
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Hiroyuki Miyachi | + 1231 word(s) | 1231 | 2021-08-31 05:26:22 | | | |
| 2 | Enzi Gong | Meta information modification | 1231 | 2021-09-07 10:12:43 | | |
Video Upload Options
Nuclear receptors (NRs) are ligand-dependent transcription factors that modulate diverse aspects of development, reproduction, and energy homeostasis. This receptor superfamily includes receptors for vitamin D, steroid hormones, thyroid hormones and retinoids, as well as a large number of orphan receptors. NRs are composed of six functionally distinct regions (termed A to F). The N-terminal AB region is highly variable and contains a constitutionally active transactivation function-1 (AF-1) motif. The central C region (a DNA-binding region) is highly conserved among NRs and contains two zinc finger motifs that make contact with specific nucleotide sequences, termed hormone response elements. The C-terminal D, E and F regions are required for ligand binding and receptor dimerization. In most NRs, these regions also contain a second highly conserved transcriptional activation function-2 (AF-2) motif, which is important for ligand-dependent transcription.
1. Overview
Progress in understanding peroxisome proliferator-activated receptor (PPAR) subtypes as nuclear receptors that have pleiotropic effects on biological responses has enabled the exploration of new subtype-selective PPAR ligands. Such ligands are useful chemical biology/pharmacological tools to investigate the functions of PPARs and are also candidate drugs for the treatment of PPAR-mediated diseases, such as metabolic syndrome, inflammation and cancer.
2. Nuclear Receptors
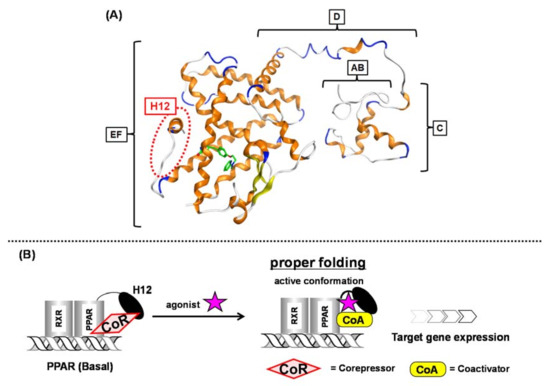
3. Peroxisome Proliferator-Activated Receptors
4. Pleiotropic Effect of PPARs
5. Working Hypothesis of the NR Ligand Superfamily
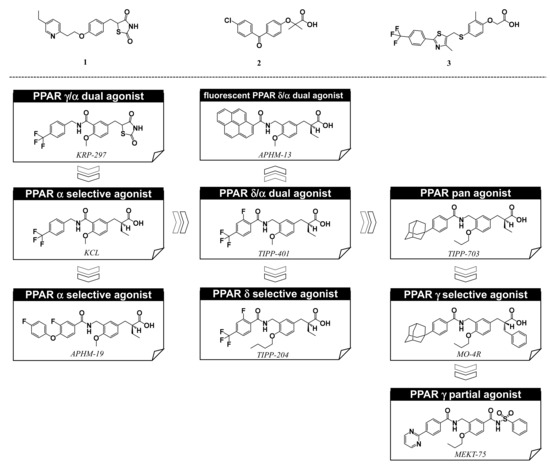
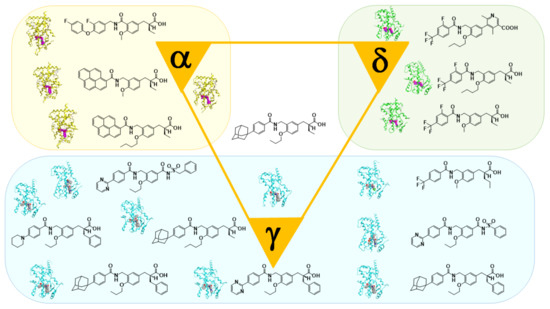
References
- Chawta, A.; Repa, J.J.; Evans, R.M.; Mangelsdorf, D.J. Nuclear receptors and lipid physiology: Opening the x-files. Science 2001, 294, 1866–1870.
- Banner, C.D.; Gottlicher, M.; Widmark, E.; Sjovall, J.; Rafter, J.J.; Gustafsson, J.A. A systematic analytical chemistry/cell assay approach to isolate activators of orphan nuclear receptors from biological extracts: Characterization of peroxisome proliferator-activated receptor activators in plasma. J. Lipid Res. 1993, 34, 1583–1591.
