
Video Upload Options
DEAD-box decapping enzyme 20 (DDX20) is a putative RNA-decapping enzyme that can be identified by the conserved motif Asp–Glu–Ala–Asp (DEAD). Cellular processes involve numerous RNA secondary structure alterations, including translation initiation, nuclear and mitochondrial splicing, and assembly of ribosomes and spliceosomes. DDX20 reportedly plays an important role in cellular transcription and post-transcriptional modifications. On the one hand, DDX20 can interact with various transcription factors and repress the transcriptional process. On the other hand, DDX20 forms the survival motor neuron complex and participates in the assembly of snRNP, ultimately affecting the RNA splicing process. Finally, DDX20 can potentially rely on its RNA-unwinding enzyme function to participate in microRNA (miRNA) maturation and act as a component of the RNA-induced silencing complex.
1. Introduction
2. Distribution, Structure, and Subcellular Localization of DDX20
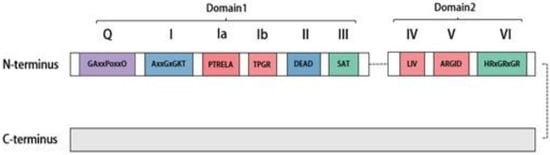
3. Splicing Features of DDX20
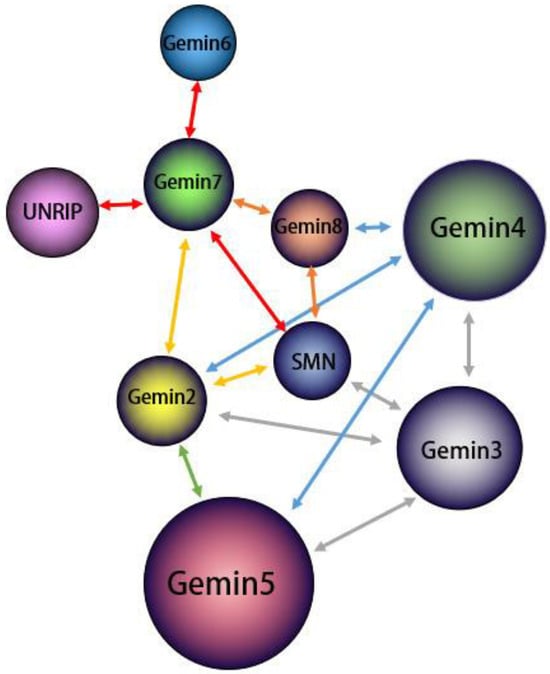
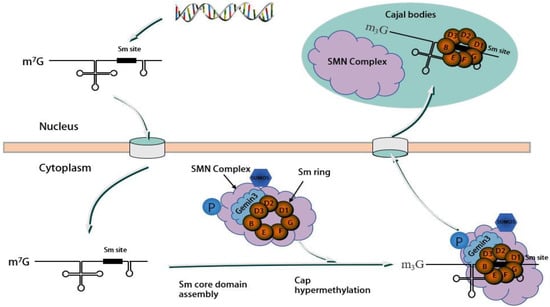
4. DDX20 Represses Transcription by Inhibiting Transcription Factors
5. Biogenesis of DDX20 and miRNA
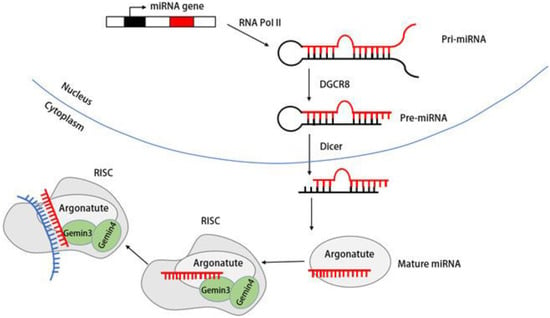
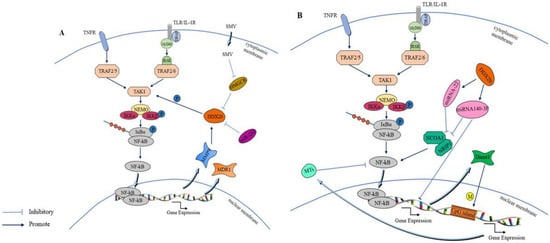
6. Functions of DDX20 in the Innate Immune Signaling Pathway
6.1. Effect of DDX20 on the NF-κB Signaling Pathway
6.2. DDX20 Affects p53 Signaling Pathway Conduction
7. DDX20 Plays Different Roles in Cancers through the NF-κB Signaling Pathway
8. Functions in Viral Infection
References
- Curmi, F.; Cauchi, R.J. The multiple lives of DEAD-box RNA helicase DP103/DDX20/Gemin3. Biochem. Soc. Trans. 2018, 46, 329–341.
