Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Roberta Fraschini | + 4033 word(s) | 4033 | 2021-05-13 12:02:36 | | | |
| 2 | Vivi Li | -4 word(s) | 4029 | 2021-05-20 05:00:41 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Fraschini, R. Cytokinesis in Eukaryotic Cells. Encyclopedia. Available online: https://encyclopedia.pub/entry/9849 (accessed on 21 May 2026).
Fraschini R. Cytokinesis in Eukaryotic Cells. Encyclopedia. Available at: https://encyclopedia.pub/entry/9849. Accessed May 21, 2026.
Fraschini, Roberta. "Cytokinesis in Eukaryotic Cells" Encyclopedia, https://encyclopedia.pub/entry/9849 (accessed May 21, 2026).
Fraschini, R. (2021, May 19). Cytokinesis in Eukaryotic Cells. In Encyclopedia. https://encyclopedia.pub/entry/9849
Fraschini, Roberta. "Cytokinesis in Eukaryotic Cells." Encyclopedia. Web. 19 May, 2021.
Copy Citation
The duplication cycle is the fascinating process that, starting from a cell, results in the formation of two daughter cells and it is essential for life. Cytokinesis is the final step of the cell cycle, it is a very complex phase, and is a concert of forces, remodeling, trafficking, and cell signaling. All of the steps of cell division must be properly coordinated with each other to faithfully segregate the genetic material and this task is fundamental for generating viable cells. Given the importance of this process, molecular pathways and proteins that are involved in cytokinesis are conserved from yeast to humans.
cell division
aneuploidy
MEN network
bud neck
model organism
tetraploidy
CIN
1. Introduction
Eukaryotic cells duplicate themselves through several rounds of a process, called mitotic cell cycle. The cell cycle is made of four phases: gap 1 (G1), DNA synthesis (S), gap 2 (G2), and mitosis (M). During mitosis, or cell division, the duplicated DNA filaments condense into chromosomes (prophase), bind the mitotic spindle in a bipolar way, and then align at the metaphase plate (metaphase); later, sister chromatids are separated and start to migrate to opposite poles of the cell (anaphase) and during telophase they conclude their movement and they decondense. After the nuclei have segregated, cytoplasm separation concludes cell division (cytokinesis). All of these events must occur in a very precise and coordinated way to preserve genetic stability, which, in turn, is essential for cell viability. Indeed, cytokinetic failure can cause multinucleate or aneuploid cells that can drive oncogenic transformation [1].
Given the importance of cytokinesis for cell viability, the basic processes of cytokinesis and several key proteins are evolutionarily conserved, for this reason, studies on model organisms are precious. The nematode C. elegans, embryos and cultured cells of the insect D. melanogaster, amphibian Xenopus eggs, and cultured mammalian cells are the mainly used animal models. The unicellular yeasts Saccharomyces cerevisiae and Schizosaccharomyces pombe are also very useful for unraveling the general principles of eukaryotic cell polarization and cytokinesis [2].
2. Cytokinesis in Animal Cells
In animal cells, at anaphase onset, the mitotic spindle forms a dense array of MTs, called central spindle. After chromosomes segregation, the plasma membrane starts to ingress at the cell equator and the central spindle and asters define this position, and then a cleavage furrow is formed that separates the daughter cells. The furrow consists of a contractile ring or cytokinetic ring (CR), a stable structure mostly made by actin and myosin II, a non-muscle motor protein, which is connected with the plasma membrane. At the molecular level, Rho proteins and their regulators, GAP and GEF, the protein kinases Aurora B and Polo accumulate at the equator (division plane) of the cell and signal formins recruitment that allows for the nucleation of filamentous actin (F-actin) and the localization of myosin II. Myosin II recruitment does not require its ATPase activity [3], but needs the phosphorylation of its light chain [4]. Microtubules spatially regulate the Rho pathway and kinases with molecular mechanisms that are not fully understood.