- Wagner, K.D.; Wagner, N. Peroxisome proliferator-activated receptor β/δ (PPARβ/δ) acts as regulator of metabolism linked to multiple cellular functions. Pharmacol. Ther. 2010, 125, 423–435.
- Keller, H.; Dreyer, C.; Medin, J.; Mahfoudi, A.; Ozato, K.; Wahli, W. Fatty acids and retinoids control lipid metabolism through activation of peroxisome proliferator-activated receptor-retinoid X receptor heterodimers. Proc. Natl. Acad. Sci. USA 1993, 90, 2160–2164.
- Staels, B.; Auwerx, J. Role of PPAR in the pharmacological regulation of lipoprotein metabolism by fibrates and thiazolidinediones. Curr. Pharm. Des. 1997, 3, 1–14.
- Lim, H.; Gupta, R.A.; Ma, W.G.; Paria, B.C.; Moller, D.E.; Morrow, J.D.; DuBois, R.N.; Trzaskos, J.M.; Dey, S.K. Cyclo-oxygenase-2-derived prostacyclin mediates embryo implantation in the mouse via PPARδ. Genes Dev. 1999, 13, 1561–1574.
- Sznaidman, M.L.; Haffner, C.D.; Maloney, P.R.; Fivush, A.; Chao, E.; Goreham, D.; Chao, E.; Goreham, D.; Sierra, M.L.; LeGrumelec, C.; et al. Novel selective small molecule agonists for peroxisome proliferator-activated receptor δ (PPARδ) synthesis and biological activity. Bioorg. Med. Chem. Lett. 2003, 13, 1517–1521.
- Oliver, W.R., Jr.; Shenk, J.L.; Snaith, M.R.; Russell, C.S.; Plunket, K.D.; Bodkin, N.L.; Lewis, M.C.; Winegar, D.A.; Sznaidman, M.L.; Lambert, M.H.; et al. A selective peroxisome proliferator-activated receptor δ agonist promotes reverse cholesterol transport. Proc. Natl. Acad. Sci. USA 2001, 98, 5306–5311.
- Tanaka, T.; Yamamoto, J.; Iwasaki, S.; Asaba, H.; Hamura, H.; Ikeda, Y.; Watanabe, M.; Magoori, K.; Ionka, R.X.; Tachibana, K.; et al. Activation of peroxisome proliferator-activated receptor δ induces fatty acid β-oxidation in skeletal muscle and attenuates metabolic syndrome. Proc. Natl. Acad. Sci. USA 2003, 100, 15924–15929.
- Okuno, A.; Tamemoto, H.; Tobe, K.; Ueki, K.; Mori, Y.; Iwamoto, K.; Umesono, K.; Akanuma, Y.; Fujiwara, T.; Horikoshi, H.; et al. Troglitazone increases the number of small adipocytes without the change of white adipose tissue mass in obese Zucker rats. J. Clin. Investig. 1998, 101, 1354–1361.
- Huang, J.-W.; Shiau, C.-W.; Yang, J.; Wang, D.-S.; Chiu, H.-C.; Chen, C.-S.; Chen, A.C.-Y. Development of small-molecule cyclin D1-ablative agents. J. Med. Chem. 2006, 49, 4684–4689.
- Duszka, K.; Gregor, A.; Guillou, H.; König, J.; Wahli, W. Peroxisome Proliferator-Activated Receptors and Caloric Restriction—Common Pathways Affecting Metabolism, Health, and Longevity. Cells 2020, 9, 1708.
- Hashimoto, Y.; Miyachi, H. Nuclear receptor antagonists designed based on the helix-folding inhibition hypothesis. Bioorg. Med. Chem. 2005, 13, 5080–5093.
- Nomura, M.; Tanase, T.; Ide, T.; Tsunoda, M.; Suzuki, M.; Uchiki, H.; Murakami, K.; Miyachi, H. Design, Synthesis and Evaluation of Substituted Phenylpropanoic Acid Derivatives as Human Peroxisome Proliferator-Activated Receptor Activators; Discovery of Potent and Human PPARα Subtype-Selective Activators. J. Med. Chem. 2003, 46, 3581–3599.
- Oyama, T.; Kamata, S.; Ishii, I.; Miyachi, H. Crystal structures of the human peroxisome proliferator-activated receptor (PPAR)a ligand-binding domain in complexes with a series of phenylpropanoic acid derivatives generated by a ligand-exchange soaking method. Bio. Pharm. Bull. 2021. accepted for publication.
