COVIDomics, namely, the proteomic and metabolomic signatures of COVID-19. Omics-based technologies have been largely adopted during the unprecedented global COVID-19 pandemic, allowing the scientific community to perform research on a large scale to understand the pathobiology of the SARS-CoV-2 infection and its replication into human cells. The application of omics techniques has been addressed to every level of application, from the detection of mutations, methods of diagnosis or monitoring, drug target discovery, and vaccine generation, to the basic definition of the pathophysiological processes and the biochemical mechanisms behind the infection and spread of SARS-CoV-2. Thus, the term COVIDomics wants to include those efforts provided by omics-scale investigations with application to the COVID-19 research.
1. Introduction
In the actual background of the world coronavirus disease 2019 (COVID-19) pandemic, scientists and researchers worldwide have made significant efforts to unravel the clinical and molecular aspects regarding the infection of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and the progression of COVID-19 disease. Meanwhile, many attempts were simultaneously made to disclose the cellular and pathophysiological processes affected in COVID-19 patients; the urgent need for pharmacological treatments prompted a rapid approach to drug repurposing and vaccines’ development, to limit the number of deaths and the spread of the infection
[1][2][3][4][5]. Classification of COVID-19 disease can be made according to the clinical characteristics of patients (
Figure 1). According to the high variability and heterogeneity of symptoms and comorbidities
[6], patients can be categorized as symptomatic or asymptomatic, and a crescent grade of severity is usually referred to as mild, moderate, or severe. The outcome of COVID-19 patients defines their status, classifying them as hospitalized, needing intensive care unit (ICU), or fatal. Very often, even after recovery, many patients can experience post-acute COVID syndrome (PACS), during which symptoms may last for several months
[7]. The World Health Organization (WHO) has organized a system score for clinical improvement and management of COVID-19 disease, based on an ordinal scale to categorize patients according to their clinical manifestations
[8].
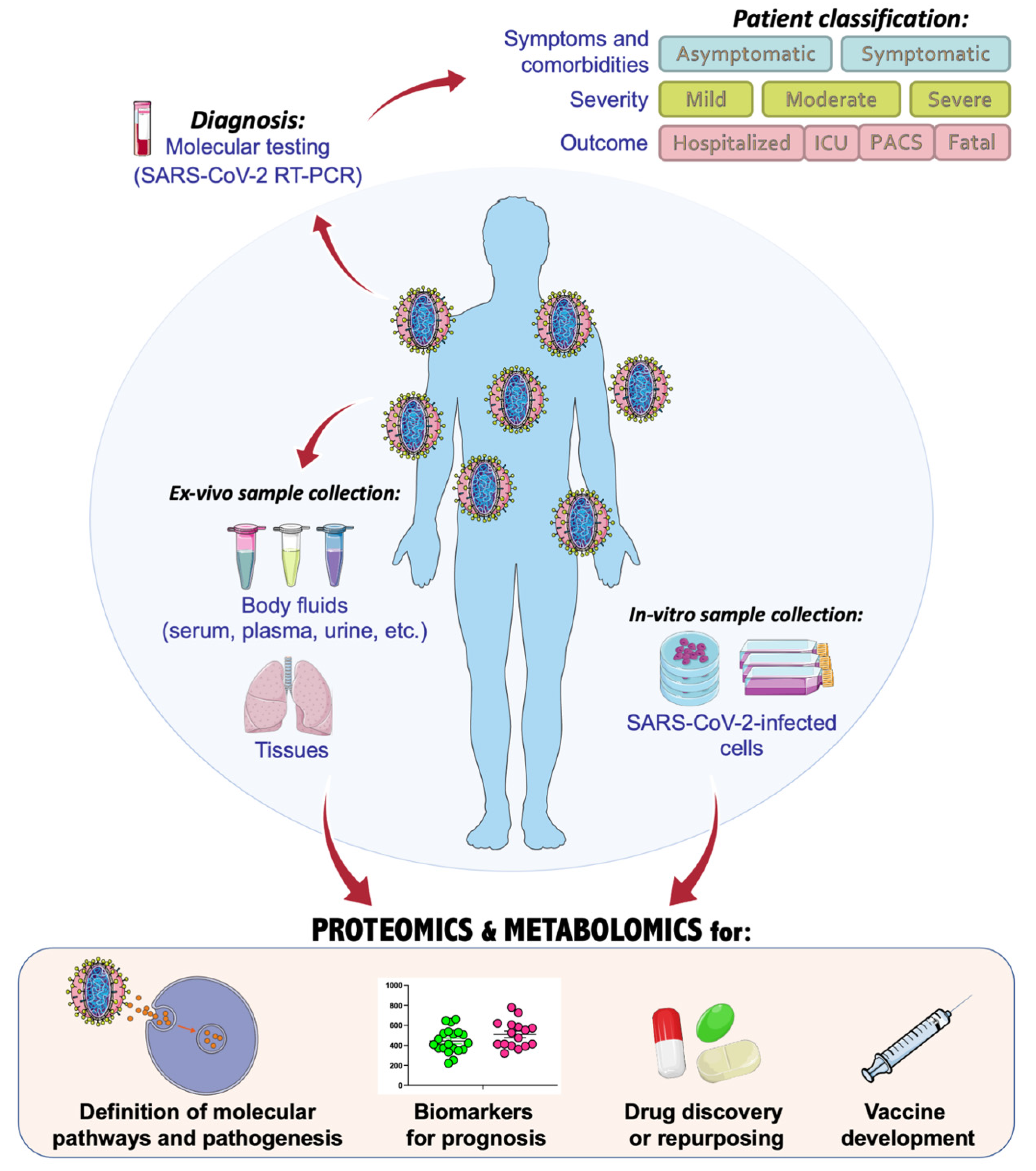
