
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Bo Sun | -- | 2637 | 2022-07-19 22:48:39 | | | |
| 2 | Peter Tang | Meta information modification | 2637 | 2022-07-20 03:47:36 | | |
Video Upload Options
Vaccines have played a significant role in protecting public and personal health against infectious diseases and proved their great potential in battling cancers as well.
1. Introduction
2. Tumor Antigens for Cancer Vaccines
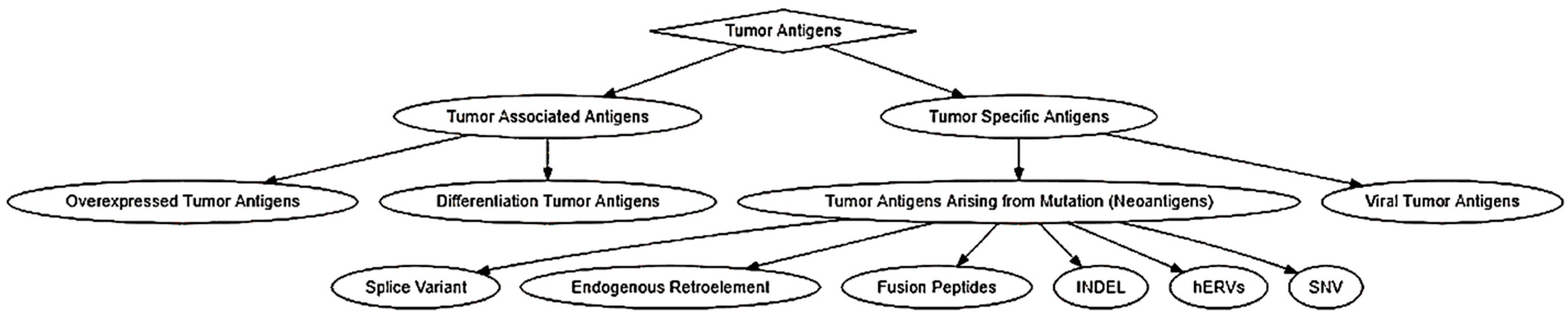
2.1. Classification of Tumor Antigens
|
Antigen |
MHC-I |
MHC-II |
|---|---|---|
|
α-actinin |
HLA-A2 [54] |
|
|
ARTC1 |
HLA-DR1 [55] |
|
|
B-RAF |
HLA-DR4 [56] |
|
|
β-catenin |
HLA-A24 [57] |
|
|
BCR-ABL |
HLA-A2 [58] HLA-A3 [59] HLA-A11 [60] HLA-B8 [58] |
HLA-DR4 [61] HLA-DR9 [62] HLA-DR11 [63] HLA-DR15 [64] HLA-DR1 [65] |
|
Caspase-5 |
HLA-A2 [66] |
|
|
Caspase-8 |
HLA-B35 [67] |
|
|
CDC27 |
HLA-DR4 [68] |
|
|
CDK-4 |
HLA-A2 [69] |
|
|
CDK-12 |
HLA-A11 [70] |
|
|
CDK2NA |
HLA-A11 [71] |
|
|
CLPP |
HLA-A2 [72] |
|
|
COA-1 |
HLA-DR4 [73] HLA-DR13 [73] |
|
|
CSNK1A1 |
HLA-A2 [70] |
|
|
dek-can |
HLA-DR53 [74] |
|
|
EFTUD2 |
HLA-A3 [75] |
|
|
ELF2M |
HLA-A68 [76] |
|
|
ETV6-AML1 |
HLA-A2 [77] |
|
|
FLT3-ITD |
HLA-A1 [78] |
|
|
fibronectin |
HLA-DR15 [79] |
|
|
FNDC3B |
HLA-A2 [80] |
|
|
GAS7 |
HLA-A2 [70] |
|
|
GPNMB |
HLA-A3 [75] |
|
|
HAUS3 |
HLA-A2 [70] |
|
|
HSDL1 |
HLA-Cw14 [81] |
|
|
HSP70-2 |
HLA-A2 [82] |
|
|
KIA A0205 |
HLA-B44 [83] |
|
|
K-ras |
HLA-B35 [84] HLA-Cw8 [85] |
|
|
LDLR-FUT |
HLA-DR1 [86] |
|
|
MART-2 |
HLA-A1 [87] |
|
|
MATN |
HLA-A11 [70] |
|
|
ME1 |
HLA-A2 [88] |
|
|
MUM-1 |
HLA-B44 [89] |
|
|
MUM-2 |
HLA-B44 [90] HLA-Cw6 [90] |
|
|
MUM-3 |
HLA-A68 [91] |
|
|
Myosin-m |
HLA-A3 [92] |
|
|
N-ras |
HLA-A1 [93] |
|
|
neo-PAP |
HLA-DR7 [94] |
|
|
NFYC |
HLA-B52 [95] |
|
|
OGT |
HLA-A2 [96] |
|
|
OS-9 |
HLA-B44 [97] |
|
|
p14ARF |
HLA-A11 [71] |
|
|
p16INK |
HLA-A11 [71] |
|
|
pml-RARalpha fusion protein |
HLA-DR11 [98] |
|
|
PPP1R3B |
HLA-A1 [70] |
|
|
PRDX5 |
HLA-A2 [99] |
|
|
PTPRK |
HLA-DR10 [100] |
|
|
RBAF600 |
HLA-B7 [75] |
|
|
SIRT2 |
HLA-A3 [75] |
|
|
SNDRP1 |
HLA-B38 |
|
|
SYT-SSX1 or -SSX2 fusion protein |
HLA-B7 [101] |
|
|
TGFBRII |
HLA-DR3 [102] |
|
|
TP53 |
HLA-A2 [103] |
|
|
TPI |
HLA-DR1 [104] |
|
|
Annexin II |
HLA-DR4 [105] |
2.2. Biochemical and Bioinformatic Approaches for the Identification of Tumor Antigens
2.2.1. Serological Analysis of Recombinant Tumor cDNA Libraries (SEREX)
2.2.2. Computational Prediction Methods for Cancer Antigens
