Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Zhang, X.; Zhu, X.; Bi, X.; Huang, J.; Zhou, L. The Insulin Receptor. Encyclopedia. Available online: https://encyclopedia.pub/entry/25663 (accessed on 29 May 2026).
Zhang X, Zhu X, Bi X, Huang J, Zhou L. The Insulin Receptor. Encyclopedia. Available at: https://encyclopedia.pub/entry/25663. Accessed May 29, 2026.
Zhang, Xiaohong, Xuezhen Zhu, Xiaoyang Bi, Jiguang Huang, Lijuan Zhou. "The Insulin Receptor" Encyclopedia, https://encyclopedia.pub/entry/25663 (accessed May 29, 2026).
Zhang, X., Zhu, X., Bi, X., Huang, J., & Zhou, L. (2022, July 29). The Insulin Receptor. In Encyclopedia. https://encyclopedia.pub/entry/25663
Zhang, Xiaohong, et al. "The Insulin Receptor." Encyclopedia. Web. 29 July, 2022.
Copy Citation
The insulin receptor (IR) is a transmembrane protein that is activated by ligands in insulin signaling pathways. The IR has been considered as a novel therapeutic target for clinical intervention, considering the overexpression of its protein and A-isoform in multiple cancers, Alzheimer’s disease, and Type 2 diabetes mellitus in humans. Meanwhile, it may also serve as a potential target in pest management due to its multiple physiological influences in insects.
insulin receptor
function
agonists
antagonists
mechanism
1. Introduction
The insulin receptor (IR) is a transmembrane protein and part of the tyrosine kinase receptors (RTK). It exists as covalently bound receptor dimers at the cell surface [1]. The IR plays essential roles in metabolism, cell growth, and development by transmitting the binding of extracellular ligands into several intracellular signaling cascades [2][3][4]. Previous studies have demonstrated that ligands and the insulin signaling IR are highly conserved among human beings and insects [5][6][7].
In human beings, the function of the IR has been studied for many years, and it has been found to play a crucial role in multiple chronic diseases, including Alzheimer’s disease (AD) [8], Type 2 diabetes mellitus (T2DM) [9][10], and various cancers [2][11][12][13], as well as neurodegenerative disorders [14] and metabolic syndromes [15]. For T2DM, the destruction and dysfunction of pancreatic β-cells are common occurrences, and insulin injection is the only choice for glycemic control [16]. The dramatic increase in T2DM over the globe has led to increasing requirements for insulin. Moreover, insulin injection may require more than one shot each day, is hazardous and inconvenient, causes tissue irritation, abscesses, discomfort, etc., and local allergic reactions, lipoatrophy, lipohypertrophy, etc., are common complications of subcutaneous injections [17][18]. Because of the multiple problems associated with insulin injection, orally active insulin-mimetic compounds would be an ideal substitute [19]. For cancer, IR makes an attractive anticancer target owing to its overexpression in a variety of cancers, especially prostate and breast cancers [20]. Therefore, regulators of the IR, such as β-site amyloid precursor protein cleaving enzyme 1 (BACE1), have been regarded as potential therapeutic target [20][21]. Similarly, IR modulators such as ceritinib and anti-idiotypic antibody AK98 (an off-target IR inhibitor) have been suggested as promising drugs for the treatment of brain tumors and breast cancer, respectively [22][23].
In insects, current evidence points to the roles of the IR in regulating development, reproduction, lifespan, caste differentiation, and wing polyphenism [24][25][26]. Neonicotinoid insecticides (e.g., imidacloprid) are selective agonists of the nicotinic acetylcholine receptor (nAChR) that have been widely used to control various insects [27]. Likewise, ryanodine and diamides are commercial insecticides that are antagonists or activators of insect ryanodine receptors (RyRs) [28]. It may be deduced that modulators of the IR that selectively activate or inhibit the IR may be of considerable value in providing promising drugs for the control of human disease, or as insecticides for the control of insects [29]. In this regard, medicines specifically targeting the IR are diverse. However, IR-targeting insecticides are still lacking. Owing to the persistent use of traditional synthetic insecticides, insect resistance has become increasingly serious. Therefore, there is a growing need for new insecticides with new mechanisms of action. Thus IR-targeting insecticides represent an opportunity in the research and development of insecticides.
2. Biology Studies of the IR
2.1. Molecular Structure of the IR
Biochemically, the IR is encoded by a single gene. The coding region of the IR gene has 22 exons and 21 introns [30]. The alternative splicing of exon 11 encodes a 12-amino-acid sequence at the C-terminus of the α-subunit of the IR gene during transcription, resulting in the formation of the isoforms IR-A and IR-B [31]. IR-B is a mature isoform due to the fact that it includes the 12-amino-acid sequence, while the fetal isoform IR-A does not [10][32]. Both isoforms are expressed in most of the cells associated with energy homeostasis, such as adipocytes, hepatocytes, myocytes, and placenta vascular endothelium; however, they present different functional features [10][33]. Several in vitro and in vivo studies have confirmed that the expression and response of the two isoforms are different in breast cancer and T2DM [11]. IR-B possesses important metabolic functions and is the dominant isoform [2]. Conversely, the less-differentiated isoform IR-A is principally expressed in cancer cells [32]. Activation of IR-A promotes the growth of the cancer cells [34].
IR structural studies have previously been described in detail [35][36][37][38] (Table 1). The IR is a glycosylated, disulfide-linked (αβ)2 transmembrane homodimer consisting of two repeated ectodomains (ECD), a single transmembrane helix, and two intracellular cytoplasmic domain that includes a tyrosine kinase domain (TKD) (Figure 1a) [38][39]. The α-subunit constitutes most of the IR-ECD, while the β-subunit is necessary for the IR-ECD, the transmembrane domain (TMD), and the intracellular TKD [38].
