
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | MENGXIAN CHI | -- | 1465 | 2024-02-29 13:21:20 | | | |
| 2 | Jessie Wu | + 6 word(s) | 1471 | 2024-03-01 04:55:22 | | |
Video Upload Options
Brain tumor segmentation plays a crucial role in the diagnosis, treatment planning, and monitoring of brain tumors. Accurate segmentation of brain tumor regions from multi-sequence magnetic resonance imaging (MRI) data is of paramount importance for precise tumor analysis and subsequent clinical decision making. The ability to delineate tumor boundaries in MRI scans enables radiologists and clinicians to assess tumor size, location, and heterogeneity, facilitating treatment planning and evaluating treatment response. Traditional manual segmentation methods are time-consuming, subjective, and prone to inter-observer variability. Therefore, the automatic segmentation algorithm has received widespread attention as an alternative solution. For instance, the self-organizing map (SOM) is an unsupervised exploratory data analysis tool that leverages principles of vector quantization and similarity measurement to automatically partition images into self-similar regions or clusters. Segmentation methods based on SOM have demonstrated the ability to distinguish high-level and low-level features of tumors, edema, necrosis, cerebrospinal fluid, and healthy tissue.
1. Attention Mechanisms in Convolutional Neural Networks
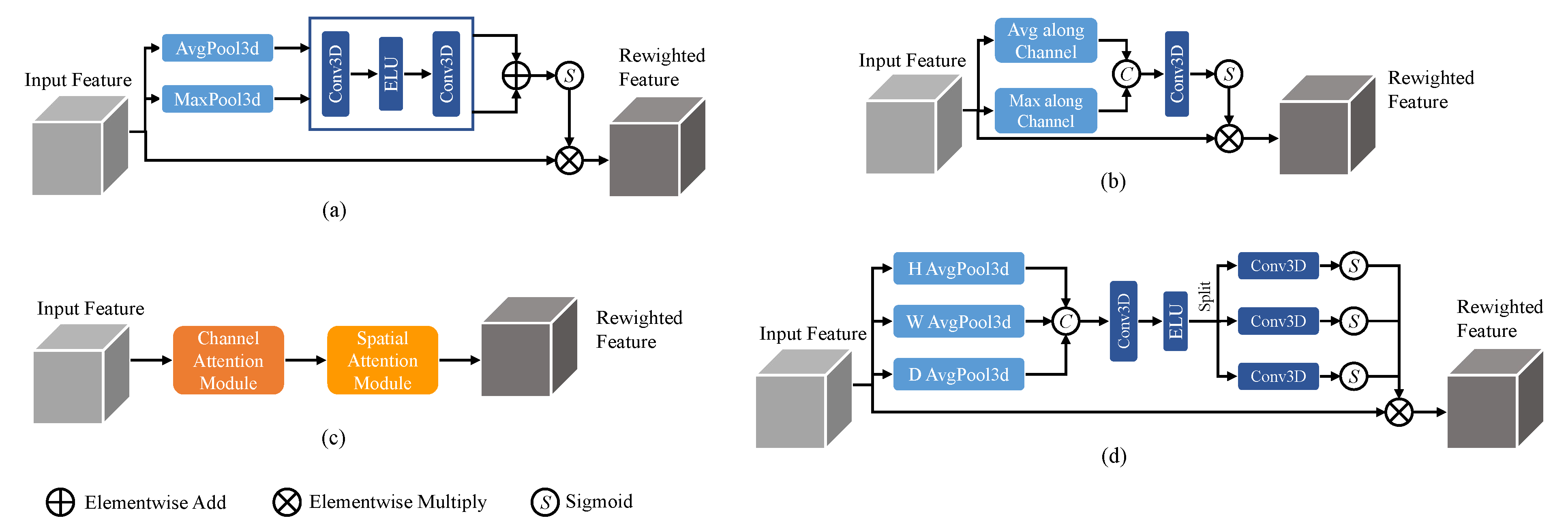
2. Single-Path and Multi-Path Convolutional Neural Networks for Brain Tumor Segmentation
3. The U-Net and Its Variants for Brain Tumor Segmentation
References
- Hu, J.; Shen, L.; Sun, G. Squeeze-and-excitation networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–23 June 2018; pp. 7132–7141.
