
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Veronica Dodero | + 2162 word(s) | 2162 | 2021-09-05 20:05:39 | | | |
| 2 | Jessie Wu | Meta information modification | 2162 | 2021-09-13 04:14:53 | | | | |
| 3 | Conner Chen | Meta information modification | 2162 | 2021-10-12 10:54:53 | | |
Video Upload Options
Gluten-related disorders (GRDs) are a group of diseases that involve the activation of the immune system triggered by the ingestion of gluten, with a worldwide prevalence of 5%. Among them, Celiac disease (CeD) is a T-cell-mediated autoimmune disease causing a plethora of symptoms from diarrhea and malabsorption to lymphoma. Even though GRDs have been intensively studied, the environmental triggers promoting the diverse reactions to gluten proteins in susceptible individuals remain elusive. It has been proposed that pathogens could act as disease-causing environmental triggers of CeD by molecular mimicry mechanisms. Additionally, it could also be possible that unrecognized molecular, structural, and physical parallels between gluten and bacteria have a relevant role.
1. Introduction
2. Primary Structure Analysis and Potential Functions
2.1. CeD-T-Cell Epitopes
2.2. SH3/WW Domains Binders
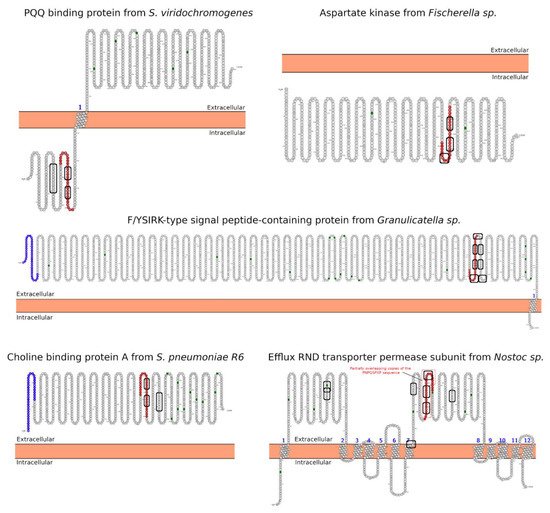
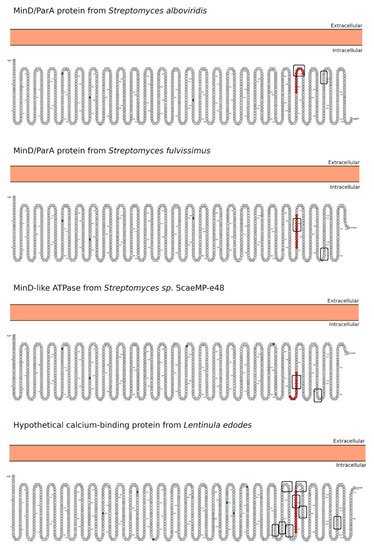
3. Morphology Mimicry
References
- Cabanillas, B. Gluten-related disorders: Celiac disease, wheat allergy, and nonceliac gluten sensitivity. Crit. Rev. Food Sci. Nutr. 2020, 60, 2606–2621.
- Taraghikhah, N.; Ashtari, S.; Asri, N.; Shahbazkhani, B.; Al-Dulaimi, D.; Rostami-Nejad, M.; Rezaei-Tavirani, M.; Razzaghi, M.R.; Zali, M.R. An updated overview of spectrum of gluten-related disorders: Clinical and diagnostic aspects. BMC Gastroenterol. 2020, 20, 258.
- Di Sabatino, A.; Corazza, G.R. Coeliac disease. Lancet 2009, 373, 1480–1493.
- Shan, L.; Molberg, Ø.; Parrot, I.; Hausch, F.; Filiz, F.; Gray, G.M.; Sollid, L.M.; Khosla, C. Structural Basis for Gluten Intolerance in Celiac Sprue. Science 2002, 297, 2275–2279.
