Of the 5–30 million species on the Earth, fewer than 2 million have been described and fewer than 1% have been explored for a vast repertoire of new natural products with socio-economic significance. Hence, it is reasonable to expect that many more natural products not only from known species, but also from unidentified organisms are yet to come to benefit humanity and the environment. Natural products offer unique structural molecules unparalleled by any other molecular family with an array of biological activities such as for drug leads. The many natural products that occupy the market today without any chemical modification are a testimony to the remarkable properties of secondary molecules produced by an array of plants, insects, animals, microbes and numerous species of marine organisms.
Secondary metabolites are small heterogenous organic molecules that display prominent ecological benefits to the host organisms in providing defense against predators, parasites, diseases, interspecies nutritional competence, and competitive edge over interaction with the environment. Extensive microbial structural diversification has led to maximizing chemical diversity in terms of the secondary metabolite resources that triggered scope for new drug leads. Since natural products have reflected a wide array of therapeutic and biological applications (antibiotic, anti-inflammatory, antimicrobial, antitumor, anticancer, antiparasitic and immunosuppressing agents as well as enzyme inhibitors), the scope for further exploration of uncharacterized molecules of plant and microbial origin has always remained a focused area for identifying new leads for pharmaceutical and agrochemical usages.
Continuously changing environmental patterns, the emergence of new diseases and resurgence of resistance towards existing drugs have led to an extensive search for novel natural metabolites at a rapid rate. Low molecular size secondary metabolites from living entities have been obtained with typical therapeutic, biological and agricultural implications including antimicrobials, growth promoters, disease suppressers, enzyme inhibitors, health stimulators, and biocontrol agents against pathogenic fungi, bacteria and insects. Systematic strategies for obtaining bioactive metabolites include isolation and identification of known secondary metabolites with biological activities unmatched with the molecular libraries or search for unknown natural molecules with versatile bioactivities. For both these options, the microbial world offers a great repository of natural molecules due to their extensive chemical diversity. However, there remains limitations of the culturability of microbial species and the expression of desired molecular traits or chemical species under isolated culture conditions. To overcome this, metagenomics has emerged to represent vast structural diversity of taxonomic communities with multi-functionalities having diverse chemical structures and functions.
Thus, secondary metabolites are supposed to be conserved in the species, evolved in a competitive environment, emerged to serve purposes other than primary metabolism, secreted for specific physiological or defense-related reasons, related with the habitat of producing organisms, blessed with complex chemical structures and clubbed with diverse bioactivity. These attributes potentiate the usefulness of structurally diverse but functionally sound microbial biomolecules in therapeutics, and agricultural, industrial and environmental applications. We discuss structural, chemical, biological and functional perspectives of one of the earliest known pyrrole antibiotic antifungal metabolite of microbial origin, pyrrolnitrin, which has witnessed laboratory to commercial implications.
2. Pyrrolnitrin (PRN)
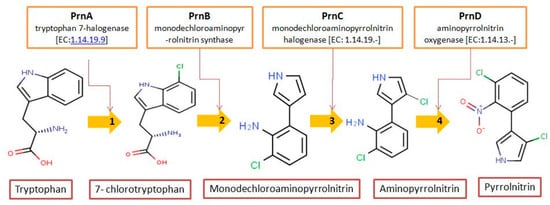
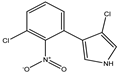
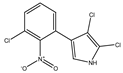
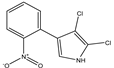
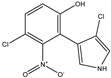
Pyrrolnitrin [3-chloro-4-(2-nitro-3-chlorophenyl) pyrrole] is a phenylpyrrole derivative containing two chlorine atoms and a nitro group. PRN, isolated from Pseudomonas pyrrocinia and various other pseudomonads, was classified as halometabolite in as early as 1964. Later, the compound was biosynthesized using tryptophan as supplement in the medium and chemically synthesized by Nakano et al. Biosynthesis of PRN in Pseudomonas aureofaciens ATCC 15926 has shown that L-tryptophan is a direct precursor (Figure 1). However, high yield of PRN was obtained in D-tryptophan amended medium. Tryptophan analogs amended in the fermentation medium can also yield a series of PRN-like derivatives (Table 1) with low antimicrobial activity than the native parent compound.
Figure 1. Biosynthetic steps in the synthesis of pyrrolnitrin. 7-chlorotryptophan is formed from tryptophan due to flavin-dependent halogenation catalyzed by the enzyme tryptophan 7-halogenase (PrnA). Further, the enzyme PrnB (monodechloroaminopyrrolnitrin synthase catalyzes formation of monodechloroaminopyrrolnitrin from 7-chlorotryptophan while the enzyme PrnC leads to catalytic reaction for the conversion of monodechloroaminopyrrolnitrin into aminopyrrolnitrin. In the last step, aminopyrrolnitrin is converted to pyrrolnitrin with the help of the enzyme PrnD (aminopyrrolnitrin oxygenase).
Structurally, PRN possesses benzene and pyrrole rings with chlorine atoms on both of them and nitro and chlorine units to form an unusual natural skeleton. It has chlorine moiety to contribute more towards biological activity in comparison to its bromine derivative. Consequently, several natural congeners of PRN such as amino-pyrrolnitrin, iso-pyrrolnitrin, 2-chloropyrrolnitrin, oxy-pyrrolnitrin, 4-fluoropyrrolnitrin, and 3-fluoro-3-dechloropyrrolnitrin have been reported. Brominated derivatives of PRN can be synthesized by replacing chlorine ion with bromine in the presence of sodium bromide.
