
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Rosina De Cario | -- | 6306 | 2022-07-29 15:49:58 | | | |
| 2 | Vivi Li | Meta information modification | 6306 | 2022-08-02 03:41:07 | | | | |
| 3 | Vivi Li | Meta information modification | 6306 | 2022-08-02 03:42:57 | | |
Video Upload Options
The main challenge in diagnosing and managing thoracic aortic aneurysm and dissection (TAA/D) is represented by the early detection of a disease that is both deadly and “elusive”, as it generally grows asymptomatically prior to rupture, leading to death in the majority of cases. Gender differences exist in aortic dissection in terms of incidence and treatment options. Efforts have been made to identify biomarkers that may help in early diagnosis and in detecting those patients at a higher risk of developing life-threatening complications. As soon as the hereditability of the TAA/D was demonstrated, several genetic factors were found to be associated with both the syndromic and non-syndromic forms of the disease, and they currently play a role in patient diagnosis/prognosis and management-guidance purposes.
1. Introduction
2. Drivers of TAA Formation: A Constant Journey through Gene Discovery
| Biological Process/Cellular Compartment | Gene | Protein | OMIM | Syndromic TAA/D | Non-Syndromic FTAA/D | Associated Syndrome/Diseases |
|---|---|---|---|---|---|---|
| Extracellular matrix/remodeling | BGN | Biglycan | 300,989 | + | − | Meester-Loeys syndrome. ARD, TAAD, pulmonary artery aneurysm, IA, arterial tortuosity [19]. |
| COL3A1 | Collagen Type III α1 Chain | 130,050 | + | − | EDS, vascular type IV. TAAD, early aortic dissection, visceral arterial dissection, vessel fragility [20]. | |
| EFEMP2 | EGF Containing Fibulin Extracellular Matrix Protein 2 | 614,437 | + | − | Cutis laxa, AR type Ib. Ascending aortic aneurysms, other arterial aneurysms, arterial tortuosity, stenosis [21]. | |
| ELN | Elastin | 123,700 185,500 |
+ | − | Cutis laxa. AD ARD, ascending aortic aneurysm and dissection [22], TAA [23][24], BAV, IA possibly associated with SVAS. | |
| FBN1 | Fibrillin-1 | 154,700 | + | + | Marfan syndrome. ARD, TAA [25], TAAD [26], AAA, other arterial aneurysms, pulmonary artery dilatation, arterial tortuosity [27]. | |
| LOX | Protein-lysine 6-oxidase | 617,168 | − | + | AAT10. AAA, hepatic artery aneurysm, BAV, CAD, TAAD [28][29]. | |
| MFAP5 | Microfibril Associated Protein 5 | 616,166 | − | + | AAT9. ARD, TAA [30][31]. | |
| Smooth muscle cells | ACTA2 | Smooth muscle α-actin | 611,788 613,834 614,042 |
+ | + | AAT6, multisystemic smooth muscle dysfunction, MYMY5. Early aortic dissection, CAD, stroke (moyamoya disease), PDA, pulmonary artery dilation, BAV, TAAD, TAA [24][32]. |
| FLNA | Filamin A | 300,049 | + | − | Periventricular nodular heterotopia and otopalatodigital syndrome. Aortic dilatation/aneurysms, peripheral arterial dilatation, PDA, IA, BAV, TAA [32][33]. | |
| MYH11 | Smooth muscle myosin heavy chain | 132,900 | − | + | AAT4. PDA, CAD, peripheral vascular occlusive disease, carotid IA, TAAD, early aortic dissection [32][34][35]. | |
| MYLK | Myosin light chain kinase | 613,780 | − | + | AAT7. TAAD, early aortic dissections [36][37]. | |
| TGF-β signaling | LTBP1 | Latent TGF-β binding protein 1 | 150,390 | + | − | Aortic dilation with associated musculoskeletal findings. Dental anomalies, short stature. TAAD, AAA, visceral and peripheral arterial aneurysm [38]. |
| LTBP3 | Latent TGF-β binding protein 3 | 602,090 | ||||
| SMAD2 | SMAD2 | 619,657 619,656 |
+ | - | Unidentified CTD with arterial aneurysm/dissections. ARD, ascending aortic aneurysms, vertebral/carotid aneurysms and dissections [39], AAA. | |
| SMAD3 | SMAD3 | 613,795 | + | + | LDS type III. ARD, TAAD [40], early aortic dissection [39], AAA, arterial tortuosity, other arterial aneurysms/dissections [9], IA, BAV. | |
| SMAD4 | SMAD4 | 175,050 | + | - | JP/HHT syndrome. ARD, TAAD [39], AVMs, IA. | |
| SMAD6 | SMAD6 | 602,931 | - | + | AOVD2. BAV/TAA [24]. | |
| TGFB2 | TGF-β2 | 614,816 | + | + | LDS type IV. ARD, TAA [40], TAAD, arterial tortuosity [39], other arterial aneurysms, BAV. | |
| TGFB3 | TGF-β3 | 615,582 | + | - | LDS type V. ARD, TAAD, AAA/dissection, other arterial aneurysms, IA/dissection [39]. | |
| TGFBR1 | TGF-β receptor type 1 |
609,192 | + | + | LDS type I+AAT5. TAAD [40], early aortic dissection, AAA, arterial tortuosity, other arterial aneurysms/dissection [9], IA, PDA, BAV. | |
| TGFBR2 | TGF-β receptor type 2 |
610,168 | + | + | LDS type II+AAT3. TAAD [40], early aortic dissection, AAA, arterial tortuosity, other arterial aneurysms/dissection [9], IA, PDA, BAV. | |
| Others | AXIN1/PDIA2 locus | − | − | + | − | BAV. BAV/TAA [41]. |
| FBN2 | Fibrillin-2 | 121,050 | + | − | Contractual arachnodactyly. Rare ARD and aortic dissection [42], BAV, PDA. | |
| FOXE3 | Forkhead box 3 | 617,349 | − | + | AAT11. TAAD [30] (primarily type A dissection). | |
| MAT2A | Methionine adenosyl-transferase II α | n.a. | − | + | FTAA Thoracic aortic aneurysms [30][43]. BAV. | |
| NOTCH1 | NOTCH1 | 109,730 | − | + | AOVD1. BAV/TAAD [24]. | |
| PRKG1 | Type 1 cGMP-dependent protein kinase | 615,436 | − | + | AAT8. TAAD [28][43], early aortic dissection, AAA, coronary artery aneurysm/dissection, aortic tortuosity, small vessel, CVD. | |
| ROBO4 | Roundabout guidance receptor 4 | 607,528 | − | + | BAV. BAV/TAA [24]. | |
| SKI | Sloan Kettering proto-oncoprotein | 182,212 | + | − | Shprintzen–Goldberg syndrome. ARD, arterial tortuosity, pulmonary artery dilation, other (splenic) arterial aneurysms [36]. | |
| SLC2A10 | Glucose transporter 10 | 208,050 | + | − | Arterial tortuosity syndrome. ARD, ascending aortic aneurysms [36], other arterial aneurysms, arterial tortuosity [44], elongated arteries, aortic/pulmonary artery stenosis. |
