
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Richard Bleisch | -- | 2782 | 2022-07-28 15:20:30 | | | |
| 2 | Beatrix Zheng | + 34 word(s) | 2816 | 2022-07-29 07:57:15 | | | | |
| 3 | Beatrix Zheng | Meta information modification | 2816 | 2022-07-29 08:00:41 | | | | |
| 4 | Beatrix Zheng | Meta information modification | 2816 | 2022-07-29 08:15:55 | | | | |
| 5 | Beatrix Zheng | Meta information modification | 2816 | 2022-07-29 08:18:41 | | | | |
| 6 | Beatrix Zheng | Meta information modification | 2816 | 2022-07-29 08:24:03 | | | | |
| 7 | Beatrix Zheng | Meta information modification | 2816 | 2022-08-01 08:36:40 | | |
Video Upload Options
Microalgal biomass and metabolites can be used as a renewable source of nutrition, pharmaceuticals and energy to maintain or improve the quality of human life. Microalgae’s high volumetric productivity and low impact on the environment make them a promising raw material in terms of both ecology and economics. To optimize biotechnological processes with microalgae, improving the productivity and robustness of the cell factories is a major step towards economically viable bioprocesses. The success of a random mutagenesis approach using microalgae is determined by multiple factors involving the treatment of the cells before, during and after the mutagenesis procedure. Using photosynthetic microalgae, the supply of light quality and quantity, as well as the supply of carbon and nitrogen, are the most important factors. Besides the environmental conditions, the type of mutagen, its concentration and exposure time are among the main factors affecting the mutation result.
1. Physical Mutagens in Microalgal Biotechnology
1.1. Ultraviolet Light
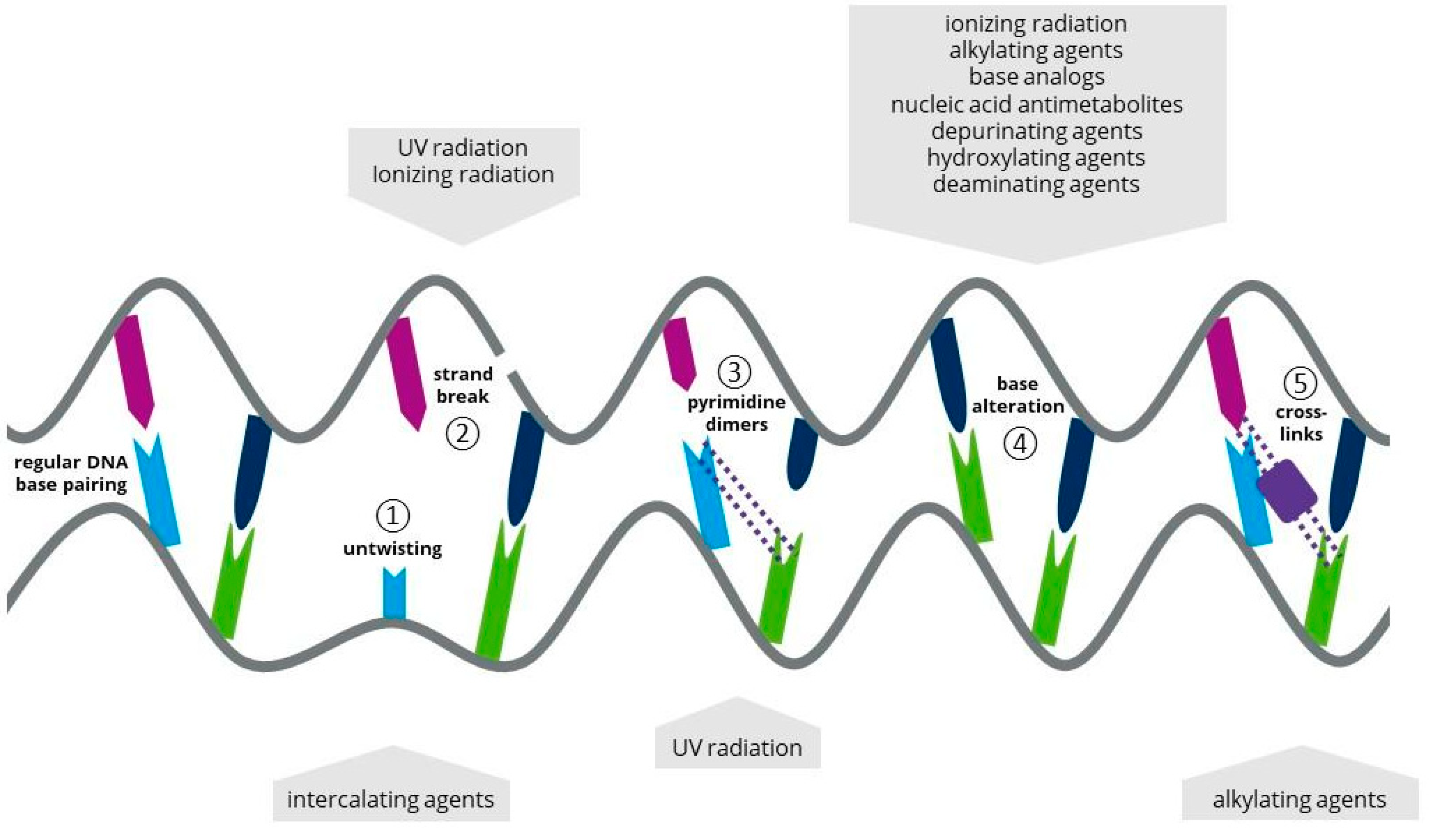
1.2. Ionizing Radiation
1.3. Atmospheric and Room Temperature Plasma
1.4. Laser Radiation
| Mutagen | Method, Exposure Time, Source, Distance, Recovery Time | Reference Microalgae | Mutation Results | References | ||
|---|---|---|---|---|---|---|
| Mutated trait |
WT * | M ** | ||||
| UV | UV 18 W, for 13 min, 15 cm, 24 h darkness | Chlorella vulgaris Y-019 |
neutral lipid accumulation [g/g dry wt] |
0.11 | 0.26 | [24] |
| UV-C | UV-C 253.7 nm, 30-W, 3–30 min, 9 cm, 24 h darkness | Chlorella sp. | protein content [g/L] | 0.0242 | 0.0688 | [25] |
| UV-C 254 nm 1.4 mW/cm2 for 60 s, 15 cm, 16 h darkness | Chlorella vulgaris | fatty acids 16:0;18:0, 20:0 [% of total fatty acids] | 27.9; 3.9; 11.9 | 47.4; 5.9; 19.9 | [26] | |
| UV-C 254 nm, 15 W, (Vilber–Lourmat, France), for 30–180 s, 5 cm, 24 h darkness | natural isolates of photosynthetic microorganism | lipid content though Nile red autofluorescence; with fluorescence emission | 35; 1081 | 983; 89,770 | [27] | |
| UV-C 40,000 μJ/cm, 254 nm, overnight darkness | Scenedesmus obliquus | trans-fatty acid productivity [g/(L·d)] |
0.095 | 0.112 | [28] | |
| UV-C 254 nm 340 mW cm2, for 3–32 min, 13.5 cm, 24 h darkness |
Isochrysis affinis galbana | total fatty acid [g/g dry wt] |
0.262 | 0.409 | [29] | |
| UV-C, for 1–10 min, 40 cm, overnight darkness | Chlorella vulgaris | lipid content [g/g] | 0.58 | 0.75 | [30] | |
| Gamma irradiation | 10 doses of irradiation 50–7000 kGy, 60Co gamma ray irradiator, room temperature |
Scenedesmus sp. | lipid productivity [g/L·d] |
