
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Daniel Guerra | + 1682 word(s) | 1682 | 2021-11-16 05:11:38 | | | |
| 2 | Yvaine Wei | Meta information modification | 1682 | 2021-11-18 05:06:01 | | | | |
| 3 | Rwik Sen | + 22 word(s) | 1704 | 2021-11-22 21:18:11 | | |
Video Upload Options
COVID-19 is caused by a coronavirus called severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). The difficulty in containing SARS-CoV-2 has underscored the need for techniques such as mass spectrometry in the diagnosis and treatment of COVID-19. Mass spectrometry-based methods have been employed in several studies to detect changes in interactions among host proteins, and between host and viral proteins in COVID-19 patients. The methods have also been used to characterize host and viral proteins, and analyze lipid metabolism in COVID-19 patients. Information obtained using the above methods are complemented by high-throughput analysis of transcriptomic and epigenomic changes associated with COVID-19, coupled with next-generation sequencing.
1. Proteomics and Mass Spectrometry in Coronavirus Disease 2019 (COVID-19)
1.1. COVID-19-Linked Host Protein Characterization Discovered through Mass Spectrometric Proteomics
1.2. Proteomics in COVID-19 Treatment Identification
2. Lipidomics and Mass Spectrometry in COVID-19
2.1. The Immunomodulatory Axis of COVID-19 Disease Burden
2.3. Mass Spectrometry-Based Studies on Spike Protein Helps in Vaccine Development
2.4. Signaling Pattern-Recognition Receptors (PRRs) Are Found on Multiple Host-Cell Surfaces
2.5. Sphingolipids at the Center of COVID-19 Infection Dynamics
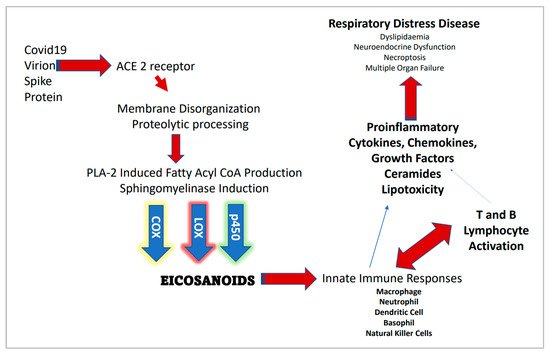
2.6. Contributions of Lipidomics in Understanding and Treating COVID-19
References
- Jannetto, P.J.; Danso, D. Mass spectrometry. Clin. Biochem. 2020, 82, 1.
- Beccaria, M.; Cabooter, D. Current developments in LC-MS for pharmaceutical analysis. Analyst 2020, 145, 1129–1157.
- Seger, C.; Salzmann, L. After another decade: LC–MS/MS became routine in clinical diagnostics. Clin. Biochem. 2020, 82, 2–11.
- Nikolaev, E.N.; Indeykina, M.I.; Brzhozovskiy, A.G.; Bugrova, A.E.; Kononikhin, A.S.; Starodubtseva, N.L.; Petrotchenko, E.V.; Kovalev, G.I.; Borchers, C.H.; Sukhikh, G.T. Mass-Spectrometric Detection of SARS-CoV-2 Virus in Scrapings of the Epithelium of the Nasopharynx of Infected Patients via Nucleocapsid N Protein. J. Proteome Res. 2020, 19, 4393–4397.
- Rybicka, M.; Milosz, E.; Bielawski, K.P. Superiority of MALDI-TOF Mass Spectrometry over Real-Time PCR for SARS-CoV-2 RNA Detection. Viruses 2021, 13, 730.
- Hober, A.; Hua, T.M.K.; Foley, D.; McDonald, T.; Vissers, J.P.; Pattison, R.; Ferries, S.; Hermansson, S.; Betner, I.; Uhlen, M.; et al. Rapid and Sensitive Detection of SARS-CoV-2 Infection Using Quantitative Peptide Enrichment LC-MS/MS Analysis. medRxiv 2021.
- Wang, W.; Li, Q.; Huang, G.; Lin, B.-Y.; Lin, D.-Z.; Ma, Y.; Zhang, Z.; Chen, T.; Zhou, J. Tandem Mass Tag-Based Proteomic Analysis of Potential Biomarkers for Hepatocellular Carcinoma Differentiation. OncoTargets Ther. 2021, 14, 1007–1020.
- Shen, B.; Yi, X.; Sun, Y.; Bi, X.; Du, J.; Zhang, C.; Quan, S.; Zhang, F.; Sun, R.; Qian, L.; et al. Proteomic and Metabolomic Characterization of COVID-19 Patient Sera. Cell 2020, 182, 59–72.e15.
- Gordon, D.E.; Jang, G.M.; Bouhaddou, M.; Xu, J.; Obernier, K.; White, K.M.; O’Meara, M.J.; Rezelj, V.V.; Guo, J.Z.; Swaney, D.L.; et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 2020, 583, 459–468.
