
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Ana Carolina Anauate | + 818 word(s) | 818 | 2020-03-10 03:14:44 | | | |
| 2 | Nicole Yin | Meta information modification | 818 | 2020-10-27 05:14:34 | | |
Video Upload Options
Despite the advancements in cancer treatments, gastric cancer is still one of the leading causes of death worldwide. In this context, it is of great interest to discover new and more effective ways of treating this disease. Accumulated evidences have demonstrated the amplification of 8q24.21 region in gastric tumors. Furthermore, this is the region where the widely known MYC oncogene and different microRNAs are located. MYC deregulation is key in tumorigenesis in various types of tissues, once it is associated with cell proliferation, survival, and drug resistance. microRNAs are a class of noncoding RNAs that negatively regulate the protein translation, and which deregulation is related with gastric cancer development. However, little is understood about the interactions between microRNAs and MYC. Here, we overview the MYC role and its relationship with the microRNAs network in gastric cancer aiming to identify potential targets useful to be used in clinic, not only as biomarkers, but also as molecules for development of promising therapies.
1. Introduction
The MYC family is a group of cellular proto-oncogenes with the following three highly related nuclear phosphoproteins: MYC, N-MYC, and L-MYC[1]. MYC has a low expression and has a short half-life in normal cells, and its mRNA level is tightly regulated by both transcriptional and post-transcriptional mechanisms[2]. However, it is overexpressed in several neoplasms.
2. Biological Significance of MYC
Our group and others have shown MYC overexpression in GC samples[3][4][5], including early stages[6][7], and reported MYC protein overexpression[6][8]. Moreover, other studies revealed the importance of the co-amplification of MYC and EGFR and FGFR2, in predicting poor survival of patients undergoing cancer therapy[9]. In tumor cells, MYC activation occurs as follows: (1) mutations in signaling pathways proteins upstream from MYC; (2) mutations and single nucleotide polymorphisms in regulatory regions that enhance the stability of these proteins[10] and (3) direct modification of MYC gene via gene amplification, mutation, chromosomal translocation and epigenetic modifications[6][10][11][12].
MYC deregulation plays an important role in neoplastic development by targeting genes involved in critical cellular functions, such as DNA metabolism and dynamics, cell cycle, apoptosis, adhesion, survival, and protein and macromolecular synthesis[2][13][14]. Moreover, it contributes to aerobic metabolism by activating the expression of several genes essential for glycolysis and mitochondrial biogenesis[15]. Additionally, its hyperactivity can allow widespread miRNAs downregulation through the regulation of transcriptional and post-transcriptional mechanisms. Indeed, MYC is known as the gene with the highest interaction with downregulated miRNAs[16][17][18] . Taken together, this scenario shows that MYC deregulation (usually overexpression) can have an impact in various cellular functions, contributing to an abnormal cell growth (Figure 1)[15][19].
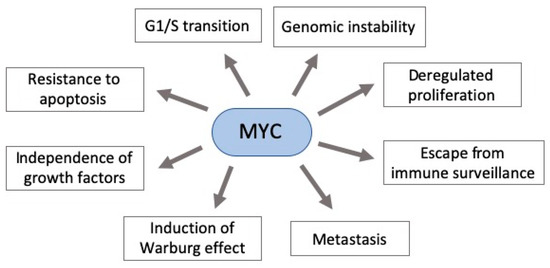
Figure 1. Pleiotropic consequence of MYC deregulation in cancer. MYC overexpression in gastric carcinogenesis affects various components of signaling pathways critical to cancer establishment. Some of these pathways’ phenotypes are shown here.