2.1. Pyrrolnitrin: Chemical Synthesis
PRN is positive towards Ehrlich’s reagent where pyrrole ring gets condensed with p-dimethylaminobenzaldehyde to form the violet color complex. Pauly’s coupling reaction yields red color and gives a negative reaction to the ferric chloride nitro group detection test. PRN can be oxidized by chromic acid to form corresponding compound which on oxidation with permanganate, yields carboxylic acid.
Modern synthetic targets for chemical synthesis require regiospecific polysubstituted aromatic or heteroaromatic components . PRN is chemically synthesized by α-block of pyrrole ring and 2-nitro-3-chloroacetophenone, and subsequent chlorination at 4-position and oxidation of methyles using sulfurilchloride followed by decarboxylation. In another approach, 2-Methyl-4-(2-nitro-3-chloro-phenyl)-5-ethoxycarbonyl-pyrrole was prepared in various steps from 2-amino-3-chloro-toluene. One of the most versatile synthetic approach for PRN allowed access to analog compounds such as monodechloroaminopyrrolnitrin and aminopyrrolnitrin. This step facilitated PRN synthesis using Suzuki–Miyaura cross-coupling of an appropriately halogenated pyrrole pinacolboronate ester with halogenated arylpyrroles using 2,6-disubstituted nitrobenzenes or 2,6-disubstituted anilines. Palladium-catalyzed coupling of 1-(triisopropylsilyl-3-substituted pyrroles with arylhaildes has also been described. However, chemical synthesis of PRN makes synthetic route cost extensive and pose threat to the environment. Furthermore, the chemical process utilizes noxious chemicals, high temperature and pressure, more energy and yield poor regioselectivity with lack of public acceptability. Thus, chemical industries prefer microbial species for more selective, greener and cost-effective approach for synthesizing PRN.
2.2. Microbial Pyrrolnitrin Production and Recovery
Microbial synthesis of PRN is easy, reliable and eco-friendly and requires low-cost medium constituents, ambient conditions for growth and production, the least additional energy requirements and minimum expensive equipment. This is the major reason microbial synthesis of PRN has become the preferred alternative to chemical processes. After initial isolation of PRN from Pseudomonas pyrrocinia and thereafter reports from different fluorescent and non-fluorescent Pseudomonas species, several strains of Burkholderia cepacia, Corallococcus exiguus, Cystobacter ferrugineus, Enterobacter agglomerans, Myxococcus fulvus, Serratia spp. and Actinosporangium vitaminophilum have been classified to produce PRN in varying quantities. Serratia plymuthica and S. ruhidaea are identified for enhanced production of PRN. Recently, a strain belonging to B. cepacia complex, JKB9, showing broad-spectrum antifungal activity, was held responsible for suppressing growth of Phytophthora capsici, Fusarium oxysporum and Rhizoctonia solani. This strain, which has shown stronger antifungal activity than Burkholderia strains KCTC2973 and ATCC25416 against Phytophthora blight, was confirmed for PRN production using thin layer chromatography (TPC), high performance liquid chromatography (HPLC) and Nuclear Magnetic Resonance (NMR) spectrometric studies. Complete genome sequencing of Burkholderia pyrrocinia 2327T revealed insights into the cells possessing antibiotic capabilities for the biosynthesis of PRN. Cloning of gene clusters responsible for encoding enzymes involved in the production of pyrrolnitrin in organisms has greatly helped in marking of the biosynthetic routes. Using an antibiotic producing strain of P. fluorescens cloned four gene clusters to elucidate biochemistry of these molecules and to link it with the enzymes that may offer the routes for the synthesis of new chemical structures. Earlier, prnABCD operon from P. protegens Pf-5 was co-expressed in tomato plants with universal vector IL-60 and successfully demonstrated resistance to damping-off disease caused by R. solani .
Microbial wild type strains secrete PRN in low quantity (Table 2) and production varies with the medium constituents. P. aureofaciens ATCC 15926 strain when grown in minimal medium, secreted PRN in low concentration (<0.3 µg mL−1). Even optimized variation of constituents in growth medium could not increase PRN production. However, the production enhanced by 30-fold when P. aureofaciens ATCC 15926 was mutated with N-methyl-N’-nitro-N-nitrosoguanidine. Addition of DL-tryptophan (1 mg mL−1) in CMM medium also doubled PRN production after 120 h but additional amount of tryptophan resulted in less yield.