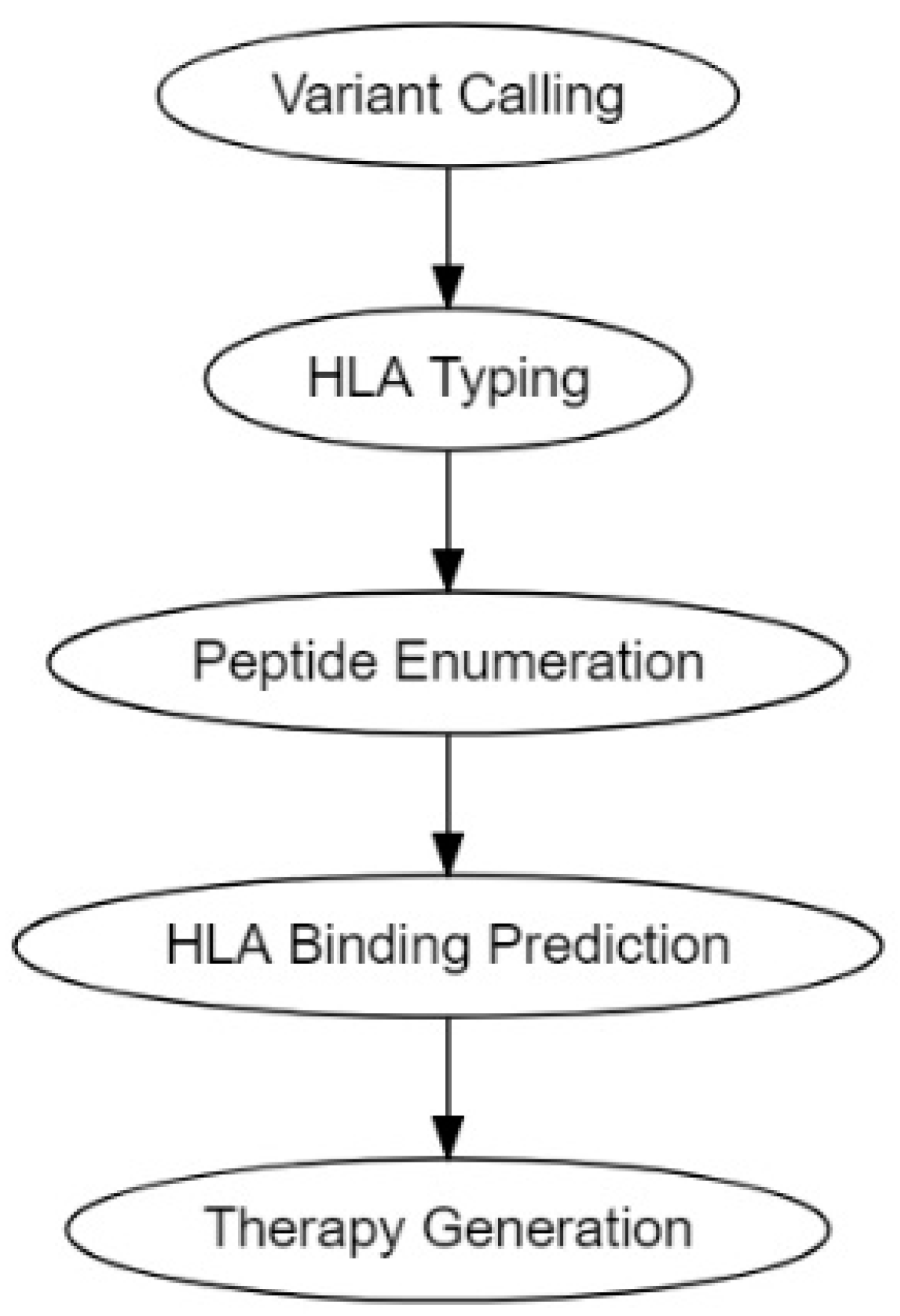
2.3. Delivery of Neoantigens
3. Vaccine Adjuvants
References
- Plotkin, S.A. Vaccines: Past, present and future. Nat. Med. 2005, 11, S5–S11.
- Riedel, S. Edward Jenner and the history of smallpox and vaccination. Proc. (Bayl. Univ. Med. Cent.) 2005, 18, 21–25.
- Tregoning, J.S.; Flight, K.E.; Higham, S.L.; Wang, Z.; Pierce, B.F. Progress of the COVID-19 vaccine effort: Viruses, vaccines and variants versus efficacy, effectiveness and escape. Nat. Rev. Immunol. 2021, 21, 626–636.
- A Phase 1 Study to Evaluate the Safety and Immunogenicity of eOD-GT8 60mer mRNA Vaccine (mRNA-1644) and Core-g28v2 60mer mRNA Vaccine (mRNA-1644v2-Core); 2021. Available online: https://clinicaltrials.gov/ct2/show/results/NCT05001373?view=results (accessed on 30 November 2021).
- Saxena, M.; van der Burg, S.H.; Melief, C.J.M.; Bhardwaj, N. Therapeutic cancer vaccines. Nat. Rev. Cancer 2021, 21, 360–378.
- Prehn, R.T.; Main, J.M. Immunity to Methylcholanthrene-Induced Sarcomas. J. Natl. Cancer Inst. 1957, 18, 769–778.
- van der Bruggen, P.; Traversari, C.; Chomez, P.; Lurquin, C.; De Plaen, E.; Van den Eynde, B.; Knuth, A.; Boon, T. A gene encoding an antigen recognized by cytolytic T lymphocytes on a human melanoma. Science 1991, 254, 1643.
- Murphy, K.; Weaver, C.; Janeway, C. Janeway’s Immunobiology, 9th ed.; Garland Science: New York, NY, USA, 2017.
- Piertney, S.B.; Oliver, M.K. The evolutionary ecology of the major histocompatibility complex. Heredity 2006, 96, 7–21.
- Klein, J.; Figueroa, F. Evolution of the major histocompatibility complex. Crit. Rev. Immunol. 1986, 6, 295–386.
- Altuvia, Y.; Margalit, H. A structure-based approach for prediction of MHC-binding peptides. Methods 2004, 34, 454–459.
- Wieczorek, M.; Abualrous, E.T.; Sticht, J.; Álvaro-Benito, M.; Stolzenberg, S.; Noé, F.; Freund, C. Major Histocompatibility Complex (MHC) Class I and MHC Class II Proteins: Conformational Plasticity in Antigen Presentation. Front. Immunol. 2017, 8, 292.
- Jeffery, K.J.M.; Bangham, C.R.M. Do infectious diseases drive MHC diversity? Microbes Infect. 2000, 2, 1335–1341.
- Dausset, J. The Major Histocompatibility Complex in Man. Science 1981, 213, 1469–1474.
- Bruggen, P.V.D.; Szikora, J.-P.; Boël, P.; Wildmann, C.; Somville, M.; Sensi, M.; Boon, T. Autologous cytolytic T lymphocytes recognize a MAGE-1 nonapeptide on melanomas expressing HLA-Cw* 1601. Eur. J. Immunol. 1994, 24, 2134–2140.