- Suvarna, K.; Biswas, D.; Pai, M.G.J.; Acharjee, A.; Bankar, R.; Palanivel, V.; Salkar, A.; Verma, A.; Mukherjee, A.; Choudhury, M.; et al. Proteomics and Machine Learning Approaches Reveal a Set of Prognostic Markers for COVID-19 Severity With Drug Repurposing Potential. Front. Physiol. 2021, 12, 652799.
- Chilosi, M.; Poletti, V.; Ravaglia, C.; Rossi, G.; Dubini, A.; Piciucchi, S.; Pedica, F.; Bronte, V.; Pizzolo, G.; Martignoni, G.; et al. The pathogenic role of epithelial and endothelial cells in early-phase COVID-19 pneumonia: Victims and partners in crime. Mod. Pathol. 2021, 34, 1444–1455.
- Chua, R.L.; Lukassen, S.; Trump, S.; Hennig, B.P.; Wendisch, D.; Pott, F.; Debnath, O.; Thürmann, L.; Kurth, F.; Völker, M.T.; et al. COVID-19 severity correlates with airway epithelium–immune cell interactions identified by single-cell analysis. Nat. Biotechnol. 2020, 38, 970–979.
- Hammock, B.D.; Wang, W.; Gilligan, M.M.; Panigrahyyz, D. Eicosanoids: The Overlooked Storm in Coronavirus Disease 2019 (COVID-19)? Am. J. Pathol. 2020, 190, 1782–1788.
- Penttilä, P.A.; van Gassen, S.; Panovska, D.; Vanderbeke, L.; van Herck, Y.; Quintelier, K.; Emmaneel, A.; Filtjens, J.; Malengier-Devlies, B.; Ahmadzadeh, K.; et al. High dimensional profiling identifies specific immune types along the recovery trajectories of critically ill COVID19 patients. Experientia 2021, 78, 3987–4002.
- Sichitiu, J.; Fakhouri, F.; Desseauve, D. Antenatal corticosteroid therapy and COVID-19: Pathophysiological considerations. Acta Obstet. Gynecol. Scand. 2020, 99, 952.
- Praissman, J.; Wells, L. Proteomics-Based Insights Into the SARS-CoV-2-Mediated COVID-19 Pandemic: A Review of the First Year of Research. Mol. Cell. Proteom. 2021, 20, 100103.
- Miller, L.M.; Barnes, L.F.; Raab, S.A.; Draper, B.E.; El-Baba, T.J.; Lutomski, C.A.; Robinson, C.V.; Clemmer, D.E.; Jarrold, M.F. Heterogeneity of Glycan Processing on Trimeric SARS-CoV-2 Spike Protein Revealed by Charge Detection Mass Spectrometry. J. Am. Chem. Soc. 2021, 143, 3959–3966.
- Roberts, D.S.; Mann, M.W.; Melby, J.A.; Larson, E.J.; Zhu, Y.; Brasier, A.R.; Jin, S.; Ge, Y. Structural O-Glycoform Heterogeneity of the SARS-CoV-2 Spike Protein Receptor-Binding Domain Revealed by Top-Down Mass Spectrometry. J. Am. Chem. Soc. 2021, 143, 12014–12024.
- Cho, B.G.; Gautam, S.; Peng, W.; Huang, Y.; Goli, M.; Mechref, Y. Direct Comparison of N-Glycans and Their Isomers Derived from Spike Glycoprotein 1 of MERS-CoV, SARS-CoV-1, and SARS-CoV-2. J. Proteome Res. 2021, 20, 4357–4365.
- Antonopoulos, A.; Broome, S.; Sharov, V.; Ziegenfuss, C.; Easton, R.L.; Panico, M.; Dell, A.; Morris, H.R.; Haslam, S.M. Site-specific characterization of SARS-CoV-2 spike glycoprotein receptor-binding domain. Glycobiology 2021, 31, 181–187.
- Witkowska, D. Mass Spectrometry and Structural Biology Techniques in the Studies on the Coronavirus-Receptor Interaction. Molecules 2020, 25, 4133.
- Krishnan, S.; Krishnan, G.P. N-Glycosylation Network Construction and Analysis to Modify Glycans on the Spike (S) Glycoprotein of SARS-CoV-2. Front. Bioinform. 2021, 1.
- Watanabe, Y.; Berndsen, Z.T.; Raghwani, J.; Seabright, G.E.; Allen, J.D.; Pybus, O.; McLellan, J.S.; Wilson, I.A.; Bowden, T.A.; Ward, A.B.; et al. Vulnerabilities in coronavirus glycan shields despite extensive glycosylation. Nat. Commun. 2020, 11, 2688.
- Casalino, L.; Gaieb, Z.; Goldsmith, J.A.; Hjorth, C.K.; Dommer, A.C.; Harbison, A.M.; Fogarty, C.A.; Barros, E.P.; Taylor, B.C.; McLellan, J.S.; et al. Beyond Shielding: The Roles of Glycans in the SARS-CoV-2 Spike Protein. ACS Cent. Sci. 2020, 6, 1722–1734.
