Rational Design of Coronavirus Vaccines. PLoS Pathog. 2007, 3, 1062–1072.
- Afsar, M.; Narayan, R.; Akhtar, M.N.; Das, D.; Rahil, H.; Nagaraj, S.K.; Eswarappa, S.M.; Tripathi, S.; Hussain, T. Drug Targeting Nsp1-Ribosomal Complex Shows Antiviral Activity against SARS-CoV-2. Elife 2022, 11, e74877.
- Kamitani, W.; Narayanan, K.; Huang, C.; Lokugamage, K.; Ikegami, T.; Ito, N.; Kubo, H.; Makino, S. Severe Acute Respiratory Syndrome Coronavirus Nsp1 Protein Suppresses Host Gene Expression by Promoting Host MRNA Degradation. Proc. Natl. Acad. Sci. USA 2006, 103, 12885–12890.
- Ma, J.; Chen, Y.; Wu, W.; Chen, Z. Structure and Function of N-Terminal Zinc Finger Domain of SARS-CoV-2 NSP2. Virol. Sin. 2021, 36, 1104–1112.
- Cornillez-Ty, C.T.; Liao, L.; Yates, J.R.; Kuhn, P.; Buchmeier, M.J. Severe Acute Respiratory Syndrome Coronavirus Nonstructural Protein 2 Interacts with a Host Protein Complex Involved in Mitochondrial Biogenesis and Intracellular Signaling. J. Virol. 2009, 83, 10314–10318.
- Angeletti, S.; Benvenuto, D.; Bianchi, M.; Giovanetti, M.; Pascarella, S.; Ciccozzi, M. COVID-2019: The Role of the Nsp2 and Nsp3 in Its Pathogenesis. J Med Virol 2020, 92, 584–588.
- Khan, M.T.; Zeb, M.T.; Ahsan, H.; Ahmed, A.; Ali, A.; Akhtar, K.; Malik, S.I.; Cui, Z.; Ali, S.; Khan, A.S.; et al. SARS-CoV-2 Nucleocapsid and Nsp3 Binding: An in Silico Study. Arch. Microbiol. 2021, 203, 59–66.
- Alsaadi, E.A.J.; Jones, I.M. Membrane Binding Proteins of Coronaviruses. Future Virol. 2019, 14, 275–286.
- Oostra, M.; te Lintelo, E.G.; Deijs, M.; Verheije, M.H.; Rottier, P.J.M.; de Haan, C.A.M. Localization and Membrane Topology of Coronavirus Nonstructural Protein 4: Involvement of the Early Secretory Pathway in Replication. J. Virol. 2007, 81, 12323–12336.
- Helmy, Y.A.; Fawzy, M.; Elaswad, A.; Sobieh, A.; Kenney, S.P.; Shehata, A.A. The COVID-19 Pandemic: A Comprehensive Review of Taxonomy, Genetics, Epidemiology, Diagnosis, Treatment, and Control. J. Clin. Med. 2020, 9, 1225.
- Stobart, C.C.; Sexton, N.R.; Munjal, H.; Lu, X.; Molland, K.L.; Tomar, S.; Mesecar, A.D.; Denison, M.R. Chimeric Exchange of Coronavirus Nsp5 Proteases (3CLpro) Identifies Common and Divergent Regulatory Determinants of Protease Activity. J. Virol. 2013, 87, 12611.
- Benvenuto, D.; Angeletti, S.; Giovanetti, M.; Bianchi, M.; Pascarella, S.; Cauda, R.; Ciccozzi, M.; Cassone, A. Evolutionary Analysis of SARS-CoV-2: How Mutation of Non-Structural Protein 6 (NSP6) Could Affect Viral Autophagy. J. Infect. 2020, 81, e24–e27.
- Cottam, E.M.; Whelband, M.C.; Wileman, T. Coronavirus NSP6 Restricts Autophagosome Expansion. Autophagy 2014, 10, 1426–1441.
- Sun, X.; Liu, Y.; Huang, Z.; Xu, W.; Hu, W.; Yi, L.; Liu, Z.; Chan, H.; Zeng, J.; Liu, X.; et al. SARS-CoV-2 Non-Structural Protein 6 Triggers NLRP3-Dependent Pyroptosis by Targeting ATP6AP1. Cell Death Differ. 2022, 29, 1240–1254.