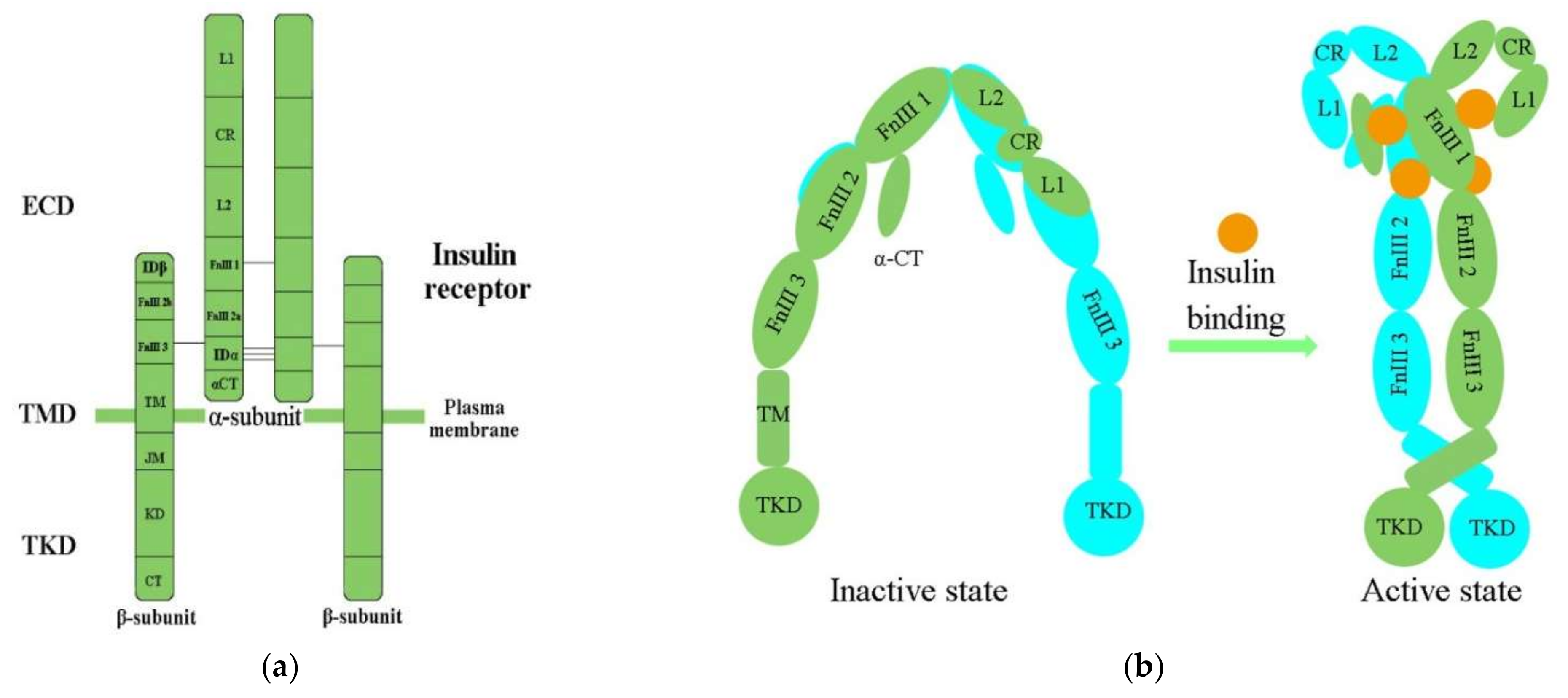
Figure 1. (a) The architectural domain of the IR (αβ) 2 homodimer. Black lines indicate the intersubunit disulfide bonds; (b) Inactive and active states of the IR; L1, L2, leucine-rich repeat domains 1, 2; CR, cysteine-rich domain; FnIII-1, 2, 3, fibronectin type-III domains 1, 2, 3; αCT, α C-terminal regions; TM, transmembrane; JM, juxtamembrane; KD, kinase domain; CT, C-terminal tail; ECD, ectodomain; TMD, transmembrane domain; TKD, tyrosine kinase domain.
Table 1. Summary of the available structures of IR.
| Classification | Structure of IR | References |
|---|---|---|
| Domain layout | an (αβ)2 disulfide-linked homodimer | [35] |
| cDNA sequenced | α chain lies on the N-terminal of the β chain | |
| 3D structure of human apo IR ectodomain | intracellular unphosphorylated from TKD (2.1 Å resolution, PDB 1IRK) | |
| receptor’s isolated L1-CR-L2 module (2.32 Å resolution, PDB 2HR7) | ||
| intact receptor ectodomain in apo form (3.8 Å resolution, PDB 2DTG) | ||
| CryoEM structures of IR | insulin holoreceptor (full-length receptor inclusive of transmembrane and cytoplasmic elements) | [40] |
| isolated receptor ectodomain | [41][42] | |
| an ectodomain construct (leucine-zippered receptor ectodomain) | [43] |
Determination of the three-dimensional (3D) crystal structure of the insulin-free IR-ECD through crystallography has revealed that the IR-ECD dimer roughly displays an inverted “U”- or “V”-shaped architecture [37][44]. Specifically, L1 and CR together with L2 form one leg, while the linearly arranged FnIII domains form the other leg [37]. However, the modular organization of the ECD, with high intrinsic flexibility and its complex ligand-binding properties, poses a challenge for structural studies of the IR. Furthermore, single-particle cryo-electron microscopy (cryo-EM) has revealed that the IR-ECD dimer converts the overall architecture from an autoinhibited inverted “V” shape into a “T”-shaped conformation, which was stabilized after binding insulin molecules to the N-terminal domains (Figure 1b) [39][40][41]. The L1, CR, and L2 domains of both IR promoters constitute the “T” horizontal part, while the FnIII-1, -2, and -3 domains of the IR dimer constitute the vertical piece of the “T” [40].
Previous biochemical and mutagenesis models of insulin binding have identified two distinct binding sites on both the IR and insulin, termed site 1 (S1) and site 2 (S2) [36]. The L1 subdomain and the α-CT helix residue have been confirmed to represent IR S1 site (IR-S1) [39][45][46]. Evidence has indicated that IR-S1 is indispensable for insulin binding, and minor modifications of it were sufficient to change the IR’s specificity for insulin [47]. Scapin (2018) has defined the full S2 binding site [39], and Gutmann (2020) first observed the connection of insulin with discrete IR-S2 [42]. Studies have demonstrated that optimal IR activation requires multiple insulin molecules bound to S1 and S2 [48][49]. A similar result has also been presented in a study of the cryo-EM structure of the IR–insulin complex at 3.2 Å resolution [40]. The binding of insulin to S1 of apo-IR could release the autoinhibited conformation, which was an essential step for IR activation, while binding to S2 was important for the IR to adopt the active T-shape [50][51][52]. Cryo-EM analysis of the insulin–IR complex has revealed that insulin binds independently to the site of S2 between the FnIII-1 and FnIII-2 domains [42]. Another study has shown that the fibronectin domain is folded inwards, in a pincer-like fashion, which brings domains FnIII-3 and FnIII-3′ into contact [43].
