Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Donatella Coradduzza | -- | 1103 | 2022-06-29 09:48:34 | | | |
| 2 | Amina Yu | -46 word(s) | 1057 | 2022-06-30 04:11:16 | | | | |
| 3 | Amina Yu | -1 word(s) | 1056 | 2022-07-01 04:22:28 | | | | |
| 4 | Amina Yu | -3 word(s) | 1053 | 2022-07-01 04:23:41 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Coradduzza, D.; Bellu, E.; Congiargiu, A.; Pashchenko, A.; Amler, E.; Necas, A.; Carru, C.; Medici, S.; Maioli, M. MiRNA Detection and Biosensing. Encyclopedia. Available online: https://encyclopedia.pub/entry/24615 (accessed on 02 April 2026).
Coradduzza D, Bellu E, Congiargiu A, Pashchenko A, Amler E, Necas A, et al. MiRNA Detection and Biosensing. Encyclopedia. Available at: https://encyclopedia.pub/entry/24615. Accessed April 02, 2026.
Coradduzza, Donatella, Emanuela Bellu, Antonella Congiargiu, Aleksei Pashchenko, Evzen Amler, Alois Necas, Ciriaco Carru, Serenella Medici, Margherita Maioli. "MiRNA Detection and Biosensing" Encyclopedia, https://encyclopedia.pub/entry/24615 (accessed April 02, 2026).
Coradduzza, D., Bellu, E., Congiargiu, A., Pashchenko, A., Amler, E., Necas, A., Carru, C., Medici, S., & Maioli, M. (2022, June 29). MiRNA Detection and Biosensing. In Encyclopedia. https://encyclopedia.pub/entry/24615
Coradduzza, Donatella, et al. "MiRNA Detection and Biosensing." Encyclopedia. Web. 29 June, 2022.
Copy Citation
MicroRNAs (miRNA) are key regulators of gene expression, controlling different biological processes such as cellular development, differentiation, proliferation, metabolism, and apoptosis. The relationships between miRNA expression and the onset and progression of different diseases, such as tumours, cardiovascular and rheumatic diseases, and neurological disorders, are well known. A nanotechnology-based approach could match miRNA delivery and detection to move beyond the proof-of-concept stage. Different kinds of nanotechnologies can have a major impact on the diagnosis and treatment of miRNA-related diseases such as cancer.
nano-microRNA
biomarcatore nanotecnologico
consegna
detection
1. Introduction
Early detection and effective treatments are crucial for disease eradication and patient healing, and for survival and/or quality of life. Cancer is one of the pathologies that can benefit from both approaches, as many other illnesses. MicroRNAs (miRNA) represent an emerging tool which could be effective in either detecting or curing several conditions. miRNAs are short (18–25 nucleotides) noncoding single-stranded endogenous oligoribonucleotides, which are relatively stable; they are predominantly secreted in vesicles, or in a complex with other proteins. MiRNA’s role is linked to the regulation of different biological processes, such as cellular development, differentiation, proliferation, metabolism, and apoptosis [1]. In fact, they regulate gene expression, and the alterations in their expression levels correlate with the onset and progression of different diseases, such as tumours, cardiovascular and rheumatic diseases, and neurological disorders [2][3][4]. Under pathological conditions, dysregulated miRNA levels are observed, but the relationship between dysregulated miRNA levels and disease is not straightforward. The oncogenic or tumour-suppressive activities of miRNAs depend on which genes are activated or inhibited through the up- or downregulation of miRNA expression. Thus, to ensure a secure diagnosis, the identification of miRNAs via expression profiling is fundamental; this is because alteration in the expression levels of a single miRNA does not have sufficient diagnostic power compared to multiplexing, that is, the parallel or simultaneous detection of known multiple miRNAs. Current quantification strategies have limitations and disadvantages. In general, all quantification methods are divided into two categories—one that utilizes direct oligo-hybridization without sample RNA amplification, and the other requiring sample amplification. Methods that do not utilize sample amplification will require a relatively larger starting amount of total RNA, while the others requiring sample amplification, with external variation as they are handling imperfections, can also be amplified. Oligo-microarray technology is relatively low-cost and readily available; a disadvantage of this method is its scale, as the resulting array will be relatively large. Alternatively, the use of synthesis and chemical modification of RNA probes is costly and often requires a large amount of total RNA. Many other methods and tools have been developed for miRNA expression profiling. Problems such as sensitivity and specificity have been addressed through various strategies; however, they are still very expensive, since answering to a variety of specific needs; an appropriate sample size, sample quantity and speed; and the requirement to identify new miRNAs can be costly. Thus, there is a great interest in developing innovative methods, and nanotechnology-based approaches are particularly sought-after. Nanotechnology will have a strong impact on delivery and diagnosis through miRNA, demonstrating that the newly developed approach works on ‘real-world’ samples under standardized conditions. The same is true also for miRNA transport and delivery, as miRNA inhibition or mimicry are strategies currently under evaluation to maintain the level of miRNA inside the cells, and nanotechnology can offer a good solution to bring the miRNA mimics to the tumour site.