2.1. Extracellular Matrix Components
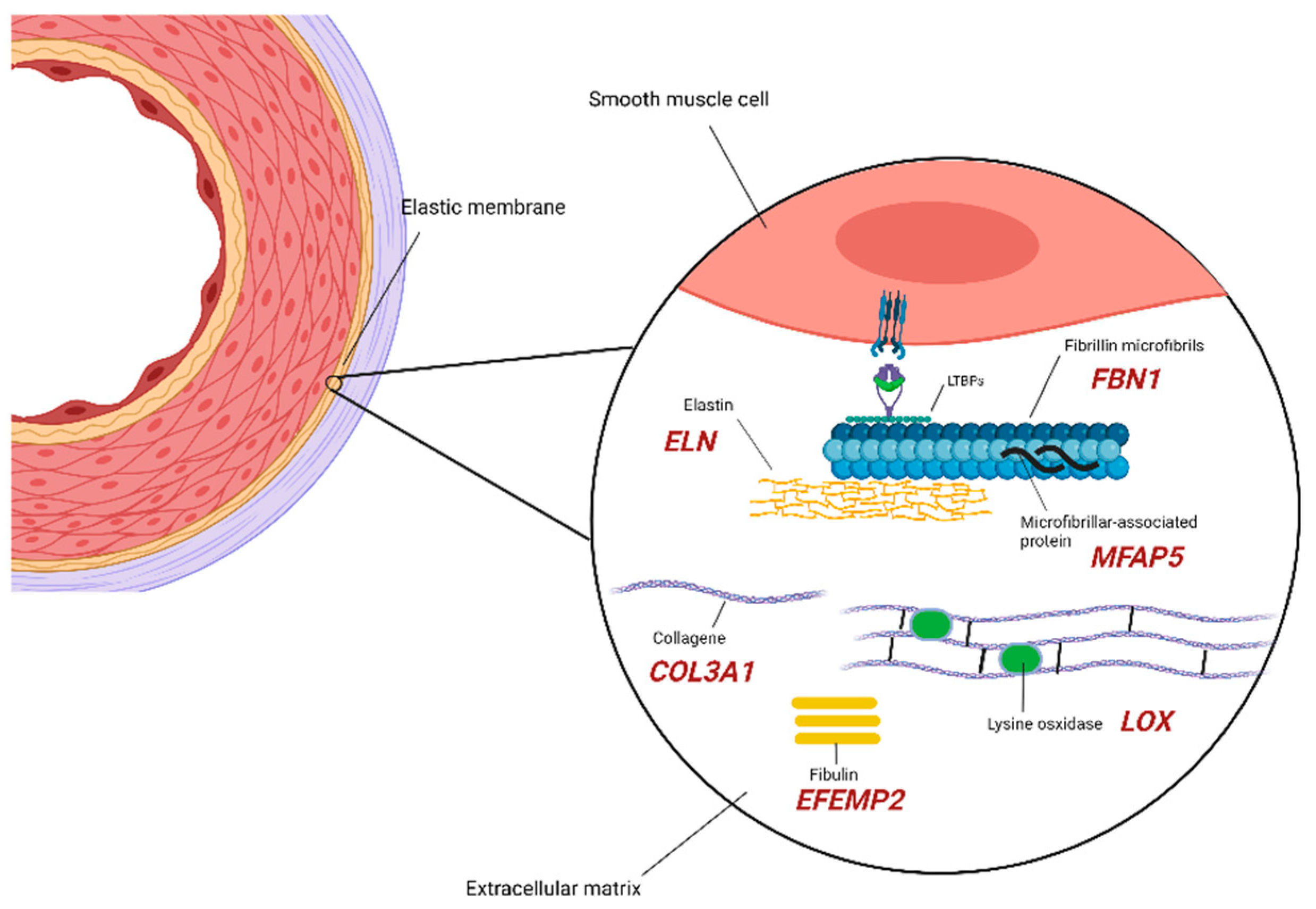
2.2. SMCs (Smooth Muscle Cells) Compartment
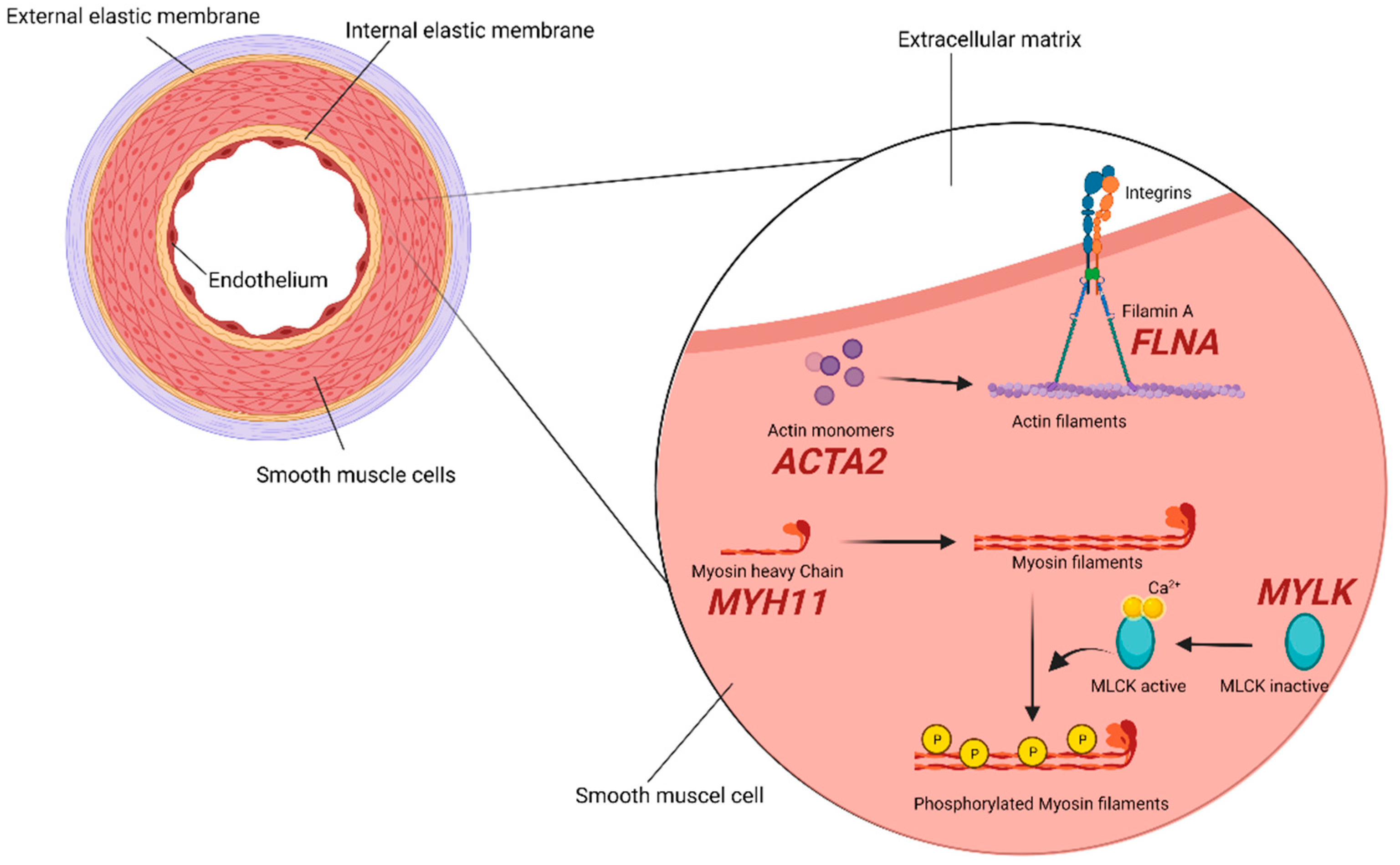
2.3. TGF-β Signaling
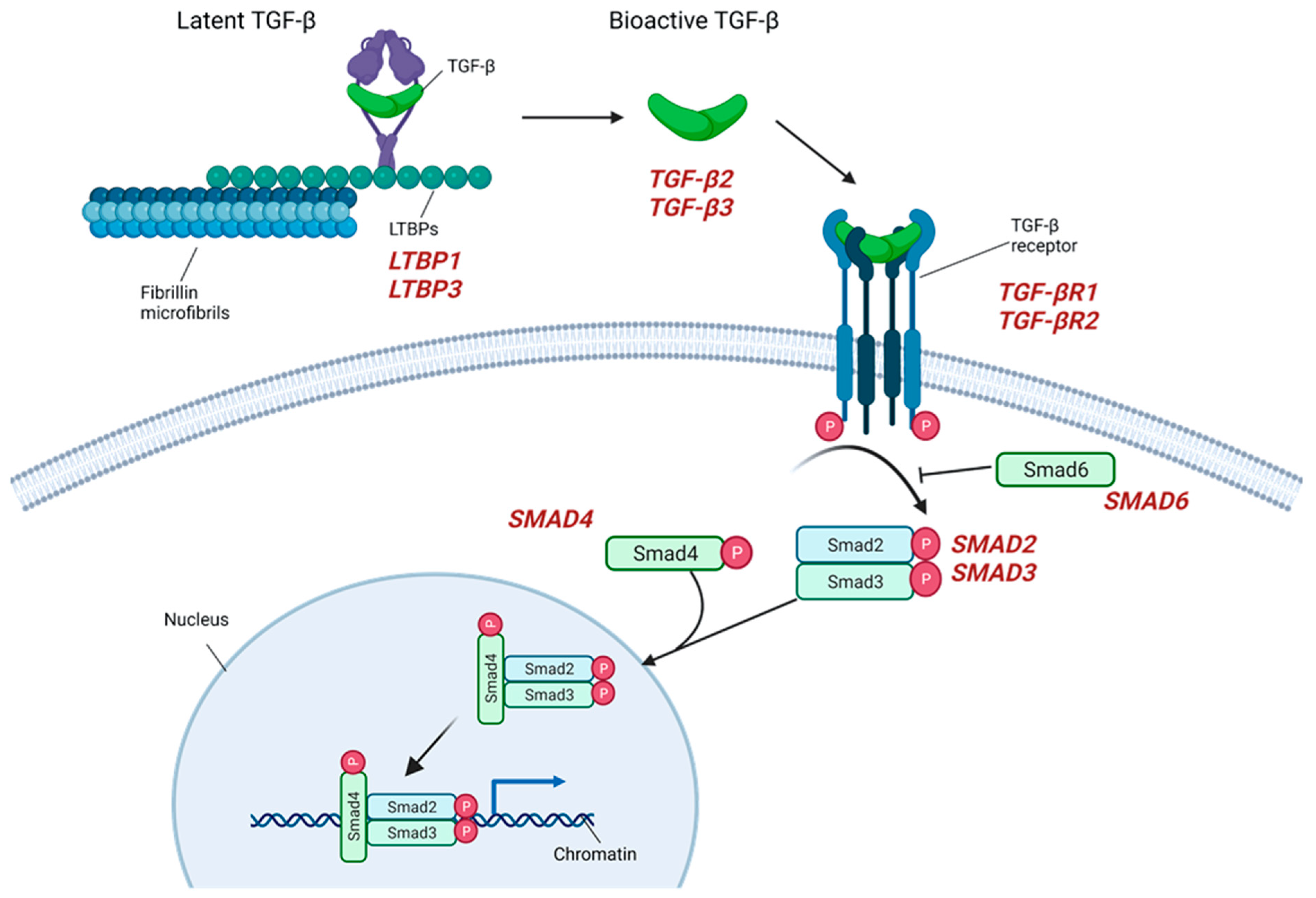
2.4. TAA in the Context of Bicuspid Aortic Valve and the Role of Proteases
3. Mechanisms of TAA Progression: The Dissection Menace
3.1. Pathophysiology and Risk Factors
3.2. Genetic Profiles of Dissection
4. Genetic Testing in Supporting TAA/D Diagnostics and In Risk Prediction: Where Do We Stand?
Although primarily considered as surgical disease, TAA’s optimal management greatly relies on an appropriate workup with the major purpose of identifying those features suggestive of a rapid progression of the aortic anomaly, thus predicting potentially life-threatening consequences of the disease. In this context, an accurate genetic evaluation/diagnosis serves different purposes: (a) guidance for overall medical management and surgical options; (b) timely evaluation of other organs that could be affected essentially in syndromic forms of TAA; (c) better definition of the prognosis; (d) identification of high-risk first-degree family members; (e) estimation of recurrence risk for future pregnancies in the prenatal diagnosis’ framework; and (f) support for imaging techniques in capturing nonsyndromic TAA patients who may be missed while developing dissection or rupture before reaching the guidelines-defined aortic diameter thresholds for aortic intervention [[118]]. As previously mentioned, syndromic and non-syndromic heritable thoracic aortic disease are, in most cases, inherited in an autosomal dominant manner except for rare X-linked and recessive conditions [119]. The accurate clinical evaluation of at-risk relatives is critical in this context, and ordinary and reproductive pre- and post-test genetic counseling allow for the early identification of an undiagnosed aortic disease in the first case and provide awareness about the risk of transmission to the offspring in the latter. Mutations are described to have variable penetrance depending on the TAA presentation, from almost 100% in MFS and 90% in LDS, to 50% in FTAAD and BAV in the presence of ascending aortic aneurysm. In fact, in the case of FTAAD, the causal mutation is found in much fewer cases (<10%) than in MFS or LDS, this discrepancy also being evident at the phenotypic level, presenting with a different severity of clinical manifestations along with age of presentation or diagnosis. When features of a connective-tissue disorder are present, patients should undergo genetic counselling and testing where appropriate [10]. The current ESC guidelines recommend genetic screening in first-degree relatives of TAA or aortic dissection and a diagnosis of familial aortic disease. In absence of a genetic diagnosis, at-risk relatives should undergo examination every 5 years. Screening should cover the entire arterial tree (including cerebral arteries) in families with nonsyndromic familial aortic disease [120]. According to the North American guidelines and related Class I recommendations, in case of identification of a mutation in one of the following genes, FBN1, TGFBR1, TGFBR2, COL3A1, ACTA2, and MYH11, which are associated with aortic aneurysm and/or dissection, first-degree relatives should undergo counseling and testing. Then, only the relatives with the genetic mutation should undergo aortic imaging. The guidelines provide some more recommendations (Class IIa and IIb): (a) ACTA2 sequencing should be considered in case of family history of thoracic aortic aneurysm and/or dissection; (b) TGFBR1, TGFBR2, and MYH11 sequencing may be considered in patients with a family history and clinical features associated with mutations in these genes; and c) if one or more first-degree relatives of a patient with known thoracic aortic aneurysm and/or dissection are found to have thoracic aortic dilatation, aneurysm, or dissection, then referral to a geneticist may be considered [121]. Following the exclusion of a syndromic condition, nonsyndromic TAA, in which mutations in genes known to be involved in syndromic forms of TAAD are rarely found, may present suggestive features of a genetic etiology, which might include young age at presentation (<50 years old), multiple aneurysms or dissections, and aortic root aneurysm [122][123]. In this scenario, genetic counseling should begin with the collection of the most detailed information of a three-generation family history, for the presence of aneurysm, dissection, sudden deaths, and syndromic features that would help in determining the inheritance pattern, identifying at-risk relatives, and recognizing syndromic signs [119]. In 2009, Ripperger and co-workers reported three cases of sudden, unexpected death due to thoracic aortic