
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Syed Nisar Hussain Bukhari | + 5411 word(s) | 5411 | 2022-02-10 11:02:12 | | | |
| 2 | Lindsay Dong | -41 word(s) | 5370 | 2022-02-16 03:18:18 | | | | |
| 3 | Lindsay Dong | -41 word(s) | 5370 | 2022-02-16 03:19:28 | | |
Video Upload Options
An antigenic determinant (AD) is a portion of an antigen molecule known as an epitope that is recognized by the human immune system, specifically by antibodies or T and B cells. Recognition of epitopes is considered important in EBPV design to contain pandemics, epidemics, and endemics due to the outbreak of infectious diseases. To design an effective and viable EBPV against different strains of a pathogen, it is important to identify the putative T- and B-cell epitopes. Using the wet-lab experimental approach to identify these epitopes is time-consuming and costly because the experimental screening of a vast number of potential epitope candidates is required. Fortunately, various available machine learning (ML)-based prediction methods have reduced the burden related to the epitope mapping process by decreasing the potential epitope candidate list for experimental trials. Moreover, these methods are also cost-effective, scalable, and fast.
1. Introduction
1.1. Epitopes and Paratopes
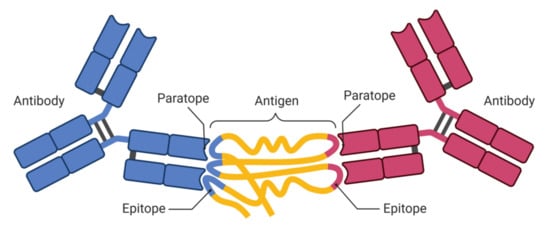
1.2. Need for T- and B-Cell Epitope Prediction
2. ML-Based Studies for the Prediction of T- and B-Cell Epitopes
| Study Conducted | Methodology Adopted | Strengths/Limitations |
|---|---|---|
| T. Liu et al. [23] | A feedforward deep neural network-based ensemble of 11 classifiers was created to predict BCEs. IEDB was used to obtain the BCE peptide dataset. On the test set, the model was evaluated using the AUROC metric. | Model reports peptide as an epitope if classified by all 11 classifiers. It would provide the best results if simple majority voting was used for classification. |
| Fatoba, A. J. et al. [24] | In [24], potential epitope-based vaccine candidates were explored. After retrieving 600 genome sequences of SARS-CoV-2 from the ViPR repository, CD8+ and CD4+ epitopes and B-cell (linear) epitopes were generated and screened for immunogenicity, antigenicity, and non-allergenicity. | The results of [25] reported 19 candidate T-cell epitopes (CD8+), which were found to overlap strongly with 8 B-cell epitopes. The results provide the basis for an experimental design for a suitable peptide vaccine against SARS-CoV-2. |
| R. Moody et al. [26] | Authors used IEDB prediction tools for predicting B-cell epitopes and those with high scores in terms of prediction were selected as candidate epitopes. The epitopes were then matched to human proteins using NCBI Blast technology. | The findings showed eleven (11) novel B-cell epitopes in the host that were capable of explaining key elements of COVID-19 extrapulmonary disease that previous research had not been able to explain. |
| Jespersen MC et al. [27] | The authors employed feedforward neural networks (FFNN) with two hidden layers, each with 25 neurons, an activation function (sigmoid) at all neurons, and an ADAM as an optimizing function to predict antibody-specific epitopes (B cell) or epitope targets of provided cognate antibodies. The dataset was obtained from the IEDB database. PCA was used for dimensionality reduction before the model was trained. | It was shown that a simple set of attributes retrieved from the cognate antibody boosted the rate of accuracy in predicting individual epitopes. Furthermore, sophisticated features such as Zernike Moments can improve the model’s predictive potential. When compared to DiscoTope 2.0, this model performs better in finding patches overlapping with an actual patch of an epitope in cross-validation and on an independent dataset. |
| Ling-yun Liu et al. [28] | The authors used PCA and RNN networks. They converted the physicochemical properties into digital vectors, intending to have high-dimensional feature space, and later PCA was applied to process them. The output from PCA was used as an input to the RNN for predicting epitopes. | Prediction results obtained by this process demonstrated that PCA reduced dimensions, but at the same time, original features of the main component were retained, and the rate of prediction was also improved. |
