Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Yuan, L.; Liangqi, C.; Xiyu, T.; Jinyao, L. Bacillus Nattokinase. Encyclopedia. Available online: https://encyclopedia.pub/entry/25520 (accessed on 06 June 2026).
Yuan L, Liangqi C, Xiyu T, Jinyao L. Bacillus Nattokinase. Encyclopedia. Available at: https://encyclopedia.pub/entry/25520. Accessed June 06, 2026.
Yuan, Li, Chen Liangqi, Tang Xiyu, Li Jinyao. "Bacillus Nattokinase" Encyclopedia, https://encyclopedia.pub/entry/25520 (accessed June 06, 2026).
Yuan, L., Liangqi, C., Xiyu, T., & Jinyao, L. (2022, July 26). Bacillus Nattokinase. In Encyclopedia. https://encyclopedia.pub/entry/25520
Yuan, Li, et al. "Bacillus Nattokinase." Encyclopedia. Web. 26 July, 2022.
Copy Citation
Bacillus nattokinase is a potential low-cost thrombolytic drug without side-effects and has been introduced into the consumer market as a functional food or dietary supplement.
Bacillus nattokinase
physiological and biochemical characteristics
molecular structure
1. Nattokinase Is a New Type of Thrombolytic Drug with Great Potential
The incidence of cardiovascular diseases has increased significantly worldwide and showed a trend of younger onset. Among cardiovascular diseases, cerebral infarction, ischemic stroke, and myocardial infarction are all related to thrombi formed by the coagulation of fibrin and platelets, and the current clinical application of thrombolytic agents including urokinase, tissue plasminogen activator (t-PA) and streptokinase all have serious side effects such as bleeding or gastric ulcer [1][2]. Therefore, the search for effective and safe thrombolytic drugs has become one of the directions in the field of cardiovascular disease research.