Table 2. Characteristics of PRN production by different microbial species inhabiting several ecohabitats
| Sr. no. |
Producer |
Habitat |
Medium |
Physical Condition |
Incubation Period (Days) |
Concentration |
Significance |
|
| 1. |
Pseudomonas pyrrocinia |
- |
Bouillon Medium |
- |
- |
ND |
Antibiotic, antifungal nature |
|
| 2. |
P. aureofaciens, P. fluorescens, P. multivorans |
- |
CMM, Synthetic C, E |
27 °C, shaker |
7 |
0.32–126 (µg mL−1) |
PRN widespread in groups of Pseudomonas |
|
| 3. |
P. aureofaciens |
- |
CMM |
27 °C, shaker |
5 |
9.5 to 50 (µg mL−1) |
Production of substituted PRN from Tryptophan analogs |
|
| 4. |
P. aureofaciens |
- |
CMM |
30 °C, shaker |
5 |
18.35–19.9 (µm) |
Possible pathway discussed |
|
| 5. |
Pseudomonas cepacia B37w (NRRL B-14858) |
Rhizosphere |
Sabouraud Maltose Broth |
- |
6 |
2.133 (mg L−1) |
Efficacy against F. Sambucinum incited potato dry rot disease |
|
| 6. |
Pseudomonas cepacia LT4-12- W |
Apple leaves |
Mineral Salt, Nutrient Broth, Kings medium B |
27 °C, 200 rpm |
7 |
1) MS: 51.50 (mg L−1) 2) NB: 7.20 (mg L−1) 3) KMB: 5.50 (mg L−1) |
Production of phenylpyrrole metabolites with respect to time |
|
| 7. |
B. cepacian |
- |
Mineral Salt |
27 °C, shaker |
7 |
ND |
Delays postharvest fruit rot in strawberries |
|
| 8. |
Enterobacter agglomerans IC1270 |
Grapes rhizosphere |
Potato Dextrose Agar |
Incubated on agar plate |
5 |
ND |
Possible role of a combination of Chitinases and pyrrolnitrin in antagonism |
|
| 9. |
B. cepacia NB-1 |
Ponds in botanical garden of Tubingen, Germany |
Minimal medium |
27 °C, aeration rate 0·5 vvm, stirrer speed 150 rev min−1, pH −7.0 |
5 |
0.54 (mg L−1) |
PRN block ETS Neurospora crassa 74 A; inhibition of Streptomycine spp. |
|
| 10. |
Burkholderia cepacia 5.5B (ATCC 55344) Wild Type |
Soil sample, North Carolina |
Nutrient broth, Mineral salt |
25 °C, at 200 rpm, pH 5.8 |
5 |
NB: 35.59; MS: 28.54 (mg 1012 cfu) |
Biocontrol of Rhizoctonia stem rot of poinsettia |
|
| 11. |
Pseudomonas fluorescens psd |
Roots of Vigna mungo |
Standard succinate medium (SSM) |
- |
- |
ND |
Biocontrol property of plants protected from strain |
|
| 12. |
Pseudomonas chlororaphis O6 |
- |
Nutrient broth, Mung bean medium |
28 °C 200 rpm |
- |
1.7 (µg mL−1) |
Regulation by glucose of PRN production influenced biocontrol of tomato leaf blight |
|
| 13. |
Acinetobacter haemolyticus A19 |
Wheat rhizosphere |
Luria broth |
- |
2 |
15 (mg L−1) |
Plasmid-mediated pyrrolnitrin production by A. Haemolyticus A19 |
|
| 14. |
Pseudomonas chlororaphis strain PA23 |
- |
M9 medium + 1 mm MgSO4 + 0.2% glucose |
- |
5 |
ND |
Nematicidal and repellent activity against Caenorhabditis elegans |
|
| 15. |
Serratia marcescens ETR17 |
Tea rhizosphere |
Semi-solid pigment producing media |
30 °C |
8 |
ND |
Effective reduction of root-rot disease tea plant on talc-based formulations; Plant growth promoting activity |
|
Besides intracellular production of PRN from Pseudomonas spp., the excretion of the compound was also detected in the supernatant of fermented medium of S. marcescens strain ETR17. B. cepacia yielded 0.54 mg L−1 of PRN in monosodium glutamate medium at 27 °C as quantified by preparative HPLC. Earlier it was reported that only 27.58% Pseudomonas spp. secreted PRN in shake flask fermentation propagated in CMM, C, or E media. The authors concluded that P. multivorans C653 (ATCC 17760) showed maximum PRN production in medium C, followed by E and then CMM. P. aureofaciens was shown to secrete moderate PRN in CMM medium (40–80 µg mL−1). The PRN concentration increased in D-tryptophan amended medium where it was incorporated in the biosynthesis of PRN.
While growing P. aureofaciens in isotopically labeled tryptophan (at different positions) containing medium, It was demonstrated that amino nitrogen of D-tryptophan became the nitro group of PRN. The two chlorine atoms in PRN, C3 of side chain became pyrrole and C2 of the indole nucleus got retained during biosynthesis (Figure 1). Furthermore, it was confirmed that H-2 and H-α of the indole and side chain give rise to H-5 and H-2 of PRN, respectively, and, thus, proposed that L-tryptophan is the immediate precursor in PRN biosynthetic pathway. PRN formation using labeled tryptophan showed that L- rather than D-tryptophan was the immediate precursor of PRN. 7-chloroindole-3-acetic acid, 3-chloroanthranilate detected in fermented medium revealed that 7-chlorotryptophan served as a common precursor for PRN.
Variety of production media and their pH remained a key parameter to influence PRN secretion. Shake flask fermentation of P. cepacia LT4-12-W revealed that the final yield (at 168 h) of PRN almost doubled at pH 5.8. Amendment of MS medium with glutamate salt of sodium yielded 60.50 mg mL−1 of PRM secretion. The effect of different physicochemical conditions on plasmid-mediated PRN secretion has also been reported from Acinetobacter haemolyticus A19 isolate from wheat rhizosphere.
Recovery strategy of PRN involves cell growth in appropriate medium, extraction in acetone followed by removal of oily matter from concentrated acetone solution using petroleum benzene. From fermented broth at pH 10 or 11 (6 mL) with NaOH, cell pellet centrifugation following sonication with acetone (600 µL) for 1 min, separation of acetone supernatant, re-extraction of pellets again in acetone (300 µL) and drying of acetone extract also yield PRN extract. Further, fermented cultures were extracted after 48 h with equal volume of ethyl acetate and centrifuged. Pellets sonicated twice with ethyl acetate (5 mL) for 3 min then recovery of organic phase resulted in PRN rich dried extract. Lysis of 18 h culture of A. haemolyticus A19 using 1% SDS followed by sonication for 5–15 min resulted in supernatant with PRN. In the case of bioactivity and characterization study, chromatographic separation techniques such as column chromatography and flash column with different mobile phases were explored (Table 3).