- Zhu, X.; Cheng, D.; Zhang, Z.; Lin, S.; Dai, J. An empirical study of spatial attention mechanisms in deep networks. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Seoul, Republic of Korea, 27 October–2 November 2019; pp. 6688–6697.
- Woo, S.; Park, J.; Lee, J.Y.; Kweon, I.S. Cbam: Convolutional block attention module. In Proceedings of the European Conference on Computer Vision (ECCV), Munich, Germany, 8–14 September 2018; pp. 3–19.
- Hou, Q.; Zhou, D.; Feng, J. Coordinate attention for efficient mobile network design. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Nashville, TN, USA, 20–25 June 2021; pp. 13713–13722.
- Sarvamangala, D.; Kulkarni, R.V. Convolutional neural networks in medical image understanding: A survey. Evol. Intell. 2022, 15, 1–22.
- Dvořák, P.; Menze, B. Local structure prediction with convolutional neural networks for multimodal brain tumor segmentation. In Proceedings of the Medical Computer Vision: Algorithms for Big Data: International Workshop, MCV 2015, Held in Conjunction with MICCAI 2015, Munich, Germany, 9 October 2015; Revised Selected Papers. Springer: Cham, Switzerland, 2016; pp. 59–71.
- Sedlar, S. Brain tumor segmentation using a multi-path CNN based method. In Proceedings of the Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries: Third International Workshop, BrainLes 2017, Held in Conjunction with MICCAI 2017, Quebec City, QC, Canada, 14 September 2017; Revised Selected Papers. Springer: Cham, Switzerland, 2018; pp. 403–422.
- Wang, G.; Li, W.; Ourselin, S.; Vercauteren, T. Automatic brain tumor segmentation using cascaded anisotropic convolutional neural networks. In Proceedings of the Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries: Third International Workshop, BrainLes 2017, Held in Conjunction with MICCAI 2017, Quebec City, QC, Canada, 14 September 2017; Revised Selected Papers. Springer: Cham, Switzerland, 2018; pp. 178–190.
- Du, G.; Cao, X.; Liang, J.; Chen, X.; Zhan, Y. Medical image segmentation based on U-Net: A review. J. Imaging Sci. Technol. 2020, 64, 020508.
- Maurya, S.; Kumar Yadav, V.; Agarwal, S.; Singh, A. Brain Tumor Segmentation in mpMRI Scans (BraTS-2021) Using Models Based on U-Net Architecture. In Proceedings of the International MICCAI Brainlesion Workshop, Online, 27 September 2021; Springer: Cham, Switzerland, 2021; pp. 312–323.
- Zhou, Z.; Rahman Siddiquee, M.M.; Tajbakhsh, N.; Liang, J. Unet++: A nested U-Net architecture for medical image segmentation. In Proceedings of the Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support: 4th International Workshop, DLMIA 2018, and 8th International Workshop, ML-CDS 2018, Held in Conjunction with MICCAI 2018, Granada, Spain, 20 September 2018; Proceedings. Springer: Cham, Switzerland, 2018; pp. 3–11.
- Myronenko, A. 3D MRI brain tumor segmentation using autoencoder regularization. In Proceedings of the Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries: 4th International Workshop, BrainLes 2018, Held in Conjunction with MICCAI 2018, Granada, Spain, 16 September 2018; Revised Selected Papers, Part II. Springer: Cham, Switzerland, 2019; pp. 311–320.
- Futrega, M.; Milesi, A.; Marcinkiewicz, M.; Ribalta, P. Optimized U-Net for brain tumor segmentation. In Proceedings of the International MICCAI Brainlesion Workshop, Online, 27 September 2021; Springer: Cham, Switzerland, 2021; pp. 15–29.
- Zhang, B.; Wang, Y.; Ding, C.; Deng, Z.; Li, L.; Qin, Z.; Ding, Z.; Bian, L.; Yang, C. Multi-scale feature pyramid fusion network for medical image segmentation. Int. J. Comput. Assist. Radiol. Surg. 2023, 18, 353–365.