- Robbins, P.F.; Kawakami, Y. Human tumor antigens recognized by T cells. Curr. Opin. Immunol. 1996, 8, 628–636.
- Boon, T.; Cerottini, J.-C.; Van den Eynde, B.; van der Bruggen, P.; Van Pel, A. Tumor Antigens Recognized by T Lymphocytes. Annu. Rev. Immunol. 1994, 12, 337–365.
- Bennett, S.R.M.; Carbone, F.R.; Karamalis, F.; Flavell, R.A.; Miller, J.F.A.P.; Heath, W.R. Help for cytotoxic-T-cell responses is mediated by CD40 signalling. Nature 1998, 393, 478–480.
- Ridge, J.P.; Di Rosa, F.; Matzinger, P. A conditioned dendritic cell can be a temporal bridge between a CD4+ T-helper and a T-killer cell. Nature 1998, 393, 474–478.
- Schoenberger, S.P.; Toes, R.E.M.; van der Voort, E.I.H.; Offringa, R.; Melief, C.J.M. T-cell help for cytotoxic T lymphocytes is mediated by CD40–CD40L interactions. Nature 1998, 393, 480–483.
- Kennedy, R.; Celis, E. Multiple roles for CD4+ T cells in anti-tumor immune responses. Immunol. Rev. 2008, 222, 129–144.
- Pardoll, D.M.; Topalian, S.L. The role of CD4+ T cell responses in antitumor immunity. Curr. Opin. Immunol. 1998, 10, 588–594.
- Valilou, S.F.; Rezaei, N. Chapter 4—Tumor Antigens. In Vaccines for Cancer Immunotherapy; Rezaei, N., Keshavarz-Fathi, M., Eds.; Academic Press: Cambridge, MA, USA, 2019; pp. 61–74.
- Boon, T.; Coulie, P.G.; Van den Eynde, B. Tumor antigens recognized by T cells. Immunol. Today 1997, 18, 267–268.
- Nagorsen, D.; Marincola, F.M. Analyzing T Cell Responses: How to Analyze Cellular Immune Responses against Tumor Associated Antigens; Springer: Berlin/Heidelberg, Germany, 2006.
- Wang, R.-F.; Rosenberg, S.A. Human tumor antigens for cancer vaccine development. Immunol. Rev. 1999, 170, 85–100.
- Richards, J.M.; Mehta, N.; Ramming, K.; Skosey, P. Sequential Chemoimmunotherapy in the Treatment of Metastatic Melanoma. In Proceedings of the Cytokines in Hemopoiesis, Oncology, and AIDS II, Berlin/Heidelberg, Germany, 1 August 1992; pp. 721–727.
- Agyemang, A.F.; Odunsi, K.O. Chapter 5—The use of immunotherapy for treatment of chemoresistant ovarian cancer. In Overcoming Ovarian Cancer Chemoresistance; Samimi, G., Annunziata, C., Eds.; Academic Press: Cambridge, MA, USA, 2021; Volume 11, pp. 79–96.
- Gjerstorff, M.F.; Andersen, M.H.; Ditzel, H.J. Oncogenic cancer/testis antigens: Prime candidates for immunotherapy. Oncotarget 2015, 6, 15772–15787.
- Smith, C.C.; Selitsky, S.R.; Chai, S.; Armistead, P.M.; Vincent, B.G.; Serody, J.S. Alternative tumour-specific antigens. Nat. Rev. Cancer 2019, 19, 465–478.
- Goodier, J.L.; Kazazian, H.H. Retrotransposons Revisited: The Restraint and Rehabilitation of Parasites. Cell 2008, 135, 23–35.
- Cherkasova, E.; Scrivani, C.; Doh, S.; Weisman, Q.; Takahashi, Y.; Harashima, N.; Yokoyama, H.; Srinivasan, R.; Linehan, W.M.; Lerman, M.I.; et al. Detection of an Immunogenic HERV-E Envelope with Selective Expression in Clear Cell Kidney Cancer. Cancer Res. 2016, 76, 2177–2185.
- Turajlic, S.; Litchfield, K.; Xu, H.; Rosenthal, R.; McGranahan, N.; Reading, J.L.; Wong, Y.N.S.; Rowan, A.; Kanu, N.; Al Bakir, M.; et al. Insertion-and-deletion-derived tumour-specific neoantigens and the immunogenic phenotype: A pan-cancer analysis. Lancet Oncol. 2017, 18, 1009–1021.
- Jayasinghe, R.G.; Cao, S.; Gao, Q.; Wendl, M.C.; Vo, N.S.; Reynolds, S.M.; Zhao, Y.; Climente-González, H.; Chai, S.; Wang, F.; et al. Systematic Analysis of Splice-Site-Creating Mutations in Cancer. Cell Rep. 2018, 23, 270–281.e273.
- Rowley, J.D. A New Consistent Chromosomal Abnormality in Chronic Myelogenous Leukaemia identified by Quinacrine Fluorescence and Giemsa Staining. Nature 1973, 243, 290–293.
- Nowell, P. The minute chromosome (Phl) in chronic granulocytic leukemia. Blut 1962, 8, 65–66.
- Shepherd, P.; Suffolk, R.; Halsey, J.; Allan, N. Analysis of molecular breakpoint and m-RNA transcripts in a prospective randomized trial of interferon in chronic myeloid leukaemia: No correlation with clinical features, cytogenetic response, duration of chronic phase, or survival. Br. J. Haematol. 1995, 89, 546–554.
- Westbrook, C.A.; Hooberman, A.L.; Spino, C.; Dodge, R.K.; Larson, R.A.; Davey, F.; Wurster-Hill, D.H.; Sobol, R.E.; Schiffer, C.; Bloomfield, C.D. Clinical Significance of the BCR-ABL Fusion Gene in Adult Acute Lymphoblastic Leukemia: A Cancer and Leukemia Group B Study (8762). Blood 1992, 80, 2983–2990.
- Russo, C.; Carroll, A.; Kohler, S.; Borowitz, M.; Amylon, M.; Homans, A.; Kedar, A.; Shuster, J.; Land, V.; Crist, W.; et al. Philadelphia Chromosome and Monosomy 7 in Childhood Acute Lymphoblastic Leukemia: A Pediatric Oncology Group Study. Blood 1991, 77, 1050–1056.
- Suryanarayan, K.; Hunger, S.P.; Kohler, S.; Carroll, A.J.; Crist, W.; Link, M.P.; Cleary, M.L. Consistent Involvement of the BCR Gene by 9;22 Breakpoints in Pediatric Acute Leukemias. Blood 1991, 77, 324–330.