- Petersen, J.; Ciacchi, L.; Tran, M.T.; Loh, K.L.; Kooy-Winkelaar, Y.; Croft, N.P.; Hardy, M.I.; Chen, Z.; McCluskey, J.; Anderson, R.P.; et al. T cell receptor cross-reactivity between gliadin and bacterial peptides in celiac disease. Nat. Struct. Mol. Biol. 2020, 27, 49–61.
- Bethune, M.T.; Khosla, C. Parallels between Pathogens and Gluten Peptides in Celiac Sprue. PLoS Pathog. 2008, 4, e34.
- Verdu, E.F.; Schuppan, D. The enemy within the gut: Bacterial pathogens in celiac autoimmunity. Nat. Struct. Mol. Biol. 2019, 27, 5–7.
- Lammers, K.M.; Herrera, M.G.; Dodero, V.I. Translational Chemistry Meets Gluten-Related Disorders. ChemistryOpen 2018, 7, 217–232.
- Doshi, N.; Mitragotri, S. Macrophages Recognize Size and Shape of Their Targets. PLoS ONE 2010, 5, e10051.
- Swartzwelter, B.; Fux, A.; Johnson, L.; Swart, E.; Hofer, S.; Hofstätter, N.; Geppert, M.; Italiani, P.; Boraschi, D.; Duschl, A.; et al. The Impact of Nanoparticles on Innate Immune Activation by Live Bacteria. Int. J. Mol. Sci. 2020, 21, 9695.
- Herrera, M.G.; Nicoletti, F.; Gras, M.; Dörfler, P.W.; Tonali, N.; Hannappel, Y.; Ennen, I.; Hütten, A.; Hellweg, T.; Lammers, K.M.; et al. Pepsin Digest of Gliadin Forms Spontaneously Amyloid-Like Nanostructures Influencing the Expression of Selected Pro-Inflammatory, Chemoattractant, and Apoptotic Genes in Caco-2 Cells: Implications for Gluten-Related Disorders. Mol. Nutr. Food Res. 2021, 65, 2100200.
- Maiuri, L.; Ciacci, C.; Ricciardelli, I.; Vacca, L.; Raia, V.; Auricchio, S.; Picard, J.; Osman, M.; Quaratino, S.; Londei, M. Association between innate response to gliadin and activation of pathogenic T cells in coeliac disease. Lancet 2003, 362, 30–37.
- Londei, M.; Ciacci, C.; Ricciardelli, I.; Vacca, L.; Quaratino, S.; Maiuri, L. Gliadin as a stimulator of innate responses in celiac disease. Mol. Immunol. 2005, 42, 913–918.
- Qiao, S.-W.; Bergseng, E.; Molberg, Ø.; Xia, J.; Fleckenstein, B.; Khosla, C.; Sollid, L.M. Antigen Presentation to Celiac Lesion-Derived T Cells of a 33-Mer Gliadin Peptide Naturally Formed by Gastrointestinal Digestion. J. Immunol. 2004, 173, 1757–1762.
- Fraser, J.S.; Engel, W.; Ellis, H.J.; Moodie, S.J.; Pollock, E.L.; Wieser, H.; Ciclitira, P.J. Coeliac disease: In vivo toxicity of the putative immunodominant epitope. Gut 2003, 52, 1698–1702.
- Sollid, L.M. Intraepithelial Lymphocytes in Celiac Disease: License to Kill Revealed. Immunity 2004, 21, 303–304.
- Herrera, M.G.; Zamarreño, F.; Costabel, M.; Ritacco, H.; Hütten, A.; Sewald, N.; Dodero, V.I. Circular dichroism and electron microscopy studies in vitro of 33-mer gliadin peptide revealed secondary structure transition and supramolecular organization. Biopolymers 2014, 101, 96–106.
- Herrera, M.G.; Benedini, L.; Lonez, C.; Schilardi, P.L.; Hellweg, T.; Ruysschaert, J.-M.; Dodero, V.I. Self-assembly of 33-mer gliadin peptide oligomers. Soft Matter 2015, 11, 8648–8660.
- Herrera, M.; Vazquez, D.; Sreij, R.; Drechsler, M.; Hertle, Y.; Hellweg, T.; Dodero, V. Insights into gliadin supramolecular organization at digestive pH 3.0. Colloids Surf. B Biointerfaces 2018, 165, 363–370.