- Biswal, M.; Diggs, S.; Xu, D.; Khudaverdyan, N.; Lu, J.; Fang, J.; Blaha, G.; Hai, R.; Song, J. Two Conserved Oligomer Interfaces of NSP7 and NSP8 Underpin the Dynamic Assembly of SARS-CoV-2 RdRP. Nucleic Acids Res. 2021, 49, 5956–5966.
- Krichel, B.; Falke, S.; Hilgenfeld, R.; Redecke, L.; Uetrecht, C. Processing of the SARS-CoV Pp1a/Ab Nsp7-10 Region. Biochem. J. 2020, 477, 1009–1019.
- Te Velthuis, A.J.W.; van den Worm, S.H.E.; Snijder, E.J. The SARS-Coronavirus Nsp7+nsp8 Complex Is a Unique Multimeric RNA Polymerase Capable of Both de Novo Initiation and Primer Extension. Nucleic Acids Res. 2012, 40, 1737–1747.
- Zeng, Z.; Deng, F.; Shi, K.; Ye, G.; Wang, G.; Fang, L.; Xiao, S.; Fu, Z.; Peng, G. Dimerization of Coronavirus Nsp9 with Diverse Modes Enhances Its Nucleic Acid Binding Affinity. J. Virol. 2018, 92, e00692-18.
- De Araújo, O.J.; Pinheiro, S.; Zamora, W.J.; Alves, C.N.; Lameira, J.; Lima, A.H. Structural, Energetic and Lipophilic Analysis of SARS-CoV-2 Non-Structural Protein 9 (NSP9). Sci. Rep. 2021, 11, 23003.
- Littler, D.R.; Gully, B.S.; Colson, R.N.; Rossjohn, J. Crystal Structure of the SARS-CoV-2 Non-Structural Protein 9, Nsp9. iScience 2020, 23, 101258.
- Rona, G.; Zeke, A.; Miwatani-Minter, B.; de Vries, M.; Kaur, R.; Schinlever, A.; Garcia, S.F.; Goldberg, H.V.; Wang, H.; Hinds, T.R.; et al. The NSP14/NSP10 RNA Repair Complex as a Pan-Coronavirus Therapeutic Target. Cell Death Differ. 2021, 29, 285–292.
- Ma, Y.; Wu, L.; Shaw, N.; Gao, Y.; Wang, J.; Sun, Y.; Lou, Z.; Yan, L.; Zhang, R.; Rao, Z. Structural Basis and Functional Analysis of the SARS Coronavirus Nsp14-Nsp10 Complex. Proc. Natl. Acad. Sci. USA 2015, 112, 9436–9441.
- Bouvet, M.; Lugari, A.; Posthuma, C.C.; Zevenhoven, J.C.; Bernard, S.; Betzi, S.; Imbert, I.; Canard, B.; Guillemot, J.C.; Lécine, P.; et al. Coronavirus Nsp10, a Critical Co-Factor for Activation of Multiple Replicative Enzymes. J. Biol. Chem. 2014, 289, 25783–25796.
- Gadhave, K.; Kumar, P.; Kumar, A.; Bhardwaj, T.; Garg, N.; Giri, R. Conformational Dynamics of 13 Amino Acids Long NSP11 of SARS-CoV-2 under Membrane Mimetics and Different Solvent Conditions. Microb. Pathog. 2021, 158, 105041.
- Elfiky, A.A. SARS-CoV-2 RNA Dependent RNA Polymerase (RdRp) Targeting: An in Silico Perspective. J. Biomol. Struct. Dyn. 2021, 39, 3204–3212.
- Khan, M.T.; Irfan, M.; Ahsan, H.; Ahmed, A.; Kaushik, A.C.; Khan, A.S.; Chinnasamy, S.; Ali, A.; Wei, D.-Q. Structures of SARS-CoV-2 RNA-Binding Proteins and Therapeutic Targets. Intervirology 2021, 64, 55–68.
- Mishra, A.; Rathore, A.S. RNA Dependent RNA Polymerase (RdRp) as a Drug Target for SARS-CoV2. J. Biomol. Struct. Dyn. 2022, 40, 6039–6051.