Figure 1. Application and integration of omics technologies to characterize the molecular biology of SARS-CoV-2 and COVID-19 pathogenic mechanisms and therapeutic approaches, with the main focus on proteomics and metabolomics investigations. This figure was drawn adapting the vector image from the Servier Medical Art bank (
http://smart.servier.com/; last accessed 2 January 2022). COVID-19 = coronavirus disease 2019; SARS-CoV-2 = severe acute respiratory syndrome coronavirus 2; ICU = intensive care unit; PACS = post-acute COVID syndrome.
In the fight against the COVID-19 pandemic, single- and multiomics-based strategies have been largely adopted with the main aim of dissecting a plethora of aspects related to the SARS-CoV-2 infection. In fact, omics-based technologies, such as genomics, transcriptomics, proteomics, and metabolomics, can serve at every stage of COVID-19 investigations, from diagnosis and progression of the disease, to the discovery of altered molecular pathways or potential drug targets and vaccines (Figure 1).
Genomics and transcriptomics have suitably allowed to promptly identify in diverse biological matrices the mutations of SARS-CoV-2, including those responsible for the occurrence of new variants, and the changes in the coronavirus genes expression
[9][10][11][12]. On the other hand, the value of proteomics and metabolomics concerns the possibility of understanding the functional alterations that depend on virus infection and its interaction with the host cell. In fact, with the increasing advance and the high throughput provided by liquid chromatography—tandem mass spectrometry (LC-MS/MS) technologies in biomedicine, the identification and quantification of several thousands of proteins and metabolites is possible in a targeted or untargeted fashion, or using the newest data-independent acquisition (DIA)-MS methods
[13][14][15][16], besides the traditional data-dependent acquisition (DDA) ones
[17][18][19][20][21]. With such sophisticated techniques, many aspects of the pathogenesis of COVID-19 can be disclosed, acquiring an important piece of knowledge to understand the molecular processes perturbed by virus infection and to predict or ameliorate the clinical outcome of patients by finding promising druggable targets. On this matter, even out of the scope here, an interactomics study performed in human cells using affinity-purification mass spectrometry found several interacting partners in 26 out of the 29 proteins of SARS-CoV-2, concluding that some of these interactors were druggable proteins
[22].