Natto, which is fermented by inoculating soybeans with Bacillus subtilis, is a traditional food with a long history in Japan. Intake of natto and other related fermented soy products is inversely related to the incidence of cardiovascular diseases, hence long-term consumption of natto is considered to be one of the important reasons for the longevity of Japanese [3]. Nattokinase is a kind of alkaline serine protease with strong fibrinolytic and thrombolytic activity, which is secreted by Bacillus natto and discovered in natto by Sumi et al. [4]. Compared with traditional thrombolytic drugs, nattokinase has a relatively lower risk of delivery, a larger tolerable dose, and lacks side effects such as gene mutation and chromosomal aberration induction [5][6]. More importantly, nattokinase also has various pharmacological effects, such as improving microcirculation and lowering blood pressure [7], anticoagulation [8], preventing atherosclerosis [9], relieving retinal angiogenesis [10], anticancer [11], inhibiting inflammation and oxidative stress, etc. [12]. In conclusion, nattokinase is a new type of thrombolytic drug with great application potential [13].
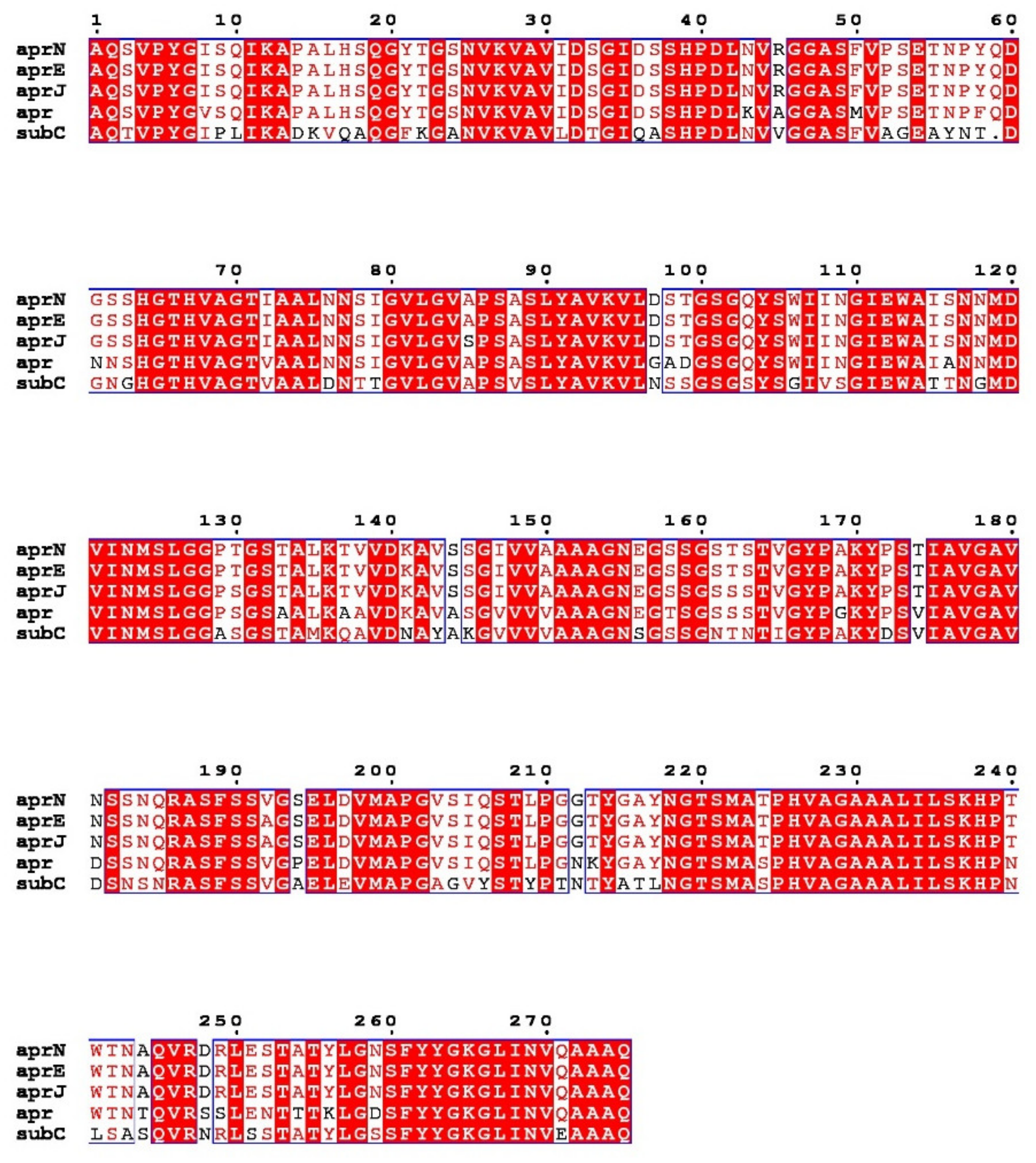 Figure 1. Multiple sequence alignment of nattokinase (AprN) compared with other serine protease homologs. Red shading and red words represent identical and similar residues, respectively. AprE: Subtilisin E; AprJ: Subtilisin BPN′; Apr: Subtilisin Carlsberg.
Figure 1. Multiple sequence alignment of nattokinase (AprN) compared with other serine protease homologs. Red shading and red words represent identical and similar residues, respectively. AprE: Subtilisin E; AprJ: Subtilisin BPN′; Apr: Subtilisin Carlsberg.
