
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Tibo De Coninck | + 1909 word(s) | 1909 | 2021-05-26 06:40:56 | | | |
| 2 | Lindsay Dong | Meta information modification | 1909 | 2021-05-28 17:45:57 | | | | |
| 3 | Lindsay Dong | + 88 word(s) | 1997 | 2021-06-03 05:45:23 | | | | |
| 4 | Tibo De Coninck | + 1 word(s) | 1998 | 2021-06-09 13:39:55 | | | | |
| 5 | Tibo De Coninck | + 2 word(s) | 1999 | 2021-06-09 14:47:41 | | | | |
| 6 | Tibo De Coninck | + 6 word(s) | 2003 | 2021-06-09 14:49:16 | | | | |
| 7 | Koen Gistelinck | + 33 word(s) | 2036 | 2021-06-14 17:35:14 | | | | |
| 8 | Els J.M. Van Damme | + 34 word(s) | 2037 | 2021-06-14 22:53:08 | | |
Video Upload Options
The development of plants and the interplay with its environment are highly linked to glycosylation of proteins and lipids as well as metabolism and signaling of sugars.
1. Plant Development and Sugars
Plant development is an overarching term for a plethora of processes, including embryonal development, seed maturation and germination, and growth of the vegetative plant with specialized roots, shoots, leaves and flowers [1][2][3]. It has been shown that sugars, glycoproteins and glycolipids play a crucial role in various pathways such as hormone signaling, cellular trafficking, development and growth [4][5][6][7][8].
Glycoproteins, glycolipids and sugars are heavily involved in mediating environmental cues [4][6][9][10][11]. During the course of a plant’s life cycle, cells undergo a multitude of changes: plant cells develop, grow and gain cellular volume [12]. As soon as germination is initiated, the cells of the growing hypocotyl, radicle and cotyledons multiply fast. This rapid growth requires intense enzymatic as well as hormonal control mechanisms [13].
In general, the addition of a carbohydrate moiety to a protein or lipid is referred to as glycosylation. The process of protein glycosylation is considered the most complicated but ubiquitous modification of secretory proteins [14]. The main types of glycosylation are N-, O-, P-, S- and C-glycosylation, referring to the atom which is involved in the glycosidic linkage [15]. However, N- and O-glycosylation are the most abundant in plants. It is estimated that approximately 50% of all proteins are glycoproteins, of which the majority is N-glycosylated [16][17]. The presence or absence of N- and O-glycans on glycoproteins has been shown to influence a proteins’ activity, stability and functionality to a large extent, and plays a critical role in cellular signaling, molecular trafficking, plant development and adaptation to biotic and abiotic stresses [7][18][19] (Figure 1).
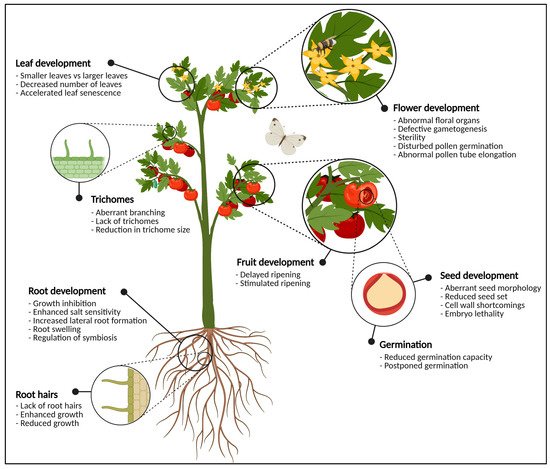
Figure 1. Graphical representation of the most important plant organs. The plant in this figure is generic and does not necessarily represent tomato. For each organ, the most significant phenotypical traits when glycosylation enzymes are knocked-out, knocked-down or over-expressed are highlighted. In the flowering parts, defects in glycosylation cause abnormal development of anthers and pistils, pollen tube germination and elongation, and defective gametogenesis which often leads to sterility. At the level of the fruit, glycosylation enzymes are important for fruit ripening and softening. Seeds show aberrant seed morphology, seed set, (embryo) lethality and cell wall defects when the expression of one or more glycosylation enzymes is disturbed. Vegetative tissues such as the leaves and the roots also experience morphological changes due to aberrant protein glycosylation. This figure was created with BioRender.com.
2. Developmental Consequences of Glycosylation: From Flowers to Germinating Seeds
2.1. Flowers Have a Sweet Tooth
A flowers’ ultimate goal is to produce viable seeds, which will generate progeny and propagate further on. However, flower development is not as straightforward as it seems, since a multitude of requirements have to be met. One of those requirements is proper protein glycosylation, as it is responsible for a plethora of developmental processes and even facilitates pivotal events during the plant’s life cycle [20] (Figure 2).