- Yin, W.; Mao, C.; Luan, X.; Shen, D.D.; Shen, Q.; Su, H.; Wang, X.; Zhou, F.; Zhao, W.; Gao, M.; et al. Structural Basis for Inhibition of the RNA-Dependent RNA Polymerase from SARS-CoV-2 by Remdesivir. Science 2020, 368, 1499.
- Wang, H.; Xue, S.; Yang, H.; Chen, C. Recent Progress in the Discovery of Inhibitors Targeting Coronavirus Proteases. Virol. Sin. 2016, 31, 24.
- Niu, X.; Kong, F.; Hou, Y.J.; Wang, Q. Crucial Mutation in the Exoribonuclease Domain of Nsp14 of PEDV Leads to High Genetic Instability during Viral Replication. Cell Biosci. 2021, 11, 106.
- Tahir, M. Coronavirus Genomic Nsp14-ExoN, Structure, Role, Mechanism, and Potential Application as a Drug Target. J. Med. Virol. 2021, 93, 4258–4264.
- Krafcikova, P.; Silhan, J.; Nencka, R.; Boura, E. Structural Analysis of the SARS-CoV-2 Methyltransferase Complex Involved in RNA Cap Creation Bound to Sinefungin. Nat. Commun. 2020, 11, 3717.
- Yoshimoto, F.K. A Biochemical Perspective of the Nonstructural Proteins (NSPs) and the Spike Protein of SARS CoV-2. Protein J. 2021, 40, 260–295.
- Vithani, N.; Ward, M.D.; Zimmerman, M.I.; Novak, B.; Borowsky, J.H.; Singh, S.; Bowman, G.R. SARS-CoV-2 Nsp16 Activation Mechanism and a Cryptic Pocket with Pan-Coronavirus Antiviral Potential. Biophys. J. 2021, 120, 2880–2889.
- Rosas-Lemus, M.; Minasov, G.; Shuvalova, L.; Inniss, N.L.; Kiryukhina, O.; Brunzelle, J.; Satchell, K.J.F. High-Resolution Structures of the SARS-CoV-2 2′-O-Methyltransferase Reveal Strategies for Structure-Based Inhibitor Design. Sci. Signal. 2020, 13, eabe1202.
- Fitzgerald, K.A.; Kagan, J.C. Toll-like Receptors and the Control of Immunity. Cell 2020, 180, 1044–1066.
- Balka, K.R.; de Nardo, D. Understanding Early TLR Signaling through the Myddosome. J. Leukoc. Biol. 2019, 105, 339–351.
- Satoh, T.; Akira, S. Toll-Like Receptor Signaling and Its Inducible Proteins. Microbiol. Spectr. 2016, 4, 4–6.
- Barton, G.M.; Kagan, J.C. A Cell Biological View of Toll-like Receptor Function: Regulation through Compartmentalization. Nat. Rev. Immunol. 2009, 9, 535–541.
- Wang, Y.; Song, E.; Bai, B.; Vanhoutte, P.M. Toll-like Receptors Mediating Vascular Malfunction: Lessons from Receptor Subtypes. Pharmacol. Ther. 2016, 158, 91–100.
- Kagan, J.C. Signaling Organelles of the Innate Immune System. Cell 2012, 151, 1168–1178.
- Yamamoto, M.; Sato, S.; Mori, K.; Hoshino, K.; Takeuchi, O.; Takeda, K.; Akira, S. Cutting Edge: A Novel Toll/IL-1 Receptor Domain-Containing Adapter That Preferentially Activates the IFN-Beta Promoter in the Toll-like Receptor Signaling. J. Immunol. 2002, 169, 6668–6672.
- Kawai, T.; Akira, S. The Role of Pattern-Recognition Receptors in Innate Immunity: Update on Toll-like Receptors. Nat. Immunol. 2010, 11, 373–384.
- Zhang, Y.; Liang, C. Innate Recognition of Microbial-Derived Signals in Immunity and Inflammation. Sci. China Life Sci. 2016, 59, 1210–1217.
- El-Zayat, S.R.; Sibaii, H.; Mannaa, F.A. Toll-like Receptors Activation, Signaling, and Targeting: An Overview. Bull. Natl. Res. Cent. 2019, 43, 187.