The cleavage furrow contains also scaffolding proteins, such as anillin and septins. Anillin associates with F-actin, myosin, septins, and activated RhoA, it localizes to the furrow at early stages of cytokinesis, and it is important for its stability, but it is not essential for its ingression [5]. Instead, it might stabilize the furrow and be important for later stages of cytokinesis. Indeed, anillin remains in the cytoplasmic bridge, even after myosin and actin have dissociated. Another class of scaffolding proteins are septins: they are GTP-binding proteins that form filaments and localize to the cytokinetic ring [6]. Several human septins localize to the central spindle and midbody during anaphase and cytokinesis. Septins participate in the regulation of actin and microtubule dynamics, directly interact with anillin, and allow for the full activation of myosin that is necessary for cytokinesis. Interestingly, septins may form a barrier that restricts the diffusion of membrane proteins in the furrow since they directly bind the plasma membrane [7].
CR are stable biochemical structures that can be isolated and are able to contract in vitro [8], however the precise ultrastructure of actin and myosin II and the mechanism of ring contraction are unsolved problems. Recent studies took advantage of super-resolution three-dimensional (3D) structured illumination light microscopy and transmission electron microscopy (TEM) technologies to reveal CR structure of sea urchin embryos. They showed that actin within the CR is organized in antiparallel linear filaments that form an array parallel to the division plane. Myosin II filaments are concomitantly reorganized with the maturation of the ring in an actin-dependent manner. Myosin II filaments are head-to-head chains that are associated laterally with their long axis parallel to the cleavage plane [9]. These findings support a mechanism of CR contraction that involves the sliding of filaments in a purse-string model: filaments sliding shortens the ring producing a force that drives furrow ingression.
Inward directed forces that are generated from the cytoskeleton and new membrane deposition promote furrow ingression. During this process, actin is specifically depolymerized at the equator due to actin-depolymerization factor (ADF) and cofilin [10]. Myosin II also has a rapid turnover, depending on the balance of recruitment and dissociation [11]. Actin and myosin II dynamics lead to a decrease of CR volume. CR assembly and contraction must be coordinated with membrane deposition. Indeed, plasma membrane deposition is essential for cleavage furrow formation and completion, and CR must be considered not only as force-generating machinery, but also as a landmark for vesicle delivery to the division site.
During furrow ingression, new membrane is added locally by vesicle trafficking and both the secretory pathway and the endocytic pathway are involved. Golgi derived vesicles bring new membrane and proteins to the furrow, whereas recycling endosomes and the endocytic pathway are important for both membrane remodeling during furrow ingression and for the final steps of cytokinesis [12][13][14]. Studies in C. elegans and Drosophila revealed that the deposition of new furrow membrane requires astral microtubules and release of calcium, indicating that astral MTs not only specify positioning of the contractile ring, but also direct the delivery of new membrane to the furrow later during cytokinesis [15].
Although CR generates force to conclude cytokinesis, in some cases cells are also able to divide in the absence of a functional CR [16][17]. For example, in animal cells, the force for cell separation can derive from pulling on the substrate [18][19], since round mitotic cells maintain some substrate attachment and, in most cases, animal cells are attached to a substrate or to other cells when they divide.
Scission or abscission is the final step of cytokinesis and requires remodeling of the plasma membrane to separate sister cells. The evolutionary conserved Endosomal Sorting Complex Required for Transport (ESCRT) proteins drive this process [20]. Abscission occurs after CR has completely contracted, myosin II and filamentous actin have dissociated, and the furrow has formed an intracellular bridge rich in antiparallel microtubules with a diameter of 1 μm, the midbody. This structure is essential for proper cytokinesis completion that requires a plasma membrane remodeling that is driven by Anillin and septins, possibly fusing transport vesicles. Interestingly, a centrosome component, centriolin, has a role in vesicle targeting and fusion in the bridge [21]. Mammalian centriolin is the homologue of budding yeast Nud1 and of fission yeast Cdc11 that are MEN/SIN components that regulate mitotic exit and cytokinesis. Likely, all of these proteins are involved in cytokinesis completion. In order to complete abscission, the activity of microtubule-severing proteins cuts the interdigitating MTs in the midbody, causing its breakage [22].