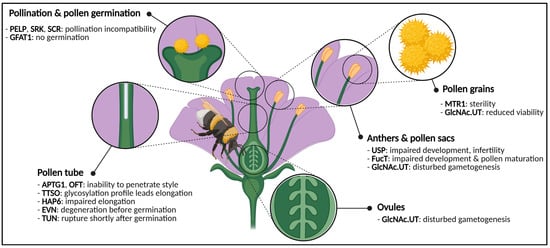
Figure 2. Graphical representation of a generic flower and phenotypes caused by impaired activity of glycosylation enzymes or glycoproteins. Disturbances cause phenotypes in both male and female gametes, anthers, pollen and pollen sacs, pollen germination and pollen tube elongation. Abbreviations: APTG1 (ABNORMAL POLLEN TUBE GUIDANCE1), EVN (EVAN), FucT (α-1,3-fucosyltransferase), GFAT1 (glutamine:fructose-6-phosphate amidotransferase), GlcNAc.UT (GlcNAc-phosphate UDP-transferase), HAP6 (HAPLESS6), MTR1 (MICROSPORE AND TAPETUM REGULATOR1), OFT (O-fucosyltransferase), PELP (Pistil-Specific Extensin-Like Proteins), SCR (S-locus Cysteine Rich), SRK (S-locus Receptor Kinase), TTSO (Transmitting Tissue Specific O-glycoprotein), TUN (TURAN), USP (UDP-sugar pyrophosphorylase). This figure was created with BioRender.com.
2.2. Eradication of Sweet Cell Walls Mediates Fruit Ripening
Fruit ripening encompasses the set of processes that facilitate fruit set, development and maturation of fruits (Figure 3). These are well-understood on a biochemical, physical and molecular level, and have been studied thoroughly. Starting from anthesis and fertilization of the ovaries, the seed as well as the surrounding fruit develops through cell division, cell enlargement and maturation [21][22]. Finally, fruits bearing mature seeds will abscise from the parental plant and start their own life cycle.
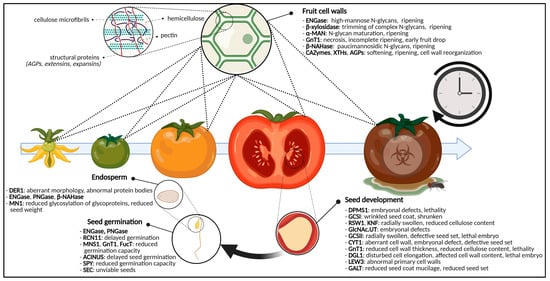
Figure 3. Graphical representation of the glycosylation-related enzymes and their role in fruit ripening and seed development and germination. For fruits, specific glycan-degrading enzymes and CAZymes are important for ripening and softening. Defects in glycosylation enzymes cause aberrant seed morphologies, cell wall shortcomings, embryonal defects, (embryo)lethality and reduced seed set. For the germinating seed, disturbed glycosylation enzymes will postpone germination, reduce germination capacity or will yield unviable seeds. Additionally, the glycosylation state of endosperm glycoproteins will cause certain developmental phenotypes. Abbreviations: ACINUS (Apoptotic Chromatin condensation Inducer in the Nucleus), AGPs (arabinogalactan proteins), α-MAN (α-mannosidase), β-NAHase (β-N-acetylhexosaminidase), CAZymes (carbohydrate-active enzymes), CYT1 (GTP:α-D-mannose-1-phosphate guanylyltransferase), DER1 (DEGRADATION IN THE ENDOPLASMIC RETICULUM1), DGL1 (DEFECTIVE IN GLYCOSYLATION1), DPMS1 (dolichol phosphate mannose synthase complex 1), ENGase (endo-N-acetyl-β-D-glucosaminidase), FucT (α-1,3-fucosyltransferase), GALT (galactosyltransferase), GCSI (α-glucosidase I), GCSII (α-glucosidase II), GlcNAc.UT (GlcNAc-phosphate UDP-transferase), GnT1 (N-acetylglucosaminyltransferase I), KNF (KNOPF), LEW3 (Leaf Wilting 3), MN1 (Miniature1), MNS1 (mannosidase I), PNGase (peptide-N4-(N-acetyl-β-D-glucosaminyl) asparagine amidase), RCN11 (Reduced Culm Number 11), RSW1 (RADIALLY SWOLLEN1), SEC (SECRET AGENT), SPY (SPINDLY), XTH (xyloglucan endotransglycosylase/hydrolase). This figure was created with BioRender.com.