| Bin Cheng et al. [29] | Authors introduced a novel scale to measure feature importance, called the relevance of amino acid pair (RAAP). RAAP was calculated by decomposing the sequences of amino acids based on their physicochemical properties. | The successful prediction rate was drastically improved here by using LSTM. It does not suffer from gradient instability and is good enough for textual classification sequences. Fivefold cross-validation was used to test and validate the models. |
| Balachandran Manavalan et al. [30] | Here, a non-redundant dataset was constructed containing 5500 BCEs experimentally validated, and 6893 non-B-cell epitopes were retrieved from IEDB. Then, an ensemble model to predict B-cell epitopes based on ERT (extremely randomized tree) and a classifier called GB (gradient boosting) was developed. The model works based on the physicochemical properties, AA composition, and combination of dipeptides and PCP as the input features. | After performing cross-validation on a benchmark dataset, it was shown that this model performed far better than the individual classifiers such as ERT and GB, with an MCC (Matthews correlation coefficient) of 0.454. |
| Yuh-Jyh Hu et al. [31] | A cost-sensitive strategy based on bagging MDT was suggested, which integrates two ensemble-based learning algorithms. Without employing the prediction of a pre-trained single predictor, it makes it independent of multiple prediction tools. It can also learn a meta-classification architecture with varied data, without being constrained by a particular hierarchy. | It was demonstrated that the performance of prediction is superior as compared to a single epitope predictor. However, epitope prediction based on meta-learning is purely dependent upon the predictive strength of various other pre-trained linear and conformational epitope prediction tools, which cannot be retained directly by users. Hence, this limits the flexibility and applicability of these meta-classifiers. |
| Jing Ren et al. [32] | The authors proposed a novel staged heterogeneity-based learning model. The model learns both heterogeneity and characteristics of data in a phased manner to identify residue of antigens of conformational B-cell type epitopes that are heterogeneous, purely based on sequences of antigens. In the first stage, the model is made to learn the generic epitope pattern with propensities, and in the second stage, the same model is made to learn the complementarity of the propensities used in the first stage, which is heterogeneous but this time on a small dataset of experimentally verified epitopes. | It was demonstrated that if heterogeneity was learned well, the transferability of the model improved remarkably in handling new data.It was tested and validated on two different datasets: one with epitopes determined experimentally and another with computationally defined. It showed outstanding performance that was around twice that of existing predictors, including CBTOPE. |
| Georgios A. et al. [33] | A novel method, “SEPIa”, has been proposed here to predict B-cell epitopes from protein sequences and is sufficiently faster, and it can also be applied to large-scale datasets. The model is the combination of two classifiers, random forest and naïve Bayes algorithm. | The average prediction accuracy of SEPIa is limited. The AUC score is 0.65 in both 10-fold cross-validation and on the independent test dataset, which is higher than other approaches tested on the same test dataset. |
| Gene Sher et al. [25] | Authors proposed a novel, analytically trained DREEP (Deep Ridge Regressed Epitope Predictor) based on string kernels using a deep neural network tailored to predict continuous epitopes. | The model was tested with input as long sequences of proteins from datasets such as AntiJen, Pellequer, and HIV. The results were compared with epitope predictors such as DMNLBE, LBtope, etc. Using the area under the curve (AUC) metric, the model achieved performance improvements over SARS by 13.7%, HIV by 8.9%, and Pellequer by 1.5%. |
| Wen Zhang et al. [34] | Authors attempted to differentiate immunogenic epitopes from non-immunogenic epitopes based purely on their primary structure. To effectively utilize various features, an ensemble method based on a genetic algorithm was proposed. | The model was tested on two benchmark datasets: IMMA2, PAAQD. The model was compared with methods such as POPI, PAAQD, and POPISK, which are considered state-of-the-art in nature. The model performed better, with an AUC score on IMMA2 of 0.846 and 0.829 on PAAQD. |