3. Cytokinesis in Budding Yeast
Cytokinesis can be studied in model organisms since several proteins and most basic cytokinesis processes are evolutionarily conserved. Table 1 shows a list of evolutionarily conserved proteins. Saccharomyces cerevisiae is a unicellular eukaryotic organism that offers many experimental advantages, among which: a fast cell cycle, easy and quick growth in solid and liquid medium, cellular morphology that changes during different cell cycle phases, haploid and diploid cycle, easy genetic manipulation, and analysis. An interesting feature is that S. cerevisiae divides asymmetrically, with the daughter cell being generated as a bud from the mother cell body. During late G1 phase of the cell cycle, an intense protein relocalization to a specific place of the cell cortex induces a strong polarization of the cells and culminates in bud emergence.
Table 1. Evolutionarily conserved proteins involved in cytokinesis.
| Generic Name | S. cerevisiae | C. elegans | D. melanogaster | Humans |
|---|---|---|---|---|
| Structural proteins | ||||
| Septins | Cdc3, Cdc10, Cdc11, Cdc12, Shs1 | UNC-59, UNC-61 | Peanut, Sep1,2,4,5 | Septin 1,4,5,6,7,8,10,11,14 |
| Cdc42 pathway | Cdc42, Cdc24 | CDC-42 | Cdc42 | Cdc42 |
| Actin | Act1 | ACT1, 3, 5 | Actin (multiple genes) | Actin (6 genes) |
| Myosin | Myo1, Mlc1, Mlc2 | NMY-2, MLC-4 | Zipper, Mlc-c, Spaghetti Squash | Myosin II, Myosin ELC, Myosin RLC |
| Rho proteins and regulators | Hof1, Rho1 | RHO-1, ECT-2 | RhoA, Pbl | RhoA, ECT2 |
| Centriolin | Nud1 | CEP110 | Centriolin | CNTRL (Cep110) |
| Formin | Bni1, Bnr1 | CYK-1 | Diaphanous | Dia1 |
| IQGAP | Iqg1 | PES-7 | - | IQGAP1, IQGAP2, IQGAP3 |
| Survivin | Bir1 | BIR-1 | Scapolo | Survivin |
| Anillin | Bud4 | ANI-1, ANI-3 | Scraps | hAnillin |
| Regulatory elements | ||||
| Aurora B | Ipl1 | AIR-2 | Aurora B | Aurora B |
| Polo kinase | Cdc5 | PLK-1 | Polo | Plk1 |
| MEN/SIN pathway | Cdc14, Cdc15, Dbf2, Cbk1 | CDC-14, WTS-1, YAP-1, EGL-44 | Cdc14, Hippo, Mats, Warts | Cdc14A, Cdc14B, MST2, LATS1, LATS2 |
The Rho-type GTPase Cdc42 concentrates at the site of bud emergence, together with its GEF Cdc24 and its GAP, and activates its effectors that organize the actin cytoskeleton to drive polarized growth and regulate the bud neck formation [23].
3.1. Bud Neck Formation
The bud neck is the site that separates the mother cell from the daughter cell, it is not only a spatial point, but it has a very complex structure (Figure 1). In late G1, the first recruited proteins are septins: 5 GTP-binding proteins (Cdc3, Cdc10, Cdc11, Cdc12, and Shs1) that form linear apolar filaments that assemble into 3D structures that are highly dynamic during mitosis [24]. Septins filaments associate with the cell membrane and they form a ring around the bud neck during bud emergence and their function is essential in maintaining bud neck structure. During the progression of the cell cycle, the septin ring changes its shape becoming a rigid hourglass structure, and then splitting into two rings at the final steps of cytokinesis [25]. Septins rearrangements are regulated by protein-protein interactions and by phosphorylation/dephosphorylation and sumoylation events [26]. Septin ring acts as a platform for other proteins and a signaling landmark for actin cytoskeleton filaments that drive mitotic spindle positioning and orientation. In addition, several protein kinases that are important for cell cycle progression and cell cycle chekpoints are associated with septins and their localization at the bud neck depends on septins [27]. It is important to point out that septins also act as a cortical barrier to block movement of membrane proteins thus helping to build cellular compartmentalization [28].