dissection, pointing out the great benefit that could be derived from alerting the at-risk relatives of the deceased about a potential heritable etiology of the disease [124]. The authors propose the development of a standard procedure which includes genetic counseling for at-risk relatives and storage of DNA or unfixed tissue for molecular investigations that would eventually allow differential diagnostic reappraisal from a genetic point of view. In any case, during genetic consultation, patients should become aware of the limitations, benefits, and personal and familial implications of genetic testing. Besides, awareness should be raised on the possibility of a negative genetic test that would not necessarily exclude a genetic etiology, thus indicating the imaging to be performed anyways in the first-degree family members in the search for aortic disease [121]. In fact, some types of genetic variants may be undetectable by standard assays and, similarly, the causative mutation may involve a gene that has not yet been associated to TAAD, due to absence of data supporting the actual pathological effect of that variant [121]. As a matter of fact, regarding the most appropriate genetic test selection, no specific indications are provided by the European guidelines. Genetic-testing panels vary significantly among laboratories and despite the enthusiasm for the so-called “exome-first” approach in diagnosing such a complex disease as TAA, its actual benefit and routine application in the diagnostic workup currently represent a matter of debate within the international scientific community.
5. Take home message
TAA’s bad reputation of “silent killer” is to be ascribed to its characteristic features, including its slow and gradual formation and the absence of visible signs, with patients remaining asymptomatic. This condition is elusive and yet potentially life-threatening, as it manifests itself only once the aneurysm is large enough to lead to an acute and devastating aortic event, with a significant percentage of patients dying before reaching the hospital. As a result, it is of the utmost relevance to identify biomarkers for the early identification of asymptomatic patients, a task which is both essential and challenging. In this regard, there’s an important distinction to be made between the TAD management within the emergency department, in which the room to maneuver is objectively limited, and those other situations in which the fatal event has not happened yet. In the first case, as Mehta and co-workers pointed out in a very recent review, the margin of intervention is essentially directed to improving the patient’s outcome by different means, including the multidisciplinary collaboration between specialists (emergency physicians, surgeons, radiologists) and identification of the optimal interventional treatment and post-operative care [125]. Traditional circulating biomarkers do not represent a satisfactory and reliable support in the initial patient screening as well, in which, on the contrary, the molecular/genetic evaluation can be diriment. Genetic testing, especially that which interrogates several genes at once in a parallel approach that is, at present, undoubtedly preferable to the cascade one, has long been included in the diagnostic flowchart for TAA diagnosis. First of all, it allows for the identification of a co-existing condition with TAA, such as MFS or LDS, thus directing the most appropriate management in terms of periodic check-ups, time of intervention, risk-recurrence calculation for pregnancies, and screening for first-degree relatives. In addition, the constant implementation of molecular methodologies allowing for the interrogation of the entire genome or transcriptome in an “omic” approach could be, undoubtedly, beneficial for patients’ stratification. In fact, the combination of the data derived from the WES/WGS/RNA-seq approaches can help define profiles that could be highly specific for subgroups of TAA patients, not to mention the potential use of those data in deepening knowledge about the disease’s onset and progression as well as for identifying new targets for therapy. Even with the considerable limitations characterizing the omic approaches (production of a large amount of bioinformatic data that need to be correctly interpreted, safely stored, and validated through functional studies; the possibility of VUSs and incidental findings), the future benefits they may represent for the improvement of the TAA diagnostic work-up have to be considered and perhaps should be addressed more closely and in greater detail in the international guidelines
References
- Bossone, E.; Eagle, K.A. Epidemiology and Management of Aortic Disease: Aortic Aneurysms and Acute Aortic Syndromes. Nat. Rev. Cardiol. 2021, 18, 331–348.