2.3. The Sugar-Craving Cell Wall of Seeds
2.3.1. The Ever-Changing Cell Wall
Plant cell walls are complex and dynamic organs which fulfil many essential functions. Cell walls are built up of O-glycosidic biomolecules, grouped in cellulose, pectic polysaccharides, hemicellulose polysaccharides and lignin. Pectin and lignin are unique to the primary cell wall and secondary cell wall, respectively, and the specific composition of the cell wall varies between different plant species [23].
2.3.2. Cellulose Biosynthesis-Related Problems Cause Cell Wall Disruptions
Two classical examples of enzymes that are of great importance for both plant N-glycosylation and seed and embryonal development are the trivalent dolichol phosphate mannose synthase (DPMS) complex [24] and α-glucosidase I (GCSI) [25]. Although DPMS and GCSI are involved in very different steps of the glycosylation pathway, plants display comparable phenotypes whenever the coding gene is interrupted or over-expressed. Whereas the DPMS complex plays a pivotal role in the synthesis of the lipid-linked glycan precursor [26], α-glucosidase activity is required during glycan trimming after en bloc transfer of the N-glycan precursor [27] (Figure 4).
Both over-expressed DPMS1 and knocked out gcsI mutants display a lethal phenotype with detrimental defects on embryonal development. One of the phenotypic particularities shared between over-expressed DPMS1 plants and mutant gcs1 plants is the aberrant seed morphology: either wrinkled or shrunken, or both. In both cases, the aberrant phenotypes can be attributed to disturbances in seed coat cell wall formation and cellulose biosynthesis [24][27][25]. Shrunken seeds with defects in the embryonal stage were also observed when two GlcNA.UT genes were knocked down in Arabidopsis. However, the observed phenotype was not linked to cell wall defects [28]. Other α-glucosidase-encoding genes are Radially Swollen (RSW) 1 and KNOPF (KNF) and both are involved in cellulose biosynthesis and morphogenesis during embryonal development. Embryos of rsw3 and knf14 mutants showed serious defects in embryogenesis, looked radially swollen and were highly reduced in cellulose content [25]. The observed phenotype was attributed to the UPR. These findings suggest that N-glycosylation is of importance for cell wall formation through cellulose biosynthesis. Additionally, gcsII mutants, also involved in glycan trimming downstream of GCSI, have been reported and linked to cellulose biosynthesis. These mutants also displayed radial swelling of the roots, lethal embryos and defects in seed setting [29][30]. Very comparable phenotypes have been observed in mutants of the GTP:α-D-mannose-1-phosphate guanylyltransferase (CYT1) enzyme in Arabidopsis which is, just like DPMS1, involved in N-glycan precursor synthesis by producing GDP-Man and GDP-Fuc (Figure 4). Distorted cell walls, disturbed embryonal development and lethality have been reported for cyt1 mutants [31]. Both GCSI and GCSII activity, and N-glycan precursor synthesis are important for cell wall construction through cellulose biosynthesis [25][32].
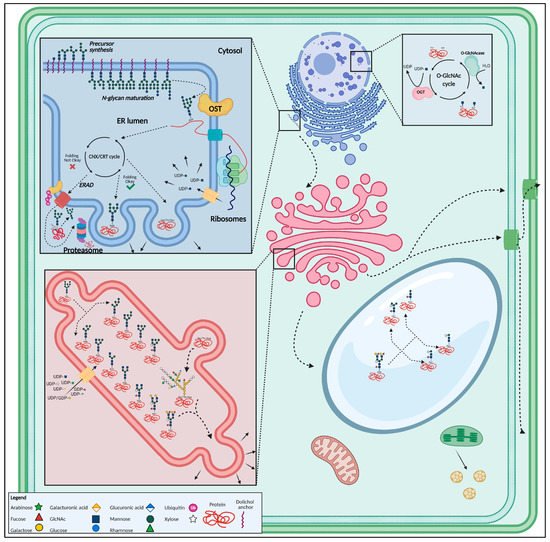
Figure 4. Overview of the most important glycosylated structures in the plant cell. N-glycan synthesis initiates in the endoplasmic reticulum, after which the glycan is added onto a polypeptide. Well-folded proteins are then guided towards the Golgi apparatus through vesicular transport, where the glycan structure is modified. After glycan maturation in the Golgi, glycoproteins are transported towards the vacuole, plasma membrane, cell wall or secreted. O-glycosylation occurs mainly in the Golgi apparatus. The O-GlcNAc modification takes place in the nucleus and cytoplasm. This figure was created with BioRender.com.