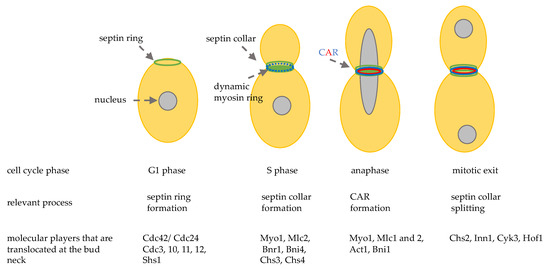
Figure 1. Schematic representation of the sequential localization at the division site of important cytokinesis players. CAR: contractile actomyosin ring.
During late G1 and S phase, septins recruit myosin II heavy chain, Myo1, and the light chain Mlc2, the formin Bnr1 [29], and the scaffold protein Bni4. After anaphase onset Hof1, Iqg1, Inn1, Cyk3, the regulatory chain Mlc1, and the formin Bni1 are localized at the bud neck and in late anaphase a functional actomyosin ring is formed, thanks to actin recruitment (Figure 1) [30].
3.2. CAR Formation and Contraction
Budding yeast cytokinesis requires the formation and contraction of an actomyosin ring, even if the bud neck is very narrow (1 μm). The contractile actomyosin ring (CAR) is a structure composed by structural and regulatory proteins, the most important being actin and myosin.
Actin is encoded by the essential gene ACT1, it is an ATP-binding protein that can form filaments that have a polarity that is built during their formation. In yeast cells, actin can form three types of structures: actin patches, actin cables, and an actin ring [23]. Actin patches are important for cell polarity formation and maintenance. Actin cables serve for intracellular transport of vesicles, organelles, and mRNA, and for mitotic spindle alignment. The actin ring is a structure that forms transiently at the mother-daughter neck, it binds myosin, and is important for cytokinesis.
The second essential element of CAR is the class II myosin heavy chain, hereafter called myosin, being encoded by MYO1 gene. Myosin is the motor that slides actin filaments and induces ring contraction [31]. The catalytic domain is located in the N-terminus, while the C-terminus contains a coiled-coiled domain that can be recruited to the site of division and it is sufficient for CAR constriction and cytokinesis. Myo1 is regulated by two light chains, an essential light chain (Mlc1) and a regulatory light chain (Mlc2). Myo1 is recruited to the emergent bud site, being present as a ring at the bud neck until cytokinesis is completed and then disassembled after contraction. In late anaphase, a complete CAR is formed around the bud neck (Figure 2A), and, after completion of previous cell cycle events, it can contract. This process does not need Myo1 motor activity and it involves a decrease in Myo1 levels and actin depolymerization regulated by cofilin, like in animal cells (see paragraph 3).
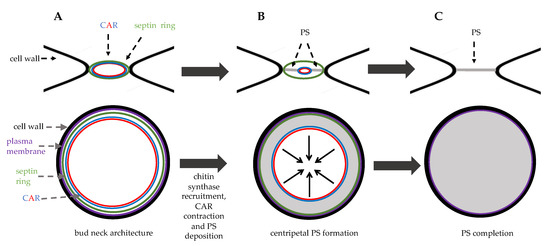
Figure 2. Bud neck structure during cytokinesis. Top panels: side view, bottom panels: top view. (A) in late anaphase septin ring and the contractile ring (CAR) are assembled at the bud neck. (B) in late anaphase the chitin synthase Chs2 is recruited to the bud neck and the primary septum (PS, grey area) is synthetized centripetally. (C) just before cell separation PS deposition is completed leading to physical dissociation of mother and daughter cytoplasms.
After mitotic exit, these events concomitantly occur: CAR contracts symmetrically, membrane invaginates, and a septum is formed in a centripetal way.
3.3. Cell Wall Deposition
Since yeast cells have a cell wall, in order to complete cell separation, two single plasma membranes and a septum must be formed. Specific proteins are expressed in order to achieve this task (Table 2). Even if budding yeast division site is very thin, cell division involves membrane deposition and vesicle recruitment at the division site. In concomitance with CAR contraction, new membrane is deposited, and this ensures that the chitin septum forms correctly and efficiently. One important role of membrane deposition is to deliver integral membrane proteins that forms the septum: the chitin synthases, transmembrane proteins that polymerize chitin chain that is extruded by the plasma membrane.