In parallel, the introduction of a metabolomic workflow in a work provides a rich source of information, allowing the phenotypic biochemical and metabolic characterization of cells, tissues, or body fluids, and discovering and monitoring biomarkers or disease signatures
[23][24][25]. What is more, the challenging advances in mass spectrometry and metabolomics research provided scientists with a powerful tool able to technically separate the analysis of small molecules from lipid molecules, diving into the deep of the metabolome and the lipidome, respectively
[26][27]. Lipids represent the structural building blocks of cell and virus membranes, thus playing an essential role for the virus life cycle, including viral invasion and replication. Since viruses are able to modulate lipid synthesis and signaling into the host cell to produce lipids for their envelopes by creating a lipid micro-environment ideal for replication, the lipid dysregulation is suggested as a drug target to prevent coronavirus infection
[28]. Furthermore, considering the high heterogeneity of the clinical manifestations of COVID-19, thus suggesting the involvement of diverse pathophysiological processes
[29], multiomics approaches and data integration analyses may result as a winning choice, contributing to a better understanding of the molecular biology of SARS-CoV-2 at every level of complexity
[30].
To unify all the efforts made by scientists at the omics-scale level in the research to combat the current pandemic and provide a comprehensive overview of the omics studies made in favor of COVID-19 research, here used the term COVIDomics. Hence, herein summarized the COVIDomics discoveries mainly based on proteomics and metabolomics studies on blood samples from patients. Many overlapping results were found in several publications from different authors for both proteomics and metabolomics applications. The relevant findings of such investigations were highlighted and, finally, summarized and analyzed with bioinformatics tools in order to capture a common signature in terms of proteins, metabolites, and pathways dysregulated in COVID-19 disease.
2. COVIDomics Data Integration