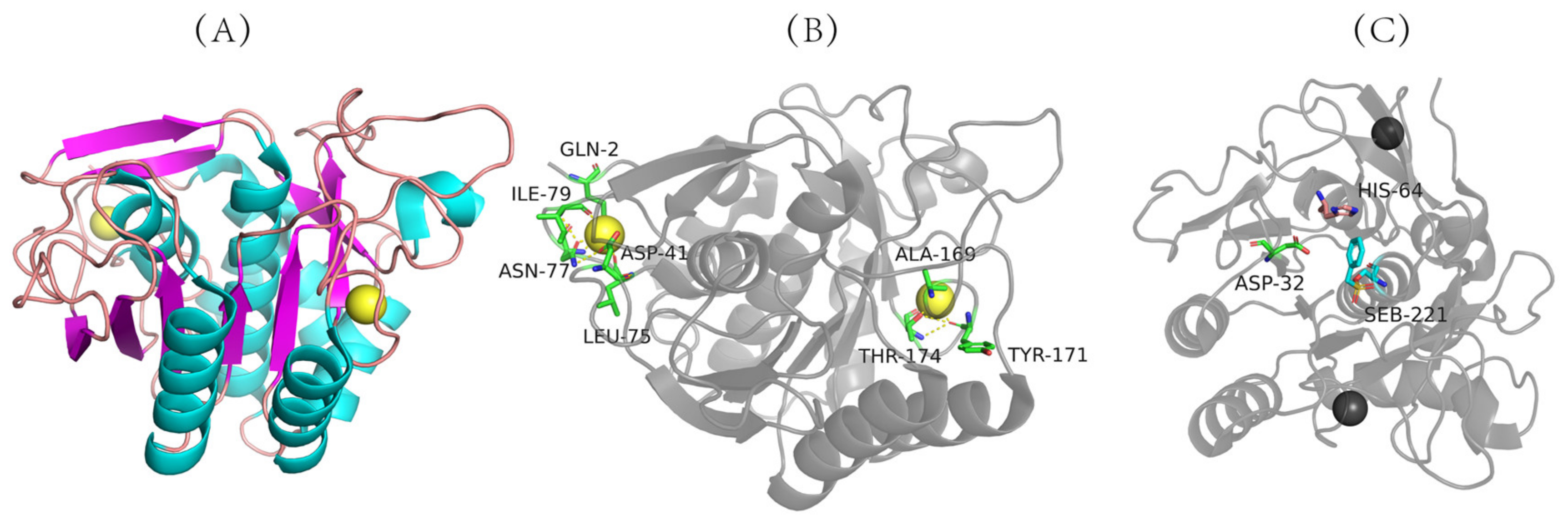 Figure 2. Three-dimensional structure of nattokinase. (A) Calcium binding site of nattokinase. (B) Three-dimensional structure of nattokinase. (C) Nattokinase triple catalyst. Nattokinase structure diagram taken from https://www.rcsb.org/structure/4DWW with modifications (accessed on 14 March 2012).
Figure 2. Three-dimensional structure of nattokinase. (A) Calcium binding site of nattokinase. (B) Three-dimensional structure of nattokinase. (C) Nattokinase triple catalyst. Nattokinase structure diagram taken from https://www.rcsb.org/structure/4DWW with modifications (accessed on 14 March 2012).
2. Bacillus Is the Main Strain for Synthesizing Nattokinase
Nattokinase is mainly produced by fermentation of Bacillus. Some marine organisms [14] and Pseudomonas sp. [15] also produce nattokinase, such as Pseudomonas aeruginosa CMSS [15] screened from cow’s milk and Pseudomonas sp. [16] obtained from the soil. Although nattokinase was first isolated from the Japanese food natto, similar fibrinolytic enzyme-producing strains are also available from other traditionally fermented foods: Bacillus subtilis Natto B-12 [17] and Bacillus subtilis JNFE0126 [18] were isolated from natto; Bacillus amyloliquefaciens DC-4 [19], Bacillus subtilis LD-8547 [20] and Bacillus sublitis DC33 [21] were obtained from Chinese traditional fermented food tempeh; Bacillus subtilis LSSE-62 [22] was obtained from Chinese soybean paste; and Bacillus sp. strain CK 11-4 [23] and Bacillus subtilis WRL101 [24] were isolated from Chungkook-Jang, a traditional Korean fermented food. In addition to fermented food sources, Bacillus cereus VITSDVM3 [25], isolated from rust, was also confirmed as a potent nattokinase producer (Table 1). Overall, most of the current research objects are mainly nattokinase derived from Bacillus subtilis natto screened in Japanese natto.
Table 1. Biodiverse sources of nattokinase.