- Herrera, M.G.; Veuthey, T.V.; Dodero, V.I. Self-organization of gliadin in aqueous media under physiological digestive pHs. Colloids Surf. B Biointerfaces 2016, 141, 565–575.
- Herrera, M.G.; Pizzuto, M.; Lonez, C.; Rott, K.; Hütten, A.; Sewald, N.; Ruysschaert, J.-M.; Dodero, V.I. Large supramolecular structures of 33-mer gliadin peptide activate toll-like receptors in macrophages. Nanomed. Nanotechnol. Biol. Med. 2018, 14, 1417–1427.
- Amundarain, M.J.; Herrera, M.G.; Zamarreño, F.; Viso, J.F.; Costabel, M.D.; Dodero, V.I. Molecular mechanisms of 33-mer gliadin peptide oligomerisation. Phys. Chem. Chem. Phys. 2019, 21, 22539–22552.
- Falcigno, L.; Calvanese, L.; Conte, M.; Nanayakkara, M.; Barone, M.V.; D’Auria, G. Structural Perspective of Gliadin Peptides Active in Celiac Disease. Int. J. Mol. Sci. 2020, 21, 9301.
- Nanayakkara, M.; Lania, G.; Maglio, M.; Auricchio, R.; De Musis, C.; Discepolo, V.; Miele, E.; Jabri, B.; Troncone, R.; Auricchio, S.; et al. P31–43, an undigested gliadin peptide, mimics and enhances the innate immune response to viruses and interferes with endocytic trafficking: A role in celiac disease. Sci. Rep. 2018, 8, 1–12.
- Herrera, M.G.; Castro, M.F.G.; Prieto, E.; Barrera, E.; Dodero, V.I.; Pantano, S.; Chirdo, F. Structural conformation and self-assembly process of p31-43 gliadin peptide in aqueous solution. Implications for celiac disease. FEBS J. 2019, 287, 2134–2149.
- Castro, M.F.G.; Miculán, E.; Herrera, M.G.; Ruera, C.; Perez, F.; Prieto, E.D.; Barrera, E.; Pantano, S.; Carasi, P.; Chirdo, F.G. p31-43 Gliadin Peptide Forms Oligomers and Induces NLRP3 Inflammasome/Caspase 1- Dependent Mucosal Damage in Small Intestine. Front. Immunol. 2019, 10, 31.
- Oldstone, M.B.A. Molecular mimicry and immune-mediated diseases. FASEB J. 1998, 12, 1255–1265.
- Kohm, A.P.; Fuller, K.G.; Miller, S.D. Mimicking the way to autoimmunity: An evolving theory of sequence and structural homology. Trends Microbiol. 2003, 11, 101–105.
- Rojas, M.; Restrepo, P.; Monsalve, D.M.; Pacheco, Y.; Acosta-Ampudia, Y.; Ramírez-Santana, C.; Leung, P.S.; Ansari, A.A.; Gershwin, M.E.; Anaya, J.-M. Molecular mimicry and autoimmunity. J. Autoimmun. 2018, 95, 100–123.
- Gómez-Rial, J.; Calle, I.R.; Salas, A.; Martinón-Torres, F. Rotavirus and autoimmunity. J. Infect. 2020, 81, 183–189.
- Cuan-Baltazar, Y.; Soto-Vega, E. Microorganisms associated to thyroid autoimmunity. Autoimmun. Rev. 2020, 19, 102614.
- Múnera, M.; Farak, J.; Pérez, M.; Rojas, J.; Villero, J.; Sánchez, A.; Emiliani, Y. Prediction of molecular mimicry between antigens from Leishmania sp. and human: Implications for autoimmune response in systemic lupus erythematosus. Microb. Pathog. 2020, 148, 104444.
- Cunningham, M.W. Streptococcus and rheumatic fever. Curr. Opin. Rheumatol. 2012, 24, 408–416.
- Cunningham, M.W. Molecular Mimicry, Autoimmunity, and Infection: The Cross-Reactive Antigens of Group A Streptococci and their Sequelae. Microbiol. Spectr. 2019, 7.
