Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Bożena Bukowska | + 2510 word(s) | 2510 | 2021-12-22 05:19:58 | | | |
| 2 | Dean Liu | Meta information modification | 2510 | 2021-12-22 10:17:32 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Bukowska, B. Influence of Benzo(a)pyrene on Different Epigenetic Processes. Encyclopedia. Available online: https://encyclopedia.pub/entry/17445 (accessed on 01 June 2026).
Bukowska B. Influence of Benzo(a)pyrene on Different Epigenetic Processes. Encyclopedia. Available at: https://encyclopedia.pub/entry/17445. Accessed June 01, 2026.
Bukowska, Bożena. "Influence of Benzo(a)pyrene on Different Epigenetic Processes" Encyclopedia, https://encyclopedia.pub/entry/17445 (accessed June 01, 2026).
Bukowska, B. (2021, December 22). Influence of Benzo(a)pyrene on Different Epigenetic Processes. In Encyclopedia. https://encyclopedia.pub/entry/17445
Bukowska, Bożena. "Influence of Benzo(a)pyrene on Different Epigenetic Processes." Encyclopedia. Web. 22 December, 2021.
Copy Citation
Benzo[a]pyrene (BaP) is a polycyclic aromatic hydrocarbon (PAH) and is used as an indicator of PAHs. It is primarily produced by burning of fossil fuels, wood and other organic materials.
The mechanism of epigenetic changes induced by BaP are mainly due to the formation of CpG-BPDE adducts, between metabolite of BaP—BPDE and CpG, which leads to changes in the level of 5-methylcytosine. BaP also acts through inhibition of DNA methyltransferases activity, as well as by increasing histone deacetylases HDACs, i.e., HDAC2 and HDAC3 activity. The aim of this entry is to discuss the mechanism of the epigenetic action of BaP on the basis of the latest publications.
benzo(a)pyrene
methylation level
lung cancer
smoke
carcinogenicity
histone modifications
1.Introduction
Human exposure to BaP is usually associated with the exposure to other PAHs [1]. BaP is present in cigarette smoke [2], but also in food products, especially in products processed at high temperatures [3][4]. BaP can enter the environment; therefore, it has been found in the atmosphere [5], surface water [6] and soil [7][8]. It has been proven that food, drinking water and the air are the prevalent sources of human exposure to BaP [2][9] (Figure 1).
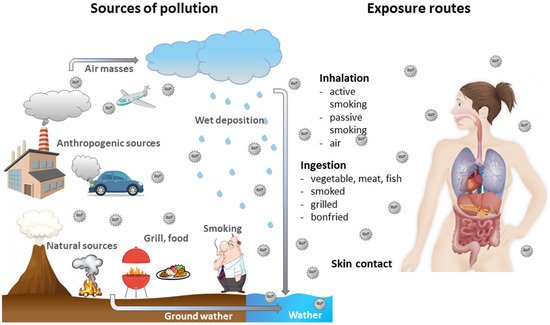
Figure 1. Sources of BaP and its movement in the environment as well as ways of entering the human body.
BaP and especially its metabolite: r-7, t-8-dihydrodiol-t-9,10-oxy-7,8,9,10-tetrahydro-benzo[a]pyrene (BPDE-I), form adducts with DNA (anti-benzo[a]pyrene-7,8-diol-9,10-oxide-DNA adducts) showing mutagenic and carcinogenic effects. BaP is a group I carcinogen [1]. BaP as a component of PAH mixtures increases the risk of cancer of the lung, skin, bladder, breast, kidney, prostate, larynx, hematopoietic system, brain and colon [9][10][11][12][13].
Epigenetic changes are one of the mechanisms responsible for adverse effects of BaP. The main mechanisms of action of BaP involve: (1) creation of stable and depurinating DNA adducts, (2) repetitive redox cycling, which generates reactive oxygen species, (3) activation of the aryl hydrocarbon receptor (AhR), (4) immunosuppression and (5) different epigenetic changes [14]. The epigenetic effect of BaP has been demonstrated in in vitro and in vivo, as well as in epidemiological studies. The ability of carcinogens to disturb epigenetic processes is one of the causes of cancer development.