Infectious agents are extremely important factors on cancer development, accounting for 16% of all new cancer cases per year worldwide[20]. Moreover, liver and gastric tumors in men account for greater than 80% of the infection-related burden cancers[20]. According to the International Agency for Research on Cancer (IARC), 78% of all GC cases are estimated to be associated to chronic H. pylori infection, a bacteria classified as a group 1 carcinogen[21]. The virulence of this bacterium is commonly determined by cagA and vacA genes. The cagA gene encodes the secretion complex, capable of introducing the cagA oncoprotein in the gastric epithelial cell, which activates mitogen-activated protein (MAP) kinases. This alteration activates cell proliferation, differentiation, and stress and inflammatory responses and inhibits programed death, leading to a precancerous process[22][23]. Especially in intestinal-type of GC, H. pylori cagA has been associated with increased MYC expression and nuclear MYC protein[24][25]. In H. pylori infected patients with active gastritis, chromosomal aneuploidy and cellular DNA damage were associated with MYC expression, leading to a chronic hyperproliferation[26]. This association may occur through H. pylori-induced activation of NF-κB and AP-1 proteins which transcriptionally regulate β-catenin expression, responsible for controlling MYC expression and consequently cell proliferation[14][27]. On the other hand, MYC overexpression was not observed in patients without H. pylori infection[26].
The alteration of the DNA methylation profile is considered to be associated with the H pylori inflammatory response, rather than the infection itself[28]. This infection participates in the regulation of MYC expression, which is necessary to gastric carcinogenesis occur (Figure 2), but its infection alone is insufficient to the disease establishment. Thus, the identification of molecules and miRNAs associated with H. pylori infection in GC can contribute to understand the key cellular and molecular processes at the beginning of carcinogenesis and how environmental factors contribute to GC etiology.
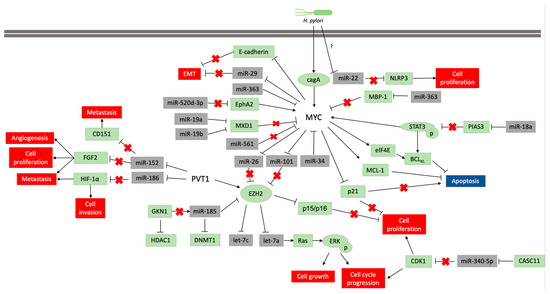
Figure 2: Pathways scheme in which miRNAs regulate MYC and PVT1 expression and vice versa in neoplastic gastric tissue samples and cell lines. The coding genes are shown in green, and the non-coding genes in grey. Lines ending with an arrow indicate activation, whereas T ending lines indicate repression. Lines with a red cross indicate that the interaction is lost due to repression or blocking of a miRNA or protein.
References
- B Mukherjee; S D Morgenbesser; R A DePinho; Myc family oncoproteins function through a common pathway to transform normal cells in culture: cross-interference by Max and trans-acting dominant mutants.. Genes & Development 1992, 6, 1480-1492, 10.1101/gad.6.8.1480.
- Jiangchuan Tao; Xiaohong Zhao; Jianguo Tao; c-MYC–miRNA circuitry. Cell Cycle 2014, 13, 191-198, 10.4161/cc.27646.
- Mariana Ferreira Leal Danielle Queiroz Calcagno; Elizabeth Suchi Chen Salim Khayat; Mario Henrique Girão Faria Ão; rcia Valé Márcia Valéria Pitombeira Ferreira; Marília De Arruda Cardoso Rommel Rodrí lia de Arruda Cardoso Smith; Rommel Rodríguez Burbano; Mariana Ferreira Leal; Aline Damaceno Seabra; Elizabeth Suchi Chen; et al.Samia DemachkiPaulo Pimentel AssumpçãoSilvia Helena Barem Rabenhorst Interrelationship between chromosome 8 aneuploidy,C-MYCamplification and increased expression in individuals from northern Brazil with gastric adenocarcinoma. World Journal of Gastroenterology 2006, 12, 6207-6211, 10.3748/wjg.v12.i38.6207.
- Park, K.U.; Lee, H.E.; Park do, J.; Jung, E.J.; Song, J.; Kim, H.H.; Choe, G.; Kim, W.H.; Lee, H.S. MYC quantitation in cell-free plasma DNA by real-time PCR for gastric cancer diagnosis. Clin. Chem. Lab. Med. 2009, 47, 530–536.
- Takahashi, Y.; Shintaku, K.; Ishii, Y.; Asai, S.; Ishikawa, K.; Fujii, M. Analysis of MYC and chromosome 8 copy number changes in gastrointestinal cancers by dual-color fluorescence in situ hybridization. Cancer Genet. Cytogenet. 1998, 107, 61–64.