- Lucchese, G.; Flöel, A. SARS-CoV-2 and Guillain-Barré syndrome: Molecular mimicry with human heat shock proteins as potential pathogenic mechanism. Cell Stress Chaperones 2020, 25, 731–735.
- Ozuna, C.V.; Iehisa, J.C.M.; Gimenez, M.J.; Alvarez, J.B.; Sousa, C.; Barro, F. Diversification of the celiac disease α-gliadin complex in wheat: A 33-mer peptide with six overlapping epitopes, evolved following polyploidization. Plant J. 2015, 82, 794–805.
- Kurochkina, N.; Guha, U. SH3 domains: Modules of protein–protein interactions. Biophys. Rev. 2013, 5, 29–39.
- Teyra, J.; Huang, H.; Jain, S.; Guan, X.; Dong, A.; Liu, Y.; Tempel, W.; Min, J.; Tong, Y.; Kim, P.M.; et al. Comprehensive Analysis of the Human SH3 Domain Family Reveals a Wide Variety of Non-canonical Specificities. Struct. 2017, 25, 1598–1610.e3.
- Chen, H.I.; Sudol, M. The WW domain of Yes-associated protein binds a proline-rich ligand that differs from the consensus established for Src homology 3-binding modules. Proc. Natl. Acad. Sci. USA 1995, 92, 7819–7823.
- Ball, L.J.; Kühne, R.; Schneider-Mergener, J.; Oschkinat, H. Recognition of Proline-Rich Motifs by Protein-Protein-Interaction Domains. Angew. Chem. Int. Ed. 2005, 44, 2852–2869.
- Bork, P.; Sudol, M. The WW domain: A signalling site in dystrophin? Trends Biochem. Sci. 1994, 19, 531–533.
- Omasits, U.; Ahrens, C.; Müller, S.; Wollscheid, B. Protter: Interactive protein feature visualization and integration with experimental proteomic data. Bioinformatics 2014, 30, 884–886.
- Méthot, P.-O.; Alizon, S. What is a pathogen? Toward a process view of host-parasite interactions. Virulence 2014, 5, 775–785.
- Ingram, J.H.; Stone, M.; Fisher, J.; Ingham, E. The influence of molecular weight, crosslinking and counterface roughness on TNF-alpha production by macrophages in response to ultra high molecular weight polyethylene particles. Biomaterials 2004, 25, 3511–3522.
- Matthews, J.; Green, T.R.; Stone, M.H.; Wroblewski, B.M.; Fisher, J.; Ingham, E. Comparison of the response of primary human peripheral blood mononuclear phagocytes from different donors to challenge with model polyethylene particles of known size and dose. Biomaterials 2000, 21, 2033–2044.
- Green, T. Polyethylene particles of a ‘critical size’ are necessary for the induction of cytokines by macrophages in vitro. Biomaterials 1998, 19, 2297–2302.
- Paul, D.; Achouri, S.; Yoon, Y.-Z.; Herre, J.; Bryant, C.E.; Cicuta, P. Phagocytosis Dynamics Depends on Target Shape. Biophys. J. 2013, 105, 1143–1150.
- Bascuñán, K.A.; Araya, M.; Roncoroni, L.; Doneda, L.; Elli, L. Dietary Gluten as a Conditioning Factor of the Gut Microbiota in Celiac Disease. Adv. Nutr. 2019, 11, 160–174.
- Akobeng, A.K.; Singh, P.; Kumar, M.; Al Khodor, S. Role of the gut microbiota in the pathogenesis of coeliac disease and potential therapeutic implications. Eur. J. Nutr. 2020, 59, 3369–3390.
- Polo, A.; Arora, K.; Ameur, H.; Di Cagno, R.; De Angelis, M.; Gobbetti, M. Gluten-free diet and gut microbiome. J. Cereal Sci. 2020, 95, 103058.
- Carding, S.; Verbeke, K.; Vipond, D.T.; Corfe, B.M.; Owen, L.J. Dysbiosis of the gut microbiota in disease. Microb. Ecol. Health Dis. 2015, 26, 26191.