2. BaP Changes Global and Gene Specific DNA Methylation
To date, numerous studies have shown in detail how the environment influences DNA methylation, leading to altered global as well as specific DNA methylation of a specific gene. These studies have focused on the exposure scenarios during the prenatal period, early life and adulthood [15].
In vitro studies have shown that BaP, a developmental and reproductive carcinogen, is an epigenetic modifier. In the early 1980s, several studies used BaP and its mutagenic metabolite—anti-7β,8α-dihydroxy-9α,10α-epoxy-7,8,9,10-tetrahydrobenzo[a]pyrene (BPDE) to study modulation of DNA methylation in vitro. BPDE was shown to bind to DNA, which resulted in the methylated DNA formation [16] and alteration of DNA methyltransferase (DNMT) [17].
Two studies have described BaP-induced hypo- and hypermethylation in in vitro cell line models [18][19]. Despite the lack of changes in expression of dnmt1, dnmt3a or dnmt13b mRNA, an increased expression of the DNMT1 protein and hypermethylation of promoter of several genes (from a panel of 30 genes analyzed) in immortalized bronchial epithelial cells was observed [20]. After treatment of immortalized bronchial epithelial cells with BPDE, the concentration of cytosine-DNA-1 methyltransferase increased, which was associated with hypermethylation of 5–10 gene promoters, including members of the cadherin gene-family [20]. However, when untransformed cells were treated with BPDE in vitro, no significant changes in methylation status were observed [21].
Studies on the effect of PAHs, and especially BaP, on global DNA methylation are quite limited and contradictory. For example, an in vitro study with TK6 cells exposed to benzo[a]fluoranthene, BaP, and benzo[a]anthracene [22], showed no global DNA methylation changes, while other studies have revealed that exposure to BaP induced global DNA hypomethylation in zebrafish (Danio rerio) embryos [23], and global DNA hypermethylation in mouse embryonic fibroblasts [24]. Similarly, primary human bronchial epithelial cell lines (16HBE cells) treated with BaP showed changes in DNA methylation, including hypermethylation and hypomethylation [20][25].
All of the above-mentioned studies have used high concentrations of BaP or BPDE. In contrast, Fang et al. [26] exposed zebrafish embryos to environmentally relevant concentrations of waterborne BaP (24 μg/L for 2.5 to 96 h) and measured both global and gene-specific DNA methylation of five developmentally important genes, namely VASA, ras-association domain family member 1 (RASSF1), telomerase reverse transcriptase (TERT), C-JUN and C-MYCA. These researchers demonstrated that BaP significantly reduced (by 44.8%) global cytosine methylation and decreased (by 17%) promoter methylation in VASA a. As a consequence, VASA expression was increased significantly (by 33%). In contrast, the exposure of zebrafish larvae to environmentally relevant concentrations of BaP did not alter methylation of CpG island nor affected gene expression in cancer genes, including ras-association domain family member 1 (RASSF1), telomerase reverse transcriptase (tert), C-JUN, and C-MYCA. Similarly, BaP made no changes in gene expression of DNMT1 and glycine N-methyltransferase (GNMT). Although total DNMT activity was not changed, the activity of the GNMT was moderately increased. Therefore, the authors suggested that BaP behaved as an epigenetic modifier of specific and global DNA methylation in zebrafish larvae.
More recently, Zhao et al. [27] investigated the mechanism of changes in the whole genomic DNA methylation of the Institute of Cancer Research (ICR) mice exposed to BaP. Blood, liver, pancreas, skin, lung, and bladder of IRC mice were removed and analyzed following a 6-h BaP exposure, and total genomic DNA methylation level was determined by high performance liquid chromatography. The results showed the tissue specificity of global DNA methylation, i.e., the total genomic DNA methylation level was significantly decreased in the blood and liver but not in the pancreas, lungs, skin and bladder of the tested animals.