Table 2. Budding yeast proteins that are involved in the final steps of cytokinesis.
| Name | Functions |
|---|---|
| Inn1 | Couples membrane ingression with CAR contraction, regulates chitin synthase Chs2 |
| Cyk3 | Interacts with Hof1 and regulates chitin synthase Chs2 |
| Chs1 | Chitin synthase, important for septum repair |
| Chs2 | Chitin synthase, mayor role in chitin primary septum synthesis |
| Chs3 | Chitin synthase |
| Chs4 | Regulator of chitin synthase Chs3, interacts with Bni4 |
| Bni4 | Required for correct septum formation |
| Fks1 | Catalytic subunit of 1,3-beta-D-glucan synthase, involved in secondary septum synthesis |
| Cts1 | Endochitinase, digestion of chitin in the primary septum |
| Eng1 | Glucanase, digestion of the secondary septum |
| Scw11 | Glucanase, digestion of the secondary septum |
| Cbk1/Mob2/Ace2 | RAM signaling pathway, required for asymmetric daughter-specific transcription of chitinase and glucanases |
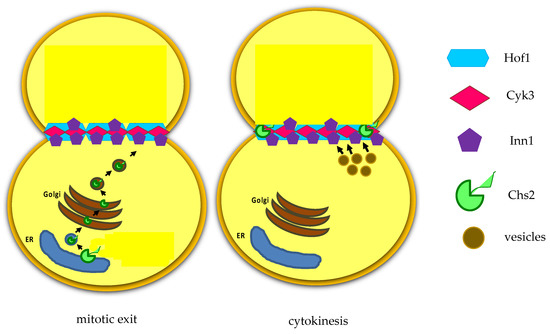
Figure 3. After mitotic exit, the chitin synthase Chs2 is translocated from endoplasmic reticulum (ER).
Recent findings indicate that Inn1 and Cyk3 regulate Chs2 catalytic domain, thus controlling chitin deposition during cytokinesis [32]. As PS is a fragile structure, after its deposition (Figure 2C), 1,3-beta-D-glucan synthases synthesize secondary septa (SS) on either side of the PS [33]. SS are glucan-rich and they have a molecular structure similar to the cell wall.
Since Chs2, Chs3, and Fks1 are localized at the division site before mitotic spindle disassembly, their activation must be tightly controlled to avoid premature spindle breakage; this task is reached by multiple regulations, among which a balance between enzyme localization and their endocytosis [34].
3.4. Cell Separation
The final step of cytokinesis is cell separation, the event that irreversibly divides the mother from the daughter cell. Cell separation requires the partial degradation of the septum, which occurs from the daughter cell side and leaves a sign, called bud scar, on the mother cell surface (Figure 4B). In order to degrade PS and SS partially, endochitinase Cts1 and glucanases Eng1 and Scw11 are activated specifically from the daughter side of the bud neck (Figure 4A) [35].
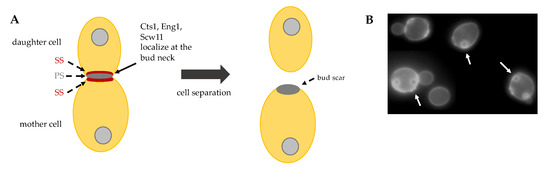
Figure 4. (A) Schematic representation of cell separation: after primary septum (PS) and secondary septa (SS) formation, Cts1, Eng1 and Scw11 activity at the daughter bud neck allows for cell separation. (B) Budding yeast exponentially growing cells stained with Calcofluor White. On the surface of mother cells these are several scars, chitin-containing rings (arrows), originated during cell division. New born daughter cells do not show bud scars.
The activity of these hydrolytic enzymes is tightly regulated: they are specifically transcribed in the daughter cell and are correctly delivered at the daughter bud neck by the Golgi pathway. The regulatory proteins Cbk1 and Mob2 localize to the neck and to the daughter nuclei at the end of mitosis [36], in particular they form a complex with the transcriptional factor Ace2 and prevent its export from the nucleus, thus ensuring daughter-specific expression of cell separation factors [37]. In addition, Cts1 is post-translationally modified and delivered to the daughter bud neck through the Golgi pathway [38]. Recently, a new level of Cts1 regulation has been described: upon incomplete septation, Fir1 inhibits Cbk1 activity, thus blocking production and secretion of septum degrading enzymes [39]. This mechanism inhibits cell separation until septation is properly executed and, therefore, enforces cytokinesis order of events.