0.0648 | 0.097 | [31] |
| ARTP | He RF power 100 W, plasma temperature 25–35 °C, for 20; 40; 60 and 80 s, 2 mm | Spirulina platensis | Carbohydrates productivity [g/L·d] |
0.0157 | 0.026 | [15] |
| He RF power 100 W, plasma temperature 25–35 °C, 20–60 s, 2 mm | Chlamydomonas reinhardtii | H2 production [mL/L] | ~16.1 | 84.1 | [32] | |
| He RF power 150 W, for 100 s | Crypthecodinium cohnii | biomass concentration [g dry wt/L] |
3.60 | 4.24 | [33] | |
| Heavy ion beam | 12 C6+ ion beam 31 keVµm−1 160 Gy, | Nannochloropsis oceanica | lipid productivity [g/L·d] | 0.211 | 0.295 | [34] |
| 12 C6+ ion beam, 90 Gy | Desmodesmus sp. | lipid productivity [g/L·d] | 0.247 | 0.298 | [35] | |
| Low-energy ion beam implementation | N+ ion beam chamber pressure 10−2 Pa Dose of implantation 0.3–3.3·1015 ions cm−2 s−1 |
Chlorella pyrenoidosa | lipid productivity [g/ L·d]; Lipid content [g/g dry wt] | 47.7; 0.337 | 64.4; 0.446 | [36] |
| laser radiation | He–Ne laser 808 nm, 6 W, 4 min, 24 h darkness | C. pyrenoidesa | lipid content [g/g dry wt] | 0.354 | 0.780 | [22] |
| Nd:YAG laser 1064 nm, 40 mW 8 min, 24 h darkness | Chlorella vulgaris | lipid content [g/g dry wt] | 0.315 | 0.525 | [22] | |
| Nd:YAG laser 1064 nm, 40 mW 2 min, 24 h darkness | Chlorella pacifica | lipid content [g L−1] |
0.033 | 0.088 | [37] | |
| semiconductor laser 632 nm, 40 mW, 4 min, 24 h darkness |
Chlorella pacifica | lipid content [g L−1] |
0.033 | 0.077 | [37] | |
2. Chemical Mutagens in Microalgal Biotechnology
2.1. Alkylating Agents as a Chemical Mutagen
2.2. Base Analogs (BAs) as a Chemical Mutagen
2.3. Antimetabolites (AMs) as a Chemical Mutagen
2.4. Intercalating Agents (IAs) as a Chemical Mutagen
2.5. Other Approaches for Chemical Mutagenesis
| Mutagen | Mutagen Concentration, Time of Exposure | Reference Microalgae | Mutation Results | References | ||
|---|---|---|---|---|---|---|
| Mutated trait |
WT * | M ** | ||||
| EMS | EMS 0.1–1.2 M for 60 min |
Nannochloropsis sp. | fatty acid methyl esters [g/g of dry wt] | 0.123 | 0.238 | [65] |
| EMS 0.4–1 g/L for 60–120 min |
Haematococcus pluvialis | total carotenoid; Astaxanthin [g/g of dry wt] |
0.02; 0.005 | 0.02; 0.019 |
[66] | |
| EMS 300 mM for 60 min | Chlorella vulgaris | protein content [g/g of dry wt] | 0.353 | 0.455 | [67] | |
| EMS 0.2–0.4 M for 2 h in darkness | Chlorella vulgaris | violaxanthin [mg/L culture] | 1.64 | 5.23 | [68] | |
| EMS 0.1–0.2 M | Phaeodactylum tricornutum | total carotenoids [g/g dry wt] | 0.009 | 0.011 | [69] | |
| EMS 0.2 M for 2 h in the dark |
Dunaliella tertiolecta | Zeaxanthin [μg/106·cells] | 0.131 | 0.359 | [70] | |
| EMS 20–40 µL/mL for 2 h | Chlamydomonas reinhardtii |
fatty acid methyl esters yield [%] | 6.53 | 7.56 | [71] | |
| EMS 0.2 M for 2 h in the dark |
Dunaliella salina |
carotenoid synthesis [Mol Car/Mol Chl] | 0.99 | 1.24 | [72] | |
| EMS 100 μ mol mL−1, for 30 min | Chlorella sp. | lipid content [g/g of dry wt]; productivity [g/(L·d)] | 0.247; 0.1536 | 0.356; 0.2487 | [73] | |
| EMS 0.4 M, for 60 min | Coelastrum sp. | Astaxanthin content [g/L] | 0.0145 | 0.0283 | [74] | |
| EMS + UV | UV + EMS 25 mM for 60 min | Chlorella vulgaris | lipid content [%] | 100 | 167 | [46] |
| UV 5–240 s, 245 nm + EMS 0.24 mol/L for 30 min | Nannochloropsis salina | fatty acid methyl ester [g/g of dry wt] | 0.175 | 0.787 | [75] | |