- Wang, X. The Role and Mechanisms of DEAD-Box RNA Helicase DDX20 on Antiviral Innate Immunity. Master’s Thesis, Soochow University, Suzhou, China, 2019.
- Mouillet, J.-F.; Yan, X.; Ou, Q.; Jin, L.; Muglia, L.J.; Crawford, P.A.; Sadovsky, Y. DEAD-box protein-103 (DP103, Ddx20) is essential for early embryonic development and modulates ovarian morphology and function. Endocrinology 2008, 149, 2168–2175.
- Bizen, N.; Bepari, A.K.; Zhou, L.; Abe, M.; Sakimura, K.; Ono, K.; Takebayashi, H. Ddx20, an Olig2 binding factor, governs the survival of neural and oligodendrocyte progenitor cells via proper Mdm2 splicing and p53 suppression. Cell Death Differ. 2022, 29, 1028–1041.
- Simankova, A.; Bizen, N.; Saitoh, S.; Shibata, S.; Ohno, N.; Abe, M.; Sakimura, K.; Takebayashi, H. Ddx20, DEAD box helicase 20, is essential for the differentiation of oligodendrocyte and maintenance of myelin gene expression. Glia 2021, 69, 2559–2574.
- Hobani, Y.H.; Almars, A.I.; Alelwani, W.; Toraih, E.A.; Nemr, N.A.; Shaalan, A.A.M.; Fawzy, M.S.; Attallah, S.M. Genetic Variation in DEAD-Box Helicase 20 as a Putative Marker of Recurrence in Propensity-Matched Colon Cancer Patients. Genes 2022, 13, 1404.
- Minasaki, R.; Puoti, A.; Streit, A. The DEAD-box protein MEL-46 is required in the germ line of the nematode Caenorhabditis elegans. BMC Dev. Biol. 2009, 9, 35.
- Takata, A.; Otsuka, M.; Kojima, K.; Yoshikawa, T.; Kishikawa, T.; Ijichi, H.; Hirata, Y.; Tateishi, K.; Yoshida, H.; Omata, M.; et al. Abstract 3978: DDX20, a suppressor of hepatocarcinogenesis, controls NF-κB activity through regulating the function of miRNA-22 and miRNA-140-3p targeting transcriptional coactivators. Cancer Res. 2011, 71, 3978.
- Takata, A.; Otsuka, M.; Yoshikawa, T.; Kishikawa, T.; Hikiba, Y.; Obi, S.; Goto, T.; Kang, Y.J.; Maeda, S.; Yoshida, H.; et al. MicroRNA-140 acts as a liver tumor suppressor by controlling NF-κB activity by directly targeting DNA methyltransferase 1 (Dnmt1) expression. Hepatology 2013, 57, 162–170.
- Ou, Q.; Mouillet, J.F.; Yan, X.; Dorn, C.; Crawford, P.A.; Sadovsky, Y. The DEAD box protein DP103 is a regulator of steroidogenic factor-1. Mol. Endocrinol. 2001, 15, 69–79.
- Chen, W.; Zhou, P.; Li, X. High expression of DDX20 enhances the proliferation and metastatic potential of prostate cancer cells through the NF-κB pathway. Int. J. Mol. Med. 2016, 37, 1551–1557.
- Yang, Y.; Yang, M.; Pang, H.; Qiu, Y.; Sun, T.; Wang, T.; Shen, S.; Wang, W. A Macrophage Differentiation-Mediated Gene: DDX20 as a Molecular Biomarker Encompassing the Tumor Microenvironment, Disease Staging, and Prognoses in Hepatocellular Carcinoma. Oxidative Med. Cell. Longev. 2022, 2022, 9971776.
- Wang, Q.; Ye, Y.; Lin, R.; Weng, S.; Cai, F.; Zou, M.; Niu, H.; Ge, L.; Lin, Y. Analysis of the expression, function, prognosis and co-expression genes of DDX20 in gastric cancer. Comput. Struct. Biotechnol. J. 2020, 18, 2453–2462.
- Vychytilova-Faltejskova, P.; Svobodova Kovarikova, A.; Grolich, T.; Prochazka, V.; Slaba, K.; Machackova, T.; Halamkova, J.; Svoboda, M.; Kala, Z.; Kiss, I.; et al. MicroRNA Biogenesis Pathway Genes Are Deregulated in Colorectal Cancer. Int. J. Mol. Sci. 2019, 20, 4460.
- Kroiss, M.; Schultz, J.; Wiesner, J.; Chari, A.; Sickmann, A.; Fischer, U. Evolution of an RNP assembly system: A minimal SMN complex facilitates formation of UsnRNPs in Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 2008, 105, 10045–10050.