| Strain | Source | References |
|---|---|---|
| Bacillus subtilis Natto B-12 | Natto | [17] |
| Bacillus sp. strain CK 11-4 | Chungkook-Jang | [23] |
| Bacillus sp. strain DJ-4 | Doen-Jang | [26] |
| Bacillus amyloliquefaciens DC-4 | Douchi | [19] |
| Bacillus subtilis DC33 | Douchi | [21] |
| Bacillus subtilis QK02 | Fermented soybeans | [27] |
| Bacillus subtilis LD-8547 | Douchi | [20] |
| Bacillus subtilis RJAS19 | Fermented soy products | [28] |
| Bacillus subtilis YJ1 | Fermented soy products | [29] |
| Bacillus subtilis TKU007 | Soil | [30] |
| Bacillus sublitis | Thailand | [31] |
| Bacillus subtilis LSSE-22 | Chinese soybean paste | [32] |
| Pseudomonas sp. TKU015 | Soil | [16] |
| Pseudomonas aeruginosa CMSS | Milk | [15] |
| Bacilluscereus VITSDVM3 | Rust | [25] |
| Bacillus subtilis WRL101 | Doen-jang | [24] |
| Bacillus velezensis KMU01 | Pickle | [33] |
| Bacillus subtilis VITMS 2 | Fermented milk of Vigna unguiculata. | [34] |
| Bacillussubtilis JNFE0126 | Natto | [18] |
| Bacillus subtilis K2 | Moromi | [35] |
| Bacillussubtilis LSSE-62 | Chinese soybean paste | [22] |
| Bacillus subtilis ICTF-1 | Ocean | [36] |
As a probiotic, Bacillus, which can synthesize nattokinase, has great potential in the fields of functional food and pharmaceutical applications. However, qualified oral nattokinase probiotics need to have the ability to overcome the special environment (gastric acid, bile salts, protease, etc.) of the human digestive system. However, none of the strains discovered so far have had their acid resistance and bile salt resistance reported. Therefore, the development of nattokinase synthetic probiotics adapted to the human digestive system has become one of the future research directions.
3. Structure and Catalytic Mechanism of Nattokinase
3.1. Nattokinase: The Only Member of the Alkaline Serine Protease Family with Thrombolytic Activity
Nattokinase (3.4.21.62) belongs to the family of alkaline serine proteases. As an endogenous fibrinolytic enzyme, nattokinase is functionally similar to human plasmin (3.4.21.7; 75 kDa). In 1992, Nakamura et al. used the shotgun method to determine that the gene encoding nattokinase (aprN) starts from GTG, has an open reading frame of 1146 bp, and encodes 381 amino acids, including a signal peptide of 29 amino acids, a propeptide of 77 amino acids, and a mature peptide of 275 amino acids with a molecular weight of 27.7 kDa. Since nattokinase is a cysteine-free protease, no disulfide bonds are observed in its structure. The open reading frame of nattokinase contains three consecutive terminators (TAATAGTAA) and is regulated by Rho-independent factors [37]. In silico analysis showed that nattokinase had 99.5%, 86%, and 72% sequence homology with subtilisin E, subtilisin BPN′, and subtilisin Carlsberg, which belong to the same alkaline serine protease family. The three amino acid residues (Ser221, His64 and Asp32) necessary for the catalytic center of serine proteases and the region near the catalytic triad are highly conserved among the above alkaline serine protease family members (Figure 1) [38]. Although nattokinase is highly homologous to many subtilisins in the serine protease family, only a few proteins, such as nattokinase, show high substrate specificity to fibrin and can directly cleave cross-linked fibrin in vitro and in vivo [39].
 Figure 1. Multiple sequence alignment of nattokinase (AprN) compared with other serine protease homologs. Red shading and red words represent identical and similar residues, respectively. AprE: Subtilisin E; AprJ: Subtilisin BPN′; Apr: Subtilisin Carlsberg.
Figure 1. Multiple sequence alignment of nattokinase (AprN) compared with other serine protease homologs. Red shading and red words represent identical and similar residues, respectively. AprE: Subtilisin E; AprJ: Subtilisin BPN′; Apr: Subtilisin Carlsberg.3.2. Structure and Reaction Mechanism of Nattokinase
The three-dimensional structure of nattokinase derived from Bacillus natto has been successfully analyzed (PDB code: 4DWW (2022) https://www.rcsb.org/structure/4DWW (accessed on 14 June 2022)) (Figure 2A), which shows that nattokinase is a single-chain polypeptide without disulfide bonds. Mature peptides consist of 9 α-helixes, 9 β-sheets and 2 Ca2+ binding sites (Gln2, Asp41, Leu75, Asn77, Ile79, Val81, Ala169, Tyr171, Thr174) for structural stability (Figure 2A,B). The catalytically active center of nattokinase consists of a conserved catalytic triad (Asp32, His64, Ser221), while its substrate-binding center contains three conserved amino acids (Ser125, Leu126, Gly127) (Figure 2B) [40]. Similarly to other subtilisin proteases, the seven typical β-sheets of nattokinase are located near the center of the enzyme molecule, and the other two β-sheets are inversely located in the domain near the C-terminus; the 9 β-sheets of nattokinase are assembled in reverse with the 9 α-helices, of which 7 α-helices are on the same surface [41].