2.4. Glycosylation Decides over Seed Germination
In germinating rice seeds, the number of N-linked glycosites equaled 242, distributed over 191 glycoproteins, highlighting the abundance and importance of (N-linked) glycosylation for the germinating seed [17].
3. Glycoproteins during Root and Leaf Development
3.1. The Deal with Glycosylation in Roots
N-glycosylation of different proteins is important for root development. The OST complex, necessary for transfer of the oligosaccharide from the lipid-linked oligosaccharide to a nascent polypeptide chain, consists of multiple subunits and mutations of some of these subunits disturb root growth (Figure 1, Figure 4 and Figure 5) [33]. The under-glycosylation due to mutations of genes involved in the OST-complex, often results in increased osmotic and salt sensitivity [34][35][36][37]. Numerous examples of plants mutated in genes encoding enzymes involved in N-glycan maturation displayed defective root growth and root architecture due to aberrant protein glycosylation [38][39][40][41] (Figure 5).
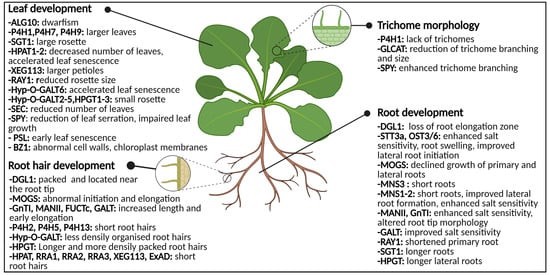
Figure 5. Graphical representation of the glycosylation-related enzymes and their role in leaf development, trichome morphology and root (hair) development. Both abnormal N-glycosylation and O-glycosylation result in changes of phenotype and morphology. Abbreviations: ALG10 (α-1,2-glucosyltransferase 10), BZ1 (Brittle stem and Zebra Leaf 1), DGL1 (DEFECTIVE IN GLYCSOYLATION1), ExAD (extensin arabinose deficient transferase), FUCTc (α1,4-fucosyltransferase), GALT (β1,3-galactosyl-transferase), GLCAT (β-glucuronosyltransferases), GnT1 (N-acetylglucosaminyltransferase I), HPAT1-2 (Hyp O-arabinosyltransferases 1 and 2), HPGT (Hydroxyproline O-galactosyltransferase), Hyp-O-GALT (hydroxyproline-O-β-galactosyltransferase), MANII (α-mannosidase II), MNS (α-mannosidase I), MOGS (mannosyloligosaccharide glucosidase), OST3/6 (oligosaccharyltransferase subunit 3/6), P4H (prolyl 4-hydroxylase), PSL (premature senescence leaf), RAY1 (REDUCED ARABINOSE YARIV1), RRA (REDUCED RESIDUAL ARABINOSE), SEC (SECRET AGENT), SGT (serine O-α-galactosyltransferase), SPY (SPINDLY), STT3a (STAUROSPORIN AND TEMPERATURE SENSITIVE3A), XEG113 (Xylo-endoglucanase113). This figure was created with BioRender.com.
Similar to N-glycosylation, O-glycosylation also affects root growth (Figure 5). O-glycans determine the role and molecular properties of the hydroxyproline-rich glycoprotein and of small hormone peptides [7]. Prior to the addition of a sugar moiety, modifying the proline to a Hyp residue is essential. The generation of Hyp residues is regulated by prolyl 4-hydroxylase (P4H) [42]. Mutational analysis of several P4Hs in Arabidopsis resulted in transgenic plants possessing short root hairs [43][44][45]. Several adverse effects were observed when O-glycosylation is altered such as defective root (hair) growth or even increased root growth [46][47].
3.2. Phenotypical Disturbances in Leaves
3.2.1. Abnormal O-Glycosylation: Leaves in Distress
Aberrant glycosylation affects several stages of plant development, resulting in a clear phenotype of the transgenic plant (Figure 1). The majority of the leaf phenotypes were reported in plants with aberrant O-glycosylation, and include abnormal leaf size, different number of leaves, enhanced leaf senescence and even altered leaf morphology [48][49][50][51]. There are also examples of leaf phenotypes in plants with abnormal N-glycosylation. For instance, the morphological analysis of transgenic plants with a mutation in the ALG10 gene, displayed inhibited growth and smaller leaves [52]. Dwarfism is observed regularly in transgenic plants with N-glycosylation defects [17][53].