The studies on the effects of paternal exposure to BaP occurring in offsprings and underlying mechanisms are very limited. Zhang et al. [28] found epigenetic changes in BaP exposed rats compared to unexposed rats. Rats in the BaP group were exposed to 0.1 mL/100 g (body weight) by intraperitoneal injection, and the BaP concentration was 0.1 mg/(kg/day) for 60 days. Rats in the control group were injected intraperitoneally with 0.5% of DMSO at 0.1 mL/100 g (body weight). Exposure to BaP resulted in hypomethylation of 3227 genes and hypermethylation of 828 genes. Kyoto Encyclopedia of Genes and Genomes pathway analysis showed that the DMGs were significantly enriched in the Ras and Rap1 signalling pathways, pancreatic secretion and neuroactive ligand-receptor interaction. Moreover, DisGeNET disease spectrum analysis showed that DMGs were associated with infertility and certain genetic diseases [28].
A growing body of human and animal studies report that exposure to BaP causes neurological abnormalities and is also associated with adverse effects, such as tumor formation, immunosuppression, teratogenicity, and hormonal disruptions. Zhang et al. [29] investigated regulatory mechanisms underlying effects of chronic BaP exposure on neurobehavioral performance in mice. The authors treated C57BL mice with various doses of BaP (1.0, 2.5, 6.25 mg/kg) dissolved in olive oil or only with olive oil. The mice were administered with BaP by intraperitoneal injections (volume 50 μL) twice a week for 12 weeks. It was found that mice that received BaP (2.5 mg/kg, 6.25 mg/kg) showed short-term memory deficits and anxiety-like behaviors. These behavioral changes were associated with downregulation of the NR2B gene and concomitant increases in DNA methylation levels in the NR2B promoter in two regions of the brain. The authors postulated that chronic exposure to BaP induced an increase in DNA methylation in the promoter of the NR2B gene and lowered NR2B expression, which may have contributed to neurotoxic effect of this compound, and the related changes in the behavior of tested animals. These results suggested that NR2B susceptibility is a target for environmental toxins in the brain.
Hypomethylation Induced by Benzo(a)pyrene and the Role of Poly(ADP-ribose) Glycohydrolase Silencing in DNA
Poly(ADP-ribose) glycohydrolase (PARG) is the primary enzyme that catalyzes the hydrolysis of poly(ADP-ribose) and participates in a number of biological processes, including the repair of DNA damage, chromatin dynamics, transcriptional regulation, and cell death. Hung et al. [25] showed that poly(ADPribosyl)ation is involved in maintenance of DNA methylation level and may prevent cancer development induced by BaP.
They used two cell lines: primary human bronchial epithelial cell lines (16HBE cells) and PARG-deficient human bronchial epithelial cell line (shPARG cell) as an in vitro model, and investigated the role of PARG silencing in DNA methylation pattern changed by BaP. They showed that BaP treatment decreased global DNA methylation level in 16HBE cells in a dose-dependent manner, but no dramatic changes were observed in shPARG cells. Further investigation revealed that PARG silencing protected DNA methyltransferases (DNMTs) activity from change by BaP exposure. These results showed an important role for PARG silencing in DNA hypomethylation induced by BaP.
3. The Intergenerational Toxic Effects on BaP Exposure Via Interference of the Circadian Rhythm
The circadian rhythm is a transcription-translation feedback oscillator loop, with a cycle of peaks and troughs in gene expression over 24 h. The circadian clock is involved in regulating life processes, such as sleep, metabolism, reproduction, development and immunity. It has been shown that circadian clock circuitry plays a role in the regulation of innate immune function, as well as in adaptive immune and allergic responses [30][31].