Cell separation is the very last event of the cell cycle and it can have great implications in cell survival. Indeed, separation of daughter from the mother must occur in correlation with other cell cycle events and a well-organized trilaminar septum structure must be formed before cell separation machinery is activated in order to obtain vital progeny. In case of overactivity of Cts1, a hole in the cell wall can be formed, in this case Chs1 is activated and repairs the cell wall at the daughter side [40].
To sum up, the successful completion of cell separation depends upon the precise construction of the trilaminar septum in coordination with the division cycle, and then on the temporal and spatial regulation of chitinase and glucanase action.
4. Checkpoint Pathways that Control Cytokinesis
Cytokinesis must be tightly coordinated with the nuclear cycle in order to maintain genetic stability. Eukaryotic cells have evolved several checkpoints and checkpoint-like mechanisms that preserve the integrity of cell division.
The mitotic checkpoint is an evolutionarily conserved signaling pathway that blocks mitosis progression in case of problems in the mitotic spindle, in chromosome attachment to MTs, or in presence of mistakes in chromosome segregation. In budding yeast, the division site is determined before mitotic spindle formation, which implies that the bipolar spindle must be properly positioned at the bud neck and aligned perpendicularly with respect to the division axis before anaphase onset. Two pathways direct spindle positioning, the dynein pathway and Kar9 pathway, and the spindle orientation checkpoint (SPOC) blocks mitotic exit and cytokinesis in the case of defects [41]. If the checkpoint fails, cytokinesis occurs, even if the nucleus divides into the mother cell, thus causing the formation of aneuploid cells. The target of the SPOC is Tem1, the G protein at the top of MEN pathway.
Mitotic Exit Network (MEN) in S. cerevisiae and Septation Initiation Network (SIN) in S. pombe are two signaling pathways that coordinate mitosis progression from anaphase onset to cell separation. MEN is a signaling cascade with the GTPase Tem1 at the top, several protein kinases, among which the Hippo-like kinase Cdc15 and the LATS-like kinase Dbf2, and the final target is the phosphatase Cdc14. MEN pathway leads to complete Cdc14 activation that causes a decrease in mitotic kinase activity (Cdk1), triggering mitotic exit [42]. Before anaphase onset, MEN components localize at the spindle pole bodies (the yeast centrosomes), but later they are found at the bud neck, where they ensure that mitotic exit only occurs after acto-myosin ring contraction and septum deposition [43]. In addition, in telophase, Cdc14 dephosphorylates cytoplasmic and bud-neck associated targets of Cdk1 and contributes to reorganize actin cytoskeleton and target secretion vesicles to the bud neck [44][45]. In S. pombe the SIN controls CAR contraction and septum deposition, rather than mitotic exit and blocks septation in the case of problems [46]. Homologs of Cdc14, MEN, and SIN components are also found in other yeasts: Candida albicans, Aspergillus nidulans, and Ashbya gossypii. Budding yeast Cdc14 is conserved in higher eukaryotes, however these homologs do not seem to control mitotic exit, rather they are involved in other processes, such as DNA replication, DNA damage repair, nuclear organization, mitotic entry, mitotic spindle assembly, and cytokinesis [47]. In Drosophila and human cells, MEN homologs are part of the Hippo pathway that controls centrosome duplication, cell proliferation, and apoptosis [48][49]. The conserved Hippo pathway inhibits cell proliferation by the activation of LATS1 and LATS2 kinases and by p53 stabilization (see below).