3.2.2. Specialized Leaf Tissues Are Annoyed by Absent Glycosylation
Trichomes are protuberances on the aerial part of the plant that differentiated from epidermal cells. Aberrant protein glycosylation can affect the architecture and development of trichomes. The majority of the trichome phenotypes were found in plants with impaired O-glycosylation. For instance, mutation of three β-glucuronosyltransferases (GLCATs) caused reduced trichome branching and size[54].
4. Perspectives
Although sugars are important during glycosylation of plant proteins by functioning directly as the substrates for glycosylation, new evidence is emerging that interplay between sugar signaling events and glycosylation might also exist. At this stage, it is not clear if there is a direct correlation between cellular sugar levels and the level of glycosylation. However, with numerous findings showing differential glycosylation patterns during stress conditions, and the importance of sugar signaling events to alleviate stresses it is likely that a high level of communication between these pathways exists. Of particular interest for future studies will be to determine how the glycosylation of proteins that are targets of sugar signalling cascades are affected, and whether such events can alter their regulation by sugar signaling kinases. Additionally, future studies should also look at glycosylation patterns in plants lacking functional components of major sugar signalling pathways under a variety of growth conditions to establish if sugar signaling events act as upstream regulators of glycosylation.
References
- Liu, C.; Xi, W.; Shen, L.; Tan, C.; Yu, H. Regulation of floral patterning by flowering time genes. Dev. Cell 2009, 16, 711–722.
- De Smet, I.; Vanneste, S.; Inze, D.; Beeckman, T. Lateral root initiation or the birth of a new meristem. Plant Mol. Biol. 2006, 60, 871–887.
- Efroni, I.; Eshed, Y.; Lifschitz, E. Morphogenesis of simple and compound leaves: A critical review. Plant Cell 2010, 22, 1019–1032.
- Eveland, A.L.; Jackson, D.P. Sugars, signalling, and plant development. J. Exp. Bot. 2012, 63, 3367–3377.
- Van den Ende, W. Sugars take a central position in plant growth, development and, stress responses. A focus on apical dominance. Front. Plant Sci. 2014, 5, 313.
- Narayanan, S.; Zoong-Lwe, Z.S.; Gandhi, N.; Welti, R.; Fallen, B.; Smith, J.R.; Rustgi, S. Comparative Lipidomic Analysis Reveals Heat Stress Responses of Two Soybean Genotypes Differing in Temperature Sensitivity. Plants 2020, 9, 457.
- Strasser, R.; Seifert, G.J.; Doblin, M.S.; Johnson, K.L.; Ruprecht, C.; Pfrengle, F.; Bacic, A.; Estevez, J.M. Cracking the “Sugar Code”: A Snapshot of N- and O-Glycosylation Pathways and Functions in Plants Cells. Front. Plant Sci. 2021, 12.
- Ciereszko, I. Regulatory roles of sugars in plant growth and development. Acta Soc. Bot. Pol. 2018, 87.
- Deepak, S.; Shailasree, S.; Kini, R.K.; Muck, A.; Mithöfer, A.; Shetty, S.H. Hydroxyproline-rich Glycoproteins and Plant Defence. J. Phytopathol. 2010, 585–593.
- Chaliha, C.; Rugen, M.D.; Field, R.A.; Kalita, E. Glycans as Modulators of Plant Defense Against Filamentous Pathogens. Front. Plant Sci. 2018, 9, 928.
- Martínez-Noël, G.M.A.; Tognetti, J.A. Sugar Signaling Under Abiotic Stress in Plants. In Plant Metabolites and Regulation Under Environmental Stress; Ahmad, P., Ahanger, M.A., Singh, V.P., Tripathi, D.K., Alam, P., Alyemeni, M.N., Eds.; Elsevier: Amsterdam, The Netherlands, 2018; pp. 397–406. ISBN 9780128126899.
- Höfte, H.; Voxeur, A. Plant cell walls. Curr. Biol. 2017, 27, 865–870.
- Yue, J.-H.; Zhang, D.; Ren, L.; Shen, X.-H. Gibberellin and auxin signals control scape cell elongation and proliferation in Agapanthus praecox ssp. orientalis. J. Plant Biol. 2016, 59, 358–368.
- Corfield, A. Eukaryotic protein glycosylation: A primer for histochemists and cell biologists. Histochem. Cell Biol. 2017, 147, 119–147.