BaP is toxic to marine animals and their offsprings, but the underlying intergenerational immunotoxic mechanism is not clearly understood. Yin et al. [32] examined offsprings of marine medaka (Oryzias melastigma), whose parents were treated with BaP at 0.5 mg/L, and found disturbances in circadian oscillations and severe DNA damage. Many clock-related genes, such as per1 were significantly modulated in the offsprings. It was found that both PER1 and TP53 were significantly inhibited, which changed cell cycle progression and inhibited DNA repair, thus possibly resulted in increased offsprings mortality. Hypermethylation of the PER1 promoter and abnormal levels of N6-methyl adenosine (m6A) suggested that the underlying mechanism was possibly related to epigenetic modification. F1 larvae from parents exposed to BaP were more sensitive to this substance, which was due to stronger expression of genes responsible for immunological and metabolic processes. Moreover, after paternal exposure to BaP, offsprings showed more severe DNA damage and a higher degree of hypermethylation than offsprings after maternal exposure. Overall, toxic effects of BaP on parents could have been passed on to the F1 generation, and the underlying mechanism was likely related to a characteristic disturbance of the circadian rhythm [32].
4. BaP Osteotoxicity and the Regulatory Roles of Epigenetic Factors; Intergenerational Osteotoxicity in Non-Exposed F3 Generation
Recent studies have shown that ancestral exposure to BaP could cause intergenerational osteotoxicity in non-exposed F3 offsprings. Consequences of environmental contamination with BaP/PAHs, can therefore be serious and require reassessment. Mo et al. [33] in their review postulated that transgenerational inheritance of osteotoxicity in fish, caused by the exposure of ancestors to BaP, was mediated by epigenetically dysregulated processes. These researchers proposed two possible epigenetic mechanisms: (i) bone miRNAs are dysregulated via altered DNA methylation and/or histone modifications, affecting target gene expression/activity; (ii) dysregulation of bone genes through altered DNA methylation and/or histone modifications.
5. Long-Term Exposure to BaP Inhibits Expression of ERα, CYP19a and VTG1 Genes and Is Toxic to Embryos and Sex Differentiation
BaP can cause endocrine disruptions in organisms. Sun et al. [34] evaluated the effect of BaP exposure on marine medaka by assessing embryonic toxicity and analysis of reproductive genes (ERα, CYP19a, and VTG1) to predict sexual differentiation of tested animals. The results showed that high BaP concentrations (200 μg/L) significantly delayed the hatching time of embryos. Moreover, medium/high concentrations of BaP (20 μg/L and 200 μg/L) prolonged the time of sexual maturity of marine medaka. The relative expression of the ERα, cyp19a and vtg1 genes was measured at 5 days post-hatching of embryos. They showed that BaP significantly inhibited expression of genes associated with development of female fish. Consequently, after BaP exposure, there were more males in the offsprings generation. Therefore, these researchers showed that BaP could have delayed the hatching time of the embryos and extended the time of sexual maturity in tested fish. BaP has also antiestrogenic and androgenic activity, and therefore can significantly inhibit the expression of ERα, CYP19a and VTG1 genes, which are associated with female reproductive development and higher production of male offsprings by marine medaka parents exposed to this compound.
6. Exposure to BaP in Mixture of PAHs Leads to Changes in DNA Modulation and RNA (hydroxy)methylation
A study by Duca, et al. [35] focused on epigenetic changes caused by exposure to a mixture of PAHs, including BaP. Female Long Evans rats were exposed to Priority 16 PAHs (US-EPA) 3 times per week for a period of 90 days. Liver samples were used to assess the (hydroxy)methylation status of genomic DNA/RNA along with reduced and oxidized forms of glutathione. The results of this research revealed that exposure to PAHs mixture caused alterations in reduced glutathione (GSH) level and induced (hydroxy)methylation of DNA and RNA, along with DNA-PAHs adducts formation. Moreover, non-monotonic associations of the response between PAHs concentration, GSH level and DNA (hydroxy)methylation level were shown at environmentally relevant doses of studied compounds.