- Lai, X.; Wang, T.; Tan, T.Z.; Casey, P.J.; Hong, W.; Sudol, M.; Tergaonkar, V.; Lee, S.C.; Kumar, A.P. Abstract A28: DEAD-box RNA helicase DP103 enhances YAP sumoylation for YAP-TEAD dependence and statin sensitivity in triple-negative breast cancer. Mol. Cancer Res. 2020, 18, A28.
- Ali, M.A.M. DEAD-box RNA helicases: The driving forces behind RNA metabolism at the crossroad of viral replication and antiviral innate immunity. Virus Res. 2021, 296, 198352.
- Schütz, P.; Karlberg, T.; van den Berg, S.; Collins, R.; Lehtiö, L.; Högbom, M.; Holmberg-Schiavone, L.; Tempel, W.; Park, H.W.; Hammarström, M.; et al. Comparative structural analysis of human DEAD-box RNA helicases. PLoS ONE 2010, 5, e12791.
- Ali, M.A.M. The DEAD-box protein family of RNA helicases: Sentinels for a myriad of cellular functions with emerging roles in tumorigenesis. Int. J. Clin. Oncol. 2021, 26, 795–825.
- Yan, X.; Mouillet, J.F.; Ou, Q.; Sadovsky, Y. A novel domain within the DEAD-box protein DP103 is essential for transcriptional repression and helicase activity. Mol. Cell Biol. 2003, 23, 414–423.
- Coady, T.H.; Lorson, C.L. SMN in spinal muscular atrophy and snRNP biogenesis. Wiley Interdiscip. Rev. RNA 2011, 2, 546–564.
- Burlet, P.; Huber, C.; Bertrandy, S.; Ludosky, M.A.; Zwaenepoel, I.; Clermont, O.; Roume, J.; Delezoide, A.L.; Cartaud, J.; Munnich, A.; et al. The distribution of SMN protein complex in human fetal tissues and its alteration in spinal muscular atrophy. Hum. Mol. Genet. 1998, 7, 1927–1933.
- Charroux, B.; Pellizzoni, L.; Perkinson, R.A.; Shevchenko, A.; Mann, M.; Dreyfuss, G. Gemin3: A novel DEAD box protein that interacts with SMN, the spinal muscular atrophy gene product, and is a component of gems. J. Cell. Biol. 1999, 147, 1181–1194.
- Pellizzoni, L.; Yong, J.; Dreyfuss, G. Essential role for the SMN complex in the specificity of snRNP assembly. Science 2002, 298, 1775–1779.
- Lemm, I.; Girard, C.; Kuhn, A.N.; Watkins, N.J.; Schneider, M.; Bordonné, R.; Lührmann, R. Ongoing U snRNP biogenesis is required for the integrity of Cajal bodies. Mol. Biol. Cell 2006, 17, 3221–3231.
- Trinkle-Mulcahy, L.; Boulon, S.; Lam, Y.W.; Urcia, R.; Boisvert, F.M.; Vandermoere, F.; Morrice, N.A.; Swift, S.; Rothbauer, U.; Leonhardt, H.; et al. Identifying specific protein interaction partners using quantitative mass spectrometry and bead proteomes. J. Cell Biol. 2008, 183, 223–239.
- Stanek, D.; Neugebauer, K.M. The Cajal body: A meeting place for spliceosomal snRNPs in the nuclear maze. Chromosoma 2006, 115, 343–354.
- Husedzinovic, A.; Oppermann, F.; Draeger-Meurer, S.; Chari, A.; Fischer, U.; Daub, H.; Gruss, O.J. Phosphoregulation of the human SMN complex. Eur. J. Cell Biol. 2014, 93, 106–117.
- Riboldi, G.M.; Faravelli, I.; Kuwajima, T.; Delestrée, N.; Dermentzaki, G.; De Planell-Saguer, M.; Rinchetti, P.; Hao, L.T.; Beattie, C.C.; Corti, S.; et al. Sumoylation regulates the assembly and activity of the SMN complex. Nat. Commun. 2021, 12, 5040.
- Lafarga, V.; Tapia, O.; Sharma, S.; Bengoechea, R.; Stoecklin, G.; Lafarga, M.; Berciano, M.T. CBP-mediated SMN acetylation modulates Cajal body biogenesis and the cytoplasmic targeting of SMN. Cell. Mol. Life Sci. CMLS 2018, 75, 527–546.
- Meier, I.D.; Walker, M.P.; Matera, A.G. Gemin4 is an essential gene in mice, and its overexpression in human cells causes relocalization of the SMN complex to the nucleoplasm. Biol. Open 2018, 7, bio032409.
- Fallini, C.; Bassell, G.J.; Rossoll, W. Spinal muscular atrophy: The role of SMN in axonal mRNA regulation. Brain Res. 2012, 1462, 81–92.
- Patterson, W.L., 3rd; Georgel, P.T. Breaking the cycle: The role of omega-3 polyunsaturated fatty acids in inflammation-driven cancers. Biochem. Cell Biol. Biochim. Biol. Cell. 2014, 92, 321–328.