| MNNG | MNNG 0.1 mM for 60 min | Haematococcus pluvialis | Total carotenoid content [g/L] | ~0.067 | 0.089 | [41] |
| MNNG 5 µg/mL for 60 min | Chlorella sp. | max. growth rate under alkaline conditions [ d−1] | 0.064 | 0.554 | [76] | |
| MNNG 0.02 mol/L for 60 min |
Nannochloropsis oceanica |
Total lipid content [g/g] Lipid productivity [g/(L·d)] |
0.241; 0.0065 | 0.299; 0.0086 | [50] | |
| MNNG 0.1–0.2 M | Phaeodactylum tricornutum | total carotenoids [g/g dry wt] | 0.009 | 0.011 | [69] | |
| MNNG 0.2 mg/mL | Chlorella sorokiniana | Lutein content [g/L] | 0.025 | 0.042 | [44] | |
| MNNG 0.25–0.5 mM | Botryosphaerella sp. | lipid [g dry wt/(m2 day)]; biomass productivity [g dry wt/(m2·day)] | 1.0; 3.2 | 1.9; 5.4 | [45] | |
| NMU | NMU 5 mM for 60–90 min |
Nannochloropsis oculata | Total fatty acid [g/g dry wt] | 0.0634 | 0.0762 | [43] |
| DES + UV | UV 7–11 min 254 nm + DES 0.1–1.5% (V/V) 40 min |
Haematococcus pluvialis | astaxanthin content [mg/L] | ~0.031 | ~0.089 | [42] |
| 5BU | 5BU 1 mM for 48 h | Chlamydomonas reinhardtii | O2 tolerance [%] | 100 | 1400 | [77] |
| 5′FDU | 5′FDU 0.25 and 0.50 mM for 1 week | Chlorella vulgaris | fatty acids 16:0; 18:0; 20:0 [% of total fatty acids] |
27.9; 3.9; 11.9 | 46.9; 5.5; 18.5 | [26] |
| Acriflavin | Acriflavin 2–8 μg/mL for 1–3 d in darkness | Chlamydomonas reinhardtii zyklo | Loss of respiratory rate [nmol O2/(min·107 cells)] through loss of mitochondrial DNA | 23.2 | 3.7 | [64] |
3. Further Approaches in Random Mutagenesis
Recently, combined mutagenesis approaches have generated high interest as results indicated that they have a higher success rate than individual approaches. For instance, Wang et al. [42] applied a two-step random mutagenesis protocol to Haematococcus pluvialis cells using first UV irradiation, then EMS and DES mutagenesis, causing astaxanthin production to increase by a factor of 1.7 compared to the wild strain. Beacham et al. [75] used a reverse protocol for Nannochloropsis salina, starting with exposure to EMS, followed by UV irradiation, yielding a three-fold increase in cellular lipid accumulation. Comparable results were achieved by Sivaramakrishnan and Incharoensakdi [78], who exposed Scenedesmus sp. to UV irradiation in combination with oxidative stress by H2O2.
Other approaches can be used to select desired microalgal cells if the results obtained by random mutagenesis are insufficient. Among them, Adaptive Laboratory Evolution (ALE) is commonly used to adapt the physiology of cells to specific process conditions, such as high temperatures [79]. Its principle is based on natural selection, as presented in the Darwinian Theory, on the laboratory bench [80], and includes extensive cultivation in a specifically designed lab environment so that enhanced phenotypes can be selected after a long period of time [81]. The environmental conditions that can be altered include light irradiation, lack of nutrients, such as nitrogen, osmotic, temperature and oxidative stress [80][82][83]. Connecting the results of ALE with whole genome sequencing and “omics” methods enables gene functions to be discovered easily [81]. However, ALE does not prevent gene instability that might occur more often than in randomly mutated cells [79][82].