- Stanley, P.; Taniguchi, N.; Aebi, M. N-glycans. In Essentials of Glycobiology; Varki, A., Cummings, R.D., Esko, J.D., Stanley, P., Hart, G.W., Aebi, M., Darvill, A.G., Kinoshita, T., Packer, N.H., Prestegard, J.H., et al., Eds.; Cold Spring Harbor Laboratory Press: New York, NY, USA, 2017; Volume 3.
- Apweiler, R.; Hermjakob, H.; Sharon, N. On the frequency of protein glycosylation, as deduced from analysis of the SWISS-PROT database. Biochim. Biophys. Acta 1999, 1473, 4–8.
- Ying, J.; Zhao, J.; Hou, Y.; Wang, Y.; Qiu, J.; Li, Z.; Tong, X.; Shi, Z.; Zhu, J.; Zhang, J. Mapping the N-linked glycosites of rice (Oryza sativa L.) germinating embryos. PLoS ONE 2017, 12, e0173853.
- Nguema-Ona, E.; Vicre-Gibouin, M.; Gotte, M.; Plancot, B.; Lerouge, P.; Bardor, M.; Driouich, A. Cell wall O-glycoproteins and N-glycoproteins: Aspects of biosynthesis and function. Front. Plant Sci. 2014, 5, 499.
- Gupta, R.; Leon, F.; Rauth, S.; Batra, S.K.; Ponnusamy, M.P. A Systematic Review on the Implications of O-linked Glycan Branching and Truncating Enzymes on Cancer Progression and Metastasis. Cells 2020, 9, 446.
- Smith, D.K.; Jones, D.M.; Lau, J.B.R.; Cruz, E.R.; Brown, E.; Harper, J.F.; Wallace, I.S. A Putative Protein O-Fucosyltransferase Facilitates Pollen Tube Penetration through the Stigma-Style Interface. Plant Physiol. 2018, 176, 2804–2818.
- Ezura, H.; Hiwasa-Tanase, K. Fruit Development. In Plant Developmental Biology; Springer International Publishing: Berlin/Heidelberg, Germany, 2010; Volume 1, pp. 301–318. ISBN 978-3-642-02300-2978-3-642-02301-9.
- Palma, J.M.; Corpas, F.J.; Freschi, L.; Valpuesta, V. Editorial: Fruit Ripening: From Present Knowledge to Future Development. Front. Plant Sci. 2019, 10, 545.
- Harholt, J.; Suttangkakul, A.; Vibe Scheller, H. Biosynthesis of pectin. Plant Physiol. 2010, 153, 384–395.
- Jadid, N.; Mialoundama, A.S.; Heintz, D.; Ayoub, D.; Erhardt, M.; Mutterer, J.; Meyer, D.; Alioua, A.; Van Dorsselaer, A.; Rahier, A.; et al. DOLICHOL PHOSPHATE MANNOSE SYNTHASE1 mediates the biogenesis of isoprenyl-linked glycans and influences development, stress response, and ammonium hypersensitivity in Arabidopsis. Plant Cell 2011, 23, 1985–2005.
- Gillmor, C.S.; Poindexter, P.; Lorieau, J.; Palcic, M.M.; Somerville, C. Alpha-glucosidase I is required for cellulose biosynthesis and morphogenesis in Arabidopsis. J. Cell Biol. 2002, 156, 1003–1013.
- Schwarz, F.; Aebi, M. Mechanisms and principles of N-linked protein glycosylation. Curr. Opin. Struct. Biol. 2011, 21, 576–582.
- Boisson, M. Arabidopsis glucosidase I mutants reveal a critical role of N-glycan trimming in seed development. EMBO J. 2001, 20, 1010–1019.
- Chen, Y.H.; Shen, H.L.; Hsu, P.J.; Hwang, S.G.; Cheng, W.H. N-acetylglucosamine-1-P uridylyltransferase 1 and 2 are required for gametogenesis and embryo development in Arabidopsis thaliana. Plant Cell Physiol. 2014, 55, 1977–1993.
- Peng, L.; Hocart, C.H.; Redmond, J.W.; Williamson, R.E. Fractionation of carbohydrates in Arabidopsis root cell walls shows that three radial swelling loci are specifically involved in cellulose production. Planta 2000, 211, 406–414.
- Burn, J.E.; Hurley, U.A.; Birch, R.J.; Arioli, T.; Cork, A.; Williamson, R.E. The cellulose-deficient Arabidopsis mutant rsw3 is defective in a gene encoding a putative glucosidase II, an enzyme processing N-glycans during ER quality control. Plant J. 2002, 32, 949–960.