- Gao, W.; Xiong, Y.; Li, Q.; Yang, H. Inhibition of Toll-like Receptor Signaling as a Promising Therapy for Inflammatory Diseases: A Journey from Molecular to Nano Therapeutics. Front. Physiol. 2017, 8, 508.
- Lushpa, V.A.; Goncharuk, M.V.; Lin, C.; Zalevsky, A.O.; Talyzina, I.A.; Luginina, A.P.; Vakhrameev, D.D.; Shevtsov, M.B.; Goncharuk, S.A.; Arseniev, A.S.; et al. Modulation of Toll-like Receptor 1 Intracellular Domain Structure and Activity by Zn2+ Ions. Commun. Biol. 2021, 4, 1003.
- Takeuchi, O.; Sato, S.; Horiuchi, T.; Hoshino, K.; Takeda, K.; Dong, Z.; Modlin, R.L.; Akira, S. Cutting Edge: Role of Toll-Like Receptor 1 in Mediating Immune Response to Microbial Lipoproteins. J. Immunol. 2002, 169, 10–14.
- Ahmed, S.; Moawad, M.; Elhefny, R.; Chest, M.A.-E.J. Is Toll like Receptor 4 a Common Pathway Hypothesis for Development of Lung Cancer and Idiopathic Pulmonary Fibrosis? Elsevier: Amsterdam, The Netherlands, 2016.
- Oliveira-Nascimento, L.; Massari, P.; Wetzler, L.M. The Role of TLR2 Ininfection and Immunity. Front. Immunol. 2012, 3, 79.
- Zainol, M.I.B.; Kawasaki, T.; Monwan, W.; Murase, M.; Sueyoshi, T.; Kawai, T. Innate Immune Responses through Toll-like Receptor 3 Require Human-Antigen-R-Mediated Atp6v0d2 MRNA Stabilization. Sci. Rep. 2019, 9, 20406.
- Chen, Y.; Lin, J.; Zhao, Y.; Ma, X.; Yi, H. Toll-like Receptor 3 (TLR3) Regulation Mechanisms and Roles in Antiviral Innate Immune Responses. J. Zhejiang Univ. Sci. B 2021, 22, 609.
- Molteni, M.; Gemma, S.; Rossetti, C. The Role of Toll-Like Receptor 4 in Infectious and Noninfectious Inflammation. Mediators Inflamm. 2016, 2016, 6978936.
- Zhang, Y.; Liang, X.; Bao, X.; Xiao, W.; Chen, G. Toll-like Receptor 4 (TLR4) Inhibitors: Current Research and Prospective. Eur. J. Med. Chem. 2022, 235, 114291.
- Caballero, I.; Boyd, J.; Alminanã, C.; Sánchez-López, J.A.; Basatvat, S.; Montazeri, M.; Maslehat Lay, N.; Elliott, S.; Spiller, D.G.; White, M.R.H.; et al. Understanding the Dynamics of Toll-like Receptor 5 Response to Flagellin and Its Regulation by Estradiol. Sci. Rep. 2017, 7, 40981.
- Zhang, W.; Wang, L.; Sun, X.H.; Liu, X.; Xiao, Y.; Zhang, J.; Wang, T.; Chen, H.; Zhan, Y.Q.; Yu, M.; et al. Toll-like Receptor 5-Mediated Signaling Enhances Liver Regeneration in Mice. Mil. Med. Res. 2021, 8, 16.
- Kim, J.H.; Kordahi, M.C.; Chac, D.; William DePaolo, R. Toll-like Receptor-6 Signaling Prevents Inflammation and Impacts Composition of the Microbiota during Inflammation-Induced Colorectal Cancer. Cancer Prev. Res. 2020, 13, 25–40.
- De Almeida, L.A.; Macedo, G.C.; Marinho, F.A.V.; Gomes, M.T.R.; Corsetti, P.P.; Silva, A.M.; Cassataro, J.; Giambartolomei, G.H.; Oliveira, S.C. Toll-like Receptor 6 Plays an Important Role in Host Innate Resistance to Brucella Abortus Infection in Mice. Infect. Immun. 2013, 81, 1654–1662.