The cryo-EM structure of the full-length human IR–insulin complex (human HEK293F cells) in the active state at an overall resolution of 3.2 Å unexpectedly revealed that a maximum of four insulin molecules can bind to the “T”-shaped IR dimer at four distinct sites [40]. Furthermore, at least one insulin molecule bound to two S2s and a maximum of four insulin molecules at four sites are required to form the “T”-shaped dimer [39][40][52]. Insulin 1 mainly binds to the primary site formed by the L1 domain, and α-CT then makes contact with a loop of the FnIII-1 domain from the IR promoter that donates α-CT [40]. During IR activation, a tripartite interface between insulin 1 and site 1 stabilizes the active IR dimer. Insulin 2 binds to a novel binding site on the FnIII-1 domain, located on the backside of the β sheet [40].
However, there is still a lack of detailed analysis of which site is connected first and how the first and second insulin binding results in different phosphorylation status of the IR [52]. The reported findings have emphasized the importance of the conformational changes of the IR-ECD and IR–insulin complex in the insulin/insulin-like growth factor signaling (IIS) pathway. Hence, the precise mechanism of how insulin binds to the IR at first remains elusive, and further research is still needed.
2.2. Activation of the IR
Physiologically, the function of the IR is activated in the insulin/IGF-1-like signal (IIS) pathway by the ligand [2]. The IIS pathway is commonly known as a significant nutrient-dependent endocrine pathway and regulates numerous physiological processes, such as metabolism, growth and development, and so on [6]. In the IIS pathway, the IR regulates two primary cell-signaling cascades (Figure 2) [53]: the phosphatidylinositol-3-kinase (PI3K)/AKT signaling pathway and the mitogen-activated protein kinase (MAPK) pathway (extracellular-signal regulated kinase signaling pathway (ERK)) [53][54][55][56]. The PI3K/AKT pathway is primarily responsible for controlling metabolic processes such as glucose transportation and the synthesis of lipids, proteins, and glycogen. In contrast, the MAPK pathway is primarily related to the mitogenic effects of insulin and is mainly responsible for cell growth and proliferation [56][57].
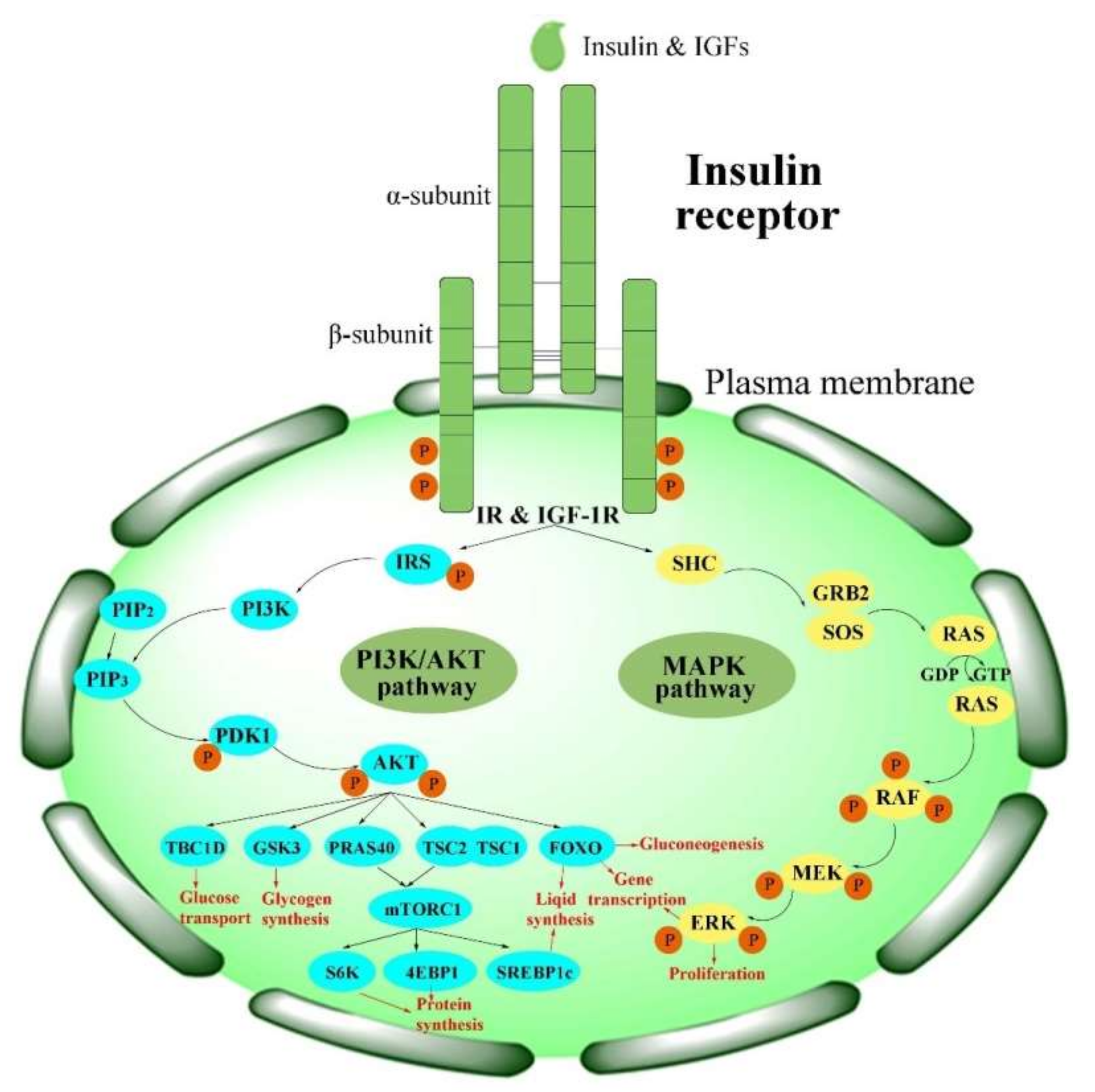
Figure 2. Activation of the IR in insulin signaling pathways. PI3K/AKT pathways: phosphatidylinositol-3-kinase signaling pathways; MAPK pathway: mitogen-activated protein kinase pathway.