- Meister, G.; Bühler, D.; Pillai, R.; Lottspeich, F.; Fischer, U. A multiprotein complex mediates the ATP-dependent assembly of spliceosomal U snRNPs. Nat. Cell Biol. 2001, 3, 945–949.
- Battle, D.J.; Kasim, M.; Wang, J.; Dreyfuss, G. SMN-independent subunits of the SMN complex. Identification of a small nuclear ribonucleoprotein assembly intermediate. J. Biol. Chem. 2007, 282, 27953–27959.
- Will, C.L.; Lührmann, R. Spliceosomal UsnRNP biogenesis, structure and function. Curr. Opin. Cell Biol. 2001, 13, 290–301.
- Massenet, S.; Pellizzoni, L.; Paushkin, S.; Mattaj, I.W.; Dreyfuss, G. The SMN complex is associated with snRNPs throughout their cytoplasmic assembly pathway. Mol. Cell. Biol. 2002, 22, 6533–6541.
- Matera, A.G.; Wang, Z. A day in the life of the spliceosome. Nat. Rev. Mol. Cell Biol. 2014, 15, 108–121.
- Martinez-Salas, E.; Embarc-Buh, A.; Francisco-Velilla, R. Emerging Roles of Gemin5: From snRNPs Assembly to Translation Control. Int. J. Mol. Sci. 2020, 21, 3868.
- Li, D.K.; Tisdale, S.; Lotti, F.; Pellizzoni, L. SMN control of RNP assembly: From post-transcriptional gene regulation to motor neuron disease. Semin. Cell Dev. Biol. 2014, 32, 22–29.
- Mouaikel, J.; Narayanan, U.; Verheggen, C.; Matera, A.G.; Bertrand, E.; Tazi, J.; Bordonné, R. Interaction between the small-nuclear-RNA cap hypermethylase and the spinal muscular atrophy protein, survival of motor neuron. EMBO Rep. 2003, 4, 616–622.
- Otter, S.; Grimmler, M.; Neuenkirchen, N.; Chari, A.; Sickmann, A.; Fischer, U. A Comprehensive Interaction Map of the Human Survival of Motor Neuron (SMN) Complex. J. Biol. Chem. 2007, 282, 5825–5833.
- Charroux, B.; Pellizzoni, L.; Perkinson, R.A.; Yong, J.; Shevchenko, A.; Mann, M.; Dreyfuss, G. Gemin4. A novel component of the SMN complex that is found in both gems and nucleoli. J. Cell Biol. 2000, 148, 1177–1186.
- Cauchi, R.J. SMN and Gemins: ‘we are family’ … or are we?: Insights into the partnership between Gemins and the spinal muscular atrophy disease protein SMN. BioEssays News Rev. Mol. Cell. Dev. Biol. 2010, 32, 1077–1089.
- Feng, W.; Gubitz, A.K.; Wan, L.; Battle, D.J.; Dostie, J.; Golembe, T.J.; Dreyfuss, G. Gemins modulate the expression and activity of the SMN complex. Hum. Mol. Genet. 2005, 14, 1605–1611.
- Cauchi, R.J.; Davies, K.E.; Liu, J.L. A motor function for the DEAD-box RNA helicase, Gemin3, in Drosophila. PLoS Genet. 2008, 4, e1000265.
- Almstead, L.L.; Sarnow, P. Inhibition of U snRNP assembly by a virus-encoded proteinase. Genes. Dev. 2007, 21, 1086–1097.
- Shpargel, K.B.; Matera, A.G. Gemin proteins are required for efficient assembly of Sm-class ribonucleoproteins. Proc. Natl. Acad. Sci. USA 2005, 102, 17372–17377.
- Borg, R.M.; Fenech Salerno, B.; Vassallo, N.; Bordonne, R.; Cauchi, R.J. Disruption of snRNP biogenesis factors Tgs1 and pICln induces phenotypes that mirror aspects of SMN-Gemins complex perturbation in Drosophila, providing new insights into spinal muscular atrophy. Neurobiol. Dis. 2016, 94, 245–258.
- Cacciottolo, R.; Ciantar, J.; Lanfranco, M.; Borg, R.M.; Vassallo, N.; Bordonné, R.; Cauchi, R.J. SMN complex member Gemin3 self-interacts and has a functional relationship with ALS-linked proteins TDP-43, FUS and Sod1. Sci. Rep. 2019, 9, 18666.
- Fuller-Pace, F.V. DExD/H box RNA helicases: Multifunctional proteins with important roles in transcriptional regulation. Nucleic Acids Res. 2006, 34, 4206–4215.
- Parker, K.L.; Schimmer, B.P. Steroidogenic factor 1: A key determinant of endocrine development and function. Endocr. Rev. 1997, 18, 361–377.