| Wei Zheng et al. [35] | The authors used ensemble learning to improve the prediction of BCEs. Their ensemble method combined twelve SVMs. To handle imbalanced datasets, resampling and AdaBoost methods were used. | The proposed ensemble model achieved an AUC score of 0.642–0.672 on the training dataset with five-fold cross-validation and an AUC score of 0.579–0.604 on the test dataset. |
| Jian Zhang et al. [36] | To predict antigenic determinants, the authors devised a cost-sensitive ensemble approach, and a spatial clustering-based algorithm was used to identify probable epitopes. | The model performed admirably in terms of prediction. AUC scores of 0.721 and 0.703 were obtained using leave-one-out cross-validation (LOOCV) on two benchmark datasets: bound and unbound. |
| Kavitha K V et al. [37] | PCA was used to reduce dimensions and to filter out the essential features; for prediction purposes, a random forest algorithm was used. | Experimental results showed that the random forest-based classifier had an improved prediction accuracy rate as compared to BCPred, AAP, etc. |
| Wen Zhang et al. [38] | The authors used sequence-derived features and developed an ensemble model based on random forest to predict epitopes accurately. | The model was evaluated using the leave-one-out cross-validation procedure, and an AUC score of 0.687 and 0.651 on bound and unbound datasets was obtained. |
| Ping Chen et al. [39] | Authors reviewed various prediction models for epitopes, such as models based on SVM, neural network, random forest, etc., to defend computational approaches in the prediction of epitopes as in silico methods require a lot of effort and time. | Apart from defending the computational approaches, it was also concluded that there is a limitation to current models as it is impossible to devise an exact model without having complete knowledge of the immune system, and current models are simply best at approximation. |
| Claus Lundegaard et al. [40] | Here, an artificial neural network was used. The standard feedforward neural network with backpropagation was employed to predict epitopes. The dataset was retrieved from the SYFPEITHI database. | The model efficiently and accurately predicts MHC class I type peptides and outperforms the existing methods. |
3. Tools for T- and B-Cell Epitope Prediction
3.1. Tools for T-Cell Epitope Prediction
3.2. Tools for B-Cell Epitope Prediction

4. Predicting SARS-CoV-2 Epitopes
| Sr. No. | Method Name | Usage |
|---|---|---|
| 01 | NetMHC [63] | To predict HLA I class or CD8+ SARS-CoV-2 T-cell epitopes |
| 02 | NetMHCpan [64] | |
| 03 | NetCTLpan_1.1 [65] | |
| 04 | NetMHC_4.0 [66] | |
| 05 | HLAthena [67] | |
| 06 | MHCflurry [68] | |
| 07 | NetHMCII_2.3 [69] | To predict HLA II class or CD4+ SARS-CoV-2 T-cell epitopes |
| 08 | NetMHCIIpan_3.0 [70] | |
| 09 | NetMHCIIpan_4.0 [71] | |
| 10 | NeonMHC2 [72] | |
| 11 | MARIA [73] |
5. Future Research Directions in T- and B-Cell Epitope Prediction
6. Conclusions
References
- Immunology Guidebook|ScienceDirect. Available online: https://www.sciencedirect.com/book/9780121983826/immunology-guidebook (accessed on 25 September 2021).
- COVID Live Update: 270,426,226 Cases and 5,321,864 Deaths from the Coronavirus—Worldometer. Available online: https://www.worldometers.info/coronavirus/ (accessed on 10 December 2021).
- Centers for Disease Control and Prevention (CDC). SARS-CoV-2 Variant Classifications and Definitions. Available online: https://www.cdc.gov/coronavirus/2019-ncov/variants/variant-info.html (accessed on 7 August 2021).
- WHO Director-General’s opening remarks at the 8th meeting of the IHR Emergency Committee on COVID-19—14 July 2021. Available online: https://www.who.int/director-general/speeches/detail/who-director-general-s-opening-remarks-at-the-8th-meeting-of-the-ihr-emergency-committee-on-covid-19-14-july-2021 (accessed on 10 December 2021).
- Coronavirus Disease 2019 (COVID-19)|CDC. Available online: https://www.cdc.gov/coronavirus/2019-ncov/index.html (accessed on 7 August 2021).
- Callaway, E. Delta coronavirus variant: Scientists brace for impact. Nature 2021, 595, 17–18.
- Li, B.; Deng, A.; Li, K.; Hu, Y.; Li, Z.; Xiong, Q.; Liu, Z.; Guo, Q.; Zou, L.; Zhang, H.; et al. Viral infection and transmission in a large, well-traced outbreak caused by the SARS-CoV-2 Delta variant. MedRxiv 2021.
- COVID-19: What Is the Mu Variant? United Nations Western Europe. Available online: https://unric.org/en/covid-19-what-is-the-mu-variant/ (accessed on 25 September 2021).
- Guruprasad, L. Human SARS CoV-2 spike protein mutations. Proteins Struct. Funct. Bioinform. 2021, 89, 569–576.
