
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Xiaohong Yan | + 392 word(s) | 392 | 2020-10-06 18:14:48 | | | |
| 2 | Vivi Li | Meta information modification | 392 | 2020-10-21 10:02:00 | | | | |
| 3 | Vivi Li | + 1 word(s) | 393 | 2020-10-21 10:39:06 | | | | |
| 4 | Xiaohong Yan | + 638 word(s) | 1031 | 2020-10-26 02:57:42 | | | | |
| 5 | Vivi Li | Meta information modification | 1031 | 2020-10-26 09:49:14 | | | | |
| 6 | Vivi Li | Meta information modification | 1031 | 2020-11-09 09:17:13 | | |
Video Upload Options
Long introm-spliced hairpin RNA (ihpRNA) constructs which contained inverted repeats of the phytoene desaturase (PDS) separated by an intron, had been shown to very effective in triggering PDS silencing in Brassica napus. Using the PDS gene as a target control, it was shown that the RCA-mediated long ihpRNA construct was signicantly effective in triggering gene silence in B. napus.
1. RCA-mediated construction of PDS ihpRNA
To investigate whether RCA effectively generates iphRNA constructs, the B. napus phytoene desaturase (PDS) gene coding sequence of 450 bp in size (from the initial codon ATG) was amplified using the primer pair PDS-F1/PDS-R1 (Fig. 1A), added an ‘A’, and inserted into the pKan-BsaⅠ using the T-A approach. Following BsaⅠ digestion, PDS fragments with two different sticky ends respectively were obtained (Fig. 1A), which then were ligated to adaptor 1 and adaptor 2, forming closed circular DNA molecules. Agarose gel electrophoresis of the ligation product showed that almost all of the BsaⅠ-digested PDS DNA fragments were connected to both adaptors, generating adaptor1::PDS DNA::adaptor2 closed circular ligation products (Fig. 1B). RCA were performed using the ligation products, Phi29, and the loop2-specific primer pair loop2-F/loop2-R. Large DNA molecules containing inverted repeat units of double-stranded DNA were observed (Fig. 1C). The RCA products were confirmed by BamHⅠ and SacⅠ digestion, which generated a 1.1-kb of linear fragment that was approximately two-fold greater in size than the unamplified PDS control, and by a further digestion for the 1.1-kb fragment with BsaⅠ, which produced a 500-bp fragment, including the PDS control and partial sequences of loop1, and a 600-bp fragment, including 450-bp PDS control fragment and 128-bp adaptor 2 (Fig. 1D). These results indicated that a pool of single inverted repeat units for PDS were formed by RCA. Finally, BamHⅠ- and SacⅠ-digested fragments of RCA products were inserted into eukaryotic expression vector pBI121, generating a 121-PDS-ihpRNA construct (Fig. 1E).