- Bortolotti, D.; Gentili, V.; Rizzo, S.; Schiuma, G.; Beltrami, S.; Strazzabosco, G.; Fernandez, M.; Caccuri, F.; Caruso, A.; Rizzo, R. TLR3 and TLR7 RNA Sensor Activation during SARS-CoV-2 Infection. Microorganisms 2021, 9, 1820.
- Macedo, A.B.; Novis, C.L.; de Assis, C.M.; Sorensen, E.S.; Moszczynski, P.; Huang, S.H.; Ren, Y.; Spivak, A.M.; Jones, R.B.; Planelles, V.; et al. Dual TLR2 and TLR7 Agonists as HIV Latency-Reversing Agents. JCI Insight 2018, 3.
- De Marcken, M.; Dhaliwal, K.; Danielsen, A.C.; Gautron, A.S.; Dominguez-Villar, M. TLR7 and TLR8 Activate Distinct Pathways in Monocytes during RNA Virus Infection. Sci. Signal. 2019, 12, eaaw1347.
- Salvi, V.; Nguyen, H.O.; Sozio, F.; Schioppa, T.; Gaudenzi, C.; Laffranchi, M.; Scapini, P.; Passari, M.; Barbazza, I.; Tiberio, L.; et al. SARS-CoV-2-Associated SsRNAs Activate Inflammation and Immunity via TLR7/8. JCI Insight 2021, 6, e150542.
- Costa, T.J.; Potje, S.R.; Fraga-Silva, T.F.C.; da Silva-Neto, J.A.; Barros, P.R.; Rodrigues, D.; Machado, M.R.; Martins, R.B.; Santos-Eichler, R.A.; Benatti, M.N.; et al. Mitochondrial DNA and TLR9 Activation Contribute to SARS-CoV-2-Induced Endothelial Cell Damage. Vascul. Pharmacol. 2022, 142, 106946.
- Khan, N.S.; Lukason, D.P.; Feliu, M.; Ward, R.A.; Lord, A.K.; Reedy, J.L.; Ramirez-Ortiz, Z.G.; Tam, J.M.; Kasperkovitz, P.V.; Negoro, P.E.; et al. CD82 Controls CpG-Dependent TLR9 Signaling. FASEB J. 2019, 33, 12500–12514.
- Lester, S.N.; Li, K. Toll-Like Receptors in Antiviral Innate Immunity. J. Mol. Biol. 2014, 426, 1246.
- Gaglia, M.M. Anti-Viral and pro-Inflammatory Functions of Toll-like Receptors during Gamma-Herpesvirus Infections. Virol. J. 2021, 18, 218.
- Shah, M.; Anwar, M.A.; Kim, J.H.; Choi, S. Advances in Antiviral Therapies Targeting Toll-like Receptors. Expert Opin. Investig. Drugs 2016, 25, 437–453.
- Vijay, K. Toll-like Receptors in Immunity and Inflammatory Diseases: Past, Present, and Future. Int. Immunopharmacol. 2018, 59, 391.
- Federico, S.; Pozzetti, L.; Papa, A.; Carullo, G.; Gemma, S.; Butini, S.; Campiani, G.; Relitti, N. Modulation of the Innate Immune Response by Targeting Toll-like Receptors: A Perspective on Their Agonists and Antagonists. J. Med. Chem. 2020, 63, 13466–13513.
- O’Neill, L.A.J.; Hennessy, E.J.; Parker, A.E. Targeting Toll-like Receptors: Emerging Therapeutics? Nat. Rev. Drug Discov. 2010, 9, 293–307.
- Zheng, W.; Xu, Q.; Zhang, Y.; Xiaofei, E.; Gao, W.; Zhang, M.; Zhai, W.; Rajkumar, R.S.; Liu, Z. Toll-like Receptor-Mediated Innate Immunity against Herpesviridae Infection: A Current Perspective on Viral Infection Signaling Pathways. Virol. J. 2020, 17, 192.
- Boehme, K.W.; Compton, T. Innate Sensing of Viruses by Toll-Like Receptors. J. Virol. 2004, 78, 7867.
- Ge, Y.; Mansell, A.; Ussher, J.E.; Brooks, A.E.S.; Manning, K.; Wang, C.J.H.; Taylor, J.A. Rotavirus NSP4 Triggers Secretion of Proinflammatory Cytokines from Macrophages via Toll-Like Receptor 2. J. Virol. 2013, 87, 11160–11167.