- Kurzrock, R.; Shtalrid, M.; Talpaz, M.; Kloetzer, W.S.; Gutterman, J.U. Expression of c-abl in Philadelphia-Positive Acute Myelogenous Leukemia. Blood 1987, 70, 1584–1588.
- Secker-Walker, L.M.; Morgan, G.J.; Min, T.; John Swansbury, G.; Craig, J.; Yamada, T.; Desalvo, L.; Medina, J.W.; Chowdhury, V.; Donahue, R.P.; et al. Inversion of chromosome 16 with the Philadelphia chromosome in acute myelomonocytic leukemia with eosinophilia: Report of two cases. Cancer Genet. Cytogenet. 1992, 58, 29–34.
- Alimena, G.; Cedrone, M.; Nanni, M.; De Cuia, M.; De Sanctis, V.; Cimino, G.; Mancini, M. Acute leukemia presenting a variant Ph chromosome with p190 expression, dup 3q and-7, developed after malignant lymphoma treated with alkylating agents and topoisomerase II inhibitors. Leukemia 1995, 9, 1483–1486.
- Fujii, H.; Yashige, H.; Misawa, S.; Tanaka, S.; Urata, Y.; Matuyama, F. Ph chromosome in a patient with non-leukemic non-hodgkin B-cell lymphoma. Am. J. Hematol. 1990, 35, 213–215.
- Mitani, K.; Sato, Y.; Tojo, A.; Ishikawa, F.; Kobayashi, Y.; Miura, Y.; Miyazono, K.; And, A.U.; Takaku, F. Philadelphia chromosome positive B-cell type malignant lymphoma expressing an aberrant 190 kDa bcr-abl protein. Br. J. Haematol. 1990, 76, 221–225.
- Juliusson, G.; Friberg, K.; Gahrton, G. Ph chromosome and B-cell malignancy-associated chromosomal aberrations in non-leukaemic immunoblastic B-cell lymphoma. Acta Haematol. 1985, 74, 171–174.
- Martiat, P.; Mecucci, C.; Nizet, Y.; Stul, M.; Philippe, M.; Cassiman, J.-J.; Michaux, J.L.; Van den Berghe, H.; Sokal, G. P190 BCR/ABL transcript in a case of Philadelphia-positive multiple myeloma. Leukemia 1990, 4, 751–754.
- Van Den Berghe, H.a.; Louwagie, A.; Broeckaert-Van Orshoven, A.; David, G.; Verwilghen, R.; Michaux, J.; Sokal, G. Philadelphia chromosome in human multiple myeloma. J. Natl. Cancer Inst. 1979, 63, 11–16.
- Karpas, A.; Fischer, P.; Swirsky, D. Human myeloma cell line carrying a Philadelphia chromosome. Science 1982, 216, 997–999.
- Bos, J.L. Ras Oncogenes in Human Cancer: A Review. Cancer Res. 1989, 49, 4682–4689.
- Bourgault Villada, I.; Bénéton, N.; Bony, C.; Connan, F.; Monsonego, J.; Bianchi, A.; Saiag, P.; Lévy, J.P.; Guillet, J.G.; Choppin, J. Identification in humans of HPV-16 E6 and E7 protein epitopes recognized by cytolytic T lymphocytes in association with HLA-B18 and determination of the HLA-B18-specific binding motif. Eur. J. Immunol. 2000, 30, 2281–2289.
- Peng, S.; Trimble, C.; Wu, L.; Pardoll, D.; Roden, R.; Hung, C.-F.; Wu, T.C. HLA-DQB1*02–Restricted HPV-16 E7 Peptide–Specific CD4+ T-Cell Immune Responses Correlate with Regression of HPV-16–Associated High-Grade Squamous Intraepithelial Lesions. Clin. Cancer Res. 2007, 13, 2479.
- Kaufmann, A.M.; Nieland, J.; Schinz, M.; Nonn, M.; Gabelsberger, J.; Meissner, H.; Müller, R.T.; Jochmus, I.; Gissmann, L.; Schneider, A.; et al. HPV16 L1E7 chimeric virus-like particles induce specific HLA-restricted T cells in humans after in vitro vaccination. Int. J. Cancer 2001, 92, 285–293.
- Echchakir, H.; Mami-Chouaib, F.; Vergnon, I.; Baurain, J.-F.; Karanikas, V.; Chouaib, S.; Coulie, P.G. A Point Mutation in the α-actinin-4 Gene Generates an Antigenic Peptide Recognized by Autologous Cytolytic T Lymphocytes on a Human Lung Carcinoma. Cancer Res. 2001, 61, 4078.
- Wang, H.Y.; Peng, G.; Guo, Z.; Shevach, E.M.; Wang, R.-F. Recognition of a New ARTC1 Peptide Ligand Uniquely Expressed in Tumor Cells by Antigen-Specific CD4+ Regulatory T Cells. J. Immunol. 2005, 174, 2661.
- Sharkey, M.S.; Lizée, G.; Gonzales, M.I.; Patel, S.; Topalian, S.L. CD4+ T-Cell Recognition of Mutated B-RAF in Melanoma Patients Harboring the V599E Mutation. Cancer Res. 2004, 64, 1595.
- Robbins, P.F.; El-Gamil, M.; Li, Y.F.; Kawakami, Y.; Loftus, D.; Appella, E.; Rosenberg, S.A. A mutated beta-catenin gene encodes a melanoma-specific antigen recognized by tumor infiltrating lymphocytes. J. Exp. Med. 1996, 183, 1185–1192.
- Yotnda, P.; Firat, H.; Garcia-Pons, F.; Garcia, Z.; Gourru, G.; Vernant, J.P.; Lemonnier, F.A.; Leblond, V.; Langlade-Demoyen, P. Cytotoxic T cell response against the chimeric p210 BCR-ABL protein in patients with chronic myelogenous leukemia. J. Clin. Investig. 1998, 101, 2290–2296.
- Greco, G.; Fruci, D.; Accapezzato, D.; Barnaba, V.; Nisini, R.; Alimena, G.; Montefusco, E.; Vigneti, E.; Butler, R.; Tanigaki, N.; et al. Two bcr-abl junction peptides bind HLA-A3 molecules and allow specific induction of human cytotoxic T lymphocytes. Leukemia 1996, 10, 693–699.
- Bocchia, M.; Korontsvit, T.; Xu, Q.; Mackinnon, S.; Yang, S.; Sette, A.; Scheinberg, D. Specific human cellular immunity to bcr-abl oncogene-derived peptides. Blood 1996, 87, 3587–3592.