 Figure 2. Three-dimensional structure of nattokinase. (A) Calcium binding site of nattokinase. (B) Three-dimensional structure of nattokinase. (C) Nattokinase triple catalyst. Nattokinase structure diagram taken from https://www.rcsb.org/structure/4DWW with modifications (accessed on 14 March 2012).
Figure 2. Three-dimensional structure of nattokinase. (A) Calcium binding site of nattokinase. (B) Three-dimensional structure of nattokinase. (C) Nattokinase triple catalyst. Nattokinase structure diagram taken from https://www.rcsb.org/structure/4DWW with modifications (accessed on 14 March 2012).The catalytic mechanism of nattokinase has not yet been reported. Since the three-dimensional structure of nattokinase highly overlaps with that of other alkaline serine proteases in its family, its molecular mechanism is similar to that of the alkaline serine protease family. First, the ring nitrogen atom of the His64 residue in the catalytically active center receives the hydroxyl proton of Ser221, which enhances the nucleophilic ability of Ser221 and attacks the hydroxyl carbon of the peptide bond of the substrate to form a tetrahedral transition state intermediate. Asp32 stabilizes the protonation state of His64 through the negative charge of the carboxyl group. Next, His64 donates a proton to the newly formed amino group to release the first product (acylation reaction) while forming a covalent acyl-enzyme complex. As the water molecule nucleophilically attacks the covalent acyl–enzyme complex to form a tetrahedral intermediate, His64 transfers the proton back to Ser221, and the transition state disintegrates to release the first product, thereby completing the deacylation reaction [40][41][42].
3.3. The Propeptide of Nattokinase Is Involved in the Correct Folding of Nattokinase as an Intramolecular Chaperone
The propeptide of nattokinase plays a key role in the correct folding of nattokinase. By comparing the thrombolytic activities of nattokinase holoenzyme (propeptide + mature peptide) and nattokinase mature peptide, Weng et al. found that only nattokinase expressing both propeptide and mature peptide has thrombolytic activity, inferring that the propeptide may be involved in the correct folding of nattokinase as an intramolecular chaperone [43]. Based on the structural similarity of serine protease family proteins, some studies have elucidated the role of the catalytic triplet in cleavage between the intramolecular chaperone and the nattokinase mature peptide. Asp32 assists in positioning the correct tautomer of His64, and Ser221 transfers its proton to His64 with increased nucleophilicity, which in turn completes substrate cleavage by nucleophilic attack on the carbonyl carbon of the propeptide’s peptide bond. However, the catalytic mechanism of this theory is being questioned [44].
The protein structure of nattokinase has now been resolved. However, the molecular mechanism of thrombolysis induced by nattokinase and the role of propeptide in the correct folding of nattokinase still need to be elucidated.
References
- Gianforcaro, A.; Kurz, M.; Guyette, F.; Callaway, C.W.; Rittenberger, J.C.; Elmer, J. Association of antiplatelet therapy with patient outcomes after out-of-hospital cardiac arrest. Resuscitation 2017, 121, 98–103.
- Furie, B.; Furie, B.C. Mechanisms of thrombus formation. N. Engl. J. Med. 2008, 359, 938–949.
- Nagata, C.; Wada, K.; Tamura, T.; Konishi, K.; Goto, Y.; Koda, S.; Kawachi, T.; Tsuji, M.; Nakamura, K. Dietary soy and natto intake and cardiovascular disease mortality in Japanese adults: The Takayama study. Am. J. Clin. Nutr. 2017, 105, 426–431.
- Sumi, H.; Hamada, H.; Tsushima, H.; Mihara, H.; Muraki, H. A novel fibrinolytic enzyme (nattokinase) in the vegetable cheese Natto; a typical and popular soybean food in the Japanese diet. Experientia 1987, 43, 1110–1111.