Additional environmental factors can be applied on microalgae; for example, Miazek et al. [84] reviewed the use of metals, metalloids and metallic nanoparticles to enhance cell characteristics. Moreover, phytohormones or chemicals acting as metabolic precursors have already been applied to microalgae [85]. A discussion of the methods used in the latter case exceeds the scope of this research.
More recently, a new technique was developed, known as Space Mutation Breeding (SMB). This technique may have direct or indirect effects on the growth and metabolic activities of microalgae, due to the unusual environment of space, characterized by high-energy ionic radiation, space’s magnetic field, ultra-high vacuum and microgravity [86]. The SMB technique provides some advantages, such as the great improvement in species’ qualities in a short time [87]. This was achieved by Chen Zishuo et al. [86], with a seawater Arthrospira platensis mutant, yielding a sugar content 62.26% higher than the wild type.
References
- Park, E.-J.; Choi, J.-I. Resistance and Proteomic Response of Microalgae to Ionizing Irradiation. Biotechnol. Bioprocess Eng. 2018, 23, 704–709.
- Chen, L.; Deng, S.; De Philippis, R.; Tian, W.; Wu, H.; Wang, J. UV-B resistance as a criterion for the selection of desert microalgae to be utilized for inoculating desert soils. J. Appl. Phycol. 2012, 25, 1009–1015.
- Rastogi, R.P.; Deng, S.; de Philippis, R.; Tian, W.; Wu, H.; Wang, J. Molecular Mechanisms of Ultraviolet Radiation-Induced DNA Damage and Repair. J. Nucleic Acids 2010, 2010, 592980.
- Tillich, U.M.; Lehmann, S.; Schulze, K.; Dühring, U.; Frohme, M. The Optimal Mutagen Dosage to Induce Point-Mutations in Synechocystis sp. PCC6803 and Its Application to Promote Temperature Tolerance. PLoS ONE 2012, 7, e49467.
- Holzinger, A.; Lütz, C. Algae and UV irradiation: Effects on ultrastructure and related metabolic functions. Micron 2006, 37, 190–207.
- Rastogi, R.P.; Sinha, R.P.; Moh, S.H.; Lee, T.K.; Kottuparambil, S.; Kim, Y.-J.; Rhee, J.-S.; Choi, E.-M.; Brown, M.; Häder, D.-P.; et al. Ultraviolet radiation and cyanobacteria. J. Photochem. Photobiol. B Biol. 2014, 141, 154–169.
- Graw, J. Genetik; Springer: Berlin/Heidelberg, Germany, 2010; ISBN 978-3-642-04998-9.
- Pfeifer, G.P.; You, Y.-H.; Besaratinia, A. Mutations induced by ultraviolet light. Mutat. Res. Mol. Mech. Mutagen. 2005, 571, 19–31.
- Yi, Z.; Xu, M.; Magnusdottir, M.; Zhang, Y.; Brynjolfsson, S.; Fu, W. Photo-Oxidative Stress-Driven Mutagenesis and Adaptive Evolution on the Marine Diatom Phaeodactylum tricornutum for Enhanced Carotenoid Accumulation. Mar. Drugs 2015, 13, 6138–6151.
- Sikder, S.; Biswas, P.; Hazra, P.; Akhtar, S.; Chattopadhyay, A.; Badigannavar, A.M.; D’Souza, S.F. Induction of mutation in tomato (Solanum lycopersicum L.) by gamma irradiation and EMS. Indian J. Genet. Plant Breed. 2013, 73, 392.
- Min, J.; Lee, C.W.; Gu, M.B. Gamma-radiation dose-rate effects on DNA damage and toxicity in bacterial cells. Radiat. Environ. Biophys. 2003, 42, 189–192.
- Klug, W.S.; Cummings, M.R.; Spencer, C.A.; Palladino, M.A. Concepts of Genetics, 11th ed.; Person Education Limited: Harlow, UK, 2016.
- Gomes, T.; Xie, L.; Brede, D.; Lind, O.-C.; Solhaug, K.A.; Salbu, B.; Tollefsen, K.E. Sensitivity of the green algae Chlamydomonas reinhardtii to gamma radiation: Photosynthetic performance and ROS formation. Aquat. Toxicol. 2017, 183, 1–10.
- Senthamilselvi, D.; Kalaiselvi, T. Gamma ray mutants of oleaginous microalga Chlorella sp. KM504965 with enhanced biomass and lipid for biofuel production. Biomass-Convers. Biorefinery 2022, 1–17.
- Fang, M.; Jin, L.; Zhang, C.; Tan, Y.; Jiang, P.; Ge, N.; Li, H.; Xing, X. Rapid Mutation of Spirulina platensis by a New Mutagenesis System of Atmospheric and Room Temperature Plasmas (ARTP) and Generation of a Mutant Library with Diverse Phenotypes. PLoS ONE 2013, 8, e77046.
- Fridman, G.; Brooks, A.D.; Balasubramanian, M.; Fridman, A.; Gutsol, A.; Vasilets, V.N.; Ayan, H.; Friedman, G. Comparison of Direct and Indirect Effects of Non-Thermal Atmospheric-Pressure Plasma on Bacteria. Plasma Process. Polym. 2007, 4, 370–375.