- Bosch, G.; Joosten, A.; Kessler, J.; Melief, C.; Leeksma, O. Recognition of BCR-ABL positive leukemic blasts by human CD4+ T cells elicited by primary in vitro immunization with a BCR-ABL breakpoint peptide. Blood 1996, 88, 3522–3527.
- Yasukawa, M.; Ohminami, H.; Kaneko, S.; Yakushijin, Y.; Nishimura, Y.; Inokuchi, K.; Miyakuni, T.; Nakao, S.; Kishi, K.; Kubonishi, I.; et al. CD4+ Cytotoxic T-Cell Clones Specific for bcr-abl b3a2 Fusion Peptide Augment Colony Formation by Chronic Myelogenous Leukemia Cells in a b3a2-Specific and HLA-DR–Restricted Manner. Blood 1998, 92, 3355–3361.
- Pawelec, G.; Max, H.; Halder, T.; Bruserud, O.; Merl, A.; da Silva, P.; Kalbacher, H. BCR/ABL leukemia oncogene fusion peptides selectively bind to certain HLA-DR alleles and can be recognized by T cells found at low frequency in the repertoire of normal donors. Blood 1996, 88, 2118–2124.
- Tanaka, Y.; Takahashi, T.; Nieda, M.; Masuda, S.; Kashiwase, K.; Ogawa, S.; Chiba, S.; Juji, T.; Hirai, H. Generation of HLA-DRB1*1501-restricted p190 minor bcr–abl (e1a2)-specific CD4+ T lymphocytes. Br. J. Haematol. 2000, 109, 435–437.
- ten Bosch, G.J.A.; Kessler, J.H.; Joosten, A.M.; Bres-Vloemans, A.A.; Geluk, A.; Godthelp, B.C.; van Bergen, J.; Melief, C.J.M.; Leeksma, O.C. A BCR-ABL Oncoprotein p210b2a2 Fusion Region Sequence Is Recognized by HLA-DR2a Restricted Cytotoxic T Lymphocytes and Presented by HLA-DR Matched Cells Transfected With an Iib2a2 Construct. Blood 1999, 94, 1038–1045.
- Schwitalle, Y.; Linnebacher, M.; Ripberger, E.; Gebert, J.; von Knebel Doeberitz, M. Immunogenic peptides generated by frameshift mutations in DNA mismatch repair-deficient cancer cells. Cancer Immun. Arch. 2004, 4, 14.
- Mandruzzato, S.; Brasseur, F.; Andry, G.; Boon, T.; van der Bruggen, P. A CASP-8 mutation recognized by cytolytic T lymphocytes on a human head and neck carcinoma. J. Exp. Med. 1997, 186, 785–793.
- Wang, R.-F.; Wang, X.; Atwood, A.C.; Topalian, S.L.; Rosenberg, S.A. Cloning Genes Encoding MHC Class II-Restricted Antigens: Mutated CDC27 as a Tumor Antigen. Science 1999, 284, 1351–1354.
- Wolfel, T.; Hauer, M.; Schneider, J.; Serrano, M.; Wolfel, C.; Klehmann-Hieb, E.; De Plaen, E.; Hankeln, T.; Meyer zum Buschenfelde, K.; Beach, D. A p16INK4a-insensitive CDK4 mutant targeted by cytolytic T lymphocytes in a human melanoma. Science 1995, 269, 1281–1284.
- Robbins, P.F.; Lu, Y.-C.; El-Gamil, M.; Li, Y.F.; Gross, C.; Gartner, J.; Lin, J.C.; Teer, J.K.; Cliften, P.; Tycksen, E.; et al. Mining exomic sequencing data to identify mutated antigens recognized by adoptively transferred tumor-reactive T cells. Nat. Med. 2013, 19, 747–752.
- Huang, J.; El-Gamil, M.; Dudley, M.E.; Li, Y.F.; Rosenberg, S.A.; Robbins, P.F. T cells associated with tumor regression recognize frameshifted products of the CDKN2A tumor suppressor gene locus and a mutated HLA class I gene product. J. Immunol. 2004, 172, 6057–6064.
- Corbière, V.; Chapiro, J.; Stroobant, V.; Ma, W.; Lurquin, C.; Lethé, B.; van Baren, N.; Van den Eynde, B.J.; Boon, T.; Coulie, P.G. Antigen Spreading Contributes to MAGE Vaccination-Induced Regression of Melanoma Metastases. Cancer Res. 2011, 71, 1253.
- Maccalli, C.; Pende, D.; Castelli, C.; Mingari, M.C.; Robbins, P.F.; Parmiani, G. NKG2D engagement of colorectal cancer-specific T cells strengthens TCR-mediated antigen stimulation and elicits TCR independent anti-tumor activity. Eur. J. Immunol. 2003, 33, 2033–2043.
- Makita, M.; Azuma, T.; Hamaguchi, H.; Niiya, H.; Kojima, K.; Fujita, S.; Tanimoto, M.; Harada, M.; Yasukawa, M. Leukemia-associated fusion proteins, dek-can and bcr-abl, represent immunogenic HLA-DR-restricted epitopes recognized by fusion peptide-specific CD4+ T lymphocytes. Leukemia 2002, 16, 2400–2407.
- Lennerz, V.; Fatho, M.; Gentilini, C.; Frye, R.A.; Lifke, A.; Ferel, D.; Wölfel, C.; Huber, C.; Wölfel, T. The response of autologous T cells to a human melanoma is dominated by mutated neoantigens. Proc. Natl. Acad. Sci. USA 2005, 102, 16013–16018.
- Hogan, K.T.; Eisinger, D.P.; Cupp, S.B.; Lekstrom, K.J.; Deacon, D.D.; Shabanowitz, J.; Hunt, D.F.; Engelhard, V.H.; Slingluff, C.L.; Ross, M.M. The Peptide Recognized by HLA-A68.2-restricted, Squamous Cell Carcinoma of the Lung-specific Cytotoxic T Lymphocytes Is Derived from a Mutated Elongation Factor 2 Gene. Cancer Res. 1998, 58, 5144.
- Yotnda, P.; Garcia, F.; Peuchmaur, M.; Grandchamp, B.; Duval, M.; Lemonnier, F.; Vilmer, E.; Langlade-Demoyen, P. Cytotoxic T cell response against the chimeric ETV6-AML1 protein in childhood acute lymphoblastic leukemia. J. Clin. Investig. 1998, 102, 455–462.
- Graf, C.; Heidel, F.; Tenzer, S.; Radsak, M.P.; Solem, F.K.; Britten, C.M.; Huber, C.; Fischer, T.; Wölfel, T. A neoepitope generated by an FLT3 internal tandem duplication (FLT3-ITD) is recognized by leukemia-reactive autologous CD8+ T cells. Blood 2006, 109, 2985–2988.