The major upstream factors of the IIS pathway are various insulin-like peptides (ILPs). Based on primary structure and receptor binding preferences, these ILPs can be subdivided into insulin, insulin-related growth factors (IGFs, including IGF-I and IGF-II) in mammals, and ILPs in insects [58]. Insulin is a peptide hormone secreted by pancreas β-cells and is one of the most conserved molecules in animals [59][60]. IGFs are peptides that have a homology of 40–80% with insulin. In humans, both insulin and IGFs can bind to the IR on the cell surface and functionally mediate cellular proliferation and differentiation, lipid metabolism, glucose homeostasis, and DNA synthesis [60][61]. Meanwhile, in insects, ILPs are the most general growth-promotion signaling factors [62][63][64][65], and evidence has suggested that ILPs are homologues of human insulin [6][66]. Insulin is the major regulatory factor in humans, but various ILPs have been identified in different insect species, ranging from one—in the Nevada dampwood termite, Zootermopsis nevadensis (Hagen)—to more than 40—in the silkworm, Bombyx mori L. [60].
ILPs first phosphorylate the IR and then activate IR signaling. The tyrosine-phosphorylated IR, in return, recruits and phosphorylates other intracellular adaptor proteins, such as IR substrate (IRS) proteins and several other substrates, including Src homology 2 domain-containing (SHC), Grb2-associated binder (GAB), APS (SHB2), and Cbl, at several tyrosine residues [56][67]. There are six isoforms (IRS1–6) in the IRS family [68]; among these, IRS1 and IRS2 are the main isoforms [68][69]. These proteins mediate the association with the Src homology 2 (SH2) domains and lead to initiation of the PI3K/AKT pathway, as well as activation of the downstream phosphoinositide-dependent kinase (PDK1) and protein kinase B (PKB, also called AKT) [4]. Phosphorylation of the IR triggers the activation of cellular signaling pathways, which play different roles in human beings and insects.
3. Functions of the IR
3.1. The Functions of the IR in Human Beings
In humans, the IR plays a crucial role in whole-body nutrient homeostasis and in various diseases, such as AD [8], T2DM [4][9][10], obesity [70], atherosclerosis [31], multiple cancers [11][12][13], and cardiovascular disease [71], as well as neurodegenerative disorders [14], metabolic syndrome [15] and polycystic ovary syndrome [72]. Thus, it is necessary to understand the cellular expression and the functions of the IR in order to propose new treatment concepts and to develop novel drugs.
The IR mediates whole-body nutrient homeostasis and is expressed ubiquitously through the classic insulin-responsive targets in the liver, muscle, and adipose tissue [3]. A has demonstrated that the IR is distributed in both dendritic shafts and spines in living hippocampal brain neurons [73]. Knockout of the IR resulted in many impaired target organs. Hepatic deletion of the IR led to hyperglycemia, disorders in fatty acid metabolism, and an increase in the expression of fatty acid oxidation enzymes [74]. In mucosal epithelial cells, the IR interacts with the voltage-dependent anion channel-1 (VDAC1) in mitochondria. Knockdown of the IR gene triggered robust mitochondrial fragmentation and altered polarization [75], while knockout of the β-cell IR gene led to impaired insulin secretion [76]. Additionally, missense mutations of the IR may cause severe inherited insulin resistance syndromes [77].
The IR is a cell-surface receptor translocating to the nucleus, and is associated strongly with RNA polymerase II in the chromatin [78]. In the cell, host cell factor-1 (HCF-1) acts as a transcriptional coregulator functionally mediating the binding of the IR to specific sites located in the gene promoters [79]. HCF-1 mediates the association between the IR and DNA. HCF-1 binds to DNA indirectly through DNA sequence-specific transcription factors, and then forms a complex with the IR and Thanatos-associated protein domain-containing protein 11 (THAP11) in the chromatin. Knockdown of HCF-1 can inhibit the binding ability of the IR to the promotors [79]. Another study indicated that the mRNA and protein levels of the IR were obviously reduced in the subcutaneous and visceral adipose tissue of women with gestational diabetes mellitus (GDMs) [80]. The decrease in IR mRNA was accompanied by a decrease in methylation levels of the IR promoter [80]. This phenomenon has also been observed in the hypothalamus [81]. The methylation degree of the IR nuclear factor I (IRNF-I) binding site within the IR promoter was dramatically inversely correlated with the gene level of the IR. These findings have opened a new avenue for further studies on the functions and mechanisms of the IR. More studies focusing on demonstrating whether epigenetic modifications in the IR sequence impact IR expression of the IR are needed [82].
3.2. The Functions of the IR in Insects
Studies of the IR in human beings have raised interest in the functions of the IR in insects and the consequent possibility for the development of new IR-targeting insecticides with high efficiency and low toxicity.
In insects, multiple functions of the IR have been revealed [83]. The IR is well-known to be implicated—either directly or by crosstalk with other major hormones such as juvenile hormone (JH) and ecdysteroids (especially 20-hydroxyecdysone, 20E)—in post-embryonic development [84][85], nutrition-based phenotypic plasticity and body size control [86][87], reproduction and diapause [55], and circadian rhythmicity and behaviors [88][89][90]. The IR is also indispensable in insect photoperiodism, lifespan, and aging due to its relation to metabolism and growth [25][91][92][93]. Overall, studies have indicated that the IR is indispensable in insect growth [94], development and reproduction [95][96], polymorphism [24], lifespan [97], and oviposition [98]. Therefore, the IR represents an important target for the management of pests and parasites.
References
- Ward, C.W.; Lawrence, M.C. Ligand-induced activation of the insulin receptor: A multi-step process involving structural changes in both the ligand and the receptor. Bioessays 2009, 31, 422–434.
- Vigneri, R.; Goldfine, I.D.; Frittitta, L. Insulin, insulin receptors, and cancer. J. Endocrinol. Investig. 2016, 39, 1365–1376.
- Haeusler, R.A.; McGraw, T.E.; Accili, D. Biochemical and cellular properties of insulin receptor signalling. Nat. Rev. Mol. Cell Biol. 2018, 19, 31–44.