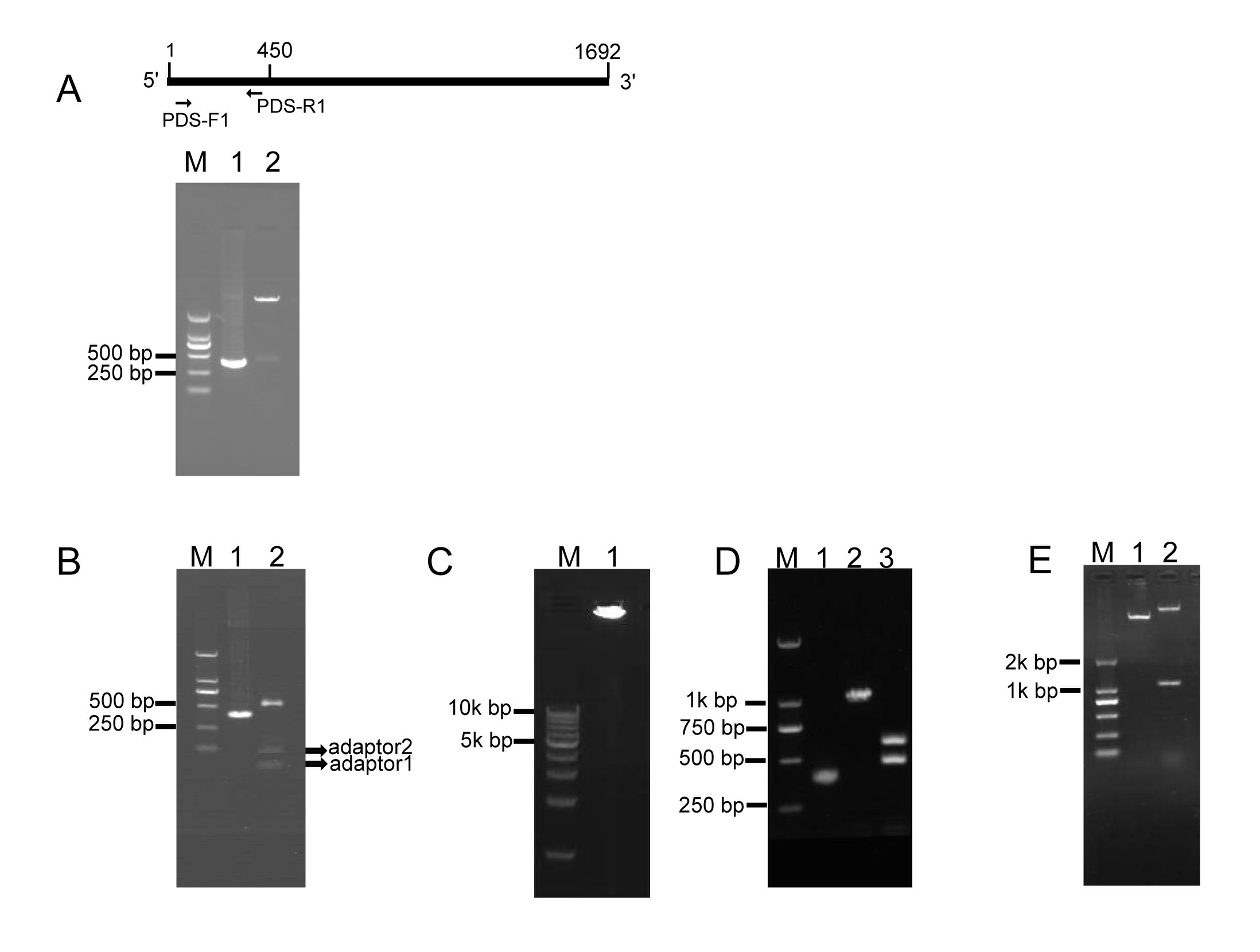
2.Testing of eukaryotic expression 121-PDS-ihpRNA construct
The 121-PDS-ihpRNA eukaryotic expressional construct was initially transferred into Agrobacterium strain GV3101 by electroporation, which was then transformed into B. napus cultivar zhongshuang 6 [1] via Agrobacterium-mediated gene transformation [2] to generate transgenic PDS-ihpRNA plants. Currently, this method of genetic transformation is considered to be of high efficiency (about 17%) [3]. The binary vector pBI121 harbors the neomycin phosphotransferase gene (NPTII) resistance selection marker (Figure 2A). The selection marker-resistant regenerated T0 transformants were initially subjected to kanamycin-based selection and then rooted well in selective medium. The selected resistant T0 transformants were then verified by PCR-based screening using the corresponding primer pairs NPTII-F and NPTII-R, which were specific for the NPTII gene in the binary construct (Figure 2A). A total of five transgenic plants (approximately 17% transformation efficiency) were obtained, all displaying photo-bleaching phenotypic characteristics (Figure 2B). To verify whether the PDS gene was silenced, qRT-PCR corresponding to the PDS target gene in five RNAi T0 lines was investigated, which confirmed that the mRNA expression level of the target PDS had significantly decreased (Fig. 2C). These results suggested that the PDS ihpRNA construct effectively silenced the PDS target gene. Due to the complex secondary hairpin structures formed by ihpRNA constructs, special primers (avoiding specificity to the stem of the hairpin structure) were designed to efficiently amplify the hpRNA target sequence in the binary pBI121 vector. The primer pairs loop2-F and 121-F or loop2-R and 121-R were used to identify the target gene sequences of the ihpRNA lines (Fig. 2D). For the five PDS RNAi lines, loop2-F and 121-F or loop2-R and 121-R primer pair successfully amplified bands of the desired size, and the amplification products were sequenced, demonstrating that the PDS ihpRNA expression cassette was successfully integrated into the B. napus plants (Fig. 2D). This result indicated that these two pairs of primers could be employed to rapidly identify the candidate target genes of the ihpRNA lines.