- Wang, H.Y.; Zhou, J.; Zhu, K.; Riker, A.I.; Marincola, F.M.; Wang, R.-F. Identification of a mutated fibronectin as a tumor antigen recognized by CD4+ T cells: Its role in extracellular matrix formation and tumor metastasis. J. Exp. Med. 2002, 195, 1397–1406.
- Rajasagi, M.; Shukla, S.A.; Fritsch, E.F.; Keskin, D.B.; DeLuca, D.; Carmona, E.; Zhang, W.; Sougnez, C.; Cibulskis, K.; Sidney, J.; et al. Systematic identification of personal tumor-specific neoantigens in chronic lymphocytic leukemia. Blood 2014, 124, 453–462.
- Wick, D.A.; Webb, J.R.; Nielsen, J.S.; Martin, S.D.; Kroeger, D.R.; Milne, K.; Castellarin, M.; Twumasi-Boateng, K.; Watson, P.H.; Holt, R.A.; et al. Surveillance of the Tumor Mutanome by T Cells during Progression from Primary to Recurrent Ovarian Cancer. Clin. Cancer Res. 2014, 20, 1125.
- Gaudin, C.; Kremer, F.; Angevin, E.; Scott, V.; Triebel, F. A hsp70-2 Mutation Recognized by CTL on a Human Renal Cell Carcinoma. J. Immunol. 1999, 162, 1730.
- Guéguen, M.; Patard, J.-J.; Gaugler, B.; Brasseur, F.; Renauld, J.-C.; Van Cangh, P.J.; Boon, T.; Van den Eynde, B.t.J. An Antigen Recognized by Autologous CTLs on a Human Bladder Carcinoma. J. Immunol. 1998, 160, 6188.
- Gjertsen, M.K.; Bjørheim, J.; Saeterdal, I.; Myklebust, J.; Gaudernack, G. Cytotoxic CD4+ and CD8+ T lymphocytes, generated by mutant p21-ras (12VAL) peptide vaccination of a patient, recognize 12VAL-dependent nested epitopes present within the vaccine peptide and kill autologous tumour cells carrying this mutation. Int. J. Cancer 1997, 72, 784–790.
- Tran, E.; Robbins, P.F.; Lu, Y.-C.; Prickett, T.D.; Gartner, J.J.; Jia, L.; Pasetto, A.; Zheng, Z.; Ray, S.; Groh, E.M.; et al. T-Cell Transfer Therapy Targeting Mutant KRAS in Cancer. N. Engl. J. Med. 2016, 375, 2255–2262.
- Wang, R.F.; Wang, X.; Rosenberg, S.A. Identification of a novel major histocompatibility complex class II-restricted tumor antigen resulting from a chromosomal rearrangement recognized by CD4(+) T cells. J. Exp. Med. 1999, 189, 1659–1668.
- Kawakami, Y.; Wang, X.; Shofuda, T.; Sumimoto, H.; Tupesis, J.P.; Fitzgerald, E.; Rosenberg, S.A. Isolation of a New Melanoma Antigen, MART-2, Containing a Mutated Epitope Recognized by Autologous Tumor-Infiltrating T Lymphocytes. J. Immunol. 2001, 166, 2871.
- Karanikas, V.; Colau, D.; Baurain, J.-F.; Chiari, R.; Thonnard, J.; Gutierrez-Roelens, I.; Goffinet, C.; Schaftingen, E.V.; Weynants, P.; Boon, T.; et al. High Frequency of Cytolytic T Lymphocytes Directed against a Tumor-specific Mutated Antigen Detectable with HLA Tetramers in the Blood of a Lung Carcinoma Patient with Long Survival. Cancer Res. 2001, 61, 3718.
- Coulie, P.G.; Lehmann, F.; Lethé, B.; Herman, J.; Lurquin, C.; Andrawiss, M.; Boon, T. A mutated intron sequence codes for an antigenic peptide recognized by cytolytic T lymphocytes on a human melanoma. Proc. Natl. Acad. Sci. USA 1995, 92, 7976–7980.
- Chiari, R.; Foury, F.; De Plaen, E.; Baurain, J.-F.; Thonnard, J.; Coulie, P.G. Two Antigens Recognized by Autologous Cytolytic T Lymphocytes on a Melanoma Result from a Single Point Mutation in an Essential Housekeeping Gene. Cancer Res. 1999, 59, 5785.
- Baurain, J.-F.; Colau, D.; van Baren, N.; Landry, C.; Martelange, V.; Vikkula, M.; Boon, T.; Coulie, P.G. High Frequency of Autologous Anti-Melanoma CTL Directed Against an Antigen Generated by a Point Mutation in a New Helicase Gene. J. Immunol. 2000, 164, 6057.
- Zorn, E.; Hercend, T. A natural cytotoxic T cell response in a spontaneously regressing human melanoma targets a neoantigen resulting from a somatic point mutation. Eur. J. Immunol. 1999, 29, 592–601.
- Linard, B.; Bézieau, S.; Benlalam, H.; Labarrière, N.; Guilloux, Y.; Diez, E.; Jotereau, F. A ras-Mutated Peptide Targeted by CTL Infiltrating a Human Melanoma Lesion. J. Immunol. 2002, 168, 4802.
- Topalian, S.L.; Gonzales, M.I.; Ward, Y.; Wang, X.; Wang, R.-F. Revelation of a Cryptic Major Histocompatibility Complex Class II-restricted Tumor Epitope in a Novel RNA-processing Enzyme. Cancer Res. 2002, 62, 5505.
- Takenoyama, M.; Baurain, J.-F.; Yasuda, M.; So, T.; Sugaya, M.; Hanagiri, T.; Sugio, K.; Yasumoto, K.; Boon, T.; Coulie, P.G. A point mutation in the NFYC gene generates an antigenic peptide recognized by autologous cytolytic T lymphocytes on a human squamous cell lung carcinoma. Int. J. Cancer 2006, 118, 1992–1997.
- Ripberger, E.; Linnebacher, M.; Schwitalle, Y.; Gebert, J.; Doeberitz, M.V.K. Identification of an HLA-A0201-Restricted CTL Epitope Generated by a Tumor-Specific Frameshift Mutation in a Coding Microsatellite of the OGT Gene. J. Clin. Immunol. 2003, 23, 415–423.
- Vigneron, N.; Ooms, A.; Morel, S.; Degiovanni, G.; Van Den Eynde, B.J. Identification of a new peptide recognized by autologous cytolytic T lymphocytes on a human melanoma. Cancer Immun. 2002, 2, 9.