- Kasuga, J.; Yamasaki, D.; Araya, Y.; Nakagawa, A.; Makishima, M.; Doi, T.; Hashimoto, Y.; Miyachi, H. Design, synthesis and evaluation of a novel series of α-substituted phenylpropanoic acid derivatives as human peroxisome proliferator-activated receptor (PPAR) α/δ dual agonists for the treatment of metabolic syndrome. Bioorg. Med. Chem. 2006, 14, 8405–8414.
- Araya, Y.; Kasuga, J.; Toyota, K.; Hirakawa, Y.; Oyama, T.; Makishima, M.; Morikawa, K.; Hashimoto, Y.; Miyachi, H. Structure-Based Design and Synthesis of Fluorescent PPARα/δ Co-agonist and Its Application as a Probe for Fluorescent Polarization Assay of PPARδ Ligands. Chem. Pharm. Bull. 2008, 56, 1357–1359.
- Kasuga, J.; Nakagome, I.; Aoyama, A.; Sako, K.; Ishizawa, M.; Ogura, M.; Makishima, M.; Hirono, S.; Hashimoto, Y.; Miyachi, H. Design, synthesis, and evaluation of potent, structurally novel peroxisome proliferator-activated receptor (PPAR) δ-selective agonists. Bioorg. Med. Chem. 2007, 15, 5177–5190.
- Kasuga, J.; Yamasaki, D.; Ogura, K.; Shimizu, M.; Sato, M.; Makishima, M.; Doi, T.; Hashimoto, Y.; Miyachi, H. SAR-oriented discovery of peroxisome proliferator-activated receptor pan agonist with a 4-adamantylphenyl group as a hydrophobic tail. Bioorg. Med. Chem. Lett. 2008, 18, 1110–1115.
- Ohashi, M.; Oyama, T.; Nakagome, I.; Satoh, M.; Nishio, Y.; Nobusada, H.; Hirono, S.; Morikawa, K.; Hashimoto, Y.; Miyachi, H. Design, Synthesis, and Structural Analysis of Phenylpropanoic Acid-Type PPARγ-Selective Agonists: Discovery of Reversed Stereochemistry-Activity Relationship. J. Med. Chem. 2011, 54, 331–341.
- Ohashi, M.; Oyama, T.; Putranto, E.W.; Waku, T.; Nobusada, H.; Kataoka, K.; Matsuno, K.; Yashiro, M.; Morikawa, K.; Huh, N.H.; et al. Design and synthesis of a series of α-benzyl phenylpropanoic acid-type peroxisome proliferator-activated receptor (PPAR) γ partial agonists with improved aqueous solubility. Bioorg. Med. Chem. 2013, 21, 2319–2332.
- Ohashi, M.; Oyama, T.; Miyachi, H. Different structures of the two peroxisome proliferator-activated receptor γ (PPARγ) ligand-binding domains in homodimeric complex with partial agonist, but not full agonist. Bioorg. Med. Chem. Lett. 2015, 25, 2639–2644.
- Oyama, T.; Toyota, K.; Waku, T.; Hirakawa, Y.; Nagasawa, N.; Kasuga, J.; Hashimoto, Y.; Miyachi, H.; Morikawa, K. Adaptability and selectivity of human peroxisome proliferator-activated receptor (PPAR) pan agonists revealed from crystal structures. Acta Crystallogr. D Biol. Crystallogr. 2009, 65, 786–795.
- Kuwabara, N.; Oyama, T.; Tomioka, D.; Ohashi, M.; Yanagisawa, J.; Shimizu, T.; Miyachi, H. Peroxisome proliferator-activated receptors (PPARs) have multiple binding points that accommodate ligands in various conformations: Phenylpropanoic acid-type PPAR ligands bind to PPAR in different conformations, depending on the subtype. J. Med. Chem. 2012, 55, 893–902.
- Ohashi, M.; Gamo, K.; Oyama, T.; Miyachi, H. Peroxisome proliferator-activated receptor γ (PPARγ) has multiple binding points that accommodate ligands in various conformations: Structurally similar PPARγ partial agonists bind to PPARγ LBD in different conformations. Bioorg. Med. Chem. Lett. 2015, 25, 2758–2762.