- Dela Justina, V.; Giachini, F.R.; Priviero, F.; Webb, R.C. COVID-19 and Hypertension: Is There a Role for DsRNA and Activation of Toll-like Receptor 3? Vascul. Pharmacol. 2021, 140, 106861.
- Lau, Y.F.; Tang, L.H.; Ooi, E.E. A TLR3 Ligand That Exhibits Potent Inhibition of Influenza Virus Replication and Has Strong Adjuvant Activity Has the Potential for Dual Applications in an Influenza Pandemic. Vaccine 2009, 27, 1354–1364.
- Miller, R.L.; Meng, T.C.; Tomai, M.A. The Antiviral Activity of Toll-like Receptor 7 and 7/8 Agonists. Drug News Perspect. 2008, 21, 69–87.
- Rose, W.A.; McGowin, C.L.; Pyles, R.B. FSL-1, a Bacterial-Derived Toll-like Receptor 2/6 Agonist, Enhances Resistance to Experimental HSV-2 Infection. Virol. J. 2009, 6, 195.
- Bowie, A.G.; Unterholzner, L. Viral Evasion and Subversion of Pattern-Recognition Receptor Signalling. Nat. Rev. Immunol. 2008, 8, 911–922.
- Duan, S.; Xu, X.; Wang, J.; Huang, L.; Peng, J.; Yu, T.; Zhou, Y.; Cheng, K.; Liu, S. TLR1/2 Agonist Enhances Reversal of HIV-1 Latency and Promotes NK Cell-Induced Suppression of HIV-1-Infected Autologous CD4 + T Cells. J. Virol. 2021, 95, e00816-21.
- Girkin, J.; Loo, S.L.; Esneau, C.; Maltby, S.; Mercuri, F.; Chua, B.; Reid, A.T.; Veerati, P.C.; Grainge, C.L.; Wark, P.A.B.; et al. TLR2-Mediated Innate Immune Priming Boosts Lung Anti-Viral Immunity. Eur. Respir. J. 2021, 58, 2001584.
- Howell, M.D.; Gallo, R.L.; Boguniewicz, M.; Jones, J.F.; Wong, C.; Streib, J.E.; Leung, D.Y.M. Cytokine Milieu of Atopic Dermatitis Skin Subverts the Innate Immune Response to Vaccinia Virus. Immunity 2006, 24, 341–348.
- Hutchens, M.A.; Luker, K.E.; Sonstein, J.; Núñez, G.; Curtis, J.L.; Luker, G.D. Protective Effect of Toll-like Receptor 4 in Pulmonary Vaccinia Infection. PLoS Pathog. 2008, 4, e1000153.
- Alexopoulou, L.; Holt, A.C.; Medzhitov, R.; Flavell, R.A. Recognition of Double-Stranded RNA and Activation of NF-KappaB by Toll-like Receptor 3. Nature 2001, 413, 732–738.
- Mazaleuskaya, L.; Veltrop, R.; Ikpeze, N.; Martin-Garcia, J.; Navas-Martin, S. Protective Role of Toll-like Receptor 3-Induced Type I Interferon in Murine Coronavirus Infection of Macrophages. Viruses 2012, 4, 901–923.
- Zhou, Y.; Wang, X.; Liu, M.; Hu, Q.; Song, L.; Ye, L.; Zhou, D.; Ho, W. A Critical Function of Toll-like Receptor-3 in the Induction of Anti-Human Immunodeficiency Virus Activities in Macrophages. Immunology 2010, 131, 40–49.
- Isogawa, M.; Robek, M.D.; Furuichi, Y.; Chisari, F.v. Toll-like Receptor Signaling Inhibits Hepatitis B Virus Replication in Vivo. J. Virol. 2005, 79, 7269–7272.
- Guillot, L.; le Goffic, R.; Bloch, S.; Escriou, N.; Akira, S.; Chignard, M.; Si-Tahar, M. Involvement of Toll-like Receptor 3 in the Immune Response of Lung Epithelial Cells to Double-Stranded RNA and Influenza A Virus. J. Biol. Chem. 2005, 280, 5571–5580.