- Gambacorti-Passerini, C.; Grignani, F.; Arienti, F.; Pandolfi, P.P.; Pelicci, P.G.; Parmiani, G. Human CD4 lymphocytes specifically recognize a peptide representing the fusion region of the hybrid protein pml/RAR alpha present in acute promyelocytic leukemia cells. Blood 1993, 81, 1369–1375.
- Sensi, M.; Nicolini, G.; Zanon, M.; Colombo, C.; Molla, A.; Bersani, I.; Lupetti, R.; Parmiani, G.; Anichini, A. Immunogenicity without Immunoselection: A Mutant but Functional Antioxidant Enzyme Retained in a Human Metastatic Melanoma and Targeted by CD8+ T Cells with a Memory Phenotype. Cancer Res. 2005, 65, 632.
- Novellino, L.; Renkvist, N.; Rini, F.; Mazzocchi, A.; Rivoltini, L.; Greco, A.; Deho, P.; Squarcina, P.; Robbins, P.F.; Parmiani, G.; et al. Identification of a Mutated Receptor-Like Protein Tyrosine Phosphatase κ as a Novel, Class II HLA-Restricted Melanoma Antigen. J. Immunol. 2003, 170, 6363–6370.
- Worley, B.S.; van den Broeke, L.T.; Goletz, T.J.; Pendleton, C.D.; Daschbach, E.M.; Thomas, E.K.; Marincola, F.M.; Helman, L.J.; Berzofsky, J.A. Antigenicity of Fusion Proteins from Sarcoma-associated Chromosomal Translocations. Cancer Res. 2001, 61, 6868.
- Sæterdal, I.; Bjørheim, J.; Lislerud, K.; Gjertsen, M.K.; Bukholm, I.K.; Olsen, O.C.; Nesland, J.M.; Eriksen, J.A.; Møller, M.; Lindblom, A.; et al. Frameshift-mutation-derived peptides as tumor-specific antigens in inherited and spontaneous colorectal cancer. Proc. Natl. Acad. Sci. USA 2001, 98, 13255–13260.
- Malekzadeh, P.; Pasetto, A.; Robbins, P.F.; Parkhurst, M.R.; Paria, B.C.; Jia, L.; Gartner, J.J.; Hill, V.; Yu, Z.; Restifo, N.P.; et al. Neoantigen screening identifies broad TP53 mutant immunogenicity in patients with epithelial cancers. J. Clin. Investig. 2019, 129, 1109–1114.
- Pieper, R.; Christian, R.E.; Gonzales, M.I.; Nishimura, M.I.; Gupta, G.; Settlage, R.E.; Shabanowitz, J.; Rosenberg, S.A.; Hunt, D.F.; Topalian, S.L. Biochemical identification of a mutated human melanoma antigen recognized by CD4(+) T cells. J. Exp. Med. 1999, 189, 757–766.
- Li, K.; Adibzadeh, M.; Halder, T.; Kalbacher, H.; Heinzel, S.; Müller, C.; Zeuthen, J.; Pawelec, G. Tumour-specific MHC-class-II-restricted responses after in vitro sensitization to synthetic peptides corresponding to gp100 and Annexin II eluted from melanoma cells. Cancer Immunol. Immunother. 1998, 47, 32–38.
- Sahin, U.; Türeci, O.; Schmitt, H.; Cochlovius, B.; Johannes, T.; Schmits, R.; Stenner, F.; Luo, G.; Schobert, I.; Pfreundschuh, M. Human neoplasms elicit multiple specific immune responses in the autologous host. Proc. Natl. Acad. Sci. USA 1995, 92, 11810–11813.
- Sahin, U.; Pfreundschuh, M. Serological analysis of human tumor antigens: Molecular definition and implications. Mol. Med. Today 1997, 3, 342–349.
- Chen, Y.-T.; Gure, A.O.; Scanlan, M.J. Serological Analysis of Expression cDNA Libraries (SEREX). In Pancreatic Cancer; Springer: Berlin/Heidelberg, Germany, 2005; pp. 207–216.
- Türeci, Ö.; Usener, D.; Schneider, S.; Sahin, U. Identification of tumor-associated autoantigens with SEREX. In Adoptive Immunotherapy: Methods and Protocols; Springer: Berlin/Heidelberg, Germany, 2005; pp. 137–154.
- Rauch, J.; Gires, O. SEREX, Proteomex, AMIDA, and beyond: Serological screening technologies for target identification. PROTEOMICS—Clin. Appl. 2008, 2, 355–371.
- Whittemore, K.; Sykes, K. A microarray method for identifying tumor antigens by screening a tumor cDNA expression library against cancer sera. Hum. Vaccines Immunother. 2013, 9, 2178–2188.
- Boncheva, V.; Bonney, S.A.; Brooks, S.E.; Tangney, M.; O’Sullivan, G.; Mirnezami, A.; Guinn, B.A. New targets for the immunotherapy of colon cancer—does reactive disease hold the answer? Cancer Gene Ther. 2013, 20, 157–168.
- Roudko, V.; Greenbaum, B.; Bhardwaj, N. Computational Prediction and Validation of Tumor-Associated Neoantigens. Front. Immunol. 2020, 11, 27.
- Koboldt, D.C. Best practices for variant calling in clinical sequencing. Genome Med. 2020, 12, 91.
- Orenbuch, R.; Filip, I.; Rabadan, R. HLA Typing from RNA Sequencing and Applications to Cancer. In Bioinformatics for Cancer Immunotherapy: Methods and Protocols; Boegel, S., Ed.; Springer: New York, NY, USA, 2020; pp. 71–92.
- Díaz-Eufracio, B.I.; Palomino-Hernández, O.; Arredondo-Sánchez, A.; Medina-Franco, J.L. D-Peptide Builder: A Web Service to Enumerate, Analyze, and Visualize the Chemical Space of Combinatorial Peptide Libraries. Mol. Inform. 2020, 39, 2000035.
- Ott, P.A.; Hu, Z.; Keskin, D.B.; Shukla, S.A.; Sun, J.; Bozym, D.J.; Zhang, W.; Luoma, A.; Giobbie-Hurder, A.; Peter, L.; et al. An immunogenic personal neoantigen vaccine for patients with melanoma. Nature 2017, 547, 217–221.
- Bjerregaard, A.-M.; Nielsen, M.; Hadrup, S.R.; Szallasi, Z.; Eklund, A.C. MuPeXI: Prediction of neo-epitopes from tumor sequencing data. Cancer Immunol. Immunother. 2017, 66, 1123–1130.