- Marshall, J.S.; Warrington, R.; Watson, W.; Kim, H.L. An introduction to immunology and immunopathology. Allergy Asthma Clin. Immunol. 2018, 14, 49.
- Abbas, A.K.; Lichtman, A.H.; Pillai, S. Cellular and Molecular Immunology; Saunders Elsevier: Philadelphia, PA, USA, 2007; p. 566.
- Doan, T.; Melvold, R.; Viselli, S. Lippincott’s Illustrated Reviews, Immunology, 2nd ed.; Wolter Kluwel: Alphen aan den Rijn, The Netherlands, 2012; ISBN 9781451109375.
- Abbas, A.K.; Lichtman, A.H.; Pillai, S. Basic Immunology: Functions and Disorders of the Immune System; Elsevier Slanders Publishing: Amsterdam, The Netherlands, 2015; ISBN 9780323400152.
- Barlow, D.J.; Edwards, M.S.; Thornton, J. Continuous and discontinuous protein antigenic determinants. Nature 1986, 322, 747–748.
- BioRender Templates. Available online: https://app.biorender.com/biorender-templates (accessed on 26 September 2021).
- Mix, E.; Goertsches, R.; Zettl, U.K. Immunoglobulins—Basic considerations. J. Neurol. 2006, 253 (Suppl. 5), V9–V17, Erratum in J. Neurol. 2008, 255, 308.
- A Compact Vocabulary of Paratope-Epitope Interactions Enables Predictability of Antibody-Antigen Binding|Elsevier Enhanced Reader. Available online: https://reader.elsevier.com/reader/sd/pii/S2211124721001704?token=74748F25258D74599D0802A9AFA03C34793008C315DF289599AE40FDBA0AF1A482C4B92C75ADC47372988E9FABB4A34B&originRegion=eu-west-1&originCreation=20210904091233 (accessed on 4 September 2021).
- Ravetch, J.V.; Bolland, S. IgG Fc Receptors. Annu. Rev. Immunol. 2001, 19, 275–290.
- Janeway, C.A., Jr.; Travers, P.; Walport, M.; Shlomchik, M.J. Immunobiology: The Immune System in Health and Disease, 5th ed.; Garland Science: New York, NY, USA, 2001; Available online: https://www.ncbi.nlm.nih.gov/books/NBK10757/ (accessed on 12 October 2021).
- Al Qaraghuli, M.M.; Kubiak-Ossowska, K.; Ferro, V.A.; Mulheran, P.A. Antibody-protein binding and conformational changes: Identifying allosteric signalling pathways to engineer a better effector response. Sci. Rep. 2020, 10, 13696.
- Introduction to Antigen-Antibody Reactions. Available online: https://microbenotes.com/introduction-to-antigen-antibody-reactions/ (accessed on 4 September 2021).
- An Introduction to Antibodies: Antibody-Antigen Interaction. Available online: https://www.sigmaaldrich.com/IN/en/technical-documents/technical-article/protein-biology/elisa/antibody-antigen-interaction (accessed on 4 September 2021).
- Roper, R.L.; Rehm, K.E. SARS vaccines: Where are we? Expert Rev. Vaccines 2009, 8, 887–898.
- Shang, W.; Yang, Y.; Rao, Y.; Rao, X. The outbreak of SARS-CoV-2 pneumonia calls for viral vaccines. NPJ Vaccines 2020, 5, 18.
- Manavalan, B.; Govindaraj, R.G.; Shin, T.H.; Kim, M.O.; Lee, G. iBCE-EL: A New Ensemble Learning Framework for Improved Linear B-Cell Epitope Prediction. Front. Immunol. 2018, 9, 1695.
- Larrañaga, P.; Calvo, B.; Santana, R.; Bielza, C.; Galdiano, J.; Inza, I.; Lozano, J.A.; Armañanzas, R.; Santafé, G.; Pérez, A.; et al. Machine learning in bioinformatics. Brief. Bioinform. 2006, 7, 86–112.
- Cunha-Neto, E.; Rosa, D.; Harris, P.; Olson, T.; Morrow, A.; Ciotlos, S.; Herst, C.V.; Rubsamen, R.M. An Approach for a Synthetic CTL Vaccine Design against Zika Flavivirus Using Class I and Class II Epitopes Identified by Computer Modeling. Front. Immunol. 2017, 8, 640.