- Locke, B.; Sato, M.; Sunka, P.; Hoffmann, M.R.; Chang, J.-S. Electrohydraulic Discharge and Nonthermal Plasma for Water Treatment. Ind. Eng. Chem. Res. 2005, 45, 882–905.
- Gaunt, L.F.; Beggs, C.B.; Georghiou, G. Bactericidal Action of the Reactive Species Produced by Gas-Discharge Nonthermal Plasma at Atmospheric Pressure: A Review. IEEE Trans. Plasma Sci. 2006, 34, 1257–1269.
- Laroussi, M.; Leipold, F. Evaluation of the roles of reactive species, heat, and UV radiation in the inactivation of bacterial cells by air plasmas at atmospheric pressure. Int. J. Mass Spectrom. 2004, 233, 81–86.
- Zhang, X.; Zhang, X.-F.; Li, H.-P.; Wang, L.-Y.; Zhang, C.; Xing, X.-H.; Bao, C.-Y. Atmospheric and room temperature plasma (ARTP) as a new powerful mutagenesis tool. Appl. Microbiol. Biotechnol. 2014, 98, 5387–5396.
- Ouf, S.A.; Alsarrani, A.Q.; Al-Adly, A.A.; Ibrahim, M.K. Evaluation of low-intensity laser radiation on stimulating the cholesterol degrading activity: Part I. Microorganisms isolated from cholesterol-rich materials. Saudi J. Biol. Sci. 2012, 19, 185–193.
- Xing, W.; Zhang, R.; Shao, Q.; Meng, C.; Wang, X.; Wei, Z.; Sun, F.; Wang, C.; Cao, K.; Zhu, B.; et al. Effects of Laser Mutagenesis on Microalgae Production and Lipid Accumulation in Two Economically Important Fresh Chlorella Strains under Heterotrophic Conditions. Agronomy 2021, 11, 961.
- Wang, K.; Lin, B.; Meng, C.; Gao, Z.; Li, Z.; Zhang, H.; Du, H.; Xu, F.; Jiang, X. Screening of three Chlorella mutant strains with high lipid production induced by 3 types of lasers. J. Appl. Phycol. 2020, 32, 1655–1668.
- Deng, X.; Li, Y.; Fei, X. Effects of Selective Medium on Lipid Accumulation of Chlorellas and Screening of High Lipid Mutants through Ultraviolet Mutagenesis. Afr. J. Agric. Res. 2011, 6, 3768–3774.
- Liu, S.; Zhao, Y.; Liu, L.; Ao, X.; Ma, L.; Wu, M.; Ma, F. Improving Cell Growth and Lipid Accumulation in Green Microalgae Chlorella sp. via UV Irradiation. Appl. Biochem. Biotechnol. 2015, 175, 3507–3518.
- Anthony, J.; Rangamaran, V.R.; Gopal, D.; Shivasankarasubbiah, K.T.; Thilagam, M.L.J.; Dhassiah, M.P.; Padinjattayil, D.S.M.; Valsalan, V.N.; Manambrakat, V.; Dakshinamurthy, S.; et al. Ultraviolet and 5′Fluorodeoxyuridine Induced Random Mutagenesis in Chlorella vulgaris and Its Impact on Fatty Acid Profile: A New Insight on Lipid-Metabolizing Genes and Structural Characterization of Related Proteins. Mar. Biotechnol. 2014, 17, 66–80.
- Ardelean, A.V.; Ardelean, I.I.; Sicuia-Boiu, O.A.; Cornea, P. Random- Mutagenesis in Photosynthetic Microorganisms Further Selected with Respect to Increased Lipid Content. Agric. Life Life Agric. Conf. Proc. 2018, 1, 501–507.
- Breuer, G.; De Jaeger, L.; Artus, V.P.G.; Martens, D.E.; Springer, J.; Draaisma, R.B.; Eggink, G.; Wijffels, R.H.; Lamers, P.P. Superior triacylglycerol (TAG) accumulation in starchless mutants of Scenedesmus obliquus: (II) evaluation of TAG yield and productivity in controlled photobioreactors. Biotechnol. Biofuels 2014, 7, 70.
- Bougaran, G.; Rouxel, C.; Dubois, N.; Kaas, R.; Grouas, S.; Lukomska, E.; Le Coz, J.-R.; Cadoret, J.-P. Enhancement of neutral lipid productivity in the microalga Isochrysis affinis Galbana (T-Iso) by a mutation-selection procedure. Biotechnol. Bioeng. 2012, 109, 2737–2745.
- Carino, J.D.; Vital, P.G. Characterization of isolated UV-C-irradiated mutants of microalga Chlorella vulgaris for future biofuel application. Environ. Dev. Sustain. 2022, 1–18.
- Liu, B.; Ma, C.; Xiao, R.; Xing, D.; Ren, H.; Ren, N. The screening of microalgae mutant strain Scenedesmus sp. Z-4 with a rich lipid content obtained by 60Co γ-ray mutation. RSC Adv. 2015, 5, 52057–52061.
- Ban, S.; Lin, W.; Luo, Z.; Luo, J. Improving hydrogen production of Chlamydomonas reinhardtii by reducing chlorophyll content via atmospheric and room temperature plasma. Bioresour. Technol. 2018, 275, 425–429.