The application of omics and multiomics technologies, enclosed here under the term COVIDomics, offers the great chance to meticulously recognize and dissect the multiple molecular aspects that characterize a complex disease such as COVID-19. The majority of patients that contract the SARS-CoV-2 infection may develop no or mild symptoms, although many others experience a severe or critical symptomatology leading to fatal cases. This high heterogeneity in the clinical manifestation is being investigated in many studies, comprising those here concluded. In fact, many comparisons investigate the differences between the severe disease toward the baseline proteome or metabolome in healthy patients; others focus mostly on the progression of the disease from a condition to another one and highlight correlations of clinical and biochemical parameters with dysregulated proteins and metabolites. Furthermore, the majority of the COVIDomics research was performed on blood, as it is an easily accessible liquid biopsy reflecting the systemic changes subsequent the strong infection of SARS-CoV-2. Thus, the comprehensive and accurate analysis of circulating molecules is helpful to unravel the systemic mechanisms of the disease. In fact, as summarized in Figure 2 and detailed in Table 4, there are several common proteins that were identified in plasma and serum as quantitatively changed between the patients and their controls, representing a relevant proteomic signature of COVID-19.
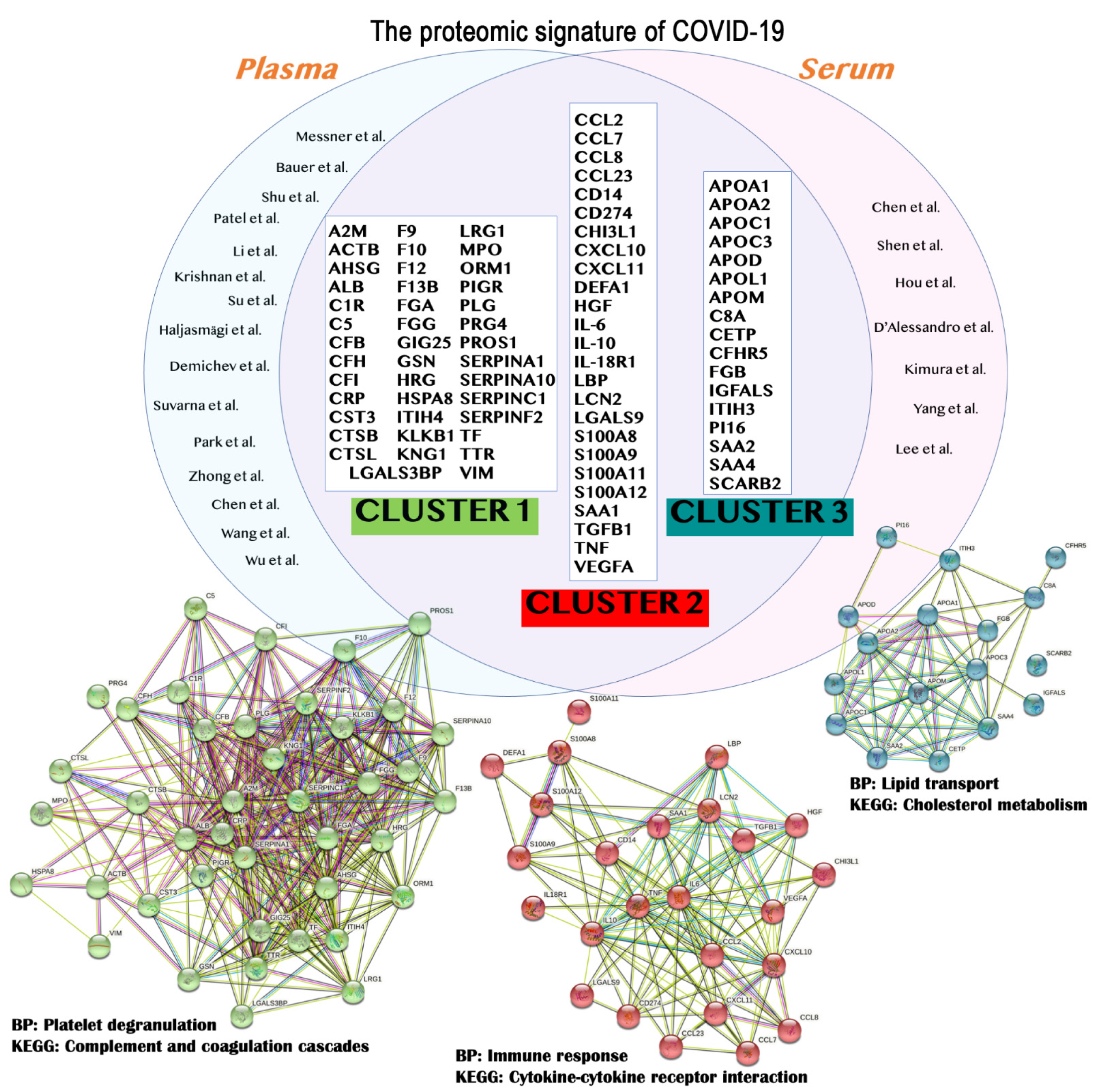
Figure 2. The main findings obtained from proteomics studies are summarized. In particular, the results from plasma
[31][32][33][34][35][36][37][38][39][40][41][42][43][44][45] and serum
[46][47][48][49][50][51][52] studies were merged to identify the common proteins (top) that should represent the proteome signature of COVID-19. These protein entries were analyzed and clustered using STRING version 11.5, revealing the formation of three main clusters (bottom). The relative biological processes (BP) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways were indicated. A detailed list for the proteins depicted in this figure is available in
Table 1.
Table 1. The list of the proteins identified in more than one study characterizing the blood proteomic signature of COVID-19.