7. Summary
BaP is an indicator for the entire group of PAHs. BaP is mainly produced as a by-product in combustion of fossil fuels and other materials, and it is also formed as a result of other technological processes. This compound contaminates the environment and is commonly found in the atmosphere, water and soil. BaP and its metabolites are used as biomarkers of PAHs exposure both in the general population and in workers occupationally exposed to this substance. BaP enters the human organisms with consumed food, water and inhaled air. The exposure to BaP concerns numerous occupational groups. It is highly toxic, and is a group I carcinogen. It induces a number of cancers, including malignancies of the respiratory and urinary systems. It exhibits toxicity by various mechanisms, including epigenetic processes. BaP exhibits epigenotoxicity (Table 1) by disrupting the methylation processes both of the entire epigenome and of promoters of individual genes. It also affects expression of histones and triggers various miRNAs expression.
Table 1. Potential epigenetic effects induced by BaP and its metabolite BPDE.
| Type of Epigenetic Change | Observed Effects | References | |
|---|---|---|---|
| Global methylation | Increased global metylation: | Mouse embryonic fibroblast cells | [24] |
| Human bronchial epithelial cells (16HBE cells) | [20] | ||
| Mouse embryonic fibroblast cells | [24] | ||
| Normal human bronchial epithelial cells (NHBE) | [16] | ||
| Decreased global methylation: | Human bronchial epithelial cell line (16HBE) | [25] | |
| Zebrafish embryos | [23] | ||
| Zebrafish embryos | [26] | ||
| IRC mice | [27] | ||
| Children, whose mothers smoked during pregnancy | [36] | ||
| No changes in global methylation: | Human cells | [21] | |
| TK6 cells | [22] | ||
| Single gene promoters methylation and gene expression |
Increased methylation in gene promoter and decreased expression in gene promoter |
IFNγ—Jurkat cells and 53 women and childrenfrom the Columbia cohort | [37] |
| NR2B—C57BL mice | [29] | ||
| H19—302 reproductive-aged males (22–46 years old) | [38] | ||
| PER1—medaka fish Oryzias melastigma | [32] | ||
| Biotinidase and holocarboxylase synthetaseHuman Bronchial Epithelial Cells (16HBE) | [39] | ||
| 828 hypermethylated genes in rats | [28] | ||
| Decreased methylation in gene promoter |
3 227 hypomethylated genes in rats | [28] | |
| Decreased gene expression | ERα, CYP19a and VTG1 expression—Oryzias Melastigma | [34] | |
| Histone modifications | Reduction in acetylation levels on H3 and H4 histones |
Human bronchial epithelial cells (16HBE) | [39] |
| Increase in histone deacetylases HDAC2 and HDAC3 | Human bronchial epithelial cells (16HBE) | [39] | |
| Changes in H3K9 histone acetylation |
Breast cancer cells (MCF7) | [40] | |
| Changes in miRNA level |
Decreased expression | miR-506 in cancer cells (16HBE-T) | [41] |
| Increased expression | miR-638 in breast cancer (BRCA1) | [42] | |
| mikroRNA-29b, mikroRNA-26a-1* i mikroRNA-122* (HepG2 cells) | [43] | ||
References
- IARC Working Groups Benzopyrene. Available online: https://monographs.iarc.who.int/wp-content/uploads/2018/06/mono100F-14.pdf (accessed on 14 December 2021).
- Susanto, A.D.; Yusril, N.; Zaini, J.; Nuwidya, F. Comparison of Serum Benzo(a)pyrene Diol Epoxide—Protein Adducts Level between Kretek Cigarette Smokers and Nonsmokers and the Related Factors. J. Nat. Sci. Biol. Med. 2021, 12, 52.
- Siddique, R.; Zahoor, A.F.; Ahmad, H.; Zahid, F.M.; Karrar, E. Impact of different cooking methods on polycyclic aromatic hydrocarbons in rabbit meat. Food Sci. Nutr. 2021, 9, 3219–3227.
- Migdał, W.; Walczycka, M.; Migdał, Ł. The Levels of Polycyclic Aromatic Hydrocarbons in Traditionally Smoked Cheeses in Poland. Polycycl. Aromat. Compd. 2020, 1–13.
- Schreiberová, M.; Vlasáková, L.; Vlček, O.; Šmejdířová, J.; Horálek, J.; Bieser, J. Benzopyrene in the Ambient Air in the Czech Republic: Emission Sources, Current and Long-Term Monitoring Analysis and Human Exposure. Atmosphere 2020, 11, 955.