- Liu, T.; Shi, K.; Li, W. Deep learning methods improve linear B-cell epitope prediction. BioData Min. 2020, 13, 1.
- Fatoba, A.J.; Maharaj, L.; Adeleke, V.T.; Okpeku, M.; Adeniyi, A.A.; Adeleke, M.A. Immunoinformatics prediction of overlapping CD8+ T-cell, IFN-γ and IL-4 inducer CD4+ T-cell and linear B-cell epitopes based vaccines against COVID-19 (SARS-CoV-2). Vaccine 2021, 39, 1111–1121.
- Moody, R.; Wilson, K.L.; Boer, J.C.; Holien, J.K.; Flanagan, K.L.; Jaworowski, A.; Plebanski, M. Predicted B Cell Epitopes Highlight the Potential for COVID-19 to Drive Self-Reactive Immunity. Front. Bioinform. 2021, 1, 31.
- Jespersen, M.C.; Mahajan, S.; Peters, B.; Nielsen, M.; Marcatili, P. Antibody Specific B-Cell Epitope Predictions: Leveraging Information from Antibody-Antigen Protein Complexes. Front. Immunol. 2019, 10, 298.
- Liu, L.-Y.; Yang, H.-G.; Cheng, B. Prediction of Linear B-cell Epitopes Based on PCA and RNN Network. In Proceedings of the 2019 IEEE 7th International Conference on Bioinformatics and Computational Biology (ICBCB), Hangzhou, China, 21–23 March 2019.
- Cheng, B.; Liu, L.-Y.; Qi, Z.-H.; Yang, H.-G. Prediction of Continuous B-cell Epitopes Using Long Short Term Memory Networks. In Proceedings of the 2018 6th International Conference on Bioinformatics and Computational Biology, Chengdu, China, 12–14 March 2018; pp. 55–59.
- Hu, Y.-J.; You, S.-N.; Ko, C.-L. Computational Ensemble Approach for Immune System Study: Conformational B-cell Epitope Prediction. Eur. J. Biomed. Inform. 2017, 14, 4–15.
- Ren, J.; Song, J.; Ellis, J.; Li, J. Staged heterogeneity learning to identify conformational B-cell epitopes from antigen sequences. BMC Genom. 2017, 18, 113.
- Georgios, A.; Rooman, D.M. SEPIa, a knowledge-driven algorithm for predicting conformational B-cell epitopes from the amino acid sequence. BMC Bioinform. 2017, 18, 95.
- Sher, G.; Zhi, D.; Zhang, S. DRREP: Deep ridge regressed epitope predictor. BMC Genom. 2017, 18, 676.
- Zhang, W.; Niu, Y.; Zou, H.; Luo, L.; Liu, Q.; Wu, W. Accurate Prediction of Immunogenic T-Cell Epitopes from Epitope Sequences Using the Genetic Algorithm-Based Ensemble Learning. PLoS ONE 2015, 10, e0128194.
- Zheng, W.; Zhang, C.; Hanlon, M.; Ruan, J.; Gao, J. An ensemble method for prediction of con-formational B-cell epitopes from antigen sequences. Comput. Biol. Chem. 2014, 49, 51–58.
- Zhang, J.; Zhao, X.; Sun, P.; Gao, B.; Ma, Z. Conformational B-Cell Epitopes Prediction from Sequences Using Cost-Sensitive Ensemble Classifiers and Spatial Clustering. BioMed Res. Int. 2014, 2014, 689219.
- Larsen, M.V.; Lundegaard, C.; Lamberth, K.; Buus, S.; Brunak, S.; Lund, O.; Nielsen, M. An integrative approach to CTL epitope prediction: A combined algorithm integrating MHC class I binding, TAP transport efficiency, and proteasomal cleavage predictions. Eur. J. Immunol. 2005, 35, 2295–2303.
- Reche, P.; Reinherz, E.L. Sequence Variability Analysis of Human Class I and Class II MHC Molecules: Functional and Structural Correlates of Amino Acid Polymorphisms. J. Mol. Biol. 2003, 331, 623–641.
- Zhang, L.; Udaka, K.; Mamitsuka, H.; Zhu, S. Toward more accurate pan-specific MHC-peptide binding prediction: A review of current methods and tools. Brief. Bioinform. 2012, 13, 350–364.
- Wang, P.; Sidney, J.; Dow, C.; Mothe, B.; Sette, A.; Peters, B. A systematic assessment of MHC class II peptide binding predictions and evaluation of a consensus approach. PLoS Comput. Biol. 2008, 4, e1000048.