- Kuzmik, G.A.; Sang, A.X.; Elefteriades, J.A. Natural History of Thoracic Aortic Aneurysms. J. Vasc. Surg. 2012, 56, 565–571.
- Clouse, W.D.; Hallett, J.W.; Schaff, H.V.; Spittell, P.C.; Rowland, C.M.; Ilstrup, D.M.; Melton, L.J. Acute Aortic Dissection: Population-Based Incidence Compared with Degenerative Aortic Aneurysm Rupture. Mayo Clin. Proc. 2004, 79, 176–180.
- Nienaber, C.A.; Clough, R.E.; Sakalihasan, N.; Suzuki, T.; Gibbs, R.; Mussa, F.; Jenkins, M.P.; Thompson, M.M.; Evangelista, A.; Yeh, J.S.M.; et al. Aortic Dissection. Nat. Rev. Dis. Primer 2016, 2, 16053.
- Smedberg, C.; Steuer, J.; Leander, K.; Hultgren, R. Sex Differences and Temporal Trends in Aortic Dissection: A Population-Based Study of Incidence, Treatment Strategies, and Outcome in Swedish Patients during 15 Years. Eur. Heart J. 2020, 41, 2430–2438.
- Faggion Vinholo, T.; Zafar, M.A.; Ziganshin, B.A.; Elefteriades, J.A. Nonsyndromic Thoracic Aortic Aneurysms and Dissections—Is Screening Possible? Semin. Thorac. Cardiovasc. Surg. 2019, 31, 628–634.
- Salameh, M.J.; Black, J.H.; Ratchford, E.V. Thoracic Aortic Aneurysm. Vasc. Med. 2018, 23, 573–578.
- Monda, E.; Fusco, A.; Della Corte, A.; Caiazza, M.; Cirillo, A.; Gragnano, F.; Giugliano, M.P.; Citro, R.; Rubino, M.; Esposito, A.; et al. Impact of Regular Physical Activity on Aortic Diameter Progression in Paediatric Patients with Bicuspid Aortic Valve. Pediatr. Cardiol. 2021, 42, 1133–1140.
- Rohde, S.; Zafar, M.A.; Ziganshin, B.A.; Elefteriades, J.A. Thoracic Aortic Aneurysm Gene Dictionary. Asian Cardiovasc. Thorac. Ann. 2021, 29, 682–696.
- Chou, E.L.; Lindsay, M.E. The Genetics of Aortopathies: Hereditary Thoracic Aortic Aneurysms and Dissections. Am. J. Med. Genet. C Semin. Med. Genet. 2020, 184, 136–148.
- Erbel, R.; Aboyans, V.; Boileau, C.; Bossone, E.; Di Bartolomeo, R.; Eggebrecht, H.; Evangelista, A.; Falk, V.; Frank, H.; Gaemperli, O.; et al. 2014 ESC Guidelines on the diagnosis and treatment of aortic diseases. Kardiol. Pol. 2014, 72, 1169–1252.
- Harris, S.L.; Lindsay, M.E. Role of Clinical Genetic Testing in the Management of Aortopathies. Curr. Cardiol. Rep. 2021, 23, 10.
- Elefteriades, J.A.; Sang, A.; Kuzmik, G.; Hornick, M. Guilt by Association: Paradigm for Detecting a Silent Killer (Thoracic Aortic Aneurysm). Open Heart 2015, 2, e000169.
- Quintana, R.A.; Taylor, W.R. Cellular Mechanisms of Aortic Aneurysm Formation. Circ. Res. 2019, 124, 607–618.
- Albornoz, G.; Coady, M.A.; Roberts, M.; Davies, R.R.; Tranquilli, M.; Rizzo, J.A.; Elefteriades, J.A. Familial Thoracic Aortic Aneurysms and Dissections—Incidence, Modes of Inheritance, and Phenotypic Patterns. Ann. Thorac. Surg. 2006, 82, 1400–1405.
- Biddinger, A.; Rocklin, M.; Coselli, J.; Milewicz, D.M. Familial Thoracic Aortic Dilatations and Dissections: A Case Control Study. J. Vasc. Surg. 1997, 25, 506–511.
- Coady, M.A.; Davies, R.R.; Roberts, M.; Goldstein, L.J.; Rogalski, M.J.; Rizzo, J.A.; Hammond, G.L.; Kopf, G.S.; Elefteriades, J.A. Familial Patterns of Thoracic Aortic Aneurysms. Arch. Surg. 1999, 134, 361–367.
- Cannon Albright, L.A.; Camp, N.J.; Farnham, J.M.; MacDonald, J.; Abtin, K.; Rowe, K.G. A Genealogical Assessment of Heritable Predisposition to Aneurysms. J. Neurosurg. 2003, 99, 637–643.
- Meester, J.A.N.; Vandeweyer, G.; Pintelon, I.; Lammens, M.; Van Hoorick, L.; De Belder, S.; Waitzman, K.; Young, L.; Markham, L.W.; Vogt, J.; et al. Loss-of-Function Mutations in the X-Linked Biglycan Gene Cause a Severe Syndromic Form of Thoracic Aortic Aneurysms and Dissections. Genet. Med. 2017, 19, 386–395.
- Takeda, N.; Komuro, I. Genetic Basis of Hereditary Thoracic Aortic Aneurysms and Dissections. J. Cardiol. 2019, 74, 136–143.
- Baldwin, A.K.; Simpson, A.; Steer, R.; Cain, S.A.; Kielty, C.M. Elastic Fibres in Health and Disease. Expert Rev. Mol. Med. 2013, 15, e8.
- Szabo, Z. Aortic Aneurysmal Disease and Cutis Laxa Caused by Defects in the Elastin Gene. J. Med. Genet. 2005, 43, 255–258.
- Guemann, A.-S.; Andrieux, J.; Petit, F.; Halimi, E.; Bouquillon, S.; Manouvrier-Hanu, S.; Van De Kamp, J.; Boileau, C.; Hanna, N.; Jondeau, G.; et al. ELN Gene Triplication Responsible for Familial Supravalvular Aortic Aneurysm. Cardiol. Young 2015, 25, 712–717.
- Kent, K.C.; Crenshaw, M.L.; Goh, D.L.M.; Dietz, H.C. Genotype-Phenotype Correlation in Patients with Bicuspid Aortic Valve and Aneurysm. J. Thorac. Cardiovasc. Surg. 2013, 146, 158–165.e1.
- Weerakkody, R.; Ross, D.; Parry, D.A.; Ziganshin, B.; Vandrovcova, J.; Gampawar, P.; Abdullah, A.; Biggs, J.; Dumfarth, J.; Ibrahim, Y.; et al. Targeted Genetic Analysis in a Large Cohort of Familial and Sporadic Cases of Aneurysm or Dissection of the Thoracic Aorta. Genet. Med. Off. J. Am. Coll. Med. Genet. 2018, 20, 1414–1422.
- Ostberg, N.P.; Zafar, M.A.; Ziganshin, B.A.; Elefteriades, J.A. The Genetics of Thoracic Aortic Aneurysms and Dissection: A Clinical Perspective. Biomolecules 2020, 10, 182.
- Franken, R.; Groenink, M.; de Waard, V.; Feenstra, H.M.A.; Scholte, A.J.; van den Berg, M.P.; Pals, G.; Zwinderman, A.H.; Timmermans, J.; Mulder, B.J.M. Genotype Impacts Survival in Marfan Syndrome. Eur. Heart J. 2016, 37, 3285–3290.
- Guo, D.; Regalado, E.S.; Gong, L.; Duan, X.; Santos-Cortez, R.L.P.; Arnaud, P.; Ren, Z.; Cai, B.; Hostetler, E.M.; Moran, R.; et al. LOX Mutations Predispose to Thoracic Aortic Aneurysms and Dissections. Circ. Res. 2016, 118, 928–934.
- Lee, V.S.; Halabi, C.M.; Hoffman, E.P.; Carmichael, N.; Leshchiner, I.; Lian, C.G.; Bierhals, A.J.; Vuzman, D.; Brigham Genomic Medicine; Mecham, R.P.; et al. Loss of Function Mutation in LOX Causes Thoracic Aortic Aneurysm and Dissection in Humans. Proc. Natl. Acad. Sci. USA 2016, 113, 8759–8764.