| Protein Symbol |
UniProt ID |
Protein Name |
STRING Cluster |
| A2M |
P01023 |
Alpha-2-macroglobulin |
Cluster 1 |
| ACTB |
P60709 |
Actin, cytoplasmic 1 |
| AHSG |
P02765 |
Alpha-2-HS-glycoprotein |
| ALB |
P02768 |
Albumin |
| C1R |
P00736 |
Complement C1r subcomponent |
| C5 |
P01031 |
Complement C5 |
| CFB |
P00751 |
Complement Factor B |
| CFH |
P08603 |
Complement factor H |
| CFI |
P05156 |
Complement factor I |
| CRP |
P02741 |
C-reactive protein |
| CST3 |
P01034 |
Cystatin-C |
| CTSB |
P07858 |
Cathepsin B |
| CTSL |
P07711 |
Procathepsin L |
| F9 |
P00740 |
Coagulation factor IX |
| F10 |
P00742 |
Coagulation factor X |
| F12 |
P00748 |
Coagulation factor XII |
| F13B |
P05160 |
Coagulation factor XIII B chain |
| FGA |
P02671 |
Fibrinogen alpha chain |
| FGG |
P02679 |
Fibrinogen gamma chain |
| GSN |
P06396 |
Gelsolin |
| HRG |
P04196 |
Histidine-rich glycoprotein |
| HSPA8 |
P11142 |
Heat shock cognate 71 kDa protein |
| ITIH4 |
Q14624 |
Inter-alpha-trypsin inhibitor heavy chain H4 |
| KLKB1 |
P03952 |
Plasma kallikrein |
| KNG1 |
P01042 |
Kininogen-1 |
| LGALS3BP |
Q08380 |
Galectin-3-binding protein |
| LRG1 |
P02750 |
Leucine-rich alpha-2-glycoprotein |
| MPO |
P05164 |
Myeloperoxidase |
| ORM1 |
P02763 |
Alpha-1-acid glycoprotein 1 |
| PIGR |
P01833 |
Polymeric immunoglobulin receptor |
| PLG |
P00747 |
Plasminogen |
| PRG4 |
Q92954 |
Proteoglycan 4 |
| PROS1 |
P07225 |
Vitamin K-dependent protein S |
| SERPINA1 |
P01009 |
Alpha-1-antitrypsin |
| SERPINA3 |
P01011 |
Alpha-1-antichymotrypsin |
| SERPINA10 |
Q9UK55 |
Protein Z-dependent protease inhibitor |
| SERPINC1 |
P01008 |
Antithrombin-III |
| SERPINF2 |
P08697 |
Alpha-2-antiplasmin |
| TF |
P02787 |
Transferrin |
| TTR |
P02766 |
Transthyretin |
| VIM |
P08670 |
Vimentin |
| CCL2 |
P13500 |
C-C motif chemokine 2 |
Cluster 2 |
| CCL7 |
P80098 |
C-C motif chemokine 7 |
| CCL8 |
P80075 |
C-C motif chemokine 8 |
| CD14 |
P08571 |
Monocyte differentiation antigen CD14 |
| CCL23 |
P55773 |
C-C motif chemokine 23 |
| CD274 |
Q9NZQ7 |
Programmed cell death 1 ligand 1 |
| CHI3L1 |
P36222 |
Chitinase-3-like protein 1 |
| CXCL10 |
P02778 |
C-X-C motif chemokine 10 |
| CXCL11 |
O14625 |
C-X-C motif chemokine 11 |
| DEFA1 |
P59665 |
Neutrophil defensin 1 |
| HGF |
P14210 |
Hepatocyte growth factor |
| IL-10 |
P22301 |
Interleukin-10 |
| IL-18R1 |
Q13478 |
Interleukin-18 receptor 1 |
| IL-6 |
P08887 |
Interleukin-6 receptor subunit alpha |
| LBP |
P18428 |
Lipopolysaccharide-binding protein |
| LCN2 |
P80188 |
Neutrophil gelatinase-associated lipocalin |
| LGALS9 |
O00182 |
Galectin-9 |
| S100A11 |
P31949 |
Protein S100-A11 |
| S100A12 |
P80511 |
Protein S100-A12 |
| S100A8 |
P05109 |
Protein S100-A8 |
| S100A9 |
P06702 |
Protein S100-A9 |
| SAA1 |
P0DJI8 |
Serum amyloid A-1 protein |
| TGFB1 |
P01137 |
Transforming growth factor beta-1 proprotein |
| TNF |
P01375 |
Tumor necrosis factor |
| VEGFA |
P15692 |
Vascular endothelial growth factor A |
| APOA1 |
P02647 |
Apolipoprotein A1 |
Cluster 3 |
| APOA2 |
P02652 |
Apolipoprotein A2 |
| APOC1 |
P02654 |
Apolipoprotein C1 |
| APOC3 |
P02656 |
Apolipoprotein C3 |
| APOD |
P05090 |
Apolipoprotein D |
| APOL1 |
O14791 |
Apolipoprotein L1 |
| APOM |
O95445 |
Apolipoprotein M |
| C8A |
P07357 |
Complement component C8 alpha chain |
| CETP |
P11597 |
Cholesteryl ester transfer protein |
| CFHR5 |
Q9BXR6 |
Complement factor H-related protein 5 |
| FGB |
P02675 |
Fibrinogen beta chain |
| IGFALS |
P35858 |
Insulin-like growth factor-binding protein complex acid labile subunit |
| ITIH3 |
Q06033 |
Inter-alpha-trypsin inhibitor heavy chain H3 |
| PI16 |
Q6UXB8 |
Peptidase inhibitor 16 |
| SAA2 |
P0DJI9 |
Serum amyloid A-2 protein |
| SAA4 |
P35542 |
Serum amyloid A-4 protein |
| SCARB2 |
Q14108 |
Lysosome membrane protein 2 |
Proteins were ordered alphabetically for each cluster according to the protein symbol.
