Long introm-spliced hairpin RNA (ihpRNA) constructs which contained inverted repeats of the phytoene desaturase (PDS) separated by an intron, had been shown to very effective in triggering PDS silencing in Brassica napus. Using the PDS gene as a target control, it was shown that the RCA-mediated long ihpRNA construct was signicantly effective in triggering gene silence in B. napus.
- Brassica napus
- RNAi
- ihpRNA
- PDS)
The 121-PDS-ihpRNA eukaryotic expressional construct was initially transferred into
strain GV3101 by electroporation, which was then transformed into
cultivar zhongshuang 6 [1] via
-mediated gene transformation [2] to generate transgenic PDS-ihpRNA plants. Currently, this method of genetic transformation is considered to be of high efficiency (about 17%) [3]. The binary vector pBI121 harbors the neomycin phosphotransferase gene (
) resistance selection marker (
1A). The selection marker-resistant regenerated T
transformants were initially subjected to kanamycin-based selection and then rooted well in selective medium. The selected resistant T
transformants were then verified by PCR-based screening using the corresponding primer pairs NPTII-F and NPTII-R, which were specific for the
gene in the binary construct (
1A). A total of five transgenic plants (approximately 17% transformation efficiency) were obtained, all displaying photo-bleaching phenotypic characteristics (
1B).
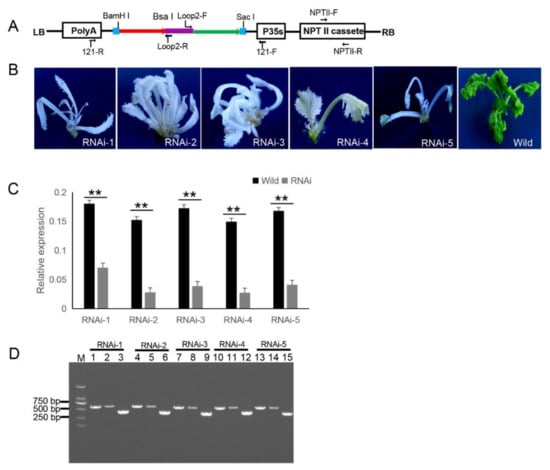
121-PDS-ihpRNA-mediated silencing of the
gene. (
) Map of the binary vector 121-PDS-ihpRNA. NPTII-F and NPTII-R primers were used in the identification of positive transgenic plants, the primer pairs loop2-F/121-F and loop2-R/121-R were utilized to rapidly screen target genes. Blue line is the adaptor1 sequence. Purple line is the adaptor2 sequence. Red line with the arrow is the sense strand of dsDNA. Green line with the arrow is the anti-sense strand of dsDNA. (
) Interfering phenotypes of 5 PDS-ihpRNA transgenic plants. RNAi-1, RNAi-2, RNAi-3, RNAi-4, and RNAi-5 were PDS-ihpRNA transgenic plants. Wild was the transgenic negative plant. (
) qRT-PCR analysis of the
target gene mRNA level in the five hpRNA lines shown in (
) using the primer pair PDS-F2/PDS-R2. Data shown as mean ± s.d.,**,
< 0.05. (
) Integration into the
genome for the
ihpRNA expression cassette. For the five PDS RNAi lines, loop2-F and 121-F or loop2-R and 121-R primer pair could amplify the bands with the desired size, demonstrating that the PDS ihpRNA expression cassette was successfully integrated into the
plants. Lanes 3, 6, 9, 12, and 15 show the 450-bp PDS control; lanes 1, 4, 7, 10, and 13 exhibit the PCR products of loop2-F and 121-F; lanes 2, 5, 8, 11, and 14 depict the PCR products of loop2-R and 121-R.
References
- Zou, C.; Li, G.; Qu, Z.; Chen, D.; Cheng, Y.; Zheng, P. Breeding of Brassica napus cultivar Zhongshuang No. 6 with double-low, higher-yield and resistance to Sclerotinia sclerotiorum. Chin. J. Oil Crop Sci. 2003, 25, 115–116.
- Yan, X.; Zhang, L.; Chen, B.; Xiong, Z.; Chen, C.; Wang, L.; Yu, J.; Lu, C.; Wei, W. Functional identification and characterization of the Brassica napus transcription factor gene BnAP2, the ortholog of Arabidopsis thaliana APETALA2. PLoS ONE 2012, 7, e33890.
- Liu, F.; Xiong, X.; Wang, P.; Lei, L.; Zeng, X.; Zhu, L.; Li, Y.; Luo, J.; Fu, D.; Fu, P. Effects of non-procedural factors in Brassica napus genetic transformation. Chin. J. Oil Crop Sci. 2017, 2, 106–121.