- Petrović, T.; Alves, I.; Bugada, D.; Pascual, J.; Vučković, F.; Skelin, A.; Gaifem, J.; Villar-Garcia, J.; Vicente, M.M.; Fernandes, Â.; et al. Composition of the immunoglobulin G glycome associates with the severity of COVID-19. Glycobiology 2020, 31, 372–377.
- Trkulja, V.; Kodvanj, I.; Homolak, J. Immunoglobulin G glycome and severity of COVID-19: More likely a quantification of bias than a true association. A comment on Petrović et al., “Composition of the immunoglobulin G glycome associates with the severity of COVID-19”. Glycobiology 2021, 31, 713–716.
- Hou, H.; Yang, H.; Liu, P.; Huang, C.; Wang, M.; Li, Y.; Zhu, M.; Wang, J.; Xu, Y.; Wang, Y.; et al. Profile of Immunoglobulin G N-Glycome in COVID-19 Patients: A Case-Control Study. Front. Immunol. 2021, 12, 748566.
- Rosenbalm, K.E.; Tiemeyer, M.; Wells, L.; Aoki, K.; Zhao, P. Glycomics-informed glycoproteomic analysis of site-specific glycosylation for SARS-CoV-2 spike protein. STAR Protoc. 2020, 1, 100214.
- Grant, O.C.; Montgomery, D.; Ito, K.; Woods, R.J. Analysis of the SARS-CoV-2 spike protein glycan shield: Implications for immune recognition. Sci. Rep. 2020, 10, 14991.
- Zhao, P.; Praissman, J.L.; Grant, O.C.; Cai, Y.; Xiao, T.; Rosenbalm, K.E.; Aoki, K.; Kellman, B.P.; Bridger, R.; Barouch, D.H.; et al. Virus-Receptor Interactions of Glycosylated SARS-CoV-2 Spike and Human ACE2 Receptor. Cell Host Microbe 2020, 28, 586–601.e6.
- Zhao, X.; Chen, H.; Wang, H. Glycans of SARS-CoV-2 Spike Protein in Virus Infection and Antibody Production. Front. Mol. Biosci. 2021, 8, 629873.
- Wang, Y.; Wu, Z.; Hu, W.; Hao, P.; Yang, S. Impact of Expressing Cells on Glycosylation and Glycan of the SARS-CoV-2 Spike Glycoprotein. ACS Omega 2021, 6, 15988–15999.
- Kasuga, Y.; Zhu, B.; Jang, K.-J.; Yoo, J.-S. Innate immune sensing of coronavirus and viral evasion strategies. Exp. Mol. Med. 2021, 53, 723–736.
- Bieberich, E. Sphingolipids and lipid rafts: Novel concepts and methods of analysis. Chem. Phys. Lipids 2018, 216, 114–131.
- Törnquist, K.; Asghar, M.Y.; Srinivasan, V.; Korhonen, L.; Lindholm, D. Sphingolipids as Modulators of SARS-CoV-2 Infection. Front. Cell Dev. Biol. 2021, 9, 689854.
- Caterino, M.; Gelzo, M.; Sol, S.; Fedele, R.; Annunziata, A.; Calabrese, C.; Fiorentino, G.; D’Abbraccio, M.; Dell’Isola, C.; Fusco, F.M.; et al. Dysregulation of lipid metabolism and pathological inflammation in patients with COVID-19. Sci. Rep. 2021, 11, 2941.
- Meoni, G.; Ghini, V.; Maggi, L.; Vignoli, A.; Mazzoni, A.; Salvati, L.; Capone, M.; Vanni, A.; Tenori, L.; Fontanari, P.; et al. Metabolomic/lipidomic profiling of COVID-19 and individual response to tocilizumab. PLoS Pathog. 2021, 17, e1009243.
- Spick, M.; Longman, K.; Frampas, C.; Lewis, H.; Costa, C.; Walters, D.D.; Stewart, A.; Wilde, M.; Greener, D.; Evetts, G.; et al. Changes to the sebum lipidome upon COVID-19 infection observed via rapid sampling from the skin. EClinicalMedicine 2021, 33, 100786.
- Bai, Y.; Huang, W.; Li, Y.; Lai, C.; Huang, S.; Wang, G.; He, Y.; Hu, L.; Chen, C. Lipidomic alteration of plasma in cured COVID-19 patients using ultra high-performance liquid chromatography with high-resolution mass spectrometry. Biosci. Rep. 2021, 41, BSR20204305.
- Bruzzone, C.; Bizkarguenaga, M.; Gil-Redondo, R.; Diercks, T.; Arana, E.; de Vicuña, A.G.; Seco, M.; Bosch, A.; Palazón, A.; San Juan, I.; et al. SARS-CoV-2 Infection Dysregulates the Metabolomic and Lipidomic Profiles of Serum. iScience 2020, 23, 101645.