- Guo, H.; Ban, Y.-H.; Cha, Y.; An, E.S.; Choi, J.; Seo, D.W.; Park, D.; Choi, E.-K.; Kim, Y.-B. Comparative anti-thrombotic activity and haemorrhagic adverse effect of nattokinase and tissue-type plasminogen activator. Food Sci. Biotechnol. 2019, 28, 1535–1542.
- Wu, H.; Wang, H.; Xu, F.; Chen, J.; Duan, L.; Zhang, F. Acute toxicity and genotoxicity evaluations of Nattokinase, a promising agent for cardiovascular diseases prevention. Regul. Toxicol. Pharmacol. 2019, 103, 205–209.
- Hsia, C.-H.; Shen, M.-C.; Lin, J.-S.; Wen, Y.-K.; Hwang, K.-L.; Cham, T.-M.; Yang, N.-C. Nattokinase decreases plasma levels of fibrinogen, factor VII, and factor VIII in human subjects. Nutr. Res. 2009, 29, 190–196.
- Jensen, G.S.; Lenninger, M.; Ero, M.P.; Benson, K.F. Consumption of nattokinase is associated with reduced blood pressure and von Willebrand factor, a cardiovascular risk marker: Results from a randomized, double-blind, placebo-controlled, multicenter North American clinical trial. Integr. Blood Press. Control. 2016, 9, 95–104.
- Chang, C.-H.; Chen, K.-T.; Lee, T.-H.; Wang, C.-H.; Kuo, Y.-W.; Chiu, Y.-H.; Hsieh, C.-L.; Wu, C.-J.; Chang, Y.-L. Effects of natto extract on endothelial injury in a rat model. Acta Med. Okayama 2010, 64, 399–406.
- Huang, Z.; Ng, T.K.; Chen, W.; Sun, X.; Huang, D.; Zheng, D.; Yi, J.; Xu, Y.; Zhuang, X.; Chen, S. Nattokinase Attenuates Retinal Neovascularization via Modulation of Nrf2/HO-1 and Glial Activation. Investig. Ophthalmol. Vis. Sci. 2021, 62, 25.
- Yan, Y.; Wang, Y.; Qian, J.; Wu, S.; Ji, Y.; Liu, Y.; Zeng, J.; Gong, A. Nattokinase Crude Extract Inhibits Hepatocellular Carcinoma Growth in Mice. J. Microbiol. Biotechnol. 2019, 29, 1281–1287.
- Wu, H.; Wang, Y.; Zhang, Y.; Xu, F.; Chen, J.; Duan, L.; Zhang, T.; Wang, J.; Zhang, F. Breaking the vicious loop between inflammation, oxidative stress and coagulation, a novel anti-thrombus insight of nattokinase by inhibiting LPS-induced inflammation and oxidative stress. Redox Biol. 2020, 32, 101500.
- Fujita, M.; Hong, K.; Ito, Y.; Fujii, R.; Kariya, K.; Nishimuro, S. Thrombolytic Effect of Nattokinase on a Chemically Induced Thrombosis Model in Rat. Biol. Pharm. Bull. 1995, 18, 1387–1391.
- Pan, S.; Chen, G.; Zeng, J.; Cao, X.; Zheng, X.; Zeng, W.; Liang, Z. Fibrinolytic enzyme production from low-cost substrates by marine Bacillus subtilis: Process optimization and kinetic modeling. Biochem. Eng. J. 2019, 141, 268–277.
- Devi, C.S.; Mohanasrinivasan, V.; Sharma, P.; Das, D.; Vaishnavi, B.; Naine, S.J. Production, Purification and Stability Studies on Nattokinase: A Therapeutic Protein Extracted from Mutant Pseudomonas aeruginosa CMSS Isolated from Bovine Milk. Int. J. Pept. Res. Ther. 2015, 22, 263–269.
- Wang, S.-L.; Chen, H.-J.; Liang, T.-W.; Lin, Y.-D. A novel nattokinase produced by Pseudomonas sp. TKU015 using shrimp shells as substrate. Process Biochem. 2009, 44, 70–76.