Table 3. Purification of pyrrolnitrin using various separation techniques with different solvent systems.
| Matrix |
Column |
Organic Phase |
Detection |
|
| Silica gel G |
35 cm × 1.5 cm |
Chloroform: methanol (9:1) |
- |
|
| Silica gel (40 μm) |
35.6 cm × 1.75 cm |
Benzene: hexane (2:1); Benzene; Benzene: acetone (1:1); Acetone; methanol |
TLC - bioautography |
|
| Silica gel (60 μm) |
- |
Chloroform: hexane (1:1, 1.5:1, 2:1, 5:1) (v/v); chloroform; chloroform-acetone (5:1, 1:1) (v/v); acetone |
Bioassay with R. solani |
|
| Sephadex LH-20 |
- |
Methanol |
pHPLC |
|
| Silica gel 60 (0.015–0.040 mm; Merck) |
- |
Dichloromethane then methanol |
TLC |
|
| Silica gel (H60) |
- |
Dichloromethane |
Bioautography |
|
| Silica gel |
(20 × 170 mm, Wakogel C-200) |
Benzene, 10% ethyl acetatobenzene, 20% ethyl acetate benzene and finally ethyl acetate |
TLC |
|
| Silicic acid |
(240 × 22 mm) |
Diethyl ether and methanol |
- |
|
| - |
RP C-18 flash |
Water and methanol |
TLC |
|
| - |
RP C-18 (MPLC) |
50% to 100% aq methanol |
HPLC |
|
| Silica gel (60 μm) |
- |
Toluene |
- |
|
2.3. Analytical Characteristics of Pyrrolnitrin
PRN is chemically substituted with 3-phenyl pyrrole derivative containing two chlorine atoms and a nitro group. The compound is a pale-yellow crystal, 3-chloro-4-(3-chloro-2-nitrophenyl)-1H-pyrrole, a phenylpyrrole molecule having a chemical formula of C10H6O2N2Cl2 and molecular weight 257.07 gmol−1. The melting point of PRN, which is sparingly soluble in water, petroleum ether and cyclohexane, but more soluble in ethanol, butanol, ethyl acetate, ethyl ether, snf chloroform, is 124.5 °C. Elemental analysis of the compound reflected C, 46.71%; H, 2.33%; O, 12.45%; N, 10.89%; and Cl, 27.68%. PRN separation was achieved by different methods like chromatography TLC and HPLC while structural features have been elucidated using Fourier transform infrared spectroscopy (FTIR), nuclear magnetic spectroscopy (NMR 1H and 13C) and mass spectroscopy (LC-MS and GC-MS).
Separation of PRN from bacterial media extract using TLC utilized various stationary phases such as silica gel G, GF254, 60 F254, KCI8 F, C18 Glass and several mobile phases. PRN can be detected on TLC under UV transilluminator and visualized by spraying diazotized sulfanilic acid (DSA) or Pauly’s, Ehrlich’s and van Urk’s reagent to develop maroon and violet color, respectively or H2SO4 on Silica Gel G plate. The Rf value for various TLC system served to identify PRN from different bacterial species. The compound has been analyzed by retention time in gradient HPLC system but isocratic solvent system of 45% water, 30% acetonitrile, and 25% methanol also separated pyrrolnitrin at 252 nm in preparative HPLC. Modifications in the polarity of solvents, mobile-stationary phase and elution methods are key strategies to quantify PRN using HPLC (Table 4). Yellow colored PRN molecule isolated from P. pyrrocinia absorbs at 252 nm with molar extinction coefficient of ε = 7500 in ethanol. Myxobacterial PRN also showed λmax at 252 nm in methanol. Functional group stretching in FTIR vary with different PRN derivatives due to its structural features. Typical bond stretching at 1530 and 1375 cm−1 characterized for nitro group while 3489 cm−1 represent pyrrole ring. Similarly, PRN isolated from supernatant of fermented medium inoculated by Myxococcus fulvus strain Mx f147 indicated infrared spectrum to confirm pyrrole ring (3460), nitro group (1530 and 1375), CH3 (stretch) (1460) and C=C aromatic weak intensity (1600). Mass spectroscopy (MS) of PRN is ascertain using different ionization techniques. MS of PRN isolated from Pseudomonas cepacia B37w showed molecular ion at m/z 256 with the formula C10H6C12N2O2. Electrospray mass spectroscopy (negative ion spectrum) of PRN further confirmed (mass-to-charge ratio; m/z) at 256. High-resolution mass spectrometry of the two molecular ions gave m/z 255.9826 and 257.9777, respectively, indicating the molecular formula C10H6N2O235C12 and C10H6N2O235Cl37Cl.
Table 4. Several HPLC methods adopted to separate and quantify pyrrolnitrin from microbes using different solvent system.