- He, Y.; Yang, C.; He, W.; Xu, F. Nationwide health risk assessment of juvenile exposure to polycyclic aromatic hydrocarbons (PAHs) in the water body of Chinese lakes. Sci. Total Environ. 2020, 723, 138099.
- Cao, H.; Wang, C.; Liu, H.; Jia, W.; Sun, H. Enzyme activities during Benzopyrene degradation by the fungus Lasiodiplodia theobromae isolated from a polluted soil. Sci. Rep. 2020, 10, 865.
- Olgun, B.; Doğan, G. Polycyclic aromatic hydrocarbon concentrations in soils of greenhouses located in Aksu Antalya, Turkey. Water Sci. Technol. 2020, 81, 283–292.
- Amadou, A.; Praud, D.; Coudon, T.; Deygas, F.; Grassot, L.; Faure, E.; Couvidat, F.; Caudeville, J.; Bessagnet, B.; Salizzoni, P.; et al. Risk of breast cancer associated with long-term exposure to benzopyrene (BaP) air pollution: Evidence from the French E3N cohort study. Environ. Int. 2021, 149, 106399.
- Kim, K.E.; Cho, D.; Park, H.J. Air pollution and skin diseases: Adverse effects of airborne particulate matter on various skin diseases. Life Sci. 2016, 152, 126–134.
- Korsh, J.; Shen, A.; Aliano, K.; Davenport, T. Polycyclic Aromatic Hydrocarbons and Breast Cancer: A Review of the Literature. Breast Care 2015, 10, 316–318.
- Hamidi, E.N.; Hajeb, P.; Selamat, J.; Razis, A.F.A. Polycyclic Aromatic Hydrocarbons (PAHs) and their Bioaccessibility in Meat: A Tool for Assessing Human Cancer Risk. Asian Pac. J. Cancer Prev. 2016, 17, 15–23.
- Petit, P.; Maître, A.; Persoons, R.; Bicout, D.J. Lung cancer risk assessment for workers exposed to polycyclic aromatic hydrocarbons in various industries. Environ. Int. 2019, 124, 109–120.
- Birkett, N.; Al-Zoughool, M.; Bird, M.; Baan, R.A.; Zielinski, J.; Krewski, D. Overview of biological mechanisms of human carcinogens. J. Toxicol. Environ. Health Part B 2019, 22, 288–359.
- Martin, E.M.; Fry, R.C. Environmental Influences on the Epigenome: Exposure- Associated DNA Methylation in Human Populations. Annu. Rev. Public Health 2018, 39, 309–333.
- Hu, W.; Feng, Z.; Tang, M.-S. Preferential Carcinogen−DNA Adduct Formation at Codons 12 and 14 in the Human K-ras Gene and Their Possible Mechanisms. Biochemistry 2003, 42, 10012–10023.
- Wojciechowski, M.F.; Meehan, T. Inhibition of DNA methyltransferases in vitro by benzopyrene diol epoxide-modified substrates. J. Biol. Chem. 1984, 259, 9711–9716.
- Sadikovic, B.; Haines, T.R.; Butcher, D.T.; Rodenhiser, D.I. Chemically induced DNA hypomethylation in breast carcinoma cells detected by the amplification of intermethylated sites. Breast Cancer Res. 2004, 6, R329–R337.
- Sadikovic, B.; Rodenhiser, D.I. Benzopyrene exposure disrupts DNA methylation and growth dynamics in breast cancer cells. Toxicol. Appl. Pharmacol. 2006, 216, 458–468.
- Damiani, L.A.; Yingling, C.M.; Leng, S.; Romo, P.E.; Nakamura, J.; Belinsky, S.A. Carcinogen-Induced Gene Promoter Hypermethylation Is Mediated by DNMT1 and Causal for Transformation of Immortalized Bronchial Epithelial Cells. Cancer Res. 2008, 68, 9005–9014.