- Wang, C.; Du, M.; Zheng, D.; Kong, F.; Zu, G.; Feng, Y. Purification and Characterization of Nattokinase from Bacillus subtilis Natto B-12. J. Agric. Food Chem. 2009, 57, 9722–9729.
- Zhang, X.; Tong, Y.; Wang, J.; Lyu, X.; Yang, R. Screening of a Bacillus subtilis strain producing both nattokinase and milk-clotting enzyme and its application in fermented milk with thrombolytic activity. J. Dairy Sci. 2021, 104, 9437–9449.
- Peng, Y.; Huang, Q.; Zhang, R.-H.; Zhang, Y.-Z. Purification and characterization of a fibrinolytic enzyme produced by Bacillus amyloliquefaciens DC-4 screened from douchi, a traditional Chinese soybean food. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2003, 134, 45–52.
- Wang, S.H.; Zhang, C.; Yang, Y.L.; Diao, M.; Bai, M.F. Screening of a high fibrinolytic enzyme producing strain and characterization of the fibrinolytic enzyme produced from Bacillus subtilis LD-8547. World J. Microbiol. Biotechnol. 2007, 24, 475–482.
- Wang, C.T.; Ji, B.P.; Li, B.; Nout, R.; Li, P.L.; Ji, H.; Chen, L.F. Purification and characterization of a fibrinolytic enzyme of Bacillus subtilis DC33, isolated from Chinese traditional Douchi. J. Ind. Microbiol. Biotechnol. 2006, 33, 750–758.
- Wei, X.; Luo, M.; Xu, L.; Zhang, Y.; Lin, X.; Kong, P.; Liu, H. Production of Fibrinolytic Enzyme from Bacillus amyloliquefaciens by Fermentation of Chickpeas, with the Evaluation of the Anticoagulant and Antioxidant Properties of Chickpeas. J. Agric. Food Chem. 2011, 59, 3957–3963.
- Kim, W.; Choi, K.; Kim, Y.; Park, H.; Choi, J.; Lee, Y.; Oh, H.; Kwon, I.; Lee, S. Purification and characterization of a fibrinolytic enzyme produced from Bacillus sp. strain CK 11-4 screened from Chungkook-Jang. Appl. Environ. Microbiol. 1996, 62, 2482–2488.
- Chang-Su, P.; Dong, H.K.; Woo-Yiel, L.; Dae-Ook, K.; Jae, J.S.; Nack-Shick, C. Identification of fibrinogen-induced nattokinase WRL101 from Bacillus subtilis WRL101 isolated from Doen-jang. Afr. J. Microbiol. Res. 2013, 7, 1983–1992.
- Vaithilingam, M.; Chandrasekaran, S.D.; Gupta, S.; Paul, D.; Sahu, P.; Selvaraj, J.N.; Babu, V. Extraction of Nattokinase Enzyme from Bacillus cereus Isolated from Rust. Natl. Acad. Sci. Lett. 2016, 39, 263–267.
- Kim, S.H.; Choi, N.S. Purification and Characterization of Subtilisin DJ-4 Secreted by Bacillus sp. Strain DJ-4 Screened from Doen-Jang. J. Agric. Chem. Soc. Jpn. 2000, 64, 1722–1725.
- Ko, J.H.; Yan, J.P.; Zhu, L.; Qi, Y.P. Identification of two novel fibrinolytic enzymes from Bacillus subtilis QK02. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2004, 137, 65–74.
- Kumar, D.M.; Rakshitha, R.; Vidhya, M.A.; Jennifer, P.S.; Prasad, S.; Kumar, M.R.; Kalaichelv, P. Production, Optimization and Characterization of Fibrinolytic Enzyme by Bacillus subtilis RJAS19. Pak. J. Biol. Sci. 2014, 17, 529–534.
- Yin, L.-J.; Lin, H.-H.; Jiang, S.-T. Bioproperties of Potent Nattokinase from Bacillus subtilis YJ1. J. Agric. Food Chem. 2010, 58, 5737–5742.
- Wang, S.-L.; Yeh, P.-Y. Production of a surfactant- and solvent-stable alkaliphilic protease by bioconversion of shrimp shell wastes fermented by Bacillus subtilis TKU007. Process Biochem. 2006, 41, 1545–1552.