- Alix, A.J.P. Predictive estimation of protein linear epitopes by using the program PEOPLE. Vaccine 1999, 18, 311–314.
- Pellequer, J.-L.; Westhof, E. PREDITOP: A program for antigenicity prediction. J. Mol. Graph. 1993, 11, 204–210.
- Blythe, M.J.; Flower, D.R. Benchmarking B cell epitope prediction: Underperformance of existing methods. Protein Sci. 2005, 14, 246–248.
- Saha, S.; Raghava, G.P.S. Prediction of continuous B-cell epitopes in an antigen using recurrent neural network. Proteins Struct. Funct. Bioinform. 2006, 65, 40–48.
- El-Manzalawy, Y.; Dobbs, D.; Honavar, V. Predicting linear B-cell epitopes using string kernels. J. Mol. Recognit. 2008, 21, 243–255.
- Singh, H.; Ansari, H.R.; Raghava, G.P.S. Improved Method for Linear B-Cell Epitope Prediction Using Antigen’s Primary Sequence. PLoS ONE 2013, 8, e62216.
- Yao, B.; Zhang, L.; Liang, S.; Zhang, C. SVMTriP: A Method to Predict Antigenic Epitopes Using Support Vector Machine to Integrate Tri-Peptide Similarity and Propensity. PLoS ONE 2012, 7, e45152.
- Jespersen, M.C.; Peters, B.; Nielsen, M.; Marcatili, P. BepiPred-2.0: Improving sequence-based B-cell epitope prediction using conformational epitopes. Nucleic Acids Res. 2017, 45, W24–W29.
- Greenbaum, J.A.; Andersen, P.H.; Blythe, M.; Bui, H.-H.; Cachau, R.E.; Crowe, J.; Davies, M.; Kolaskar, A.S.; Lund, O.; Morrison, S.; et al. Towards a consensus on datasets and evaluation metrics for developing B-cell epitope prediction tools. J. Mol. Recognit. 2007, 20, 75–82.
- Levitt, M. Nature of the protein universe. Proc. Natl. Acad. Sci. USA 2009, 106, 11079–11084.
- Su, S.; Wong, G.; Shi, W.; Liu, J.; Lai, A.C.K.; Zhou, J.; Liu, W.; Bi, Y.; Gao, G.F. Epidemiology, Genetic Recombination, and Pathogenesis of Coronaviruses. Trends Microbiol. 2016, 24, 490–502.
- Lineburg, K.E.; Grant, E.J.; Swaminathan, S.; Chatzileontiadou, D.S.; Szeto, C.; Sloane, H.; Panikkar, A.; Raju, J.; Crooks, P.; Rehan, S.; et al. CD8+ T cells specific for an immunodominant SARS-CoV-2 nucleocapsid epitope cross-react with selective seasonal coronaviruses. Immunity 2021, 54, 1055–1065.e5.
- Zhang, X.; Tan, Y.; Ling, Y.; Lu, G.; Liu, F.; Yi, Z.; Jia, X.; Wu, M.; Shi, B.; Xu, S.; et al. Viral and host factors related to the clinical outcome of COVID-19. Nature 2020, 583, 437–440.
- Schmidt, M.E.; Varga, S.M. The CD8 T Cell Response to Respiratory Virus Infections. Front. Immunol. 2018, 9, 678.
- Ng, O.-W.; Chia, A.; Tan, A.T.; Jadi, R.S.; Leong, H.N.; Bertoletti, A.; Tan, Y.-J. Memory T cell responses targeting the SARS coronavirus persist up to 11 years post-infection. Vaccine 2016, 34, 2008–2014.
- Channappanavar, R.; Perlman, S. Pathogenic human coronavirus infections: Causes and consequences of cytokine storm and immunopathology. Semin. Immunopathol. 2017, 39, 529–539.
- Huber, S.E.; Beek, J.E.; de Jonge, J.; Eluytjes, W.; Baarle, D.E. T Cell Responses to Viral Infections—Opportunities for Peptide Vaccination. Front. Immunol. 2014, 5, 171.
- Seder, R.A.; Darrah, P.A.; Roederer, M. T-cell quality in memory and protection: Implications for vaccine design. Nat. Rev. Immunol. 2008, 8, 247–258.