- Yunn, N.O.; Kim, J.; Kim, Y.; Leibiger, I.; Berggren, P.O.; Ryu, S.H. Mechanistic understanding of insulin receptor modulation: Implications for the development of anti-diabetic drugs. Pharmacolo. Therapeut. 2018, 185, 86–98.
- Brogiolo, W.; Stocker, H.; Ikeya, T.; Rintelen, F.; Fernandez, R.; Hafen, E. An evolutionarily conserved function of the Drosophila insulin receptor and insulin-like peptides in growth control. Curr. Biol. 2001, 11, 213–221.
- Lin, X.; Smagghe, G. Roles of the insulin signaling pathway in insect development and organ growth. Peptides 2019, 122, 169923.
- Liu, C.Y.; Zhao, W.L.; Wang, J.X.; Zhao, X.F. Cyclin-dependent kinase regulatory subunit 1 promotes cell proliferation by insulin regulation. Cell Cycle 2015, 14, 3045–3057.
- Craft, S.; Cholerton, B.; Baker, L.D. Insulin and Alzheimer’s disease: Untangling the web. J. Alzheimer’s Dis. 2013, 33 (Suppl. S1), S263–S275.
- Czech, M.P. Insulin action and resistance in obesity and type 2 diabetes. Nat. Med. 2017, 23, 804–814.
- Westermeier, F.; Saez, T.; Arroyo, P.; Toledo, F.; Gutierrez, J.; Sanhueza, C.; Pardo, F.; Leiva, A.; Sobrevia, L. Insulin receptor isoforms: An integrated view focused on gestational diabetes mellitus. Diabetes Metab. Res. Rev. 2016, 32, 350–365.
- Vella, V.; Milluzzo, A.; Scalisi, N.; Vigneri, P.; Sciacca, L. Insulin receptor isoforms in cancer. Int. J. Mol. Sci. 2018, 19, 3615.
- Sciacca, L.; Le Moli, R.; Vigneri, R. Insulin analogs and cancer. Front. Endocrinol. 2012, 3, 21.
- Malaguarnera, R.; Belfiore, A. The insulin receptor: A new target for cancer therapy. Front. Endocrinol. 2011, 2, 93.
- Stoeckel, L.E.; Arvanitakis, Z.; Gandy, S.; Small, D.; Kahn, C.R.; Pascual-Leone, A.; Pawlyk, A.; Sherwin, R.; Smith, P. Complex mechanisms linking neurocognitive dysfunction to insulin resistance and other metabolic dysfunction. F1000Res 2016, 5, 353.
- Titchenell, P.M.; Lazar, M.A.; Birnbaum, M.J. Unraveling the regulation of hepatic metabolism by insulin. Trends Endocrinol Metab. 2017, 28, 497–505.
- Campbell, R.K. Type 2 diabetes: Where we are today: An overview of disease burden, current treatments, and treatment strategies. J. Am. Pharm. Assoc. 2009, 49, S3–S9.
- Ermakova, A.; Stauffer, M.E.; Sieradzan, R.; Taylor, S. Characterizing the clinical and economic burden of type 2 diabetes (T2DM) patients on multiple daily injections (MDI) of insulin: A systematic literature review. Value Health 2018, 21, S70.
- Buyruk, B.A.; Kebapci, N.; Yorulmaz, G.; Alaguney, E.S.; Akalin, A.; Efe, B. Prevalence and risk factors of lipohypertrophy and lipoatrophy in diabetes patients receiving insulin therapy. Diabetes 2019, 68, 59.
- Brietzke, S.A. Oral antihyperglycemic treatment options for type 2 diabetes mellitus. Med. Clin. North Am. 2015, 99, 87–106.
- Singh, P.; Alex, J.M.; Bast, F. Insulin receptor (IR) and insulin-like growth factor receptor 1 (IGF-1R) signaling systems: Novel treatment strategies for cancer. Med. Oncol. 2014, 31, 805.
- Meakin, P.J.; Mezzapesa, A.; Benabou, E.; Haas, M.E.; Bonardo, B.; Grino, M.; Brunel, J.M.; Desbois-Mouthon, C.; Biddinger, S.B.; Govers, R.; et al. The beta secretase BACE1 regulates the expression of insulin receptor in the liver. Nat. Commun. 2018, 9, 1306.
- Russo, A.; Paret, C.; Alt, F.; Burhenne, J.; Fresnais, M.; Wagner, W.; Glaser, M.; Bender, H.; Huprich, S.; Harter, P.N.; et al. Ceritinib-induced regression of an insulin-like growth factor-driven neuroepithelial brain tumor. Int. J. Mol. Sci. 2019, 20, 4267.
- Wenbin, K.; Xiaoqin, L.; Qiuchan, D.; Xinwen, Z.; Xiaoqin, X.; Fangyuan, S.; Dabao, H.; Shuangjiu, Z. Development of a novel insulin receptor (IR) antagonist that exhibits anti-breast tumor activity. Human Cell 2020, 33, 1204–1217.
- Lin, X.; Lavine, L.C. Endocrine regulation of a dispersal polymorphism in winged insects: A short review. ScienceDirect 2018, 25, 20–24.
- Guo, S.; Wang, X.; Kang, L. Special significance of non-Drosophila insects in aging. Front. Cell Dev. Biol. 2020, 8, 576571.
- Alvarez-Rendon, J.P.; Salceda, R.; Riesgo-Escovar, J.R. Drosophila melanogaster as a model for diabetes type 2 progression. Biomed. Res. Int. 2018, 2018, 1417528.
- Houchat, J.N.; Cartereau, A.; Le Mauff, A.; Taillebois, E.; Thany, S.H. An overview on the effect of neonicotinoid insecticides on mammalian cholinergic functions through the activation of neuronal nicotinic acetylcholine receptors. Int. J. Environ. Res. Public Health 2020, 17, 3222.
- Sun, Z.; Xu, H. Ryanodine receptors for drugs and insecticides: An overview. Mini. Rev. Med. Chem. 2019, 19, 22–33.
- Sun, Z.; Lv, M.; Huang, W.; Li, T.; Xu, H. Development of botanical pesticides: Exploration on the phenotype of vestigial wings of insect pests induced by plant natural products or their derivatives by blocking tyrosine phosphorylation of insulin receptor 1. J. Agric. Food Chem. 2022, 70, 2117–2126.