Both approaches (detection and treatment) using nanotechnology-based strategies have been analysed, to point out the current trends in this promising field of bio-medical applications (Figure 1 and Figure 2).
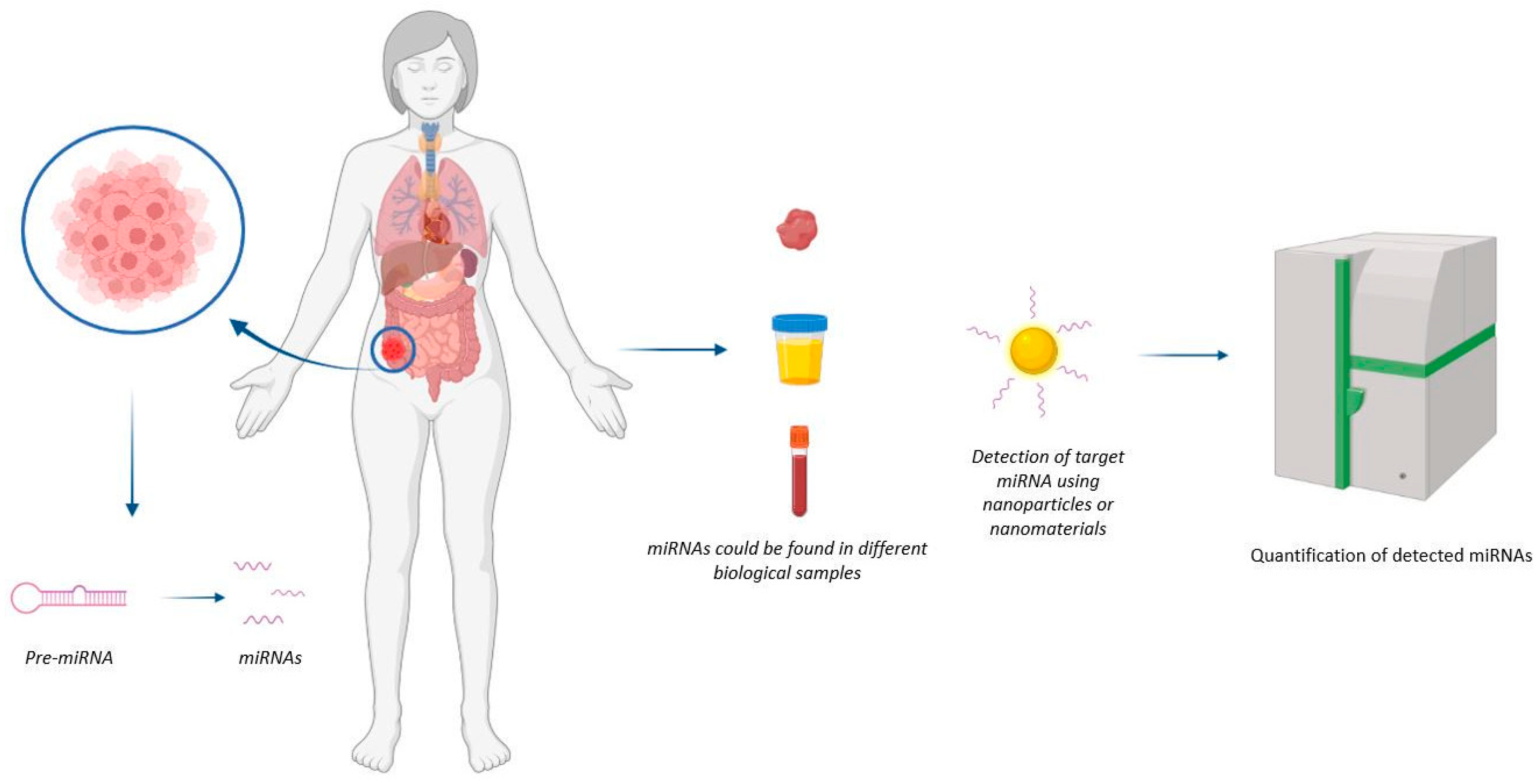
Figure 1. Representation of mechanism of action of nanodevices for miRNA detection. The nanodevice can detect the miRNA of interest at the tumour site.