- Zhang, G.L.; DeLuca, D.S.; Keskin, D.B.; Chitkushev, L.; Zlateva, T.; Lund, O.; Reinherz, E.L.; Brusic, V. MULTIPRED2: A computational system for large-scale identification of peptides predicted to bind to HLA supertypes and alleles. J. Immunol. Methods 2011, 374, 53–61.
- Nielsen, M.; Lundegaard, C.; Worning, P.; Lauemøller, S.L.; Lamberth, K.; Buus, S.; Brunak, S.; Lund, O. Reliable prediction of T-cell epitopes using neural networks with novel sequence representations. Protein Sci. 2003, 12, 1007–1017.
- Stranzl, T.; Larsen, M.V.; Lundegaard, C.; Nielsen, M. NetCTLpan: Pan-specific MHC class I pathway epitope predictions. Immunogenetics 2010, 62, 357–368.
- Paul, S.; Croft, N.P.; Purcell, A.W.; Tscharke, D.C.; Sette, A.; Nielsen, M.; Peters, B. Benchmarking predictions of MHC class I restricted T cell epitopes in a comprehensively studied model system. PLoS Comput. Biol. 2020, 16, e1007757.
- Abelin, J.; Keskin, D.B.; Sarkizova, S.; Hartigan, C.R.; Zhang, W.; Sidney, J.; Stevens, J.; Lane, W.; Zhang, G.L.; Eisenhaure, T.M.; et al. Mass Spectrometry Profiling of HLA-Associated Peptidomes in Mono-allelic Cells Enables More Accurate Epitope Prediction. Immunity 2017, 46, 315–326.
- O’Donnell, T.J.; Rubinsteyn, A.; Bonsack, M.; Riemer, A.B.; Laserson, U.; Hammerbacher, J. MHCflurry: Open-Source Class I MHC Binding Affinity Prediction. Cell Syst. 2018, 7, 129–132.e4.
- Jensen, K.K.; Andreatta, M.; Marcatili, P.; Buus, S.; Greenbaum, J.A.; Yan, Z.; Sette, A.; Peters, B.; Nielsen, M. Improved methods for predicting peptide binding affinity to MHC class II molecules. Immunology 2018, 154, 394–406.
- Karosiene, E.; Rasmussen, M.; Blicher, T.; Lund, O.; Buus, S.; Nielsen, M. NetMHCIIpan-3.0, a common pan-specific MHC class II prediction method including all three human MHC class II isotypes, HLA-DR, HLA-DP and HLA-DQ. Immunogenetics 2013, 65, 711–724.
- Reynisson, B.; Alvarez, B.; Paul, S.; Peters, B.; Nielsen, M. NetMHCpan-4.1 and NetMHCIIpan-4.0: Improved predictions of MHC antigen presentation by concurrent motif deconvolution and integration of MS MHC eluted ligand data. Nucleic Acids Res. 2020, 48, W449–W454.
- Abelin, J.; Harjanto, D.; Malloy, M.; Suri, P.; Colson, T.; Goulding, S.P.; Creech, A.L.; Serrano, L.R.; Nasir, G.; Nasrullah, Y.; et al. Defining HLA-II Ligand Processing and Binding Rules with Mass Spectrometry Enhances Cancer Epitope Prediction. Immunity 2019, 51, 766–779.e17.
- Chen, B.; Khodadoust, M.S.; Olsson, N.; Wagar, L.; Fast, E.; Liu, C.L.; Muftuoglu, Y.; Sworder, B.; Diehn, M.; Levy, R.; et al. Predicting HLA class II antigen presentation through integrated deep learning. Nat. Biotechnol. 2019, 37, 1332–1343.
- Nielsen, M.; Lundegaard, C.; Lund, O.; Keşmir, C. The role of the proteasome in generating cytotoxic T-cell epitopes: Insights obtained from improved predictions of proteasomal cleavage. Immunogenetics 2005, 57, 33–41.
- Larsen, M.V.; Lundegaard, C.; Lamberth, K.; Buus, S.; Lund, O.; Nielsen, M. Large-scale validation of methods for cytotoxic T-lymphocyte epitope prediction. BMC Bioinform. 2007, 8, 424.
- Reche, P.A.; Glutting, J.P.; Reinherz, E.L. Prediction of MHC class I binding peptides using profile motifs. Hum. Immunol. 2002, 63, 701–709.