- Belfiore, A.; Frasca, F.; Pandini, G.; Sciacca, L.; Vigneri, R. Insulin receptor isoforms and insulin receptor/insulin-like growth factor receptor hybrids in physiology and disease. Endocr. Rev. 2009, 30, 586–623.
- Beneit, N.; Fernandez-Garcia, C.E.; Martin-Ventura, J.L.; Perdomo, L.; Escribano, O.; Michel, J.B.; Garcia-Gomez, G.; Fernandez, S.; Diaz-Castroverde, S.; Egido, J.; et al. Expression of insulin receptor (IR) A and B isoforms, IGF-IR, and IR/IGF-IR hybrid receptors in vascular smooth muscle cells and their role in cell migration in atherosclerosis. Cardiovasc. Diabetol. 2016, 15, 161.
- Belfiore, A.; Malaguarnera, R.; Vella, V.; Lawrence, M.C.; Sciacca, L.; Frasca, F.; Morrione, A.; Vigneri, R. Insulin receptor isoforms in physiology and disease: An updated view. Endocr. Rev. 2017, 38, 379–431.
- Huang, J.; Morehouse, C.; Streicher, K.; Higgs, B.W.; Gao, J.; Czapiga, M.; Boutrin, A.; Zhu, W.; Brohawn, P.; Chang, Y.; et al. Altered expression of insulin receptor isoforms in breast cancer. PLoS ONE. 2011, 6, e26177.
- Heidegger, I.; Kern, J.; Ofer, P.; Klocker, H.; Massoner, P. Oncogenic functions of IGF1R and INSR in prostate cancer include enhanced tumor growth, cell migration and angiogenesis. Oncotarget 2014, 5, 2723–2735.
- Lawrence, M.C. Understanding insulin and its receptor from their three-dimensional structures. Mol Metab. 2021, 52, 101255.
- De Meyts, P. Insulin/receptor binding: The last piece of the puzzle? What recent progress on the structure of the insulin/receptor complex tells us (or not) about negative cooperativity and activation. Bioessays 2015, 37, 389–397.
- Croll, T.I.; Smith, B.J.; Margetts, M.B.; Whittaker, J.; Weiss, M.A.; Ward, C.W.; Lawrence, M.C. Higher-resolution structure of the human insulin receptor ectodomain: Multi-modal inclusion of the insert domain. Structure 2016, 24, 469–476.
- Ye, L.; Maji, S.; Sanghera, N.; Gopalasingam, P.; Gorbunov, E.; Tarasov, S.; Epstein, O.; Klein-Seetharaman, J. Structure and dynamics of the insulin receptor: Implications for receptor activation and drug discovery. Drug Discov. Today 2017, 22, 1092–1102.
- Scapin, G.; Dandey, V.P.; Zhang, Z.; Prosise, W.; Hruza, A.; Kelly, T.; Mayhood, T.; Strickland, C.; Potter, C.S.; Carragher, B. Structure of the insulin receptor-insulin complex by single-particle cryo-EM analysis. Nature 2018, 556, 122–125.
- Uchikawa, E.; Choi, E.; Shang, G.; Yu, H.; Bai, X.C. Activation mechanism of the insulin receptor revealed by cryo-EM structure of the fully liganded receptor-ligand complex. Elife 2019, 8, e48630.
- Gutmann, T.; Kim, K.H.; Grzybek, M.; Walz, T.; Coskun, U. Visualization of ligand-induced transmembrane signaling in the full-length human insulin receptor. J. Cell Biol. 2018, 217, 1643–1649.
- Gutmann, T.; Schafer, I.B.; Poojari, C.; Brankatschk, B.; Vattulainen, I.; Strauss, M.; Coskun, U. Cryo-EM structure of the complete and ligand-saturated insulin receptor ectodomain. J. Cell Biol. 2020, 219, e201907210.
- Weis, F.; Menting, J.G.; Margetts, M.B.; Chan, S.J.; Xu, Y.; Tennagels, N.; Wohlfart, P.; Langer, T.; Muller, C.W.; Dreyer, M.K.; et al. The signalling conformation of the insulin receptor ectodomain. Nat. Commun. 2018, 9, 4420.
- McKern, N.M.; Lawrence, M.C.; Streltsov, V.A.; Lou, M.Z.; Adams, T.E.; Lovrecz, G.O.; Elleman, T.C.; Richards, K.M.; Bentley, J.D.; Pilling, P.A.; et al. Structure of the insulin receptor ectodomain reveals a folded-over conformation. Nature 2006, 443, 218–221.
- Whittaker, J.; Whittaker, L. Characterization of the functional insulin binding epitopes of the full-length insulin receptor. J. Biol. Chem. 2005, 280, 20932–20936.
- Menting, J.G.; Whittaker, J.; Margetts, M.B.; Whittaker, L.J.; Kong, G.K.; Smith, B.J.; Watson, C.J.; Zakova, L.; Kletvikova, E.; Jiracek, J.; et al. How insulin engages its primary binding site on the insulin receptor. Nature 2013, 493, 241–245.
- Chrudinova, M.; Zakova, L.; Marek, A.; Socha, O.; Budesinsky, M.; Hubalek, M.; Picha, J.; Machackova, K.; Jiracek, J.; Selicharova, I. A versatile insulin analog with high potency for both insulin and insulin-like growth factor 1 receptors: Structural implications for receptor binding. J. Biol. Chem. 2018, 293, 16818–16829.
- Kavran, J.M.; McCabe, J.M.; Byrne, P.O.; Connacher, M.K.; Wang, Z.; Ramek, A.; Sarabipour, S.; Shan, Y.; Shaw, D.E.; Hristova, K.; et al. How IGF-1 activates its receptor. Elife 2014, 3, e03772.
- Xu, Y.; Kong, G.K.; Menting, J.G.; Margetts, M.B.; Delaine, C.A.; Jenkin, L.M.; Kiselyov, V.V.; De Meyts, P.; Forbes, B.E.; Lawrence, M.C. How ligand binds to the type 1 insulin-like growth factor receptor. Nat. Commun. 2018, 9, 821.
- Yunn, N.-O.; Park, M.; Noh, J.; Ryu, S.H. Stepwise autophosphorylation regulates biased agonism of the insulin receptor. Faseb. J. 2019, 34, s1.08794.
- Hernandez-Sanchez, C.; Mansilla, A.; Pablo, F.d.; Zardoya, R. Evolution of the insulin receptor family and receptor isoform expression in vertebrates. Mol. Biol. Evol. 2008, 25, 1043–1053.