- Zhang, M.; Yan, Z.; Wang, J.; Yao, X. Toll-like Receptors 7 and 8 Expression Correlates with the Expression of Immune Biomarkers and Positively Predicts the Clinical Outcome of Patients with Melanoma. Onco Targets Ther. 2017, 10, 4339.
- Drake, M.G.; Evans, S.E.; Dickey, B.F.; Fryer, A.D.; Jacoby, D.B. Toll-like Receptor-2/6 and Toll-like Receptor-9 Agonists Suppress Viral Replication but Not Airway Hyperreactivity in Guinea Pigs. Am J Respir Cell Mol. Biol. 2013, 48, 790–796.
- Proud, P.C.; Tsitoura, D.; Watson, R.J.; Chua, B.Y.; Aram, M.J.; Bewley, K.R.; Cavell, B.E.; Cobb, R.; Dowall, S.; Fotheringham, S.A.; et al. Prophylactic Intranasal Administration of a TLR2/6 Agonist Reduces Upper Respiratory Tract Viral Shedding in a SARS-CoV-2 Challenge Ferret Model. EBioMedicine 2021, 63, 103153.
- Lau, Y.F.; Tang, L.H.; Ooi, E.E.; Subbarao, K. Activation of the Innate Immune System Provides Broad-Spectrum Protection against Influenza A Viruses with Pandemic Potential in Mice. Virology 2010, 406, 80–87.
- Christopher, M.; Wong, J. Use of Toll-Like Receptor 3 Agonists Against Respiratory Viral Infections. Antiinflamm. Antiallergy Agents Med. Chem. 2011, 10, 327.
- Kim, M.; Lee, J.E.; Cho, H.; Jung, H.G.; Lee, W.; Seo, H.Y.; Lee, S.H.; Ahn, D.G.; Kim, S.J.; Yu, J.W.; et al. Antiviral Efficacy of Orally Delivered Neoagarohexaose, a Nonconventional TLR4 Agonist, against Norovirus Infection in Mice. Biomaterials 2020, 263, 120391.
- Luchner, M.; Reinke, S.; Milicic, A. TLR Agonists as Vaccine Adjuvants Targeting Cancer and Infectious Diseases. Pharmaceutics 2021, 13, 142.
- Georgel, A.F.; Cayet, D.; Pizzorno, A.; Rosa-Calatrava, M.; Paget, C.; Sencio, V.; Dubuisson, J.; Trottein, F.; Sirard, J.C.; Carnoy, C. Toll-like Receptor 5 Agonist Flagellin Reduces Influenza A Virus Replication Independently of Type I Interferon and Interleukin 22 and Improves Antiviral Efficacy of Oseltamivir. Antiviral Res. 2019, 168, 28–35.
- Melin, N.; Sánchez-Taltavull, D.; Fahrner, R.; Keogh, A.; Dosch, M.; Büchi, I.; Zimmer, Y.; Medová, M.; Beldi, G.; Aebersold, D.M.; et al. Synergistic Effect of the TLR5 Agonist CBLB502 and Its Downstream Effector IL-22 against Liver Injury. Cell Death Dis. 2021, 12, 366.
- SenGupta, D.; Brinson, C.; DeJesus, E.; Mills, A.; Shalit, P.; Guo, S.; Cai, Y.; Wallin, J.J.; Zhang, L.; Humeniuk, R.; et al. The TLR7 Agonist Vesatolimod Induced a Modest Delay in Viral Rebound in HIV Controllers after Cessation of Antiretroviral Therapy. Sci. Transl. Med. 2021, 13, eabg3071.
- Vanwalscappel, B.; Tada, T.; Landau, N.R. Toll-like Receptor Agonist R848 Blocks Zika Virus Replication by Inducing the Antiviral Protein Viperin. Virology 2018, 522, 199–208.
- Amin, O.E.; Colbeck, E.J.; Daffis, S.; Khan, S.; Ramakrishnan, D.; Pattabiraman, D.; Chu, R.; Micolochick Steuer, H.; Lehar, S.; Peiser, L.; et al. Therapeutic Potential of TLR8 Agonist GS-9688 (Selgantolimod) in Chronic Hepatitis B: Remodeling of Antiviral and Regulatory Mediators. Hepatology 2021, 74, 55–71.