- Zhao, L.; Kim, K.W.; Ikeda, Y.; Anderson, K.K.; Beck, L.; Chase, S.; Tobet, S.A.; Parker, K.L. Central nervous system-specific knockout of steroidogenic factor 1 results in increased anxiety-like behavior. Mol. Endocrinol. 2008, 22, 1403–1415.
- Lee, J.; Yang, D.J.; Lee, S.; Hammer, G.D.; Kim, K.W.; Elmquist, J.K. Nutritional conditions regulate transcriptional activity of SF-1 by controlling sumoylation and ubiquitination. Sci. Rep. 2016, 6, 19143.
- Lee, M.B.; Lebedeva, L.A.; Suzawa, M.; Wadekar, S.A.; Desclozeaux, M.; Ingraham, H.A. The DEAD-box protein DP103 (Ddx20 or Gemin-3) represses orphan nuclear receptor activity via SUMO modification. Mol. Cell. Biol. 2005, 25, 1879–1890.
- Treuter, E.; Venteclef, N. Transcriptional control of metabolic and inflammatory pathways by nuclear receptor SUMOylation. Biochim. Biophys. Acta 2011, 1812, 909–918.
- Hoivik, E.A.; Lewis, A.E.; Aumo, L.; Bakke, M. Molecular aspects of steroidogenic factor 1 (SF-1). Mol. Cell. Endocrinol. 2010, 315, 27–39.
- Yao, C.; Sun, Y.; Zhang, Z.; Jia, X.; Zou, P.; Wang, Y. Integration of RNAi and RNA-seq uncovers the regulation mechanism of DDX20 on vitellogenin expression in Scylla paramamosain. Comp. Biochem. Physiol. Part D Genom. Proteom. 2022, 44, 101028.
- Li, Q.; Xie, J.; He, L.; Wang, Y.; Yang, H.; Duan, Z.; Wang, Q. FOXL2 down-regulates vitellogenin expression at mature stage in Eriocheir sinensis. Biosci. Rep. 2015, 35, e00278.
- Pisarska, M.D.; Bae, J.; Klein, C.; Hsueh, A.J. Forkhead l2 is expressed in the ovary and represses the promoter activity of the steroidogenic acute regulatory gene. Endocrinology 2004, 145, 3424–3433.
- Lee, K.; Pisarska, M.D.; Ko, J.J.; Kang, Y.; Yoon, S.; Ryou, S.M.; Cha, K.Y.; Bae, J. Transcriptional factor FOXL2 interacts with DP103 and induces apoptosis. Biochem. Biophys. Res. Commun. 2005, 336, 876–881.
- Sawka-Verhelle, D.; Escoubet-Lozach, L.; Fong, A.L.; Hester, K.D.; Herzig, S.; Lebrun, P.; Glass, C.K. PE-1/METS, an antiproliferative Ets repressor factor, is induced by CREB-1/CREM-1 during macrophage differentiation. J. Biol. Chem. 2004, 279, 17772–17784.
- Klappacher, G.W.; Lunyak, V.V.; Sykes, D.B.; Sawka-Verhelle, D.; Sage, J.; Brard, G.; Ngo, S.D.; Gangadharan, D.; Jacks, T.; Kamps, M.P.; et al. An induced Ets repressor complex regulates growth arrest during terminal macrophage differentiation. Cell 2002, 109, 169–180.
- Hester, K.D.; Verhelle, D.; Escoubet-Lozach, L.; Luna, R.; Rose, D.W.; Glass, C.K. Differential repression of c-myc and cdc2 gene expression by ERF and PE-1/METS. Cell Cycle 2007, 6, 1594–1604.
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297.
- Gregory, R.I.; Yan, K.P.; Amuthan, G.; Chendrimada, T.; Doratotaj, B.; Cooch, N.; Shiekhattar, R. The Microprocessor complex mediates the genesis of microRNAs. Nature 2004, 432, 235–240.
- Lund, E.; Güttinger, S.; Calado, A.; Dahlberg, J.E.; Kutay, U. Nuclear export of microRNA precursors. Science 2004, 303, 95–98.
- Chendrimada, T.P.; Gregory, R.I.; Kumaraswamy, E.; Norman, J.; Cooch, N.; Nishikura, K.; Shiekhattar, R. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature 2005, 436, 740–744.
- Mourelatos, Z.; Dostie, J.; Paushkin, S.; Sharma, A.; Charroux, B.; Abel, L.; Rappsilber, J.; Mann, M.; Dreyfuss, G. miRNPs: A novel class of ribonucleoproteins containing numerous microRNAs. Genes. Dev. 2002, 16, 720–728.
- Dostie, J.; Mourelatos, Z.; Yang, M.; Sharma, A.; Dreyfuss, G. Numerous microRNPs in neuronal cells containing novel microRNAs. RNA 2003, 9, 180–186.