- Li, J.; Park, J.; Mayer, J.P.; Webb, K.J.; Uchikawa, E.; Wu, J.Y.; Liu, S.; Zhang, X.W.; Stowell, M.H.B.; Choi, E.; et al. Synergistic activation of the insulin receptor via two distinct sites. Nat. Struct. Mol. Biol. 2022, 29, 357–368.
- Das, D.; Arur, S. Conserved insulin signaling in the regulation of oocyte growth, development, and maturation. Mol. Reprod. Dev. 2017, 84, 444–459.
- Lawrence, M.C.; McKern, N.M.; Ward, C.W. Insulin receptor structure and its implications for the IGF-1 receptor. Curr. Opin. Struct Biol. 2007, 17, 699–705.
- Sim, C.; Denlinger, D.L. Insulin signaling and the regulation of insect diapause. Front. Physiol. 2013, 4, 189.
- Hall, C.; Yu, H.; Choi, E. Insulin receptor endocytosis in the pathophysiology of insulin resistance. Exp. Mol. Med. 2020, 52, 911–920.
- Cai, W.; Sakaguchi, M.; Kleinridders, A.; Gonzalez-Del Pino, G.; Dreyfuss, J.M.; O’Neill, B.T.; Ramirez, A.K.; Pan, H.; Winnay, J.N.; Boucher, J.; et al. Domain-dependent effects of insulin and IGF-1 receptors on signalling and gene expression. Nat. Commun. 2017, 8, 14892.
- Okada, Y.; Katsuki, M.; Okamoto, N.; Fujioka, H.; Okada, K. A specific type of insulin-like peptide regulates the conditional growth of a beetle weapon. PLoS Biol. 2019, 17, e3000541.
- Laustsen, P.G.; Russell, S.J.; Cui, L.; Entingh-Pearsall, A.; Holzenberger, M.; Liao, R.; Kahn, C.R. Essential role of insulin and insulin-like growth factor 1 receptor signaling in cardiac development and function. Mol. Cell Biol. 2007, 27, 1649–1664.
- Nassel, D.R.; Vanden Broeck, J. Insulin/IGF signaling in Drosophila and other insects: Factors that regulate production, release and post-release action of the insulin-like peptides. Cell Mol. Life Sci. 2016, 73, 271–290.
- Inoue, H. Central insulin-mediated regulation of hepatic glucose production. Endocr. J. 2016, 63, EJ15-0540.
- Nijhout, H.F.; Smith, W.A.; Schachar, I.; Subramanian, S.; Tobler, A.; Grunert, L.W. The control of growth and differentiation of the wing imaginal disks of Manduca sexta. Dev. Biol. 2007, 302, 569–576.
- Chen, C.H.; Pan, J.; Di, Y.Q.; Liu, W.; Hou, L.; Wang, J.X.; Zhao, X.F. Protein kinase C delta phosphorylates ecdysone receptor B1 to promote gene expression and apoptosis under 20-hydroxyecdysone regulation. Proc. Natl. Acad. Sci. USA 2017, 114, E7121–E7130.
- Ajaha, A.; Bouayad, N.; Aarab, A.; Rharrabe, K. Effect of 20-hydroxyecdysone, a phytoecdysteroid, on development, digestive, and detoxification enzyme activities of Tribolium castaneum (Coleoptera: Tenebrionidae). J. Insect Sci. 2019, 19, 18.
- Wu, Q.; Brown, M.R. Signaling and function of insulin-like peptides in insects. Annu. Rev. Entomol. 2006, 51, 1–24.
- Nagasawa, H.; Kataoka, H.; Isogai, A.; Tamura, S.; Suzuki, A.; Ishizaki, H.; Mizoguchi, A.; Fujiwara, Y.; Suzuki, A. Amino-terminal amino acid sequence of the silkworm prothoracicotropic hormone: Homology with insulin. Science 1984, 226, 1344–1345.
- Siddle, K. Molecular basis of signaling specificity of insulin and IGF receptors: Neglected corners and recent advances. Front. Endocrinol. 2012, 3, 34.
- Boucher, J.; Kleinridders, A.; Kahn, C.R. Insulin receptor signaling in normal and insulin-resistant states. Cold Spring Harb Perspect Biol. 2014, 6, a009191.
- Versteyhe, S.; Blanquart, C.; Hampe, C.; Mahmood, S.; Christeff, N.; Meyts1, P.D.; Gray, S.G.; Issad, T. Insulin receptor substrates-5 and -6 are poor substrates for the insulin receptor. Mol. Med. Rep. 2010, 3, 189–193.
- Schmidt, V.; Schulz, N.; Yan, X.; Schurmann, A.; Kempa, S.; Kern, M.; Bluher, M.; Poy, M.N.; Olivecrona, G.; Willnow, T.E. SORLA facilitates insulin receptor signaling in adipocytes and exacerbates obesity. J. Clin. Investig. 2016, 126, 2706–2720.
- Rask-Madsen, C.; Kahn, C.R. Tissue-specific insulin signaling, metabolic syndrome, and cardiovascular disease. Arterioscler. Thromb. Vasc. Biol. 2012, 32, 2052–2059.
- Shi, X.; Xie, X.; Jia, Y.; Li, S. Associations of insulin receptor and insulin receptor substrates genetic polymorphisms with polycystic ovary syndrome: A systematic review and meta-analysis. J. Obstet. Gynaecol. Res. 2016, 42, 844–854.
- Gralle, M.; Labrecque, S.; Salesse, C.; De Koninck, P. Spatial dynamics of the insulin receptor in living neurons. J. Neurochem. 2020, 156, 88–105.
- Ling, A.V.; Gearing, M.E.; Semova, I.; Shin, D.J.; Clements, R.; Lai, Z.W.; Biddinger, S.B. FoxO1 is required for most of the metabolic and hormonal perturbations produced by hepatic insulin receptor deletion in male mice. Endocrinology 2018, 159, 1253–1263.
- Titone, R.; Robertson, D.M. Insulin receptor preserves mitochondrial function by binding VDAC1 in insulin insensitive mucosal epithelial cells. FASEB J. 2019, 34, 754–775.
- Oakie, A.; Zhou, L.; Rivers, S.; Cheung, C.; Li, J.; Wang, R. Postnatal knockout of beta cell insulin receptor impaired insulin secretion in male mice exposed to high-fat diet stress. Mol. Cell Endocrinol. 2020, 499, 110588.