- Láuren Cláudia Costa Raiol; Evely Cristina Figueira Silva; Diana Mendes Da Fonseca; Mariana Ferreira Leal; Adriana Costa Guimarães; Danielle Queiroz Calcagno; André Salim Khayat; Paulo Pimentel Assumpção; Marília Cardoso Smith; Rommel Rodríguez Burbano; et al. Interrelationship between MYC gene numerical aberrations and protein expression in individuals from northern Brazil with early gastric adenocarcinoma. Cancer Genetics and Cytogenetics 2008, 181, 31-35, 10.1016/j.cancergencyto.2007.10.011.
- Zheng Chen; Mohammed Soutto; Bushra Rahman; Muhammad W. Fazili; DunFa Peng; Maria Blanca Piazuelo; Heidi Chen; M. Kay Washington; Yu Shyr; Wael M. El-Rifai; et al. Integrated expression analysis identifies transcription networks in mouse and human gastric neoplasia.. Genes, Chromosomes and Cancer 2017, 56, 535-547, 10.1002/gcc.22456.
- Carolina Rosal Teixeira De Souza; Mariana Ferreira Leal; Danielle Queiroz Calcagno; Eliana Kelly Costa Sozinho; B. N. Borges; Raquel Carvalho Montenegro; Ândrea Kely Campos Ribeiro Dos Santos; Sidney Emanuel Batista Dos Santos; Helem Ferreira Ribeiro; Paulo Pimentel Assumpção; et al.Marília Cardoso SmithRommel Rodriguez Burbano MYC Deregulation in Gastric Cancer and Its Clinicopathological Implications. PLoS ONE 2013, 8, e64420, 10.1371/journal.pone.0064420.
- Hyong Kyu Kim; I J Choi; C G Kim; Akira Oshima; Yuki Yamada; Telma Cristina Arao; Kazuto Nishio; Aleksandra M Michalowski; Jeffrey E Green; Three-gene predictor of clinical outcome for gastric cancer patients treated with chemotherapy. The Pharmacogenomics Journal 2010, 12, 119-127, 10.1038/tpj.2010.87.
- Fernanda Wisnieski; Danielle Queiroz Calcagno; Mariana Ferreira Leal; Elizabeth Suchi Chen; Carolina Oliveira Gigek; Leonardo Caires Santos; Thaís Brilhante Pontes; Lucas T. Rasmussen; Spencer Luiz Marques Payão; Paulo Pimentel Assumpção; et al.Laercio Gomes LourençoSamia DemachkiRicardo ArtigianiRommel Rodríguez BurbanoMarília Cardoso Smith Differential expression of histone deacetylase and acetyltransferase genes in gastric cancer and their modulation by trichostatin A. Tumor Biology 2014, 35, 6373-6381, 10.1007/s13277-014-1841-0.
- Dang, C.V. MYC on the path to cancer. Cell 2012, 149, 22–35.
- Lin, C.Y.; Loven, J.; Rahl, P.B.; Paranal, R.M.; Burge, C.B.; Bradner, J.E.; Lee, T.I.; Young, R.A. Transcriptional amplification in tumor cells with elevated c-Myc. Cell 2012, 151, 56–67.
- Dang, C.V.; O’Donnell, K.A.; Zeller, K.I.; Nguyen, T.; Osthus, R.C.; Li, F. The c-Myc target gene network. Semin. Cancer Biol. 2006, 16.
- Dang, C.V. c-Myc target genes involved in cell growth, apoptosis, and metabolism. Mol. Cell. Biol. 1999, 19, 1–11.
- Ga Young Lee; Yang-Sook Chun; Hyun-Woo Shin; Jong-Wan Park; Potential role of the N-MYC downstream-regulated gene family in reprogramming cancer metabolism under hypoxia. Oncotarget 2016, 7, 57442-57451, 10.18632/oncotarget.10684.
- Lima, V.P.; de Lima, M.A.; Andre, A.R.; Ferreira, M.V.; Barros, M.A.; Rabenhorst, S.H. H pylori (CagA) and Epstein-Barr virus infection in gastric carcinomas: Correlation with p53 mutation and c-Myc, Bcl-2 and Bax expression. World J. Gastroenterol. 2008, 14, 884–891.