- Tommasi, S.; Kim, S.-I.; Zhong, X.; Wu, X.; Pfeifer, G.P.; Besaratinia, A. Investigating the Epigenetic Effects of a Prototype Smoke-Derived Carcinogen in Human Cells. PLoS ONE 2010, 5, e10594.
- Tabish, A.M.; Poels, K.; Hoet, P.; Godderis, L. Epigenetic Factors in Cancer Risk: Effect of Chemical Carcinogens on Global DNA Methylation Pattern in Human TK6 Cells. PLoS ONE 2012, 7, e34674.
- Corrales, J.; Fang, X.; Thornton, C.; Mei, W.; Barbazuk, W.; Duke, M.; Scheffler, B.; Willett, K. Effects on specific promoter DNA methylation in zebrafish embryos and larvae following benzopyrene exposure. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2014, 163, 37–46.
- Yauk, C.L.; Polyzos, A.; Rowan-Carroll, A.; Kortubash, I.; Williams, A.; Kovalchuk, O. Tandem repeat mutation, global DNA methylation, and regulation of DNA methyltransferases in cultured mouse embryonic fibroblast cells chronically exposed to chemicals with different modes of action. Environ. Mol. Mutagen. 2008, 49, 26–35.
- Huang, H.; Hu, G.; Cai, J.; Xia, B.; Liu, J.; Li, X.; Gao, W.; Zhang, J.; Liu, Y.; Zhuang, Z. Role of poly(ADP-ribose) glycohydrolase silencing in DNA hypomethylation induced by benzo(a)pyrene. Biochem. Biophys. Res. Commun. 2014, 452, 708–714.
- Fang, X.; Thornton, C.; Scheffler, B.E.; Willett, K.L. Benzopyrene decreases global and gene specific DNA methylation during zebrafish development. Environ. Toxicol. Pharmacol. 2013, 36, 40–50.
- Zhao, L.; Zhang, S.; An, X.; Tan, W.; Pang, D.; Ouyang, H. Toxicological effects of benzopyrene on DNA methylation of whole genome in ICR mice. Cell. Mol. Boil. 2015, 61, 115–119.
- Zhang, C.M.; Sun, Z.X.; Wang, Z.L.; Chen, J.S.; Chang, Z.; Zhu, L.; Ma, Z.H.; Peng, Y.J.; Xu, Z.A.; Wang, S.Q. Abnormal methylation of spermatozoa induced by benzo(a)pyrene in rats. Hum. Exp. Toxicol. 2019, 38, 846–856.
- Zhang, W.; Tian, F.; Zheng, J.; Li, S.; Qiang, M. Chronic Administration of Benzo(a)pyrene Induces Memory Impairment and Anxiety-Like Behavior and Increases of NR2B DNA Methylation. PLoS ONE 2016, 11, e0149574.
- Krishnaiah, S.Y.; Wu, G.; Altman, B.; Growe, J.; Rhoades, S.D.; Coldren, F.; Venkataraman, A.; Olarerin-George, A.O.; Francey, L.J.; Mukherjee, S.; et al. Clock Regulation of Metabolites Reveals Coupling between Transcription and Metabolism. Cell Metab. 2017, 25, 961–974.e4.
- Patke, A.; Murphy, P.J.; Onat, O.E.; Krieger, A.C.; Özçelik, T.; Campbell, S.S.; Young, M.W. Mutation of the Human Circadian Clock Gene CRY1 in Familial Delayed Sleep Phase Disorder. Cell 2017, 169, 203–215.e13.
- Yin, X.-H.; Liu, Y.; Zeb, R.; Chen, F.-Y.; Chen, H.-Y.; Wang, K.-J. The intergenerational toxic effects on offspring of medaka fish Oryzias melastigma from parental benzopyrene exposure via interference of the circadian rhythm. Environ. Pollut. 2020, 267, 115437.
- Mo, J.; Au, D.W.-T.; Guo, J.; Winkler, C.; Kong, R.Y.-C.; Seemann, F. Benzopyrene osteotoxicity and the regulatory roles of genetic and epigenetic factors: A review. Crit. Rev. Environ. Sci. Technol. 2021, 1–39.