As for proteins, herein have also summarized the common findings obtained analyzing metabolomics studies in terms of metabolites and metabolic pathways highlighted in plasma and serum from COVID-19 patients (
Figure 3). In particular, herein have focused the analysis on small molecules, since metabolomics software only recognize well-annotated HMDB (Human Metabolome Database) compounds, and lipids may be not correctly mapped. Indeed, the enrichment analysis of the metabolites associated with COVID-19 was performed using MetaboAnalyst 5.0 software
[53][54][55][56].
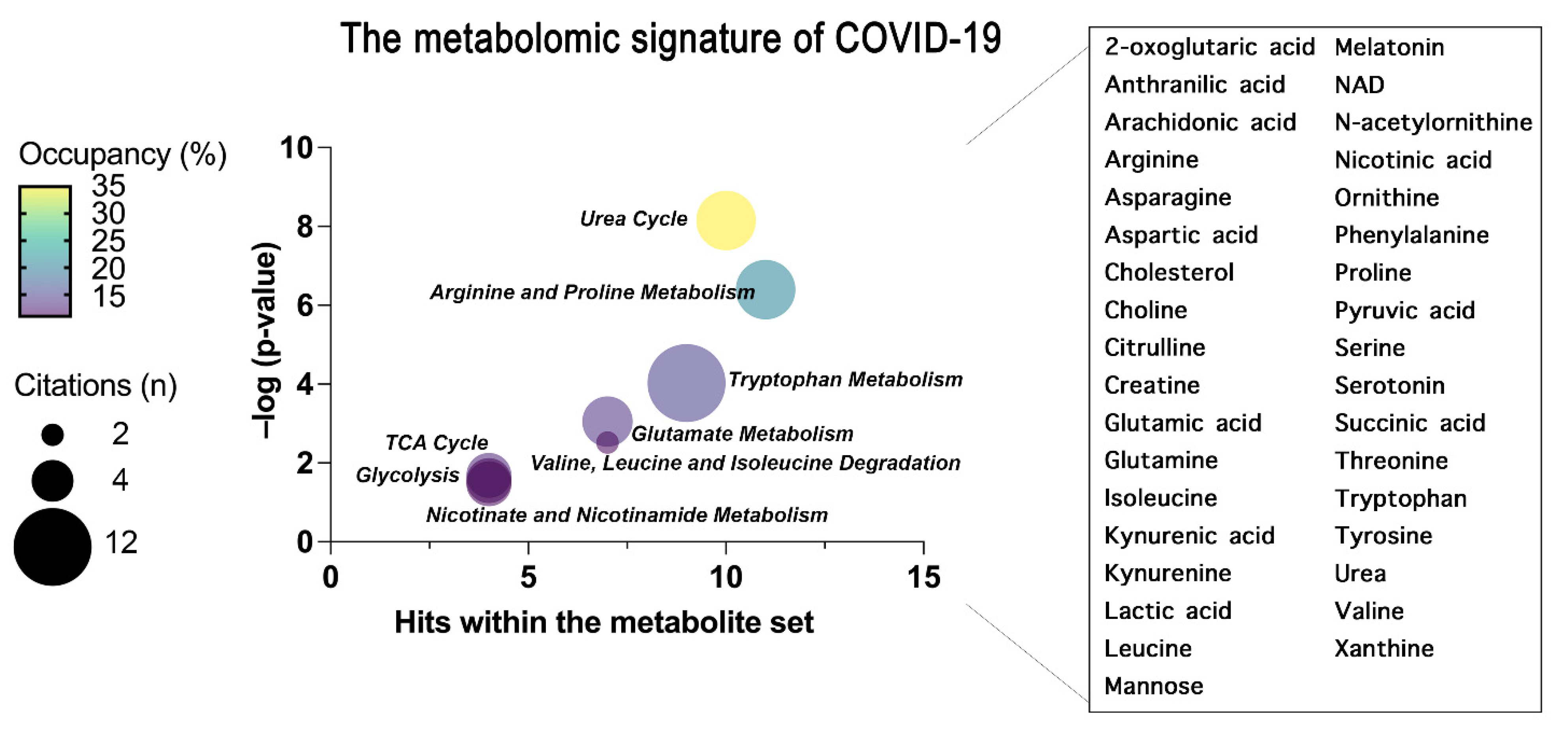 Figure 3.