- Liu, B.; Sun, Z.; Ma, X.; Yang, B.; Jiang, Y.; Wei, D.; Chen, F. Mutation Breeding of Extracellular Polysaccharide-Producing Microalga Crypthecodinium cohnii by a Novel Mutagenesis with Atmospheric and Room Temperature Plasma. Int. J. Mol. Sci. 2015, 16, 8201–8212.
- Ma, Y.; Wang, Z.; Zhu, M.; Yu, C.; Cao, Y.; Zhang, D.; Zhou, G. Increased lipid productivity and TAG content in Nannochloropsis by heavy-ion irradiation mutagenesis. Bioresour. Technol. 2013, 136, 360–367.
- Hu, G.; Fan, Y.; Zhang, L.; Yuan, C.; Wang, J.; Li, W.; Hu, Q.; Li, F.-L. Enhanced Lipid Productivity and Photosynthesis Efficiency in a Desmodesmus sp. Mutant Induced by Heavy Carbon Ions. PLoS ONE 2013, 8, e60700.
- Tu, R.; Jin, W.; Wang, M.; Han, S.; Abomohra, A.E.-F.; Wu, W.-M. Improving of lipid productivity of the biodiesel promising green microalga Chlorella pyrenoidosa via low-energy ion implantation. J. Appl. Phycol. 2016, 28, 2159–2166.
- Zhang, H.; Gao, Z.; Li, Z.; Du, H.; Lin, B.; Cui, M.; Yin, Y.; Lei, F.; Yu, C.; Meng, C. Laser Radiation Induces Growth and Lipid Accumulation in the Seawater Microalga Chlorella pacifica. Energies 2017, 10, 1671.
- Drabløs, F.; Feyzi, E.; Aas, P.A.; Vaagbø, C.B.; Kavli, B.; Bratlie, M.S.; Peña-Diaz, J.; Otterlei, M.; Slupphaug, G.; Krokan, H.E. Alkylation damage in DNA and RNA—Repair mechanisms and medical significance. DNA Repair 2004, 3, 1389–1407.
- Slameňová, D.; Gábelová, A.; Ružeková, L.; Chalupa, I.; Horváthová, E.; Farkašová, T.; Bozsakyová, E.; Štětina, R. Detection of MNNG-induced DNA lesions in mammalian cells; validation of comet assay against DNA unwinding technique, alkaline elution of DNA and chromosomal aberrations. Mutat. Res. Repair 1997, 383, 243–252.
- Engelward, B.P.; Allan, J.M.; Dreslin, A.J.; Kelly, J.D.; Wu, M.M.; Gold, B.; Samson, L.D. A Chemical and Genetic Approach Together Define the Biological Consequences of 3-Methyladenine Lesions in the Mammalian Genome. J. Biol. Chem. 1998, 273, 5412–5418.
- Kamath, B.S.; Vidhyavathi, R.; Sarada, R.; Ravishankar, G. Enhancement of carotenoids by mutation and stress induced carotenogenic genes in Haematococcus pluvialis mutants. Bioresour. Technol. 2008, 99, 8667–8673.
- Wang, N.; Guan, B.; Kong, Q.; Sun, H.; Geng, Z.; Duan, L. Enhancement of astaxanthin production from Haematococcus pluvialis mutants by three-stage mutagenesis breeding. J. Biotechnol. 2016, 236, 71–77.
- Chaturvedi, R.; Uppalapati, S.R.; Alamsjah, M.A.; Fujita, Y. Isolation of quizalofop-resistant mutants of Nannochloropsis oculata (Eustigmatophyceae) with high eicosapentaenoic acid following N-methyl-N-nitrosourea-induced random mutagenesis. J. Appl. Phycol. 2004, 16, 135–144.
- Cordero, B.F.; Obraztsova, I.; Couso, I.; Leon, R.; Vargas, M.A.; Rodriguez, H. Enhancement of Lutein Production in Chlorella sorokiniana (Chorophyta) by Improvement of Culture Conditions and Random Mutagenesis. Mar. Drugs 2011, 9, 1607–1624.
- Nojima, D.; Ishizuka, Y.; Muto, M.; Ujiro, A.; Kodama, F.; Yoshino, T.; Maeda, Y.; Matsunaga, T.; Tanaka, T. Enhancement of Biomass and Lipid Productivities of Water Surface-Floating Microalgae by Chemical Mutagenesis. Mar. Drugs 2017, 15, 151.
- Sarayloo, E.; Simsek, S.; Ünlü, Y.S.; Cevahir, G.; Erkey, C.; Kavakli, I.H. Enhancement of the lipid productivity and fatty acid methyl ester profile of Chlorella vulgaris by two rounds of mutagenesis. Bioresour. Technol. 2018, 250, 764–769.
- Lian, M.; Huang, H.; Ren, L.; Ji, X.; Zhu, J.; Jin, L. Increase of Docosahexaenoic Acid Production by Schizochytrium sp. Through Mutagenesis and Enzyme Assay. Appl. Biochem. Biotechnol. 2009, 162, 935–941.
- Khromov-Borisov, N.N. Naming the mutagenic nucleic acid base analogs: The Galatea syndrome. Mutat. Res. Mol. Mech. Mutagen. 1997, 379, 95–103.
- Psoda, A.; Kierdaszuk, B.; Pohorille, A.; Geller, M.; Kusmierek, J.T.; Shugar, D. Interaction of the mutagenic base analogs O6-methylguanine and N4-hydroxycytosine with potentially complementary bases. Int. J. Quantum Chem. 1981, 20, 543–554.