- Bueno, M.J.; Gómez de Cedrón, M.; Gómez-López, G.; Pérez de Castro, I.; Di Lisio, L.; Montes-Moreno, S.; Martínez, N.; Guerrero, M.; Sánchez-Martínez, R.; Santos, J.; et al. Combinatorial effects of microRNAs to suppress the Myc oncogenic pathway. Blood 2011, 117, 6255–6266.
- Alzahrani, S.; Lina, T.T.; Gonzalez, J.; Pinchuk, I.V.; Beswick, E.J.; Reyes, V.E. Effect of Helicobacter pylori on gastric epithelial cells. World J. Gastroenterol. 2014, 20, 12767–12780.
- German Ott; Andreas Rosenwald; Elias Campo; Understanding MYC-driven aggressive B-cell lymphomas: pathogenesis and classification. Blood 2013, 122, 3884-3891, 10.1182/blood-2013-05-498329.
- Catherine De Martel; Jacques Ferlay; Silvia Franceschi; Jérôme Vignat; Freddie Bray; David Forman; Martyn Plummer; Global burden of cancers attributable to infections in 2008: a review and synthetic analysis. The Lancet Oncology 2012, 13, 607-615, 10.1016/s1470-2045(12)70137-7.
- IARC. Helicobacter pylori Eradication as a Strategy for Preventing Gastric Cancer. Helicobacter pylori Working Group. International Agency for Research on Cancer. IARC Work. Group Rep. 2014, 8.
- Correa, P. New strategies for the prevention of gastric cancer: Helicobacter pylori and genetic susceptibility. J. Surg. Oncol. 2005, 90, 134–138.
- Keates, S.; Keates, A.C.; Warny, M.; Peek, R.M.; Murray, P.G.; Kelly, C.P. Differential Activation of Mitogen-Activated Protein Kinases in AGS Gastric Epithelial Cells by cag+ and cag− Helicobacter pylori. J. Immunol. 1999, 163, 5552–5559.
- Hayashi, Y.; Tsujii, M.; Wang, J.; Kondo, J.; Akasaka, T.; Jin, Y.; Li, W.; Nakamura, T.; Nishida, T.; Iijima, H.; et al. CagA mediates epigenetic regulation to attenuate let-7 expression in Helicobacter pylori-related carcinogenesis. Gut 2013, 62, 1536–1546.
- Silva-Fernandes, I.d.J.L.; Alves, M.K.S.; Lima, V.P.; de Lima, M.A.P.; Barros, M.A.P.; Ferreira, M.V.P.; Rabenhorst, S.H.B. Differential expression of MYC in H. pylori-related intestinal and diffuse gastric tumors. Virchows Arch. 2011, 458, 725–731.
- G Nardone; S Staibano; A Rocco; E Mezza; F P D'armiento; L Insabato; A Coppola; G Salvatore; A Lucariello; N Figura; et al.G De RosaG Budillon Effect of Helicobacter pylori infection and its eradication on cell proliferation, DNA status, and oncogene expression in patients with chronic gastritis. Gut 1999, 44, 789-799.
- Eunyoung Byun; Bohye Park; Joo Weon Lim; Hye-Young Kim; Activation of NF-κB and AP-1 Mediates Hyperproliferation by Inducing β-Catenin and c-Myc in Helicobacter pylori-Infected Gastric Epithelial Cells. Yonsei Medical Journal 2016, 57, 647-651, 10.3349/ymj.2016.57.3.647.
- Tohru Niwa; Tetsuya Tsukamoto; Takeshi Toyoda; Akiko Mori; Harunari Tanaka; Takao Maekita; Masao Ichinose; Masae Tatematsu; Toshikazu Ushijima; Inflammatory Processes Triggered by Helicobacter pylori Infection Cause Aberrant DNA Methylation in Gastric Epithelial Cells. Cancer Research 2010, 70, 1430-1440, 10.1158/0008-5472.can-09-2755.