- Inatsu, Y.; Nakamura, N.; Yuriko, Y.; Fushimi, T.; Watanasiritum, L.; Kawamoto, S. Characterization of Bacillus subtilis strains in Thua nao, a traditional fermented soybean food in northern Thailand. Lett. Appl. Microbiol. 2006, 43, 237–242.
- Wei, X.; Luo, M.; Xie, Y.; Yang, L.; Li, H.; Xu, L.; Liu, H. Strain Screening, Fermentation, Separation, and Encapsulation for Production of Nattokinase Functional Food. Appl. Biochem. Biotechnol. 2012, 168, 1753–1764.
- Heo, S.; Kim, J.-H.; Kwak, M.-S.; Sung, M.-H.; Jeong, D.-W. Functional Annotation Genome Unravels Potential Probiotic Bacillus velezensis Strain KMU01 from Traditional Korean Fermented Kimchi. Foods 2021, 10, 563.
- Keziah, S.M.; Devi, C.S. Fibrinolytic and ACE Inhibitory Activity of Nattokinase Extracted from Bacillus subtilis VITMS 2: A Strain Isolated from Fermented Milk of Vigna unguiculata. J. Protein Chem. 2021, 40, 876–890.
- Syahbanu, F.; Giriwono, P.E.; Tjandrawinata, R.R.; Suhartono, M.T. Molecular analysis of a fibrin-degrading enzyme from Bacillus subtilis K2 isolated from the Indonesian soy-bean-based fermented food moromi. Mol. Biol. Rep. 2020, 47, 8553–8563.
- Mahajan, P.M.; Nayak, S.; Lele, S.S. Fibrinolytic enzyme from newly isolated marine bacterium Bacillus subtilis ICTF-1: Media optimization, purification and characterization. J. Biosci. Bioeng. 2012, 113, 307–314.
- Nakamura, T.; Yamagata, Y.; Ichishima, E. Nucleotide Sequence of the Subtilisin NAT Gene, aprN, of Bacillus subtilis(natto). Biosci. Biotechnol. Biochem. 1992, 56, 1869–1871.
- Kobayashi, T.; Hakamada, Y.; Adachi, S.; Hitomi, J.; Yoshimatsu, T.; Koike, K.; Kawai, S.; Ito, S. Purification and properties of an alkaline protease from alkalophilic Bacillus sp. KSM-K16. Appl. Micro-Biol. Biotechnol. 1995, 43, 473–481.
- Song, X.; Yang, Y.; Liang, N.; Yang, F.; Chen, S.; Zhou, L.; Zhou, K.; Wang, Y. Quantitative pharmacokinetic evaluation of Subtilisin QK-2 after a bolus IV injection in a rat model using a novel sandwich enzyme-linked immunosorbent assay. J. Pharm. Biomed. Anal. 2020, 186, 113264.
- Carter, P.; Wells, J.A. Dissecting the catalytic triad of a serine protease. Nature 1988, 332, 564–568.
- Yanagisawa, Y.; Chatake, T.; Chiba-Kamoshida, K.; Naito, S.; Ohsugi, T.; Sumi, H.; Yasuda, I.; Morimoto, Y. Purification, crystallization and preliminary X-ray diffraction experiment of nattokinase from Bacillus subtilis natto. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 2010, 66 Pt 12, 1670–1673.
- Carter, P.; Wells, J.A. Functional interaction among catalytic residues in subtilisin BPN′. Proteins Struct. Funct. Bioinform. 1990, 7, 335–342.
- Weng, M.; Zheng, Z.; Bao, W.; Cai, Y.; Yin, Y.; Zou, G. Enhancement of oxidative stability of the subtilisin nattokinase by site-directed mutagenesis expressed in Escherichia coli. Biochim. Biophys. Acta 2009, 1794, 1566–1572.
- Yang, M.; Wu, J.; Huang, Q.; Jia, Y. Probing the Role of Catalytic Triad on the Cleavage between Intramolecular Chaperone and NK Mature Peptide. J. Agric. Food Chem. 2021, 69, 2348–2353.
More
Information
Subjects:
Food Science & Technology
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.4K
Revisions:
2 times
(View History)
Update Date:
28 Jul 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