- Kuang, S.-Q.; Medina-Martinez, O.; Guo, D.-C.; Gong, L.; Regalado, E.S.; Reynolds, C.L.; Boileau, C.; Jondeau, G.; Prakash, S.K.; Kwartler, C.S.; et al. FOXE3 Mutations Predispose to Thoracic Aortic Aneurysms and Dissections. J. Clin. Investig. 2016, 126, 948–961.
- Schubert, J.A.; Landis, B.J.; Shikany, A.R.; Hinton, R.B.; Ware, S.M. Clinically Relevant Variants Identified in Thoracic Aortic Aneurysm Patients by Research Exome Sequencing. Am. J. Med. Genet. A 2016, 170, 1288–1294.
- Milewicz, D.M.; Guo, D.-C.; Tran-Fadulu, V.; Lafont, A.L.; Papke, C.L.; Inamoto, S.; Kwartler, C.S.; Pannu, H. Genetic Basis of Thoracic Aortic Aneurysms and Dissections: Focus on Smooth Muscle Cell Contractile Dysfunction. Annu. Rev. Genom. Hum. Genet. 2008, 9, 283–302.
- Chen, M.H.; Choudhury, S.; Hirata, M.; Khalsa, S.; Chang, B.; Walsh, C.A. Thoracic Aortic Aneurysm in Patients with Loss of Function Filamin A Mutations: Clinical Characterization, Genetics, and Recommendations. Am. J. Med. Genet. A. 2018, 176, 337–350.
- Zhu, L.; Vranckx, R.; Khau Van Kien, P.; Lalande, A.; Boisset, N.; Mathieu, F.; Wegman, M.; Glancy, L.; Gasc, J.-M.; Brunotte, F.; et al. Mutations in Myosin Heavy Chain 11 Cause a Syndrome Associating Thoracic Aortic Aneurysm/Aortic Dissection and Patent Ductus Arteriosus. Nat. Genet. 2006, 38, 343–349.
- Luyckx, I.; Loeys, B.L. Curriculum topic: Disease of the aorta and trauma to the aorta and heart The Genetic Architecture of Non-Syndromic Thoracic Aortic Aneurysm. Heart Br. Card. Soc. 2015, 101, 1678–1684.
- Renard, M.; Francis, C.; Ghosh, R.; Scott, A.F.; Witmer, P.D.; Adès, L.C.; Andelfinger, G.U.; Arnaud, P.; Boileau, C.; Callewaert, B.L.; et al. Clinical Validity of Genes for Heritable Thoracic Aortic Aneurysm and Dissection. J. Am. Coll. Cardiol. 2018, 72, 605–615.
- Ponińska, J.K.; Bilińska, Z.T.; Truszkowska, G.; Michalak, E.; Podgórska, A.; Stępień-Wojno, M.; Chmielewski, P.; Lutyńska, A.; Płoski, R. Good Performance of the Criteria of American College of Medical Genetics and Genomics/Association for Molecular Pathology in Prediction of Pathogenicity of Genetic Variants Causing Thoracic Aortic Aneurysms and Dissections. J. Transl. Med. 2022, 20, 42.
- Quiñones-Pérez, B.; VanNoy, G.E.; Towne, M.C.; Shen, Y.; Singh, M.N.; Agrawal, P.B.; Smith, S.E. Three-Generation Family with Novel Contiguous Gene Deletion on Chromosome 2p22 Associated with Thoracic Aortic Aneurysm Syndrome. Am. J. Med. Genet. A 2018, 176, 560–569.
- MacCarrick, G.; Black, J.H.; Bowdin, S.; El-Hamamsy, I.; Frischmeyer-Guerrerio, P.A.; Guerrerio, A.L.; Sponseller, P.D.; Loeys, B.; Dietz, H.C. Loeys–Dietz Syndrome: A Primer for Diagnosis and Management. Genet. Med. 2014, 16, 576–587.
- Milewicz, D.M.; Regalado, E. Heritable Thoracic Aortic Disease Overview. In GeneReviews®; Adam, M.P., Ardinger, H.H., Pagon, R.A., Wallace, S.E., Bean, L.J., Gripp, K.W., Mirzaa, G.M., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993.
- Wooten, E.C.; Iyer, L.K.; Montefusco, M.C.; Hedgepeth, A.K.; Payne, D.D.; Kapur, N.K.; Housman, D.E.; Mendelsohn, M.E.; Huggins, G.S. Application of Gene Network Analysis Techniques Identifies AXIN1/PDIA2 and Endoglin Haplotypes Associated with Bicuspid Aortic Valve. PLoS ONE 2010, 5, e8830.
- Takeda, N.; Morita, H.; Fujita, D.; Inuzuka, R.; Taniguchi, Y.; Imai, Y.; Hirata, Y.; Komuro, I. Congenital Contractural Arachnodactyly Complicated with Aortic Dilatation and Dissection: Case Report and Review of Literature. Am. J. Med. Genet. A 2015, 167, 2382–2387.
- Guo, D.; Gong, L.; Regalado, E.S.; Santos-Cortez, R.L.; Zhao, R.; Cai, B.; Veeraraghavan, S.; Prakash, S.K.; Johnson, R.J.; Muilenburg, A.; et al. MAT2A Mutations Predispose Individuals to Thoracic Aortic Aneurysms. Am. J. Hum. Genet. 2015, 96, 170–177.
- Callewaert, B.; De Paepe, A.; Coucke, P. Arterial Tortuosity Syndrome. In GeneReviews®; Adam, M.P., Ardinger, H.H., Pagon, R.A., Wallace, S.E., Bean, L.J., Gripp, K.W., Mirzaa, G.M., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993.
- Lillie, M.A.; David, G.J.; Gosline, J.M. Mechanical Role of Elastin-Associated Microfibrils in Pig Aortic Elastic Tissue. Connect. Tissue Res. 1998, 37, 121–141.
- Creamer, T.J.; Bramel, E.E.; MacFarlane, E.G. Insights on the Pathogenesis of Aneurysm through the Study of Hereditary Aortopathies. Genes 2021, 12, 183.
- Dietz, H.C.; Cutting, C.R.; Pyeritz, R.E.; Maslen, C.L.; Sakai, L.Y.; Corson, G.M.; Puffenberger, E.G.; Hamosh, A.; Nanthakumar, E.J.; Curristin, S.M.; et al. Marfan Syndrome Caused by a Recurrent de Novo Missense Mutation in the Fibrillin Gene. Nature 1991, 352, 337–339.
- Csiszar, K. Lysyl Oxidases: A Novel Multifunctional Amine Oxidase Family. In Progress in Nucleic Acid Research and Molecular Biology; Elsevier: Amsterdam, The Netherlands, 2001; Volume 70l, pp. 1–32. ISBN 978-0-12-540070-1.
- Zentner, D.; James, P.; Bannon, P.; Jeremy, R. Familial Aortopathies—State of the Art Review. Heart Lung Circ. 2020, 29, 607–618.
- Papke, C.L.; Yanagisawa, H. Fibulin-4 and Fibulin-5 in Elastogenesis and beyond: Insights from Mouse and Human Studies. Matrix Biol. 2014, 37, 142–149.
- De Figueiredo Borges, L.; Jaldin, R.G.; Dias, R.R.; Stolf, N.A.G.; Michel, J.-B.; Gutierrez, P.S. Collagen Is Reduced and Disrupted in Human Aneurysms and Dissections of Ascending Aorta. Hum. Pathol. 2008, 39, 437–443.
- Combs, M.D.; Knutsen, R.H.; Broekelmann, T.J.; Toennies, H.M.; Brett, T.J.; Miller, C.A.; Kober, D.L.; Craft, C.S.; Atkinson, J.J.; Shipley, J.M.; et al. Microfibril-Associated Glycoprotein 2 (MAGP2) Loss of Function Has Pleiotropic Effects in Vivo. J. Biol. Chem. 2013, 288, 28869–28880.
- Kolb, M.; Margetts, P.J.; Sime, P.J.; Gauldie, J. Proteoglycans Decorin and Biglycan Differentially Modulate TGF-β-Mediated Fibrotic Responses in the Lung. Am. J. Physiol. Lung Cell. Mol. Physiol. 2001, 280, L1327–L1334.
- Guo, D.-C.; Pannu, H.; Tran-Fadulu, V.; Papke, C.L.; Yu, R.K.; Avidan, N.; Bourgeois, S.; Estrera, A.L.; Safi, H.J.; Sparks, E.; et al. Mutations in Smooth Muscle α-Actin (ACTA2) Lead to Thoracic Aortic Aneurysms and Dissections. Nat. Genet. 2007, 39, 1488–1493.