- Wang, S.; Zhang, L.; Yang, G.; Han, J.; Thomsen, L.; Pan, K. Breeding 3 elite strains of Nannochloropsis oceanica by nitrosoguanidine mutagenesis and robust screening. Algal Res. 2016, 19, 104–108.
- Becket, E.; Tse, L.; Yung, M.; Cosico, A.; Miller, J.H. Polynucleotide Phosphorylase Plays an Important Role in the Generation of Spontaneous Mutations in Escherichia coli. J. Bacteriol. 2012, 194, 5613–5620.
- Cupples, C.G.; Miller, J.H. A set of lacZ mutations in Escherichia coli that allow rapid detection of each of the six base substitutions. Proc. Natl. Acad. Sci. USA 1989, 86, 5345–5349.
- Jackson-Grusby, L.; Laird, P.W.; Magge, S.N.; Moeller, B.J.; Jaenisch, R. Mutagenicity of 5-aza-2′-deoxycytidine is mediated by the mammalian DNA methyltransferase. Proc. Natl. Acad. Sci. USA 1997, 94, 4681–4685.
- Ang, J.; Song, L.Y.; D’Souza, S.; Hong, I.L.; Luhar, R.; Yung, M.; Miller, J.H. Mutagen Synergy: Hypermutability Generated by Specific Pairs of Base Analogs. J. Bacteriol. 2016, 198, 2776–2783.
- Azin, M.; Noroozi, E. Random mutagenesis and use of 2-deoxy-D-glucose as an antimetabolite for selection of α-amylase-overproducing mutants of Aspergillus oryzae. World J. Microbiol. Biotechnol. 2001, 17, 747–750.
- PubChem Floxuridine|C9H11FN2O5|CID 5790. Available online: https://pubchem.ncbi.nlm.nih.gov/compound/5790 (accessed on 3 June 2020).
- Sibirnyĭ, A.A.; Shavlovskiĭ, G.M.; Goloshchapova, G.V. Mutants of Pichia guilliermondii yeasts with multiple sensitivity to antibiotics and antimetabolites. I. The selection and properties of the mutants. Genetika 1977, 13, 872–879.
- Streisinger, G.; Okada, Y.; Emrich, J.; Newton, J.; Tsugita, A.; Terzaghi, E.; Inouye, M. Frameshift Mutations and the Genetic Code. Cold Spring Harb. Symp. Quant. Biol. 1966, 31, 77–84.
- Ferguson, L.R.; Denny, W.A. Genotoxicity of non-covalent interactions: DNA intercalators. Mutat. Res. Mol. Mech. Mutagen. 2007, 623, 14–23.
- Ferguson, L.R.; Turner, P.M.; Denny, W.A. The mutagenic effects of diacridines and diquinolines in microbial systems. Mutat. Res. Mol. Mech. Mutagen. 1990, 232, 337–343.
- Shafer, R.H.; Waring, M.J. DNA bis-intercalation: Application of theory to the binding of echinomycin to DNA. Biopolymers 1980, 19, 431–443.
- Ephrussi, B. Action de l’ac-Riflavine Sur Res Levures. I. La Mutation “Petite Colonie”. Ann. Inst. Pasteur 1949, 76, 351–367.
- Ferguson, L.R.; von Borstel, R. Induction of the cytoplasmic ‘petite’ mutation by chemical and physical agents in Saccharomyces cerevisiae. Mutat. Res. Mol. Mech. Mutagen. 1992, 265, 103–148.
- Matagne, R.F.; Michel-Wolwertz, M.-R.; Munaut, C.; Duyckaerts, C.; Sluse, F. Induction and characterization of mitochondrial DNA mutants in Chlamydomonas reinhardtii. J. Cell Biol. 1989, 108, 1221–1226.
- Doan, T.T.Y.; Obbard, J.P. Enhanced intracellular lipid in Nannochloropsis sp. via random mutagenesis and flow cytometric cell sorting. Algal Res. 2012, 1, 17–21.
- Gómez, P.I.; Inostroza, I.; Pizarro, M.; Pérez, J. From genetic improvement to commercial-scale mass culture of a Chilean strain of the green microalga Haematococcus pluvialis with enhanced productivity of the red ketocarotenoid astaxanthin. AoB Plants 2013, 5, plt026.
- Schüler, L.M.; de Morais, E.G.; Dos Santos, M.; Machado, A.; Carvalho, B.; Carneiro, M.; Maia, I.B.; Soares, M.; Duarte, P.; Barros, A.; et al. Isolation and Characterization of Novel Chlorella Vulgaris Mutants with Low Chlorophyll and Improved Protein Contents for Food Applications. Front. Bioeng. Biotechnol. 2020, 8, 469.
- Kim, J.; Kim, M.; Lee, S.; Jin, E. Development of a Chlorella vulgaris mutant by chemical mutagenesis as a producer for natural violaxanthin. Algal Res. 2020, 46, 101790.
- Yi, Z.; Su, Y.; Xu, M.; Bergmann, A.; Ingthorsson, S.; Rolfsson, O.; Salehi-Ashtiani, K.; Brynjolfsson, S.; Fu, W. Chemical Mutagenesis and Fluorescence-Based High-Throughput Screening for Enhanced Accumulation of Carotenoids in a Model Marine Diatom Phaeodactylum tricornutum. Mar. Drugs 2018, 16, 272.