- Ardon, O.; Procter, M.; Tvrdik, T.; Longo, N.; Mao, R. Sequencing analysis of insulin receptor defects and detection of two novel mutations in INSR gene. Mol. Genet. Metab. Rep. 2014, 1, 71–84.
- Greenhill, C. Insulin and the insulin receptor regulate gene expression. Nat. Rev. Endocrinol. 2019, 15, 315.
- Hancock, M.L.; Meyer, R.C.; Mistry, M.; Khetani, R.S.; Wagschal, A.; Shin, T.; Ho Sui, S.J.; Naar, A.M.; Flanagan, J.G. Insulin receptor associates with promoters genome-wide and regulates gene expression. Cell 2019, 177, 722–736.
- Ott, R.; Melchior, K.; Stupin, J.H.; Ziska, T.; Schellong, K.; Henrich, W.; Rancourt, R.C.; Plagemann, A. Reduced insulin receptor expression and altered DNA methylation in fat tissues and blood of women with GDM and offspring. J. Clin. Endocrinol. Metab. 2019, 104, 137–149.
- Schellong, K.; Melchior, K.; Ziska, T.; Ott, R.; Henrich, W.; Rancourt, R.C.; Plagemann, A. Hypothalamic insulin receptor expression and DNA promoter methylation are sex-specifically altered in adult offspring of high-fat diet (HFD)-overfed mother rats. J. Nutr. Biochem. 2019, 67, 28–35.
- Stolzenbach, F.; Valdivia, S.; Ojeda-Provoste, P.; Toledo, F.; Sobrevia, L.; Kerr, B. DNA methylation changes in genes coding for leptin and insulin receptors during metabolic-altered pregnancies. Biochim. Biophys. Acta Mol. Basis Dis. 2020, 1866, 165465.
- Pan, J.; Di, Y.Q.; Li, Y.B.; Chen, C.H.; Wang, J.X.; Zhao, X.F. Insulin and 20-hydroxyecdysone oppose each other in the regulation of phosphoinositide-dependent kinase-1 expression during insect pupation. J. Biol. Chem. 2018, 293, 18613–18623.
- Liu, Y.; Zhou, S.; Ma, L.; Tian, L.; Wang, S.; Sheng, Z.; Jiang, R.J.; Bendena, W.G.; Li, S. Transcriptional regulation of the insulin signaling pathway genes by starvation and 20-hydroxyecdysone in the Bombyx fat body. J. Insect Physiol. 2010, 56, 1436–1444.
- Li, Y.L.; Yao, Y.X.; Zhao, Y.M.; Di, Y.Q.; Zhao, X.F. The steroid hormone 20-hydroxyecdysone counteracts insulin signaling via insulin receptor dephosphorylation. J. Biol. Chem. 2021, 296, 100318.
- Emlen, D.J.; Warren, I.A.; Johns, A.; Dworkin, I.; Lavine, L.C. A mechanism of extreme growth and reliable signaling in sexually selected ornaments and weapons. Science 2012, 337, 860–864.
- Casasa, S.; Moczek, A.P. Insulin signalling’s role in mediating tissue-specific nutritional plasticity and robustness in the horn-polyphenic beetle Onthophagus taurus. Proc. Biol. Sci. 2018, 285, 20181631.
- Eichler, K.; Li, F.; Litwin-Kumar, A.; Park, Y.; Andrade, I.; Schneider-Mizell, C.M.; Saumweber, T.; Huser, A.; Eschbach, C.; Gerber, B.; et al. The complete connectome of a learning and memory centre in an insect brain. Nature 2017, 548, 175–182.
- Thum, A.S.; Gerber, B. Connectomics and function of a memory network: The mushroom body of larval Drosophila. Curr. Opin. Neurobiol. 2019, 54, 146–154.
- Eschment, M.; Franz, H.R.; Gullu, N.; Holscher, L.G.; Huh, K.E.; Widmann, A. Insulin signaling represents a gating mechanism between different memory phases in Drosophila larvae. PLoS Genet. 2020, 16, e1009064.
- Toprak, U. The role of peptide hormones in insect lipid metabolism. Front. Physiol. 2020, 11, 434.
- Smykal, V.; Pivarci, M.; Provaznik, J.; Bazalova, O.; Jedlicka, P.; Luksan, O.; Horak, A.; Vaneckova, H.; Benes, V.; Fiala, I.; et al. Complex evolution of insect insulin receptors and homologous decoy receptors, and functional significance of their multiplicity. Mol. Biol. Evol. 2020, 37, 1775–1789.
- Barbera, M.; Canas-Canas, R.; Martinez-Torres, D. Insulin-like peptides involved in photoperiodism in the aphid Acyrthosiphon pisum. Insect Biochem. Mol. Biol. 2019, 112, 103185.
- Lin, X.; Yu, N.; Smagghe, G. Insulin receptor regulates food intake through sulfakinin signaling in the red flour beetle, Tribolium castaneum. Peptides 2016, 80, 89–95.
- Sang, M.; Li, C.; Wu, W.; Li, B. Identification and evolution of two insulin receptor genes involved in Tribolium castaneum development and reproduction. Gene 2016, 585, 196–204.
- Nuss, A.B.; Brown, M.R.; Murty, U.S.; Gulia-Nuss, M. Insulin receptor knockdown blocks filarial parasite development and alters egg production in the southern house mosquito, Culex quinquefasciatus. PLoS Negl. Trop. Dis. 2018, 12, e0006413.
- Ihle, K.E.; Mutti, N.S.; Kaftanoglu, O.; Amdam, G.V. Insulin receptor substrate gene knockdown accelerates behavioural maturation and shortens lifespan in honeybee workers. Insects 2019, 10, 390.
- Han, B.; Zhang, T.; Feng, Y.; Liu, X.; Zhang, L.; Chen, H.; Zeng, F.; Wang, M.; Liu, C.; Li, Y.; et al. Two insulin receptors coordinate oogenesis and oviposition via two pathways in the green lacewing, Chrysopa pallens. J. Insect Physiol. 2020, 123, 104049.
More
Information
Subjects:
Chemistry, Medicinal
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
2.0K
Revisions:
2 times
(View History)
Update Date:
01 Aug 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