- Gong, Y.; Yu, X.; Ding, Y.; Peng, X.; Zhao, J.; Han, Z. Effective fusion factor in FPN for tiny object detection. In Proceedings of the IEEE/CVF Winter Conference on Applications of Computer Vision, Waikoloa, HI, USA, 3–8 January 2021; pp. 1160–1168.
- Chen, M.; Wu, Y.; Wu, J. Aggregating multi-scale prediction based on 3D U-Net in brain tumor segmentation. In Proceedings of the Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries: 5th International Workshop, BrainLes 2019, Held in Conjunction with MICCAI 2019, Shenzhen, China, 17 October 2019; Revised Selected Papers, Part I. Springer: Cham, Switzerland, 2020; pp. 142–152.
- Chi, M.; An, H.; Jin, X.; Wen, K.; Nie, Z. SCAR U-Net: A 3D Spatial-Channel Attention ResU-Net for Brain Tumor Segmentation. In Proceedings of the 3rd International Symposium on Artificial Intelligence for Medicine Sciences, Amsterdam, The Netherlands, 13–15 October 2022; pp. 497–501.
- Hua, R.; Huo, Q.; Gao, Y.; Sui, H.; Zhang, B.; Sun, Y.; Mo, Z.; Shi, F. Segmenting brain tumor using cascaded V-Nets in multimodal MR images. Front. Comput. Neurosci. 2020, 14, 9.
- Avesta, A.; Hossain, S.; Lin, M.; Aboian, M.; Krumholz, H.M.; Aneja, S. Comparing 3D, 2.5 D, and 2D Approaches to Brain Image Auto-Segmentation. Bioengineering 2023, 10, 181.
- Xu, C.; Howey, J.; Ohorodnyk, P.; Roth, M.; Zhang, H.; Li, S. Segmentation and quantification of infarction without contrast agents via spatiotemporal generative adversarial learning. Med. Image Anal. 2020, 59, 101568.
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, Ł.; Polosukhin, I. Attention is all you need. Adv. Neural Inf. Process. Syst. 2017, 30, 5998–6008.
- He, K.; Gan, C.; Li, Z.; Rekik, I.; Yin, Z.; Ji, W.; Gao, Y.; Wang, Q.; Zhang, J.; Shen, D. Transformers in medical image analysis. Intell. Med. 2023, 3, 59–78.
- Wang, T.; Nie, Z.; Wang, R.; Xu, Q.; Huang, H.; Xu, H.; Xie, F.; Liu, X.J. PneuNet: Deep learning for COVID-19 pneumonia diagnosis on chest X-ray image analysis using Vision Transformer. Med. Biol. Eng. Comput. 2023, 61, 1395–1408.
- Hatamizadeh, A.; Tang, Y.; Nath, V.; Yang, D.; Myronenko, A.; Landman, B.; Roth, H.R.; Xu, D. UNETR: Transformers for 3d medical image segmentation. In Proceedings of the IEEE/CVF Winter Conference on Applications of Computer Vision, Waikoloa, HI, USA, 4–8 January 2022; pp. 574–584.
- Hatamizadeh, A.; Nath, V.; Tang, Y.; Yang, D.; Roth, H.R.; Xu, D. Swin UNETR: Swin transformers for semantic segmentation of brain tumors in mri images. In Proceedings of the International MICCAI Brainlesion Workshop, Online, 27 September 2021; Springer: Cham, Switzerland, 2021; pp. 272–284.
- Zhou, H.Y.; Guo, J.; Zhang, Y.; Han, X.; Yu, L.; Wang, L.; Yu, Y. nnFormer: Volumetric medical image segmentation via a 3D transformer. IEEE Trans. Image Process. 2023, 32, 4036–4045.
- Jain, J.; Singh, A.; Orlov, N.; Huang, Z.; Li, J.; Walton, S.; Shi, H. Semask: Semantically masked transformers for semantic segmentation. In Proceedings of the IEEE/CVF International Conference on Computer Vision, New Orleans, LA, USA, 18–24 June 2022; pp. 752–761.