| Column |
Flow Rate (mL/min−1) |
Solvent System |
Detector |
Retention Time (min) |
|
| RP 18 |
2 |
Methanol: water (70:30; v/v) |
- |
- |
|
| 50 mm × 4.6 mm I.D. guard |
1.0 |
Acetonitrile: methanol: water (1:1:1) |
UV (254 nm) |
10 |
|
| Rainin Dynamax C18 (21.4 × 250 mm) |
- |
Acetonitrile: water (3:2; v/v) fractions collected at 9.5 to 12.5 min and re-chromatographed on a silica column eluted with chloroform: hexane (1:1; v/v) |
- |
13.5 |
|
| C-18 column, 5 µm |
- |
Isocratic acetonitrile: methanol: water (1:1:1) |
- |
- |
|
| Hypersil octyldecyl saline (2.1 mm diameter by 10 cm) |
- |
Water: methanol from 0%: 100 % and gradually changing up to 100%: 0% |
- |
between 10-15 |
|
| Reverse phase 18 |
0.7 |
0 min 50% methanol in water
15 min 100 % methanol in water
17 min 100% methanol in water
20 min 50% methanol in water
25 min 50% methanol in water |
UV (252 nm) |
15.4 |
|
| C-18 reverse phase (125 × 4.6 mm) |
- |
Methanol: water (70:30; v/v) |
UV (252 nm) |
- |
|
| - |
1.0 |
2-min initialization at 10% ACN: 0.1% TFA; 20-min linear gradient to 100% ACN: 0.1% TFA |
990-photodiode array detector |
- |
|
| Nucleosil C-18 |
|
Acetonitrile: water (20 to 100%) |
- |
27.5 |
|
| RP C-18 column |
1.0 |
Isocratically 45% water: 30% acetonitrile: 25% methanol |
- |
- |
|
| C-18 RP column |
|
10% acetonitrile: water (v/v) (both acidified with 0.1% amino acid) run for 2min linear gradient 100% acetonitrile (acidified with 0.1% amino acid) |
- |
18 |
|
| - |
- |
30 ~ 60% aq acetonitrile (for 70 min) |
- |
68.9 |
|
| Gemini C18 column (100 × 4.6 mm; 5mm particle diameter) |
1.0 |
Isocratically 45% acetonitrile: 35% water: 20% methanol |
Dionex AD20 (Dionex,Sunnyvale, CA) (225 nm) |
- |
|
| Cosmosil C18 |
0.7 |
18 min linear gradient from 50 to 100% methanol and 0.1% trifluoracetic acid in methanol |
- |
- |
|
NMR spectroscopy is widely used for analytical measurement of microbial metabolites. The PRN is confirmed by NMR spectrum with values: (i) 1H NMR: H-2 and H-5: 6.82 (m, 2H); H-5′: 7.41-7.53 (m, 3H); NH: 8.29 (br s, 1H); and (ii) 13C NMR δ value: 111.9 indicated C-3, 115.4 for C-4, 116.6 for C-5, 117.4 meant for C-2, while 124.8, 127.6, 128.6, 130.1, and 130.3 designated for C-3′, C-1′, C-6′, C-4′ and C-5′, respectively. Chemical shift (δ) values at 6.81 (m, 2H) indicate the presence of H-2 and H-5, 7.41: H-6′, 7.43: H-5′, 7.52: H-4′, 8.38: NH. NMR spectrum of purified PRN secreted by plasmid-mediated A. haemolyticus A19 revealed the values δ: 6.2–6.6 (m, 2H, H-2, H-5), 6.77 (q 1H, H6), 7.03 (m, 1H, H-4), 7.38 (m2H, Ha, Hc) compared with standard 1H NMR spectrum of. PRN synthesized from Myxococcus fulvus strain Mx f147 showed 13C NMR spectrum (in acetone-d6; Bruker 400 MHz). Structural investigation of PRN with X-ray analysis revealed the presence of two molecules with observed density of 1.74 g/cm3 that lie opposite to each other about the center of symmetry. It further confirmed the location of two Cl atoms in the asymmetric unit with 3D Patterson function, dihedral angle of the pyrrole, the benzene rings and chlorine substitution on pyrrole ring located apart from the nitro group.
2.4. Biochemistry of Pyrrolnitrin
Microbial synthesis of PRN requires D-tryptophan, but cost of precursor amino acid and intracellular secretion limits its large-scale production. The NO2 group is derived from anthranilic acid, phenylalanine and tryptophan that could serve as a precursor for PRN secretion. However, anthranilic acid and L-phenylalanine usually decrease PRN secretion in P. aureofaciens and B. cepacian, while tryptophan stimulates PRN production. In the medium, L-tryptophan gets quick intake within the cells than the D- isomer but addition of L-isomer could not yield more PRN secretion. In actinomycetes, D-tryptophan enhances secretion of PRN when added separately in the culture medium and maximum accumulation was observed at stationary phase after 120 h. It indicated that the L-isomer of tryptophan enter cells quickly and participate in the protein synthesis, while D-tryptophan enter slowly and available at the time of antibiotic secretion. Besides, L-glutamic acid amended medium showed maximum antifungal activity, which substantially declined with the addition of L-tryptophan, L-valine, L-serine, L-phenylalanine and L-cysteine. In brief, D-tryptophan and L-glutamic acid are more direct precursors of PRN than any other amino acids.
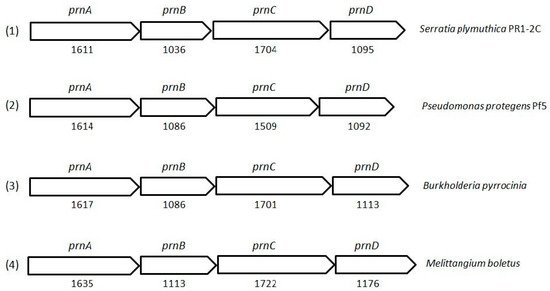
PRN biosynthesis was unraveled in P. aureofaciens. Later, genes (prnABCD operon) and corresponding enzymes involved were delineated in P. fluorescens BL915 (Figure 2). The biosynthesis of PRN occurs in four sequential steps: chlorination by prnA, rearrangement and decarboxylation by prnB, chlorination by prnC and oxidation by prnD enzyme (Figure 1). This involves regioselective halogenation of tryptophan through the addition of chlorine into D-tryptophan by tryptophan 7-halogenase (prnA) following nucleophilic and electrophilic reactions and activation of intermediate lysine-chloramine species as the first step. Further, the reaction catalyzed by prnB shows structural similarity with two-domain indoleamine 2,3-dioxygenase enzyme (IDO) and involves several intermediary steps. The second step forms a binary complex that combines with L-tryptophan or 7-Cl-L-tryptophan to create a ternary complex. The third step in PRN biosynthetic pathway of P. fluorescens leads to catalytic conversion of mono-chloro-deamino-pyrrolnitrin into amino-pyrrolnitrin by regioselectivity using halogenating and chlorinating enzyme. In the last step, prnD catalyzes the oxidation of amino group of aminopyrrolnitrin to nitro group and thus forms PRN.