- Nelson, P.T.; Hatzigeorgiou, A.G.; Mourelatos, Z. miRNP:mRNA association in polyribosomes in a human neuronal cell line. RNA 2004, 10, 387–394.
- Hutvágner, G.; Zamore, P.D. A microRNA in a multiple-turnover RNAi enzyme complex. Science 2002, 297, 2056–2060.
- Meister, G.; Landthaler, M.; Patkaniowska, A.; Dorsett, Y.; Teng, G.; Tuschl, T. Human Argonaute2 mediates RNA cleavage targeted by miRNAs and siRNAs. Mol. Cell 2004, 15, 185–197.
- Pillai, R.S.; Bhattacharyya, S.N.; Filipowicz, W. Repression of protein synthesis by miRNAs: How many mechanisms? Trends Cell Biol. 2007, 17, 118–126.
- Behm-Ansmant, I.; Rehwinkel, J.; Doerks, T.; Stark, A.; Bork, P.; Izaurralde, E. mRNA degradation by miRNAs and GW182 requires both CCR4:NOT deadenylase and DCP1:DCP2 decapping complexes. Genes. Dev. 2006, 20, 1885–1898.
- Murashov, A.K.; Chintalgattu, V.; Islamov, R.R.; Lever, T.E.; Pak, E.S.; Sierpinski, P.L.; Katwa, L.C.; Van Scott, M.R. RNAi pathway is functional in peripheral nerve axons. FASEB J. 2007, 21, 656–670.
- Bollrath, J.; Greten, F.R. IKK/NF-kappaB and STAT3 pathways: Central signalling hubs in inflammation-mediated tumour promotion and metastasis. EMBO Rep. 2009, 10, 1314–1319.
- Sun, S.C. The non-canonical NF-κB pathway in immunity and inflammation. Nat. Rev. Immunol. 2017, 17, 545–558.
- Gilmore, T.D. Introduction to NF-kappaB: Players, pathways, perspectives. Oncogene 2006, 25, 6680–6684.
- Guttridge, D.C.; Albanese, C.; Reuther, J.Y.; Pestell, R.G.; Baldwin, A.S., Jr. NF-kappaB controls cell growth and differentiation through transcriptional regulation of cyclin D1. Mol. Cell. Biol. 1999, 19, 5785–5799.
- Patel, M.; Horgan, P.G.; McMillan, D.C.; Edwards, J. NF-κB pathways in the development and progression of colorectal cancer. Transl. Res. J. Lab. Clin. Med. 2018, 197, 43–56.
- Verzella, D.; Pescatore, A.; Capece, D.; Vecchiotti, D.; Ursini, M.V.; Franzoso, G.; Alesse, E.; Zazzeroni, F. Life, death, and autophagy in cancer: NF-κB turns up everywhere. Cell Death Dis. 2020, 11, 210.
- Motolani, A.; Martin, M.; Sun, M.; Lu, T. 6.19—NF-κB and Cancer Therapy Drugs. In Comprehensive Pharmacology; Kenakin, T., Ed.; Elsevier: Oxford, UK, 2022; pp. 351–363.
- Tao, Y.; Zhou, J.; Wang, Z.; Tao, H.; Bai, J.; Ge, G.; Li, W.; Zhang, W.; Hao, Y.; Yang, X.; et al. Human bone mesenchymal stem cells-derived exosomal miRNA-361-5p alleviates osteoarthritis by downregulating DDX20 and inactivating the NF-κB signaling pathway. Bioorganic Chem. 2021, 113, 104978.
- Cai, W.; Wang, C.; Goh, J.N.; Loo, S.Y.; Yap, C.T.; Kumar, A.P. DEAD-box RNA Helicases: The microRNA managers of breast cancer. RNA Dis. 2015, 2, e846.
- Takata, A.; Otsuka, M.; Kojima, K.; Yoshikawa, T.; Kishikawa, T.; Yoshida, H.; Koike, K. MicroRNA-22 and microRNA-140 suppress NF-κB activity by regulating the expression of NF-κB coactivators. Biochem. Biophys. Res. Commun. 2011, 411, 826–831.
- Takata, A.; Otsuka, M.; Kudo, Y.; Kojima, K.; Maeda, S.; Tateishi, K.; Ikenoue, T.; Ijichi, H.; Hirata, Y.; Yoshida, H.; et al. Abstract 2065: DDX20 deficiency enhances NF-κB by impairing NF-κB suppressive-microRNA function and leads to hepatocarcinogenesis. Cancer Res. 2010, 70, 2065.
- Huang, Y.; Wang, C.; Li, K.; Ye, Y.; Shen, A.; Guo, L.; Chen, P.; Meng, C.; Wang, Q.; Yang, X.; et al. Death-associated protein kinase 1 suppresses hepatocellular carcinoma cell migration and invasion by upregulation of DEAD-box helicase 20. Cancer Sci. 2020, 111, 2803–2813.