- Morisaki, H.; Akutsu, K.; Ogino, H.; Kondo, N.; Yamanaka, I.; Tsutsumi, Y.; Yoshimuta, T.; Okajima, T.; Matsuda, H.; Minatoya, K.; et al. Mutation of ACTA2 Gene as an Important Cause of Familial and Nonfamilial Nonsyndromatic Thoracic Aortic Aneurysm and/or Dissection (TAAD). Hum. Mutat. 2009, 30, 1406–1411.
- Disabella, E.; Grasso, M.; Gambarin, F.I.; Narula, N.; Dore, R.; Favalli, V.; Serio, A.; Antoniazzi, E.; Mosconi, M.; Pasotti, M.; et al. Risk of Dissection in Thoracic Aneurysms Associated with Mutations of Smooth Muscle Alpha-Actin 2 (ACTA2). Heart Br. Card. Soc. 2011, 97, 321–326.
- Lu, H.; Fagnant, P.M.; Bookwalter, C.S.; Joel, P.; Trybus, K.M. Vascular Disease-Causing Mutation R258C in ACTA2 Disrupts Actin Dynamics and Interaction with Myosin. Proc. Natl. Acad. Sci. USA 2015, 112, E4168–E4177.
- Kim, H.; McCulloch, C.A. Filamin A Mediates Interactions between Cytoskeletal Proteins That Control Cell Adhesion. FEBS Lett. 2011, 585, 18–22.
- Siddiqui, S.T.; Fisher, S.D. Heritable FLNA Gene Mutation in a Patient with Thoracic Aortic Aneurysm. JACC Case Rep. 2022, 4, 87–90.
- Pomianowski, P.; Elefteriades, J.A. The Genetics and Genomics of Thoracic Aortic Disease. Ann. Cardiothorac. Surg. 2013, 2, 271–279.
- Daugherty, A.; Chen, Z.; Sawada, H.; Rateri, D.L.; Sheppard, M.B. Transforming Growth Factor-β in Thoracic Aortic Aneurysms: Good, Bad, or Irrelevant? J. Am. Heart Assoc. 2017, 6, e005221.
- Michel, J.-B.; Jondeau, G.; Milewicz, D.M. From Genetics to Response to Injury: Vascular Smooth Muscle Cells in Aneurysms and Dissections of the Ascending Aorta. Cardiovasc. Res. 2018, 114, 578–589.
- MacFarlane, E.G.; Parker, S.J.; Shin, J.Y.; Ziegler, S.G.; Creamer, T.J.; Bagirzadeh, R.; Bedja, D.; Chen, Y.; Calderon, J.F.; Weissler, K.; et al. Lineage-Specific Events Underlie Aortic Root Aneurysm Pathogenesis in Loeys-Dietz Syndrome. J. Clin. Investig. 2019, 129, 659–675.
- Lindsay, M.E.; Dietz, H.C. Lessons on the Pathogenesis of Aneurysm from Heritable Conditions. Nature 2011, 473, 308–316.
- Guo, D.-C.; Regalado, E.S.; Pinard, A.; Chen, J.; Lee, K.; Rigelsky, C.; Zilberberg, L.; Hostetler, E.M.; Aldred, M.; Wallace, S.E.; et al. LTBP3 Pathogenic Variants Predispose Individuals to Thoracic Aortic Aneurysms and Dissections. Am. J. Hum. Genet. 2018, 102, 706–712.
- Schepers, D.; Tortora, G.; Morisaki, H.; MacCarrick, G.; Lindsay, M.; Liang, D.; Mehta, S.G.; Hague, J.; Verhagen, J.; van de Laar, I.; et al. A Mutation Update on the LDS-Associated Genes TGFB2/3 and SMAD2/3. Hum. Mutat. 2018, 39, 621–634.
- Cecconi, M.; Manfrin, M.; Moraca, A.; Zanoli, R.; Colonna, P.L.; Bettuzzi, M.G.; Moretti, S.; Gabrielli, D.; Perna, G.P. Aortic Dimensions in Patients with Bicuspid Aortic Valve without Significant Valve Dysfunction. Am. J. Cardiol. 2005, 95, 292–294.
- Pepe, G.; Nistri, S.; Giusti, B.; Sticchi, E.; Attanasio, M.; Porciani, C.; Abbate, R.; Bonow, R.O.; Yacoub, M.; Gensini, G.F. Identification of Fibrillin 1 Gene Mutations in Patients with Bicuspid Aortic Valve (BAV) without Marfan Syndrome. BMC Med. Genet. 2014, 15, 23.
- Boyum, J.; Fellinger, E.K.; Schmoker, J.D.; Trombley, L.; McPartland, K.; Ittleman, F.P.; Howard, A.B. Matrix Metalloproteinase Activity in Thoracic Aortic Aneurysms Associated with Bicuspid and Tricuspid Aortic Valves. J. Thorac. Cardiovasc. Surg. 2004, 127, 686–691.
- Rabkin, S.W. Differential Expression of MMP-2, MMP-9 and TIMP Proteins in Thoracic Aortic Aneurysm - Comparison with and without Bicuspid Aortic Valve: A Meta-Analysis. Vasa 2014, 43, 433–442.
- Martin-Blazquez, A.; Heredero, A.; Aldamiz-Echevarria, G.; Martin-Lorenzo, M.; Alvarez-Llamas, G. Non-Syndromic Thoracic Aortic Aneurysm: Cellular and Molecular Insights. J. Pathol. 2021, 254, 229–238.
- Geng, L.; Wang, W.; Chen, Y.; Cao, J.; Lu, L.; Chen, Q.; He, R.; Shen, W. Elevation of ADAM10, ADAM17, MMP-2 and MMP-9 Expression with Media Degeneration Features CaCl2-Induced Thoracic Aortic Aneurysm in a Rat Model. Exp. Mol. Pathol. 2010, 89, 72–81.
- Ren, P.; Zhang, L.; Xu, G.; Palmero, L.C.; Albini, P.T.; Coselli, J.S.; Shen, Y.H.; LeMaire, S.A. ADAMTS-1 and ADAMTS-4 Levels Are Elevated in Thoracic Aortic Aneurysms and Dissections. Ann. Thorac. Surg. 2013, 95, 570–577.
- Wilton, E.; Bland, M.; Thompson, M.; Jahangiri, M. Matrix Metalloproteinase Expression in the Ascending Aorta and Aortic Valve. Interact. Cardiovasc. Thorac. Surg. 2008, 7, 37–40.
- Evangelista, A.; Isselbacher, E.M.; Bossone, E.; Gleason, T.G.; Eusanio, M.D.; Sechtem, U.; Ehrlich, M.P.; Trimarchi, S.; Braverman, A.C.; Myrmel, T.; et al. Insights from the International Registry of Acute Aortic Dissection: A 20-Year Experience of Collaborative Clinical Research. Circulation 2018, 137, 1846–1860.
- Silaschi, M.; Byrne, J.; Wendler, O. Aortic Dissection: Medical, Interventional and Surgical Management. Heart 2017, 103, 78–87.
- Hagan, P.G.; Nienaber, C.A.; Isselbacher, E.M.; Bruckman, D.; Karavite, D.J.; Russman, P.L.; Evangelista, A.; Fattori, R.; Suzuki, T.; Oh, J.K.; et al. The International Registry of Acute Aortic Dissection (IRAD): New Insights into an Old Disease. JAMA 2000, 283, 897.
- Vilacosta, I.; San Román, J.A.; di Bartolomeo, R.; Eagle, K.; Estrera, A.L.; Ferrera, C.; Kaji, S.; Nienaber, C.A.; Riambau, V.; Schäfers, H.-J.; et al. Acute Aortic Syndrome Revisited: JACC State-of-the-Art Review. J. Am. Coll. Cardiol. 2021, 78, 2106–2125.
- White, A.; Broder, J.; Mando-Vandrick, J.; Wendell, J.; Crowe, J. Acute Aortic Emergencies—Part 2 Aortic Dissections. Adv. Emerg. Nurs. J. 2013, 35, 28–52.
- Von Kodolitsch, Y.; Csösz, S.K.; Koschyk, D.H.; Schalwat, I.; Loose, R.; Karck, M.; Dieckmann, C.; Fattori, R.; Haverich, A.; Berger, J.; et al. Intramural Hematoma of the Aorta: Predictors of Progression to Dissection and Rupture. Circulation 2003, 107, 1158–1163.