Figure 2. Cluster organization of pyrrolnitrin biosynthetic genes in S. plymuthica PR1-2C (1), P. protegens Pf5 (2), B. pyrrocinia (3) and myxobacterium Melittangium boletus (4). Nucleotide sequence (NT size indicated for prnA–D genes) of the bacterial species was obtained from KEGG database.
Aminopyrrolnitrin oxidase or arylamine oxygenase (rieske N-oxygenase) catalyzes oxidation of an arylamine into the arylnitro group. Except prnB, trptophan-7-halogenase (prnA), monodechloroaminopyrrolnitrin (prnC) and aminopyrrolnitrin oxidase (prnD) enzymes require flavin reductase (prnF) gene located close to the prnABCD operon which is considered as a part of the cluster. Bioinformatics clubbed with the biochemical tools identified the role of prnF gene in prnD-catalyzed unusual arylamine oxidation in P. fluorescens Pf-5. The prnF and prnD genes form a two-component oxygenase system, in which the gene product enzyme prnF supplies the reduced flavin to prnD. The prnF requires NADH as an electron donor to reduce FAD so that reduced FAD supplies electrons from NADPH to the prnD oxygenase component through protein-protein interactions in order to protect the flavin from oxidation.
The prnF gene having molecular mass of 17kD with GC content of 62%, encodes for a polypeptide chain of 160 amino acids. The enzyme belongs to flavin:NAD(P)H reductases family with part of two-component monooxygenase systems and its C-terminal region possesses highly conserved GDH motif for NAD(P)H binding]. It resembles with PheA2, SnaC, VlmR, ActVB and HpaC with 31.5%, 28.6%, 26.4%, 25.6%, and 25.5% amino acid identity, respectively.
2.5. Pyrrolnitrin Derivatives
Several halogen variations of the PRN molecule have been isolated in the past in the form of bromo-analogs of pyrrolnitrin from fermentation of P. aureofaciens in sodium bromide with low antifungal activity. In addition, 2-chloropyrrolnitrin contain an additional chlorine atom which possesses about 10% of the antifungal activity of PRN. The pyrrolomycin (B, C, D, E, F1, F2a, F2b, and F3) derivatives encompass a chlorine or bromine atom at the 3-position of the pyrrole ring, and either two chlorine atoms at positions 4 and 5 or one chlorine and one bromine at any of these positions have shown significant antifungal activity. Novel oxidized derivatives of pyrrolnitrin including two new pyrrolnitrin analogs, namely 3-chloro-4-(3-chloro-2-nitrophenyl)-5-methoxy-3-pyrrolin-2-oneand4-chloro-3-(3-chloro-2-nitrophenyl)-5-methoxy-3-pyrrolin-2-onehave, were reported from B. cepacia K87.
Furthermore, number of de-chloro and de-nitro derivatives of PRN and the isomers were synthesized by cyclization of enamine reaction, hydrolysis, carboxylation and Mannich’s reaction. The strongest antifungal activity of PRN and its analogs resulted when it got unsubstituted by ester group at any position. The antifungal activity become more stronger when shift of NO2 group was increased. Few PRN derivatives such as denitropyrrolnitrin (3-chloro-4-(3-chlorophenyl), bromo analog: 3-chloro-4-(3-bromophenyl)pyrrole) and trifluoromethyl derivative (3-chloro-4-(3-trifiuorornethyl) pyrrole) were strong antimicrobials. However, among all the analogs homologous to NO2 group of pyrrolnitrin, PRN has remained the strongest biologically active compound. The UV irradiation of 2-(pyrrol-3-yl)nitrobenzene moiety of PRN in an anhydrous aprotic solvent yielded 7,4′-dichlorospiro(1,3-dihydrobenzo(c)isoxazole-3,3′-pyrrolin-2′-one) by the intramolecular oxidation. Hence, the photodegradation of PRN depends on aqueous reaction media and the nature of its excited state.
3. Applications of Pyrrolnitrin
3.1. Biological Activity
Structure–activity mechanism reveals that the primary target of PRN lies in the cell membrane to impede protein, RNA, DNA synthesis and uncouple the normal electron flow in the respiratory electron transport chain. The metabolite has demonstrated biological activity at low concentration and act as an uncoupler of oxidative phosphorylation in N. crassa. High concentration of PRN causes impairment of electron transport in flavin region and cytochrome c oxidase; accumulation of glycerol; synthesis of triacyl glycerol leading to leakage of cell membrane and inhibition of cell growth; in vitro activity against bacteria and fungi in the range of 1–100 µg mL−1; in vitro activity against leukemia and melanoma cell lines; and moderate antimycobacterial activity at 8 µg mL−1. The halometabolite was used as a drug lead for fenipoclonil and fludioxonil synthesis. The amino derivative of PRN was identified as an androgen receptor antagonist. PRN has the unique property to persist actively in the soil over a month, and can be readily diffused and slowly released after lysis of host bacterial cell. However, the compound is sensitive to decomposition due to light.