- Shi, D.; Jiang, P. A Different Facet of p53 Function: Regulation of Immunity and Inflammation During Tumor Development. Front. Cell Dev. Biol. 2021, 9, 762651.
- Bordon, Y. Cell death: Tumour suppressor p53 helps phagocytes clean up. Nat. Rev. Immunol. 2015, 15, 525.
- Aylon, Y.; Oren, M. Living with p53, dying of p53. Cell 2007, 130, 597–600.
- Cooks, T.; Harris, C.C.; Oren, M. Caught in the cross fire: p53 in inflammation. Carcinogenesis 2014, 35, 1680–1690.
- Cai, Q.; Guo, Y.; Xiao, B.; Banerjee, S.; Saha, A.; Lu, J.; Glisovic, T.; Robertson, E.S. Epstein-Barr virus nuclear antigen 3C stabilizes Gemin3 to block p53-mediated apoptosis. PLoS Pathog. 2011, 7, e1002418.
- Mirzaei, S.; Zarrabi, A.; Hashemi, F.; Zabolian, A.; Saleki, H.; Ranjbar, A.; Seyed Saleh, S.H.; Bagherian, M.; Sharifzadeh, S.O.; Hushmandi, K.; et al. Regulation of Nuclear Factor-KappaB (NF-κB) signaling pathway by non-coding RNAs in cancer: Inhibiting or promoting carcinogenesis? Cancer Lett. 2021, 509, 63–80.
- Ramadass, V.; Vaiyapuri, T.; Tergaonkar, V. Small Molecule NF-κB Pathway Inhibitors in Clinic. Int. J. Mol. Sci. 2020, 21, 5164.
- Cai, W.; Xiong Chen, Z.; Rane, G.; Satendra Singh, S.; Choo, Z.; Wang, C.; Yuan, Y.; Zea Tan, T.; Arfuso, F.; Yap, C.T.; et al. Wanted DEAD/H or Alive: Helicases Winding Up in Cancers. J. Natl. Cancer Inst. 2017, 109, djw278.
- Shin, E.M.; Osato, M.; Kumar, A.P.; Tergaonkar, V. RNA helicase DP103 and TAK1: A new connection in cancer. Mol. Cell Oncol. 2015, 2, e985911.
- Bentires-Alj, M.; Barbu, V.; Fillet, M.; Chariot, A.; Relic, B.; Jacobs, N.; Gielen, J.; Merville, M.P.; Bours, V. NF-kappaB transcription factor induces drug resistance through MDR1 expression in cancer cells. Oncogene 2003, 22, 90–97.
- Shin, E.M.; Hay, H.S.; Lee, M.H.; Goh, J.N.; Tan, T.Z.; Sen, Y.P.; Lim, S.W.; Yousef, E.M.; Ong, H.T.; Thike, A.A.; et al. DEAD-box helicase DP103 defines metastatic potential of human breast cancers. J. Clin. Investig. 2014, 124, 3807–3824.
- Casey, P.; Yap, C. RNA helicase DDX20 as a surrogate marker of statin activity in invasive breast cancer. Chin. J. Pharmacol. Toxicol. 2015, S1, 84.
- Nee, G.J. Characterization of Statins-Induced DDX20 Silencing in Invasive Breast Cancers. Master’s Thesis, Yong Loo Lin School of Medicine, National University of Singapore, Singapore, 2013.
- Chao, W. Interplay between Mevalonate and Hippo Pathways Regulates Ddx20 Transciption via Yao-Tead Complex in Triple Negative Breast Cancers. Ph.D. Thesis, National University of Singapore, Singapore, 2017.
- Pohl, S.Ö.-G. Mediation of Triple-Negative Breast Cancer Cell Fate via Cellular Redox and Wnt Signalling. Ph.D. Thesis, Curtin University, Bentley, WA, Australia, 2018.
- Zender, L.; Xue, W.; Zuber, J.; Semighini, C.P.; Krasnitz, A.; Ma, B.; Zender, P.; Kubicka, S.; Luk, J.M.; Schirmacher, P.; et al. An oncogenomics-based in vivo RNAi screen identifies tumor suppressors in liver cancer. Cell 2008, 135, 852–864.
- Chang, T.C.; Yu, D.; Lee, Y.S.; Wentzel, E.A.; Arking, D.E.; West, K.M.; Dang, C.V.; Thomas-Tikhonenko, A.; Mendell, J.T. Widespread microRNA repression by Myc contributes to tumorigenesis. Nat. Genet. 2008, 40, 43–50.
- Bartel, D.P. MicroRNAs: Target recognition and regulatory functions. Cell 2009, 136, 215–233.
- Thomson, J.M.; Newman, M.; Parker, J.S.; Morin-Kensicki, E.M.; Wright, T.; Hammond, S.M. Extensive post-transcriptional regulation of microRNAs and its implications for cancer. Genes. Dev. 2006, 20, 2202–2207.