- Manabe, T.; Imoto, K.; Uchida, K.; Doi, C.; Takanashi, Y. Decreased Tissue Inhibitor of Metalloproteinase-2/Matrix Metalloproteinase Ratio in the Acute Phase of Aortic Dissection. Surg. Today 2004, 34, 220–225.
- Koullias, G.J.; Ravichandran, P.; Korkolis, D.P.; Rimm, D.L.; Elefteriades, J.A. Increased Tissue Microarray Matrix Metalloproteinase Expression Favors Proteolysis in Thoracic Aortic Aneurysms and Dissections. Ann. Thorac. Surg. 2004, 78, 2106–2110.
- Del Porto, F.; di Gioia, C.; Tritapepe, L.; Ferri, L.; Leopizzi, M.; Nofroni, I.; De Santis, V.; Della Rocca, C.; Mitterhofer, A.P.; Bruno, G.; et al. The Multitasking Role of Macrophages in Stanford Type A Acute Aortic Dissection. Cardiology 2014, 127, 123–129.
- Landenhed, M.; Engström, G.; Gottsäter, A.; Caulfield, M.P.; Hedblad, B.; Newton-Cheh, C.; Melander, O.; Smith, J.G. Risk Profiles for Aortic Dissection and Ruptured or Surgically Treated Aneurysms: A Prospective Cohort Study. J. Am. Heart Assoc. 2015, 4, e001513.
- Hahn, A.W.A.; Jonas, U.; Bühler, F.R.; Resink, T.J. Activation of Human Peripheral Monocytes by Angiotensin II. FEBS Lett. 1994, 347, 178–180.
- Stumpf, C.; Jukic, J.; Yilmaz, A.; Raaz, D.; Schmieder, R.E.; Daniel, W.G.; Garlichs, C.D. Elevated VEGF-Plasma Levels in Young Patients with Mild Essential Hypertension. Eur. J. Clin. Investig. 2009, 39, 31–36.
- Derosa, G.; D’Angelo, A.; Ciccarelli, L.; Piccinni, M.N.; Pricolo, F.; Salvadeo, S.; Montagna, L.; Gravina, A.; Ferrari, I.; Galli, S.; et al. Matrix Metalloproteinase-2, -9, and Tissue Inhibitor of Metalloproteinase-1 in Patients with Hypertension. Endothel. J. Endothel. Cell Res. 2006, 13, 227–231.
- Elefteriades, J.A. Natural History of Thoracic Aortic Aneurysms: Indications for Surgery, and Surgical versus Nonsurgical Risks. Ann. Thorac. Surg. 2002, 74, S1877–S1880, discussion S1892–S1898.
- Pape, L.A.; Tsai, T.T.; Isselbacher, E.M.; Oh, J.K.; O’gara, P.T.; Evangelista, A.; Fattori, R.; Meinhardt, G.; Trimarchi, S.; Bossone, E.; et al. Aortic Diameter > or =5.5 Cm Is Not a Good Predictor of Type A Aortic Dissection: Observations from the International Registry of Acute Aortic Dissection (IRAD). Circulation 2007, 116, 1120–1127.
- Davies, R.R.; Goldstein, L.J.; Coady, M.A.; Tittle, S.L.; Rizzo, J.A.; Kopf, G.S.; Elefteriades, J.A. Yearly Rupture or Dissection Rates for Thoracic Aortic Aneurysms: Simple Prediction Based on Size. Ann. Thorac. Surg. 2002, 73, 17–27, discussion 27–28.
- Gawinecka, J.; Schönrath, F.; von Eckardstein, A. Acute Aortic Dissection: Pathogenesis, Risk Factors and Diagnosis. Swiss Med. Wkly. 2017, 147, w14489.
- Pyeritz, R.E. Recent Progress in Understanding the Natural and Clinical Histories of the Marfan Syndrome. Trends Cardiovasc. Med. 2016, 26, 423–428.
- Weinsaft, J.W.; Devereux, R.B.; Preiss, L.R.; Feher, A.; Roman, M.J.; Basson, C.T.; Geevarghese, A.; Ravekes, W.; Dietz, H.C.; Holmes, K.; et al. Aortic Dissection in Patients with Genetically Mediated Aneurysms. J. Am. Coll. Cardiol. 2016, 67, 2744–2754.
- Den Hartog, A.W.; Franken, R.; Zwinderman, A.H.; Timmermans, J.; Scholte, A.J.; van den Berg, M.P.; de Waard, V.; Pals, G.; Mulder, B.J.M.; Groenink, M. The Risk for Type B Aortic Dissection in Marfan Syndrome. J. Am. Coll. Cardiol. 2015, 65, 246–254.
- Pape, L.A.; Awais, M.; Woznicki, E.M.; Suzuki, T.; Trimarchi, S.; Evangelista, A.; Myrmel, T.; Larsen, M.; Harris, K.M.; Greason, K.; et al. Presentation, Diagnosis, and Outcomes of Acute Aortic Dissection: 17-Year Trends from the International Registry of Acute Aortic Dissection. J. Am. Coll. Cardiol. 2015, 66, 350–358.
- Tran-Fadulu, V.; Pannu, H.; Kim, D.H.; Vick, G.W.; Lonsford, C.M.; Lafont, A.L.; Boccalandro, C.; Smart, S.; Peterson, K.L.; Hain, J.Z.; et al. Analysis of Multigenerational Families with Thoracic Aortic Aneurysms and Dissections Due to TGFBR1 or TGFBR2 Mutations. J. Med. Genet. 2009, 46, 607–613.
- Jondeau, G.; Ropers, J.; Regalado, E.; Braverman, A.; Evangelista, A.; Teixedo, G.; De Backer, J.; Muiño-Mosquera, L.; Naudion, S.; Zordan, C.; et al. International Registry of Patients Carrying TGFBR1 or TGFBR2 Mutations: Results of the MAC (Montalcino Aortic Consortium). Circ. Cardiovasc. Genet. 2016, 9, 548–558.
- Van de Laar, I.M.B.H.; Oldenburg, R.A.; Pals, G.; Roos-Hesselink, J.W.; de Graaf, B.M.; Verhagen, J.M.A.; Hoedemaekers, Y.M.; Willemsen, R.; Severijnen, L.-A.; Venselaar, H.; et al. Mutations in SMAD3 Cause a Syndromic Form of Aortic Aneurysms and Dissections with Early-Onset Osteoarthritis. Nat. Genet. 2011, 43, 121–126.
- Laterza, D.; Ritelli, M.; Zini, A.; Colombi, M.; Dell’Acqua, M.L.; Vandelli, L.; Bigliardi, G.; Verganti, L.; Vallone, S.; Vincenzi, C.; et al. Novel Pathogenic TGFBR1 and SMAD3 Variants Identified after Cerebrovascular Events in Adult Patients with Loeys-Dietz Syndrome. Eur. J. Med. Genet. 2019, 62, 103727.
- Zhao, H.; Yang, Y.; Pan, X.; Li, W.; Sun, L.; Guo, J. Identification of Clinically Relevant Variants by Whole Exome Sequencing in Chinese Patients with Sporadic Non-Syndromic Type A Aortic Dissection. Clin. Chim. Acta 2020, 506, 160–165.
- Hostetler, E.M.; Regalado, E.S.; Guo, D.-C.; Hanna, N.; Arnaud, P.; Muiño-Mosquera, L.; Callewaert, B.L.; Lee, K.; Leal, S.M.; Wallace, S.E.; et al. SMAD3 Pathogenic Variants: Risk for Thoracic Aortic Disease and Associated Complications from the Montalcino Aortic Consortium. J. Med. Genet. 2019, 56, 252–260.
- Hilhorst-Hofstee, Y.; Scholte, A.J.H.A.; Rijlaarsdam, M.E.B.; van Haeringen, A.; Kroft, L.J.; Reijnierse, M.; Ruivenkamp, C.A.L.; Versteegh, M.I.M.; Pals, G.; Breuning, M.H. An Unanticipated Copy Number Variant of Chromosome 15 Disrupting SMAD3 Reveals a Three-Generation Family at Serious Risk for Aortic Dissection. Clin. Genet. 2013, 83, 337–344.
- Lee, S.-T.; Kim, J.-A.; Jang, S.-Y.; Kim, D.-K.; Kim, J.-W.; Ki, C.-S. A Novel COL3A1 Gene Mutation in Patient with Aortic Dissected Aneurysm and Cervical Artery Dissections. Heart Vessels 2008, 23, 144–148.