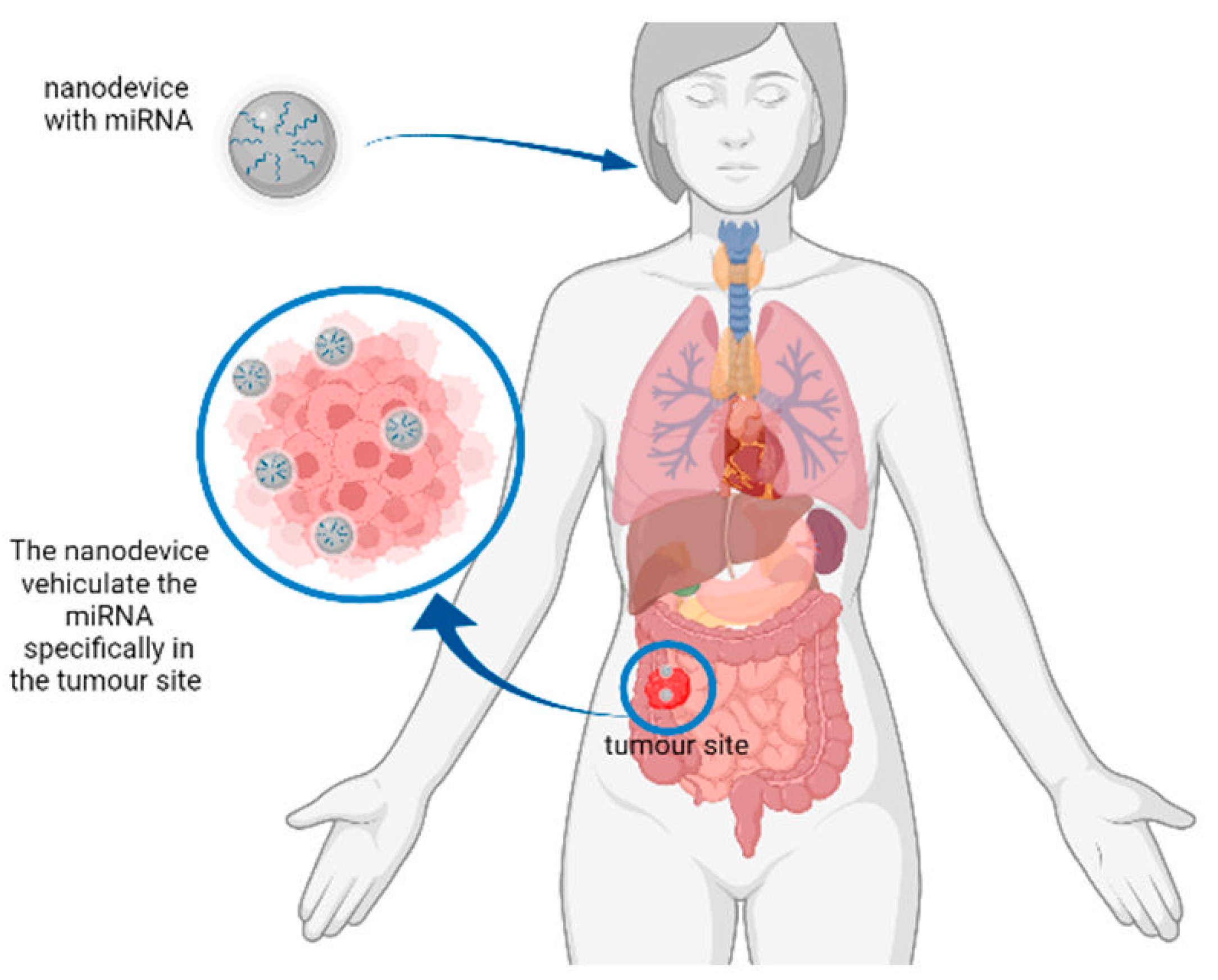
Figure 2. Representation of a hypothetical mechanism of action of nanodevices for miRNA delivery. The nanodevice can deliver the miRNA of interest to the tumour site to reduce the expression of tumour-related genes.