- Zhang, J.; Xiao, T.; Cai, Y.; Chen, B. Structure of SARS-CoV-2 spike protein. Curr. Opin. Virol. 2021, 50, 173–182.
- Crooke, S.; Ovsyannikova, I.G.; Kennedy, R.B.; Poland, G.A. Immunoinformatic identification of B cell and T cell epitopes in the SARS-CoV-2 proteome. Sci. Rep. 2020, 10, 14179.
- Bukhari, S.N.H.; Jain, A.; Haq, E. A Novel Ensemble Machine Learning Model for Prediction of Zika Virus T-Cell Epitopes. In Lecture Notes on Data Engineering and Communications Technologies; Gupta, D., Polkowski, Z., Khanna, A., Bhattacharyya, S., Castillo, O., Eds.; Springer: Singapore, 2022; Volume 91, pp. 275–292.
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273.
- The Effects of Virus Variants on COVID-19 Vaccines. Available online: https://www.who.int/news-room/feature-stories/detail/the-effects-of-virus-variants-on-covid-19-vaccines (accessed on 7 August 2021).
- Wee, L.J.; Simarmata, D.; Kam, Y.W.; Ng, L.F.; Tong, J.C. SVM-based prediction of linear B-cell epitopes using Bayes Feature Extraction. BMC Genom. 2010, 11, S21.
- Nisar, S.; Bukhari, H.; Dar, M.A. Using Random Forest to Predict T -Cell Epitopes of Dengue Virus. Dengue Virus 2021, 20, 2543–2547.
- Artificial Neural Network Disadvantages. Retrieved 4 September 2021. Available online: https://www.datascienceexamples.com/artificial-neural-network-disadvantages/ (accessed on 10 December 2021).
- Linardatos, P.; Papastefanopoulos, V.; Kotsiantis, S. Explainable AI: A Review of Machine Learning Interpretability Methods. Entropy 2021, 23, 18.
- Gagniuc, P.A.; Ionescu-Tirgoviste, C.; Gagniuc, E.; Militaru, M.; Nwabudike, L.C.; Pavaloiu, B.I.; Vasilăţeanu, A.; Goga, N.; Drăgoi, G.; Popescu, I.; et al. Spectral forecast: A general purpose prediction model as an alternative to classical neural networks. Chaos Interdiscip. J. Nonlinear Sci. 2020, 30, 033119.
- Bukhari, S.N.H.; Jain, A.; Haq, E.; Khder, M.A.; Neware, R.; Bhola, J.; Najafi, M.L. Machine Learning-Based Ensemble Model for Zika Virus T-Cell Epitope Prediction. J. Health Eng. 2021, 2021, 9591670.
- Huang, F.; Xie, G.; Xiao, R. Research on Ensemble Learning. In Proceedings of the 2009 International Conference on Artificial Intelligence and Computational Intelligence, Shanghai, China, 7–8 November 2009; Volume 3, pp. 249–252.
- A Gentle Introduction to Ensemble Learning Algorithms. Available online: https://machinelearningmastery.com/tour-of-ensemble-learning-algorithms (accessed on 8 September 2021).
- Dong, X.; Yu, Z.; Cao, W.; Shi, Y.; Ma, Q. A survey on ensemble learning. Front. Comput. Sci. 2020, 14, 241–258.
- Why Use Ensemble Learning? Available online: https://machinelearningmastery.com/why-use-ensemble-learning/ (accessed on 10 July 2021).
- Osorio, D.; Rondón-Villarreal, P.; Torres, R.T.R. Peptides: A Package for Data Mining of Antimicrobial Peptides. Small 2015, 12, 44–444.
- Hofmann, H.; Hare, E.; GGobi Foundation. Peptider: Evaluation of Diversity in Nucleotide Libraries. R Package Version 0.2.2. 2015. Available online: https://CRAN.R-project.org/package=peptider (accessed on 10 September 2021).
- Jain, P.; Chawla, P. A Novel Smart Healthcare System Design for Internet of Health Things. In Proceedings of the 2021 International Conference on Innovative Computing, Intelligent Communication and Smart Electrical Systems (ICSES), Chennai, India, 24–25 September 2021; pp. 1–8.
- Bukhari, S.N.H.; Jain, A.; Haq, E.; Mehbodniya, A.; Webber, J. Ensemble Machine Learning Model to Predict SARS-CoV-2 T-Cell Epitopes as Potential Vaccine Targets. Diagnostics 2021, 11, 1990.