- Chang, T.-C.; Carter, R.A.; Li, Y.; Li, Y.; Wang, H.; Edmonson, M.N.; Chen, X.; Arnold, P.; Geiger, T.L.; Wu, G.; et al. The neoepitope landscape in pediatric cancers. Genome Med. 2017, 9, 78.
- Hundal, J.; Carreno, B.M.; Petti, A.A.; Linette, G.P.; Griffith, O.L.; Mardis, E.R.; Griffith, M. pVAC-Seq: A genome-guided in silico approach to identifying tumor neoantigens. Genome Med. 2016, 8, 11.
- Kim, S.; Kim, H.S.; Kim, E.; Lee, M.G.; Shin, E.C.; Paik, S.; Kim, S. Neopepsee: Accurate genome-level prediction of neoantigens by harnessing sequence and amino acid immunogenicity information. Ann. Oncol. 2018, 29, 1030–1036.
- Rech, A.J.; Balli, D.; Mantero, A.; Ishwaran, H.; Nathanson, K.L.; Stanger, B.Z.; Vonderheide, R.H. Tumor Immunity and Survival as a Function of Alternative Neopeptides in Human Cancer. Cancer Immunol. Res. 2018, 6, 276–287.
- Rubinsteyn, A.; Kodysh, J.; Hodes, I.; Mondet, S.; Aksoy, B.A.; Finnigan, J.P.; Bhardwaj, N.; Hammerbacher, J. Computational Pipeline for the PGV-001 Neoantigen Vaccine Trial. Front. Immunol. 2018, 8, 1807.
- Zhang, J.; Mardis, E.R.; Maher, C.A. INTEGRATE-neo: A pipeline for personalized gene fusion neoantigen discovery. Bioinformatics 2017, 33, 555–557.
- Zhou, Z.; Lyu, X.; Wu, J.; Yang, X.; Wu, S.; Zhou, J.; Gu, X.; Su, Z.; Chen, S. TSNAD: An integrated software for cancer somatic mutation and tumour-specific neoantigen detection. R. Soc. Open Sci. 2017, 4, 170050.
- Smit, A.; Hubley, R.; Green, P. RepeatMasker Open-4.0. Available online: http://www.repeatmasker.org (accessed on 30 November 2021).
- Smith, C.C.; Beckermann, K.E.; Bortone, D.S.; De Cubas, A.A.; Bixby, L.M.; Lee, S.J.; Panda, A.; Ganesan, S.; Bhanot, G.; Wallen, E.M.; et al. Endogenous retroviral signatures predict immunotherapy response in clear cell renal cell carcinoma. J. Clin. Investig. 2018, 128, 4804–4820.
- Lee, S.-H.; Danishmalik, S.N.; Sin, J.-I. DNA vaccines, electroporation and their applications in cancer treatment. Hum. Vaccines Immunother. 2015, 11, 1889–1900.
- Zhao, Z.; Zheng, L.; Chen, W.; Weng, W.; Song, J.; Ji, J. Delivery strategies of cancer immunotherapy: Recent advances and future perspectives. J. Hematol. Oncol. 2019, 12, 126.
- Pardi, N.; Hogan, M.J.; Porter, F.W.; Weissman, D. mRNA vaccines—A new era in vaccinology. Nat. Rev. Drug Discov. 2018, 17, 261–279.
- Guo, Y.; Lei, K.; Tang, L. Neoantigen Vaccine Delivery for Personalized Anticancer Immunotherapy. Front. Immunol. 2018, 9, 1499.
- Um, W.; Gupta, A.; Song, S.H.; Kim, C.H.; Park, J.H. Biomaterials as Antigen Delivery Carrier for Cancer Immunotherapy. Macromol. Res. 2021, 29, 834–842.
- Cox, J.C.; Coulter, A.R. Adjuvants—A classification and review of their modes of action. Vaccine 1997, 15, 248–256.
- Banzhoff, A.; Gasparini, R.; Laghi-Pasini, F.; Staniscia, T.; Durando, P.; Montomoli, E.; Capecchi, P.L.; di Giovanni, P.; Sticchi, L.; Gentile, C.; et al. MF59-adjuvanted H5N1 vaccine induces immunologic memory and heterotypic antibody responses in non-elderly and elderly adults. PLoS ONE 2009, 4, e4384.
- Coffman, R.L.; Sher, A.; Seder, R.A. Vaccine Adjuvants: Putting Innate Immunity to Work. Immunity 2010, 33, 492–503.
- Didierlaurent, A.M.; Morel, S.; Lockman, L.; Giannini, S.L.; Bisteau, M.; Carlsen, H.; Kielland, A.; Vosters, O.; Vanderheyde, N.; Schiavetti, F.; et al. AS04, an Aluminum Salt- and TLR4 Agonist-Based Adjuvant System, Induces a Transient Localized Innate Immune Response Leading to Enhanced Adaptive Immunity. J. Immunol. 2009, 183, 6186–6197.
- Garçon, N.; Di Pasquale, A. From discovery to licensure, the Adjuvant System story. Hum. Vaccines Immunother. 2016, 13, 19–33.
- Ammi, R.; De Waele, J.; Willemen, Y.; Van Brussel, I.; Schrijvers, D.M.; Lion, E.; Smits, E.L.J. Poly(I:C) as cancer vaccine adjuvant: Knocking on the door of medical breakthroughs. Pharmacol. Ther. 2015, 146, 120–131.
- Apostólico, J.d.S.; Lunardelli, V.A.S.; Coirada, F.C.; Boscardin, S.B.; Rosa, D.S. Adjuvants: Classification, Modus Operandi, and Licensing. J. Immunol. Res. 2016, 2016, 1459394.
- Bowen, W.S.; Svrivastava, A.K.; Batra, L.; Barsoumian, H.; Shirwan, H. Current challenges for cancer vaccine adjuvant development. Expert Rev. Vaccines 2018, 17, 207–215.
- Vermaelen, K. Vaccine Strategies to Improve Anti-cancer Cellular Immune Responses. Front. Immunol. 2019, 10, 8.
- Buonsanti, C.; D’Oro, U. Chapter 5—Discovery of Immune Potentiators as Vaccine Adjuvants. In Immunopotentiators in Modern Vaccines, 2nd ed.; Schijns, V.E.J.C., O’Hagan, D.T., Eds.; Academic Press: Cambridge, MA, USA, 2017; pp. 85–104.
- Olive, C. Pattern recognition receptors: Sentinels in innate immunity and targets of new vaccine adjuvants. Expert Rev. Vaccines 2012, 11, 237–256.