Inhibitory effect of PRN is seen on the mitochondrial electron transport system of Neurospora crassa 74A. Studies using N,N,N’,N’-tetramethyl-p-phenylenediamine dihydrochloride (TMPD) confirmed that PRN block transfer of electron between the dehydrogenases and cytochrome c-oxidase components of the respiratory chain. At low concentrations, PRN uncouples oxidative phosphorylation in Neurospora mitochondria and impedes electron transport in both the Flavin region and cytochrome C oxidase at high concentration. PRN also function as a signal molecule, beyond its role as a bioactive molecule to suppress fungal and affected cell motility. Antifungal activity of the compound increased at pH 6.0, became maximum at pH 10 or 11 and declined after pH 11. Temperature influence on antifungal activity was maximum at 28 °C. Similarly, 2% NaCl content in the medium showed maximum activity. Such studies indicated more scope for medium modifications for obtaining maximum PRN production followed by maximizing biological activity of the compound.
The quorum-sensing system related regulation of PRN is reported in a chitinolytic bacterium S. plymuthica strain HRO-C48, that protects oilseed rape crop from Verticillium wilt. The mutant deficient in PRN production shown the ability to produce the compound in the medium supplemented with chemically synthesized N-hexanoyl-HSL and N-3-oxo-hexanoyl-HSL (OHHL) (100 µM), thus suggesting that quorum sensing (QS) ability regulated PRN biosynthesis. While investigating the role of N-acylhomoserine lactone (AHL)-dependent quorum sensing for expressing antifungal traits, It was found that PRN expression was positively regulated by CepR gene at transcription level. PRN is reported to have significant antimicrobial potential (Table 5) against Streptomyces antibioticus, S. violaceoruber, Paecilomyces variotii and Penicillium puberulum. However, S. prasinus, S. ramulosus, Aspergillus proliferans and A. terreus showed tolerance to pyrrolnitrin. PRN displayed activity against Ustilago maydis, Candida albicans, Hansenula anomala, Arthrobacter oxidans, Bacillus coagulans, B. lichenifernis, B. subtilis and B. thuriengiensis at low concentration. Pyrrolnitrin produced by Pseudomonas chlororaphis strain PA23 exhibited nematicidal and repellent activity against Caenorhabditis elegans. Co-culturing P. chlororaphis and C. elegans enhanced expression of biocontrol-related phzA, hcnA, phzR, phzl, rpoS and gacS genes and, thus, contributed to the fast killing of nematode in bacterial interaction.
Table 5. Bioactivity spectrum of pyrrolnitrin against bacteria, fungi and nematodes.
| Sr. No. |
Name of Test Microorganism |
PRN (µg mL−1) |
|
| Bacteria |
| 1. |
Staphylococcus aureus 209-P |
6.2 |
|
| 2. |
Escherichia coli |
250 |
|
| 3. |
M. tuberculosis CIP 103471 |
4.0 |
|
| 4. |
M. avium CIP 103317 |
8.0 |
|
| 5. |
M. smegmatis CIP 103599 |
16.0 |
|
| 6. |
M. gordonae CIP 6427 |
>16.0 |
|
| 7. |
M. marinum CIP 6423 |
>16.0 |
|
| Yeast |
| 8. |
Candida albicans |
1.0 |
|
| 9. |
Saccharomyces cerevisiae |
10.0 |
|
| 10. |
Cryptococcus neoformans |
< 0.78 |
|
| 11. |
Candida albicans |
12.5 |
|
| 12. |
Cryptococcus neoformans |
0.78 |
|
| 13. |
Candida utilis |
10.0 |
] |
| Fungi |
| 14. |
Trichophyton asteroids |
0.05 |
|
| 15. |
Sporotrichum schenckii |
< 0.78 |
|
| 16. |
Penicillium atrovenetwn |
10.0 |
|
| 17. |
P. oxalicwn |
10.0 |
|
| 18. |
Sporotrichum schenckii |
3.12 |
|
| 19. |
Blastomyces dermatitidis |
< 0.78 |
|
| 20. |
Histoplasma capsulatu |
0.156 |
|
| 21. |
Sclerotinia sclerotiorum |
0.01 |
|
| 22. |
Rhizoctonia solani |
50 (µg/disc) |
|
| Nematode |
| 23. |
Caenorhabditis elegans |
0.1 |
|
Bacterial growth inhibition by PRN forms complex with phospholipids of cell membranes that eventually cease cellular respiration. Furthermore, PRN causes leakage of A260 mµ absorbing material inside the cells and impairs synthesis of protein, DNA and RNA. However, in vitro protein synthesis in PRN treated Rhizoctonia solani and Escherichia coli remained unaffected. It bursts protoplast of B. megaterium KM at growth inhibitory concentration. The multitudes and range of activity of PRN makes it a preferred bioactive compound for agricultural chemical sector.
3.2. Agricultural Applications
Phenylpyrroles were proven and effective agents against Trichophyton, Microsporium, Epidermophyton, Penicillium, Candida spp. and several Gram-positive bacteria. Besides, PRN showed activity against soilborne fungal phytopathogens Rhizoctonia solani and Fusarium sambucinum and foliar fungal pathogens Fusarium graminearum, F. culmorum, Pyrenophora triticirepentis, Thielaviopsis brasicola, Verticillium dahlia and Alternaria spp. The compound inhibited Gaeumannomyces graminis of wheat, Alternaria brassicae and Botrytis cinerea, and partially inhibited Fusarium roseum. Remarkable inhibition of mycelial growth and conidial germination was observed at a PRN concentration of 0.4 µg mL−1 compared to phenazine-1-carboxylic acid at 50 μg mL−1. Results suggest strong possibility of the compound being a prospective biocontrol agent in the agriculture.