- Koitabashi, N.; Yamaguchi, T.; Fukui, D.; Nakano, T.; Umeyama, A.; Toda, K.; Funada, R.; Ishikawa, M.; Kawamura, R.; Okada, K.; et al. Peripartum Iliac Arterial Aneurysm and Rupture in a Patient with Vascular Ehlers-Danlos Syndrome Diagnosed by Next-Generation Sequencing. Int. Heart J. 2018, 59, 1180–1185.
- Makrygiannis, G.; Loeys, B.; Defraigne, J.-O.; Sakalihasan, N. Cervical Artery Dissections and Type A Aortic Dissection in a Family with a Novel Missense COL3A1 Mutation of Vascular Type Ehlers–Danlos Syndrome. Eur. J. Med. Genet. 2015, 58, 634–636.
- Nakamura, M.; Yajima, J.; Oikawa, Y.; Ogasawara, K.; Uejima, T.; Abe, K.; Aizawa, T. Vascular Ehlers-Danlos Syndrome. J. Cardiol. 2009, 53, 458–462.
- Shields, L.B.E.; Rolf, C.M.; Davis, G.J.; Hunsaker III, J.C. Sudden and Unexpected Death in Three Cases of Ehlers-Danlos Syndrome Type IV*: SUDDEN DEATH IN EHLERS-DANLOS TYPE IV. J. Forensic Sci. 2010, 55, 1641–1645.
- Schwarze, U.; Goldstein, J.A.; Byers, P.H. Splicing Defects in the COL3A1 Gene: Marked Preference for 5′ (Donor) Spice-Site Mutations in Patients with Exon-Skipping Mutations and Ehlers-Danlos Syndrome Type IV. Am. J. Hum. Genet. 1997, 61, 1276–1286.
- Sakai, K.; Toda, M.; Kyoyama, H.; Nishimura, H.; Kojima, A.; Kuwabara, Y.; Kobayashi, Y.; Kikuchi, S.; Hirata, Y.; Moriyama, G.; et al. Vascular Ehlers-Danlos Syndrome with a Novel Missense Mutation in COL3A1: A Man in His 50s with Aortic Dissection after Interventional Treatment for Hemothorax as the First Manifestation. Intern. Med. Tokyo Jpn. 2019, 58, 3441–3447.
- Shalhub, S.; Black, J.H.; Cecchi, A.C.; Xu, Z.; Griswold, B.F.; Safi, H.J.; Milewicz, D.M.; McDonnell, N.B. Molecular Diagnosis in Vascular Ehlers-Danlos Syndrome Predicts Pattern of Arterial Involvement and Outcomes. J. Vasc. Surg. 2014, 60, 160–169.
- Meienberg, J.; Rohrbach, M.; Neuenschwander, S.; Spanaus, K.; Giunta, C.; Alonso, S.; Arnold, E.; Henggeler, C.; Regenass, S.; Patrignani, A.; et al. Hemizygous Deletion of COL3A1, COL5A2, and MSTN Causes a Complex Phenotype with Aortic Dissection: A Lesson for and from True Haploinsufficiency. Eur. J. Hum. Genet. 2010, 18, 1315–1321.
- Wang, Z.; Zhuang, X.; Chen, B.; Wen, J.; Peng, F.; Liu, X.; Wei, M. 99-Case Study of Sporadic Aortic Dissection by Whole Exome Sequencing Indicated Novel Disease-Associated Genes and Variants in Chinese Population. BioMed Res. Int. 2020, 2020, 7857043.
- Chen, Y.; Sun, Y.; Li, Z.; Li, C.; Xiao, L.; Dai, J.; Li, S.; Liu, H.; Hu, D.; Wu, D.; et al. Identification of COL3A1 Variants Associated with Sporadic Thoracic Aortic Dissection: A Case-Control Study. Front. Med. 2021, 15, 438–447.
- Frank, M.; Albuisson, J.; Ranque, B.; Golmard, L.; Mazzella, J.-M.; Bal-Theoleyre, L.; Fauret, A.-L.; Mirault, T.; Denarié, N.; Mousseaux, E.; et al. The Type of Variants at the COL3A1 Gene Associates with the Phenotype and Severity of Vascular Ehlers–Danlos Syndrome. Eur. J. Hum. Genet. 2015, 23, 1657–1664.
- Pepin, M.G.; Schwarze, U.; Rice, K.M.; Liu, M.; Leistritz, D.; Byers, P.H. Survival Is Affected by Mutation Type and Molecular Mechanism in Vascular Ehlers-Danlos Syndrome (EDS Type IV). Genet. Med. Off. J. Am. Coll. Med. Genet. 2014, 16, 881–888.
- Saratzis, A.; Bown, M.J. The Genetic Basis for Aortic Aneurysmal Disease. Heart Br. Card. Soc. 2014, 100, 916–922.
- Guo, D.; Regalado, E.; Casteel, D.E.; Santos-Cortez, R.L.; Gong, L.; Kim, J.J.; Dyack, S.; Horne, S.G.; Chang, G.; Jondeau, G.; et al. Recurrent Gain-of-Function Mutation in PRKG1 Causes Thoracic Aortic Aneurysms and Acute Aortic Dissections. Am. J. Hum. Genet. 2013, 93, 398–404.
- Teodoro Jerves; Andrea Beaton; Paul Kruszka; The genetic workup for structural congenital heart disease. American Journal of Medical Genetics Part C: Seminars in Medical Genetics 2019, 184, 178-186, 10.1002/ajmg.c.31759.
- Christina M. Rigelsky; Rocio T. Moran; Genetics of syndromic and nonsyndromic aortopathies. Current Opinion in Pediatrics 2019, 31, 694-701, 10.1097/mop.0000000000000836.
- Jj Gómez de Diego; Comments on the 2014 ESC Guidelines on the Diagnosis and Treatment of Aortic Diseases. Revista Española de Cardiología (English Edition) 2015, 68, 179-184, 10.1016/j.rec.2014.12.003.
- John Augoustides; Michael Andritsos; Hiratzka Lf; Bakris Gl; Beckman Ja; Bersin Rm; Carr Vf; Eagle Ka; Hermann Lk; Isselbacher Em; et al.Kazerooni EaKouchoukos NtLytle BwMilewicz DmReich DlSen SShinn JaSvensson LgWilliams DmAmerican College of Cardiology Foundation/American Heart Association Task Force on Practice GuidelinesAmerican Association for Thoracic SurgeryAmerican College of RadiologyAmerican Stroke AssociationSociety Of Cardiovascular AnesthesiologistsSociety For Cardiovascular Angiography And InterventionsSociety Of Interventional RadiologySociety Of Thoracic SurgeonsSociety for Vascular Medicine 2010 ACCF/AHA/AATS/ACR/ASA/SCA/SCAI/SIR/STS/SVM guidelines for the diagnosis and management of patients with Thoracic Aortic Disease: a report of the American College of Cardiology Foundation/American Heart Association Task Force on Practice Guidelines, American Association for Thoracic Surgery, American College of Radiology, American Stroke Association, Society of Cardiovascular Anesthesiologists, Society for Cardiovascular Angiography and Interventions, Society of Interventional Radiology, Society of Thoracic Surgeons, and Society for Vascular Medicine.. Faculty Opinions – Post-Publication Peer Review of the Biomedical Literature 2010, 121, e266–e369, 10.3410/f.2671959.2334057.
- Dong-Chuan Guo; Ellen M. Hostetler; Yuxin Fan; Richard J. Kulmacz; Di Zhang; Deborah A. Nickerson; Suzanne M. Leal; Scott A. LeMaire; Ellen S. Regalado; Dianna M. Milewicz; et al. Heritable Thoracic Aortic Disease Genes in Sporadic Aortic Dissection. Journal of the American College of Cardiology 2017, 70, 2728-2730, 10.1016/j.jacc.2017.09.1094.
- Kelli L. Hicks; Peter H. Byers; Elina Quiroga; Melanie G. Pepin; Sherene Shalhub; Testing patterns for genetically triggered aortic and arterial aneurysms and dissections at an academic center. Journal of Vascular Surgery 2018, 68, 701-711, 10.1016/j.jvs.2017.12.023.
- Tim Ripperger; Hans Dieter Tröger; Jörg Schmidtke; The genetic message of a sudden, unexpected death due to thoracic aortic dissection. Forensic Science International 2009, 187, 1-5, 10.1016/j.forsciint.2009.01.020.
- Christopher K. Mehta; Andre Y. Son; Matthew C. Chia; Ashley N. Budd; Bradley D. Allen; Patricia Vassallo; Andrew W. Hoel; William J. Brady; Jose V. Nable; Management of acute aortic syndromes from initial presentation to definitive treatment. The American Journal of Emergency Medicine 2021, 51, 108-113, 10.1016/j.ajem.2021.10.016.