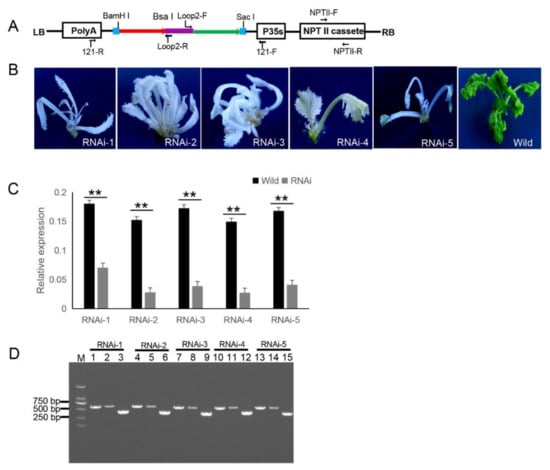
Figure 2. 121-PDS-ihpRNA-mediated silencing of the PDS gene. (A) Map of the binary vector 121-PDS-ihpRNA. NPTII-F and NPTII-R primers were used in the identification of positive transgenic plants, the primer pairs loop2-F/121-F and loop2-R/121-R were utilized to rapidly screen target genes. Blue line is the adaptor1 sequence. Purple line is the adaptor2 sequence. Red line with the arrow is the sense strand of dsDNA. Green line with the arrow is the anti-sense strand of dsDNA. (B) Interfering phenotypes of 5 PDS-ihpRNA transgenic plants. RNAi-1, RNAi-2, RNAi-3, RNAi-4, and RNAi-5 were PDS-ihpRNA transgenic plants. Wild was the transgenic negative plant. (C) qRT-PCR analysis of the PDS target gene mRNA level in the five hpRNA lines shown in (B) using the primer pair PDS-F2/PDS-R2. Data shown as mean ± s.d.,**, p < 0.05. (D) Integration into the Brassica napus genome for the PDS ihpRNA expression cassette. For the five PDS RNAi lines, loop2-F and 121-F or loop2-R and 121-R primer pair could amplify the bands with the desired size, demonstrating that the PDS ihpRNA expression cassette was successfully integrated into the B. napus plants. Lanes 3, 6, 9, 12, and 15 show the 450-bp PDS control; lanes 1, 4, 7, 10, and 13 exhibit the PCR products of loop2-F and 121-F; lanes 2, 5, 8, 11, and 14 depict the PCR products of loop2-R and 121-R.
References
- Zou, C.; Li, G.; Qu, Z.; Chen, D.; Cheng, Y.; Zheng, P. Breeding of Brassica napus cultivar Zhongshuang No. 6 with double-low, higher-yield and resistance to Sclerotinia sclerotiorum. Chin. J. Oil Crop Sci. 2003, 25, 115–116.
- Yan, X.; Zhang, L.; Chen, B.; Xiong, Z.; Chen, C.; Wang, L.; Yu, J.; Lu, C.; Wei, W. Functional identification and characterization of the Brassica napus transcription factor gene BnAP2, the ortholog of Arabidopsis thaliana APETALA2. PLoS ONE 2012, 7, e33890.
- Liu, F.; Xiong, X.; Wang, P.; Lei, L.; Zeng, X.; Zhu, L.; Li, Y.; Luo, J.; Fu, D.; Fu, P. Effects of non-procedural factors in Brassica napus genetic transformation. Chin. J. Oil Crop Sci. 2017, 2, 106–121.