The fungistatic effect of PRN was most distinct against Alternaria sp., Botrytis cinerea, Pythium aphanidermatum, P. ultimum, Rhizoctonia solani, Rhizopus sp. Aspergillus niger, Fusarium oxysporum, Penicillium expansum and Sclerotium rolfsii. Antibacterial activity was also recorded against Agrobacterium tumefaciens, Corynebacterium insidiousum, Pseudomonas syringae pathovar syringae, and Xanthomonas campestris (Minimum Inhibitory Concentration (MIC) ≥1 μg mL−1). Organisms such as Clavibacterium michiganense and S. marcescens were suppressed at MIC ≥ 10 μg mL−1. Moderate activity against Gram-positive and Gram-negative bacteria was seen at 12.5–100 mg mL−1 (MIC). Strong toxicity was noticed against fungi, especially trichophytes, Trichophyton asteroids, at MIC of 0.05 mg mL−1. In addition, PRN as a nitro-heterocyclic chemotherapeutic agent exhibited antimycobacterial activity against M. tuberculosis and M. avium. At present, only two synthesized derivatives of PRN, namely fludioxonil and fenpiclonil analogs, were registered as agricultural fungicides in France and Switzerland, respectively. Commercial products of fenpiclonil and fludioxonil include BERET, GALBAS and SAPHIRE, CELEST and MAXIM sold by Syngenta, respectively.
PRN found most prolific applications in controlling damping-off disease of cotton and cucumber, tan spot of wheat, storage molds of pome fruits, seedling disease of cotton, dry rot of potato and sclerotinia wilt of sunflower. More usage of the compound lies in its significant antibiotic activity and low toxicity to mammalian species. Wounds on apple and pear were challenged with a conidial suspension of antagonist grey mold B. cinerea and blue mold Penicillium expansum to investigate the efficacy of pyrrolnitrin (6–200 µg mL−1) to control diseases at 2 and 24 °C after harvest. High concentrations of PRN proved effective at 24 °C on both diseases of apple and pear, while low concentrations appeared effective at cold temperature. Hence, PRN is an attractive strategy to control postharvest diseases on fruits, vegetables and other agricultural products being produced at low temperature conditions. In a preliminary field experiment on strawberries, postharvest treatment with PRN (250 mg L−1) at low storage temperature delayed development of post-harvest rot by 2–4 days, but did not reduce rate of development and spoilage to acceptable levels.
In greenhouse studies, PRN showed prominent activity against Pyricularia oryzae and Botrytis cinerea. The PRN producer P. chlororaphis O6 has shown antifungal activities both in vitro and in planta on tomato against late blight disease and demonstrated major antagonism. In addition, biocontrol of fungal disease Fusarium Head Blight (FHB) caused by F. graminearum on wheat heads in growth chamber conditions was studied using strain P. chlororaphis G05 co-treated with: (i) wild-type strain G05; (ii) phz-deleted mutant G05Δphz; and (iii) mutant G05Δprn. The experiment showed wheat heads were infected with F. graminearum at rates of 5–8% and 80–90%, respectively, when co-sprayed with wild-type strain G05 and mutant G05Δprn, and PRN of wild type strain was found to be vigorously active against FHB disease.
The glasshouse experiments with talc-based formulation of S. marcescens ETR17 were similar to in vitro studies. Incidence of root rot in bacteria treated tea plants were considerably lower in comparison to untreated control as well as the fungicide treated sets. Additionally, ETR17 formulation also increased the root and shoot length of the tea seedlings under both sterile and unsterile soil conditions in comparison to the untreated controls.
3.3. Pharmaceutical Applications
Pyrrolnitrin demonstrated strong protecting activity against various pathogenic fungi, especially against dermatophytosis. It has been recommended for the treatment of superficial fungal infection of dermatophytic Trychophyton in Japan. A patent has been granted on antifungal composition containing pyrrolnitrin and antimycotic imidazole compound in 1987. The product was commercialized under trade name Pyro-Ace W powder Spray by Fujisawa Pharmaceutical Company Ltd., Osaka. This was marketed by Pharmacia in Italy as “Micutrin” and, in combination with betamethasone valerate, it was formulated as “Beta Micutrin” for athlete’s foot and ring worm diseases. The derivative, 3-cyanopyrroles, is more biologically active as pyrrolnitrin and very stable under light. Jespers and co-workers (1993) reported a Fenpiclonil (CGA 142705) with more cytotoxicity for the representatives of Ascomycetes, Basidiomycetes, and Deuteromycetes. PRN formulated with carboxymethyl cellulose (5%) was injected intraperitoneally into mice and LD50 was observed at a dosage of 500 mg Kg−1.
In pharmacology, in vitro radioactive studies of pyrrolnitrin reflected that pyrrole ring is readily oxidized by enzymes undetected in urine and bile after administration. Along with this, surface antigens of Candida albicans were released after treatment with PRN. It also showed cytotoxicity at 10 µg mL−1 after 24 h and highest after 72 h on rat clonal pancreatic β-lines, INS-1. Thus, the compound becomes diabetogenic but appears nontoxic and insulinotropic at lower concentration. PRN affected physiology of Caenorhabditis elegans, acted as repellent for adult nematodes to lower egg hatching by almost <50% at higher concentrations of PRN (1, 5, and 10 μg mL−1) after 24 h of exposure.
The review article and the statements therein, is Referenced as per the citations available with the MDPI Biomolecule original MS published in Biomolecules in 2020.
Authors : Shraddha P., Chaudhary A. L., Prabha, Ratna, Renu, S., Singh, Dhananjaya P.
Submitted by : Dhananjaya P. Singh