- Sun, D.; Chen, Q.; Zhu, B.; Lan, Y.; Duan, S. Long-Term Exposure to BenzoPyrene Affects Sexual Differentiation and Embryos Toxicity in Three Generations of Marine Medaka (Oryzias Melastigma). Int. J. Environ. Res. Public Health 2020, 17, 970.
- Duca, R.-C.; Grova, N.; Ghosh, M.; Do, J.-M.; Hoet, P.H.M.; Vanoirbeek, J.A.J.; Appenzeller, B.M.R.; Godderis, L. Exposure to Polycyclic Aromatic Hydrocarbons Leads to Non-monotonic Modulation of DNA and RNA (hydroxy)methylation in a Rat Model. Sci. Rep. 2018, 8, 10577.
- Joubert, B.; Håberg, S.E.; Nilsen, R.M.; Wang, X.; Vollset, S.E.; Murphy, S.; Huang, Z.; Hoyo, C.; Midttun, Ø.; Uicab, L.C.; et al. 450K Epigenome-Wide Scan Identifies Differential DNA Methylation in Newborns Related to Maternal Smoking during Pregnancy. Environ. Health Perspect. 2012, 120, 1425–1431.
- Tang, W.-Y.; Levin, L.; Talaska, G.; Cheung, Y.Y.; Herbstman, J.; Tang, D.; Miller, R.L.; Perera, F.; Ho, S.-M. Maternal Exposure to Polycyclic Aromatic Hydrocarbons and 5’-CpG Methylation of Interferon-γ in Cord White Blood Cells. Environ. Health Perspect. 2012, 120, 1195–1200.
- Yang, J.; Lu, Z.; Liu, Z.; Wang, L.; Qiang, M. Methylation of Imprinted Genes in Sperm DNA Correlated to Urinary Polycyclic Aromatic Hydrocarbons (PAHs) Exposure Levels in Reproductive-Aged Men and the Birth Outcomes of the Offspring. Front. Genet. 2021, 11, 1706.
- Xia, B.; Yang, L.Q.; Huang, H.Y.; Pang, L.; Yang, X.F.; Yi, Y.J.; Ren, X.H.; Li, J.; Zhuang, Z.X.; Liu, J.J. Repression of Bio-tin-Related Proteins by BenzoPyrene-Induced Epigenetic Modifications in Human Bronchial Epithelial Cells. Int. J. Toxicol. 2016, 35, 336–343.
- Sadikovic, B.; Andrews, J.; Carter, D.; Robinson, J.; Rodenhiser, D.I. Genome-wide H3K9 Histone Acetylation Profiles Are Altered in Benzopyrene-treated MCF7 Breast Cancer Cells. J. Biol. Chem. 2008, 283, 4051–4060.
- Zhao, Y.; Liu, H.; Li, Y.; Wu, J.; Greenlee, A.R.; Yang, C.; Jiang, Y. The role of miR-506 in transformed 16HBE cells induced by anti-benzopyrene-trans-7,8-dihydrodiol-9,10-epoxide. Toxicol. Lett. 2011, 205, 320–326.
- Li, D.; Wang, Q.; Liu, C.; Duan, H.; Zeng, X.; Zhang, B.; Li, X.; Zhao, J.; Tang, S.; Li, Z.; et al. Aberrant Expression of miR-638 Contributes to Benzo(a)pyrene-Induced Human Cell Transformation. Toxicol. Sci. 2012, 125, 382–391.
- Lizárraga, D.; Gaj, S.; Brauers, K.J.; Timmermans, L.; Kleinjans, J.C.; Van Delft, J.H.M. Benzopyrene-Induced Changes in MicroRNA–mRNA Networks. Chem. Res. Toxicol. 2012, 25, 838–849.
More
Information
Subjects:
Environmental Sciences
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.8K
Entry Collection:
Environmental Sciences
Revisions:
2 times
(View History)
Update Date:
22 Dec 2021
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