- Lukowitz, W.; Nickle, T.C.; Meinke, D.W.; Last, R.L.; Conklin, P.L.; Somerville, C.R. Arabidopsis cyt1 mutants are deficient in a mannose-1-phosphate guanylyltransferase and point to a requirement of N-linked glycosylation for cellulose biosynthesis. Proc. Natl. Acad. Sci. USA 2001, 98, 2262–2267.
- Gillmor, C.S.; Lukowitz, W.; Brininstool, G.; Sedbrook, J.C.; Hamann, T.; Poindexter, P.; Somerville, C. Glycosylphosphatidylinositol-anchored proteins are required for cell wall synthesis and morphogenesis in Arabidopsis. Plant Cell 2005, 17, 1128–1140.
- Lannoo, N.; Van Damme, E.J.M. Review/N-glycans: The making of a varied toolbox. Plant Sci. 2015, 239, 67–83.
- Zhang, M.; Henquet, M.; Chen, Z.; Zhang, H.; Zhang, Y.; Ren, X.; van der Krol, S.; Gonneau, M.; Bosch, D.; Gong, Z.; et al. LEW3, encoding a putative alpha-1,2-mannosyltransferase (ALG11) in N-linked glycoprotein, plays vital roles in cell-wall biosynthesis and the abiotic stress response in Arabidopsis thaliana. Plant J. 2009, 60, 983–999, 10.1111/j.1365-313X.2009.04013.x.
- Farid, A.; Malinovsky, F.G.; Veit, C.; Schoberer, J.; Zipfel, C.; Strasser, R.; Specialized roles of the conserved subunit OST3/6 of the oligosaccharyltransferase complex in innate immunity and tolerance to abiotic stresses. Plant Physiol. 2013, 162, 24–38, 10.1104/pp.113.215509.
- Koiwa, H.; Li, F.; McCully, M.G.; Mendoza, I.; Koizumi, N.; Manabe, Y.; Nakagawa, Y.; Zhu, J.; Rus, A.; Pardo, J.M.; et al.Bressan, R.Hasegawa, P. The STT3a subunit isoform of the Arabidopsis oligosaccharyltransferase controls adaptive responses to salt/osmotic stress. Plant Cell 2003, 15, 2273–2284, 10.1105/tpc.013862.
- Frank, J.; Kaulfurst-Soboll, H.; Rips, S.; Koiwa, H.; von Schaewen, A.; Comparative analyses of Arabidopsis complex glycan1 mutants and genetic interaction with staurosporin and temperature sensitive3a. Plant Physiol. 2008, 148, 1354–1367, 10.1104/pp.108.127027.
- Zhang, M.; Henquet, M.; Chen, Z.; Zhang, H.; Zhang, Y.; Ren, X.; van der Krol, S.; Gonneau, M.; Bosch, D.; Gong, Z.; et al. LEW3, encoding a putative alpha-1,2-mannosyltransferase (ALG11) in N-linked glycoprotein, plays vital roles in cell-wall biosynthesis and the abiotic stress response in Arabidopsis thaliana.. Plant J. 2009, 60, 983-999, 10.1111/j.1365-313X.2009.04013.x.
- Farid, A.; Malinovsky, F.G.; Veit, C.; Schoberer, J.; Zipfel, C.; Strasser, R.; Specialized roles of the conserved subunit OST3/6 of the oligosaccharyltransferase complex in innate immunity and tolerance to abiotic stresses. . Plant Physiol. 2013, 162, 24-38, 10.1104/pp.113.215509.
- Koiwa, H.; Li, F.; McCully, M.G.; Mendoza, I.; Koizumi, N.; Manabe, Y.; Nakagawa, Y.; Zhu, J.; Rus, A.; Pardo, J.M.; et al.Bressan, R.Hasegawa, P. The STT3a subunit isoform of the Arabidopsis oligosaccharyltransferase controls adaptive responses to salt/osmotic stress.. Plant Cell 2003, 15, 2273-2284, 10.1105/tpc.013862.
- Frank, J.; Kaulfurst-Soboll, H.; Rips, S.; Koiwa, H.; von Schaewen, A.; Comparative analyses of Arabidopsis complex glycan1 mutants and genetic interaction with staurosporin and temperature sensitive3a.. Plant Physiol. 2008, 148, 1354-1367, 10.1104/pp.108.127027.
- Wang, S.; Xu, Y.; Li, Z.; Zhang, S.; Lim, J.-M.; Lee, K.O.; Li, C.; Qian, Q.; Jiang, D.A.; Qi, Y. OsMOGS is required forN-glycan formation and auxin-mediated root development in rice (Oryza sativaL.). Plant J. 2014, 78, 632–645.