- Kim, M.; Ahn, J.; Jeon, H.; Jin, E. Development of a Dunaliella tertiolecta Strain with Increased Zeaxanthin Content Using Random Mutagenesis. Mar. Drugs 2017, 15, 189.
- Lee, B.; Choi, G.-G.; Choi, Y.-E.; Sung, M.; Park, M.S.; Yang, J.-W. Enhancement of lipid productivity by ethyl methane sulfonate-mediated random mutagenesis and proteomic analysis in Chlamydomonas reinhardtii. Korean J. Chem. Eng. 2014, 31, 1036–1042.
- Jin, E.; Feth, B.; Melis, A. A mutant of the green algaDunaliella salina constitutively accumulates zeaxanthin under all growth conditions. Biotechnol. Bioeng. 2002, 81, 115–124.
- Nayak, M.; Suh, W.I.; Oh, Y.T.; Ryu, A.J.; Jeong, K.J.; Kim, M.; Mohapatra, R.K.; Lee, B.; Chang, Y.K. Directed evolution of Chlorella sp. HS2 towards enhanced lipid accumulation by ethyl methanesulfonate mutagenesis in conjunction with fluorescence-activated cell sorting based screening. Fuel 2022, 316, 123410.
- Tharek, A.; Yahya, A.; Salleh, M.; Jamaluddin, H.; Yoshizaki, S.; Hara, H.; Iwamoto, K.; Suzuki, I.; Mohamad, S.E. Improvement and screening of astaxanthin producing mutants of newly isolated Coelastrum sp. using ethyl methane sulfonate induced mutagenesis technique. Biotechnol. Rep. 2021, 32, e00673.
- Beacham, T.; Macia, V.M.; Rooks, P.; White, D.; Ali, S. Altered lipid accumulation in Nannochloropsis salina CCAP849/3 following EMS and UV induced mutagenesis. Biotechnol. Rep. 2015, 7, 87–94.
- Kuo, C.-M.; Lin, T.-H.; Yang, Y.-C.; Zhang, W.-X.; Lai, J.-T.; Wu, H.-T.; Chang, J.-S.; Lin, C.-S. Ability of an alkali-tolerant mutant strain of the microalga Chlorella sp. AT1 to capture carbon dioxide for increasing carbon dioxide utilization efficiency. Bioresour. Technol. 2017, 244, 243–251.
- Seibert, M.; Flynn, T.Y.; Ghirardi, M.L. Strategies for Improving Oxygen Tolerance of Algal Hydrogen Production. In Biohydrogen II; Elsevier: Amsterdam, The Netherlands, 2001; pp. 67–77. ISBN 978-0-08-043947-1.
- Sivaramakrishnan, R.; Incharoensakdi, A. Enhancement of lipid production in Scenedesmus sp. by UV mutagenesis and hydrogen peroxide treatment. Bioresour. Technol. 2017, 235, 366–370.
- Hu, X.; Tang, X.; Bi, Z.; Zhao, Q.; Ren, L. Adaptive evolution of microalgae Schizochytrium sp. under high temperature for efficient production of docosahexaeonic acid. Algal Res. 2021, 54, 102212.
- Mavrommati, M.; Daskalaki, A.; Papanikolaou, S.; Aggelis, G. Adaptive laboratory evolution principles and applications in industrial biotechnology. Biotechnol. Adv. 2021, 54, 107795.
- Dragosits, M.; Mattanovich, D. Adaptive laboratory evolution—Principles and applications for biotechnology. Microb. Cell Fact. 2013, 12, 64.
- Sun, X.-M.; Ren, L.-J.; Bi, Z.-Q.; Ji, X.-J.; Zhao, Q.-Y.; Huang, H. Adaptive evolution of microalgae Schizochytrium sp. under high salinity stress to alleviate oxidative damage and improve lipid biosynthesis. Bioresour. Technol. 2018, 267, 438–444.
- Pinheiro, M.J.; Bonturi, N.; Belouah, I.; Miranda, E.A.; Lahtvee, P.-J. Xylose Metabolism and the Effect of Oxidative Stress on Lipid and Carotenoid Production in Rhodotorula toruloides: Insights for Future Biorefinery. Front. Bioeng. Biotechnol. 2020, 8, 1008.
- Miazek, K.; Iwanek, W.; Remacle, C.; Richel, A.; Goffin, D. Effect of Metals, Metalloids and Metallic Nanoparticles on Microalgae Growth and Industrial Product Biosynthesis: A Review. Int. J. Mol. Sci. 2015, 16, 23929–23969.
- Yu, X.; Chen, L.; Zhang, W. Chemicals to enhance microalgal growth and accumulation of high-value bioproducts. Front. Microbiol. 2015, 6, 56.
- Chen, Z.; Tan, L.; Yang, B.; Wu, J.; Li, T.; Wu, H.; Wu, H.; Xiang, W. A mutant of seawater Arthrospira platensis with high polysaccharides production induced by space environment and its application potential. Algal Res. 2021, 61, 102562.
- Liu, L.X.; Guo, H.J.; Zhao, L.S.; Wang, J.; Gu, J.Y.; Zhao, S.R. Achievements and Perspectives of Crop Space Breeding in China. Induced Plant Mutations in the Genomics Era; FAO: Rome, Italy, 2009; pp. 213–215.