- Takata, A.; Otsuka, M.; Yoshikawa, T.; Kishikawa, T.; Kudo, Y.; Goto, T.; Yoshida, H.; Koike, K. A miRNA machinery component DDX20 controls NF-κB via microRNA-140 function. Biochem. Biophys. Res. Commun. 2012, 420, 564–569.
- Ye, Y.; Spitz, M.R.; Yang, H.; Wu, X. Abstract 1174: Genetic variations in microRNA biogenesis pathway genes as susceptibility loci for lung cancer risk. Cancer Res. 2011, 71, 1174.
- Grundhoff, A.T.; Kremmer, E.; Türeci, O.; Glieden, A.; Gindorf, C.; Atz, J.; Mueller-Lantzsch, N.; Schubach, W.H.; Grässer, F.A. Characterization of DP103, a novel DEAD box protein that binds to the Epstein-Barr virus nuclear proteins EBNA2 and EBNA3C. J. Biol. Chem. 1999, 274, 19136–19144.
- Thorley-Lawson, D.A. Epstein-Barr virus: Exploiting the immune system. Nat. Rev. Immunol. 2001, 1, 75–82.
- Kawa, K. Epstein-Barr virus-associated diseases in humans. Int. J. Hematol. 2000, 71, 108–117.
- Hille, A.; Badu-Antwi, A.; Holzer, D.; Grässer, F.A. Lysine residues of Epstein-Barr virus-encoded nuclear antigen 2 do not confer secondary modifications via ubiquitin or SUMO-like proteins but modulate transcriptional activation. J. Gen. Virol. 2002, 83, 1037–1042.
- Strobl, L.J.; Höfelmayr, H.; Stein, C.; Marschall, G.; Brielmeier, M.; Laux, G.; Bornkamm, G.W.; Zimber-Strobl, U. Both Epstein-Barr Viral Nuclear Antigen 2 (EBNA2) and Activated Notch1 Transactivate Genes by Interacting with the Cellular Protein RBP-Jκ. Immunobiology 1997, 198, 299–306.
- Ling, P.D.; Rawlins, D.R.; Hayward, S.D. The Epstein-Barr virus immortalizing protein EBNA-2 is targeted to DNA by a cellular enhancer-binding protein. Proc. Natl. Acad. Sci. USA 1993, 90, 9237–9241.
- Harada, S.; Kieff, E. Epstein-Barr virus nuclear protein LP stimulates EBNA-2 acidic domain-mediated transcriptional activation. J. Virol. 1997, 71, 6611–6618.
- Voss, M.D.; Hille, A.; Barth, S.; Spurk, A.; Hennrich, F.; Holzer, D.; Mueller-Lantzsch, N.; Kremmer, E.; Grässer, F.A. Functional cooperation of Epstein-Barr virus nuclear antigen 2 and the survival motor neuron protein in transactivation of the viral LMP1 promoter. J. Virol. 2001, 75, 11781–11790.
- West, M.J. Structure and function of the Epstein-Barr virus transcription factor, EBNA 3C. Curr. Protein Pept. Sci. 2006, 7, 123–136.
- Subramanian, C.; Knight, J.S.; Robertson, E.S. The Epstein Barr nuclear antigen EBNA3C regulates transcription, cell transformation and cell migration. Front. Biosci. J. Virtual Libr. 2002, 7, d704–d716.
- Styles, C.T.; Paschos, K.; White, R.E.; Farrell, P.J. The Cooperative Functions of the EBNA3 Proteins Are Central to EBV Persistence and Latency. Pathogens 2018, 7, 31.
- Robert, F.; Pelletier, J. Perturbations of RNA helicases in cancer. Wiley Interdiscip. Rev. RNA 2013, 4, 333–349.
- Jäger, S.; Cimermancic, P.; Gulbahce, N.; Johnson, J.R.; McGovern, K.E.; Clarke, S.C.; Shales, M.; Mercenne, G.; Pache, L.; Li, K.; et al. Global landscape of HIV–human protein complexes. Nature 2012, 481, 365–370.
- Yang, Z.; Yang, J.; Wang, J.; Lu, X.; Jin, C.; Xie, T.; Wu, N. Identify Potential Regulators in HIV-1 Latency by Joint microRNA and mRNA Analysis. Cell. Physiol. Biochem. Int. J. Exp. Cell. Physiol. Biochem. Pharmacol. 2015, 36, 569–584.
- Greenwood, E.J.D.; Williamson, J.C.; Sienkiewicz, A.; Naamati, A.; Matheson, N.J.; Lehner, P.J. Promiscuous Targeting of Cellular Proteins by Vpr Drives Systems-Level Proteomic Remodeling in HIV-1 Infection. Cell Rep. 2019, 27, 1579–1596.e7.