- Frank, M.; Kaulfürst-Soboll, H.; Fischer, K.; Von Schaewen, A. Complex-Type N-Glycans Influence the Root Hair Landscape of Arabidopsis Seedlings by Altering the Auxin Output. Front. Plant Sci. 2021, 12.
- Liebminger, E.; Hüttner, S.; Vavra, U.; Fischl, R.; Schoberer, J.; Grass, J.; Blaukopf, C.; Seifert, G.J.; Altmann, F.; Mach, L.; et al. Class I α-Mannosidases Are Required for N-Glycan Processing and Root Development in Arabidopsis thaliana. Plant Cell 2009, 21, 3850–3867.
- Velasquez, S.M.; Ricardi, M.M.; Poulsen, C.P.; Oikawa, A.; Dilokpimol, A.; Halim, A.; Mangano, S.; Juarez, S.P.D.; Marzol, E.; Salter, J.D.S.; et al. Complex Regulation of Prolyl-4-Hydroxylases Impacts Root Hair Expansion. Mol. Plant 2014, 8, 734–746.
- Showalter, A.M.; Basu, D.; Extensin and Arabinogalactan-Protein Biosynthesis: Glycosyltransferases, Research Challenges, and Biosensors. Plant Sci. 2016, 7, 814, 10.3389/fpls.2016.00814.
- Gille, S.; Sharma, V.; Baidoo, E.E.K.; Keasling, J.D.; Scheller, H.V.; Pauly, M.; Arabinosylation of a Yariv-Precipitable Cell Wall Polymer Impacts Plant Growth as Exemplified by the Arabidopsis Glycosyltransferase Mutant ray1. Mol. Plant 2013, 6, 1369–1372, 10.1093/mp/sst029.
- Saito, F.; Suyama, A.; Oka, T.; Yoko-o, T.; Matsuoka, K.; Jigami, Y.; Shimma, Y.-I.; Identification of Novel Peptidyl Serine α-Galactosyltransferase Gene Family in Plants. J. Biol. Chem. 2014, 289, 20405–20420, 10.1074/jbc.M114.553933.
- Gille, S.; Hansel, U.; Ziemann, M.; Pauly, M.; Identification of plant cell wall mutants by means of a forward chemical genetic approach using hydrolases. Proc. Natl. Acad. Sci. USA 2009, 106, 14699–14704, 10.1073/pnas.0905434106.
- Basu, D.; Wang, W.; Ma, S.; Debrosse, T.; Poirier, E.; Emch, K.; Soukup, E.; Tian, L.; Showalter, A.M.; Two Hydroxyproline Galactosyltransferases, GALT5 and GALT2, Function in Arabinogalactan-Protein Glycosylation, Growth and Development in Arabidopsis. PLoS ONE 2015, 10, e0125624, 10.1371/journal.pone.0125624.
- Zhang, Y.; Held, M.A.; Kaur, D.; Showalter, A.M.; CRISPR-Cas9 multiplex genome editing of the hydroxyproline-O-galactosyltransferase gene family alters arabinogalactan-protein glycosylation and function in Arabidopsis. BMC Plant Biol. 2021, 21, 16, 10.1186/s12870-020-02791-9.
- Mócsai, R.; Göritzer, K.; Stenitzer, D.; Maresch, D.; Strasser, R.; Altmann, F. Prolyl Hydroxylase Paralogs in Nicotiana benthamiana Show High Similarity With Regard to Substrate Specificity. Front. Plant Sci. 2021, 12.
- Velasquez, S.M.; Ricardi, M.M.; Dorosz, J.G.; Fernandez, P.V.; Nadra, A.D.; Pol-Fachin, L.; Egelund, J.; Gille, S.; Harholt, J.; Ciancia, M.; et al. O-Glycosylated Cell Wall Proteins Are Essential in Root Hair Growth. Science 2011, 332, 1401–1403.
- Zhang, Y.; Held, M.A.; Showalter, A.M.; Elucidating the roles of three β-glucuronosyltransferases (GLCATs) acting on arabinogalactan-proteins using a CRISPR-Cas9 multiplexing approach in Arabidopsis. BMC Plant Biol. 2020, 20, 221, 10.1186/s12870-020-02420-5.
- Zhang, Y.; Held, M.A.; Showalter, A.M.; Elucidating the roles of three β-glucuronosyltransferases (GLCATs) acting on arabinogalactan-proteins using a CRISPR-Cas9 multiplexing approach in Arabidopsis. BMC Plant Biol. 2020, 20, 221, 10.1186/s12870-020-02420-5.