2. MiRNA Detection and Biosensing
Conventionally, miRNA detection is based on real-time polymerase chain reaction (qRT-PCR), microarrays, Northern blotting, and Next-generation sequencing (NGS). These measurement methods just reflect the average gene expression level and cannot provide the heterogeneity and transient spatiotemporal variations of miRNAs in living cells [5]. The main difference between nanotechnology-based and conventional methods lies in the transduction mechanism. The peculiar physicochemical properties of nanostructured materials are essential to enhancing signal readout [6]. Several nanoparticles (NPs) have excellent optical properties, making them ideally suited for the development of sensing strategies. Some NPs are bright and stable fluorescence emitters, such as silver nanoclusters (AgNCs) and quantum dots (QDs), and can be used either directly or in fluorescence resonance energy transfer strategies. Other NPs, such as gold NPs (AuNPs) and carbonaceous NPs, can be used as efficient fluorescence quenchers in fluorescence recovery approaches. In the presence of target miRNA, the fluorophores, physically or chemically released from the NP, can emit fluorescence in a concentration-dependent manner.
These few examples indicate that nanomaterials can have great potential in miRNA detection and biosensing. Biosensors represent innovative analytical tools for clinical diagnosis as well as for a better understanding of the molecular mechanisms involved in pathophysiology, revealing new biomarkers that are useful for the evaluation of appropriate pharmaceutical treatments [7]; moreover, miRNAs are surely promising and effective biomarkers. Biosensors can help in the early diagnosis and monitoring of pathological conditions, particularly for oncological diseases, and are useful in prognosis, surveilling the evolution of the disease, and opening the door to access to global health care. Among the different biosensing techniques, plasmonic sensor platforms are able to analyse different classes of biomolecules of clinical interest [8]. Different classes of biomolecules can be quantitatively detected in real time via high-throughput exploitation of (Localized) the surface plasmon resonance (SPR or LSPR) of metal nanoparticles and nanofilms, able to monitor and also perform label-free interactions. In fact, surface plasmons have been employed to enhance the surface sensitivity of different spectroscopic techniques, such as fluorescence and Raman scattering, applied to the detection of biomolecules. In the case of metal nanoparticles, LSP oscillations are responsible for the deep colors of their suspensions or sols, due to strong absorption bands in the ultraviolet–visible region, that are not present in the bulk material. The surface interaction of metal nanoparticles with biopolymers such as proteins, DNA, and RNA causes shifts in this resonance that can be used to detect and quantify their presence.
References
- Revythis, A.; Shah, S.; Kutka, M.; Moschetta, M.; Ozturk, M.A.; Pappas-Gogos, G.; Ioannidou, E.; Sheriff, M.; Rassy, E.; Boussios, S. Unraveling the Wide Spectrum of Melanoma Biomarkers. Diagnostics 2021, 11, 1341.
- De Planell-Saguer, M.; Rodicio, M.C. Analytical aspects of microRNA in diagnostics: A review. Anal. Chim. Acta 2011, 699, 134–152.
- Li, Y.; Kowdley, K.V. MicroRNAs in common human diseases. Genom. Proteom. Bioinform. 2012, 10, 246–253.
- Wark, A.W.; Lee, H.J.; Corn, R.M. Multiplexed detection methods for profiling microRNA expression in biological samples. Angew. Chem. Int. Ed. Engl. 2008, 47, 644–652.
- Hunt, E.A.; Broyles, D.; Head, T.; Deo, S.K. MicroRNA Detection: Current Technology and Research Strategies. Annu. Rev. Anal. Chem. 2015, 8, 217–237.
- Fiammengo, R. Can nanotechnology improve cancer diagnosis through miRNA detection? Biomark. Med. 2017, 11, 69–86.
- Bellassai, N.; Spoto, G. Biosensors for liquid biopsy: Circulating nucleic acids to diagnose and treat cancer. Anal. Bioanal. Chem. 2016, 408, 7255–7264.
- Mariani, S.; Minunni, M. Surface plasmon resonance applications in clinical analysis. Anal. Bioanal. Chem. 2014, 406, 2303–2323.
More
Information
Subjects:
Biochemistry & Molecular Biology
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
778
Revisions:
4 times
(View History)
Update Date:
01 Jul 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