Figure 3. The common metabolites and metabolic pathways obtained from the conclusion of plasma and serum metabolomics studies are summarized. A specific signature of the metabolome of COVID-19 patients was obtained representing the most enriched pathways constructed through an over-representation analysis (ORA) within MetaboAnalyst 5.0 and represented as bubble plot. The bubble plot takes into account the statistical significance (–log
p-value) of each metabolite set identified through the ORA analysis. Where the size of each bubble refers to the number of times the metabolite set is cited here by each scientist, the color instead refers to the number of metabolites detected over the total number of metabolites within that pathway (occupancy). On the right, the metabolites common to all the concluded papers that have generated the pathway enrichment are grouped.
3. Concluding Remarks
During the current COVID-19 pandemic, unprecedented efforts have been made by the scientific community to dissect the molecular bases of SARS-CoV-2 infection, spreading, and pathogenicity. Omics scientists deserved particular merits for performing several proteomics- and metabolomics-based investigations, but also integrative multiomics analyses, using samples derived from patients. Thus, the application of COVIDomics strategies has certainly empowered the knowledge relative to COVID-19 in terms of biomarkers of disease and specific physiological mechanisms disturbed upon SARS-CoV-2 infection. The study of coronavirus structure, replication, and infection, together with the virus-host interaction and the host response, have remarkably provided us with efficient tools to diagnose the disease and ameliorate the fate of COVID-19 patients. In general, a huge improvement in patients’ outcomes is correlated to the development of vaccines as prophylactic therapeutic approaches, despite it remaining not currently possible to foresee the prognosis of infected patients by a single analysis or through the detection of a single and specific marker. Nonetheless, in this scenario, the use of omics sciences and their integration have provided us with relevant proteomics and metabolomics signatures, as a point of junction between SARS-CoV-2 infection/pathobiology with the subsequent clinical outcome of affected patients. Thus, COVIDomics is contributing to a better comprehension of the molecular intricacy of COVID-19, in the view of identifying new molecular actors involved in the pathogenesis of the disease to be used for efficient therapeutic approaches in combatting the COVID-19 pandemic.



 Figure 3. The common metabolites and metabolic pathways obtained from the conclusion of plasma and serum metabolomics studies are summarized. A specific signature of the metabolome of COVID-19 patients was obtained representing the most enriched pathways constructed through an over-representation analysis (ORA) within MetaboAnalyst 5.0 and represented as bubble plot. The bubble plot takes into account the statistical significance (–log p-value) of each metabolite set identified through the ORA analysis. Where the size of each bubble refers to the number of times the metabolite set is cited here by each scientist, the color instead refers to the number of metabolites detected over the total number of metabolites within that pathway (occupancy). On the right, the metabolites common to all the concluded papers that have generated the pathway enrichment are grouped.
Figure 3. The common metabolites and metabolic pathways obtained from the conclusion of plasma and serum metabolomics studies are summarized. A specific signature of the metabolome of COVID-19 patients was obtained representing the most enriched pathways constructed through an over-representation analysis (ORA) within MetaboAnalyst 5.0 and represented as bubble plot. The bubble plot takes into account the statistical significance (–log p-value) of each metabolite set identified through the ORA analysis. Where the size of each bubble refers to the number of times the metabolite set is cited here by each scientist, the color instead refers to the number of metabolites detected over the total number of metabolites within that pathway (occupancy). On the right, the metabolites common to all the concluded papers that have generated the pathway enrichment are grouped.