Mammalian cells have a checkpoint that arrest cells in mitosis in response to several problems in order to preserve genetic stability and cell survival, rarely cells can escape this arrest and enter mitosis without chromosome segregation, thus becoming tetraploid [50][51]. Importantly, another checkpoint control is active in human cells to recognize cytokinesis failure and induce proliferation arrest in G1. Indeed, in tetraploid cells, arising either by failure of mitotic spindle or of cytokinesis, the checkpoint induces a p53-dependent cell cycle arrest, thus preventing the proliferation of aneuploid cells and carcinogenesis [52]. However it is not clear how, at which stage and by which signal (from extra chromosomes or extra centrosomes) p53 is activated by tetraploidy. Interestingly, cytokinesis failure activates the Hippo pathway: LATS2, which is an important kinase of Hippo pathway, translocates from centrosomes to the nucleus and stabilizes p53 in the presence of additional centrosomes [53][54]. In addition, extra centrosomes can activate RAC1, which is known to antagonize RhoA, which leads to LATS2 activation [55]. If p53 function is lost, cells can override the cell cycle block that is induced by tetraploidization, accumulate chromosome aberrations, and start to proliferate without control.
Cytokinesis can be blocked even when the furrow started to ingress: the abscission checkpoint delays cytokinesis completion and can even drive the regression of the cleavage furrow in the presence of chromatin bridges or lagging chromosomes [56]. The abscission delay is dependent upon the localization of the Aurora B kinase at the midbody. Aurora B translocates from kinetochores to the division site after anaphase onset. At the midbody, activated Aurora B phosphorylates the ESCRT-III subunit Chmp4c, but the molecular mechanism that delays abscission is not completely understood.
Human cells are well equipped with several pathways that block tetraploid cells proliferation since cytokinesis failure can be tumorigenic. However tetraploidization is used by evolution and it can be important for certain tissues to gain new advantageous traits, such as resistance to drugs or stimuli (see also paragraph 6).
Interesting data suggest new connections between cytokinesis and DNA damage, in particular cytokinesis might be regulated in response to DNA damage. Damaged DNA should not be segregated, so, in this case, cytokinesis must be blocked to prevent the cleavage furrow from cutting damaged DNA. Proteins that are involved in DNA repair, such as BRCA2 and BCCIP, may be directly involved in cytokinesis, as their deficiency induces cytokinetic abnormalities [57]. The polo kinase Plk1 is an important regulator of mitosis and cytokinesis and it is also a modulator of the DNA damage checkpoint [58]. The budding yeast DNA damage checkpoint kinase Rad53 localizes at the division site and associates with septins [59]. Other experiments suggest that DNA damage pathways may regulate cytokinesis proteins by modulating their the expression or their post-translational modifications, as p53 and Rb pathways inhibit the expression of cytokinetic proteins, such as Plk1, ECT2, anillin, and survivin [60][61], and BRCT inhibits Aurora B kinase activity by Poly(ADP-ribosyl)ation in response to DNA strand breaks [62]. It is reasonable that DNA damage, and perhaps incompletely replicated DNA, activates pathways that promote DNA repair, arrests cell cycle progression, and blocks cytokinesis completion.
References
- Storchova, Z.; Pellman, D. From polyploidy to aneuploidy, genome instability and cancer. Nat. Rev. Mol. Cell Biol. 2004, 5, 45–54.
- Fraschini, R. Divide Precisely and Proliferate Safely: Lessons from Budding Yeast. Front. Genet. 2019, 9, 738.
- Guha, M.; Zhou, M.; Wang, Y.L. Cortical actin turnover during cytokinesis requires myosin II. Curr. Biol. 2005, 15, 732–736.
- Matsumura, F.; Ono, S.; Yamakita, Y.; Totsukawa, G.; Yamashiro, S. Specific localization of serine 19 phosphorylated myosin II during cell locomotion and mitosis of cultured cells. J. Cell Biol. 1998, 140, 119–129.
- Piekny, A.J.; Glotzer, M. Anillin is a Scaffold Protein that Links RhoA, Actin, and Myosin during Cytokinesis. Curr. Biol. 2008, 18, 30–36.
- Kinoshita, M.; Field, C.M.; Coughlin, M.L.; Straight, A.F.; Mitchison, T.J. Self- and actintemplated assembly of Mammalian septins. Dev. Cell 2002, 3, 791–802.
- Dobbelaere, J.; Barral, Y. Spatial coordination of cytokinetic events by compartmentalization of the cell cortex. Science 2004, 305, 393–396.
- Mabuchi, I.; Tsukita, S.; Tsukita, S.; Sawai, T. Cleavage furrow isolated from newt eggs: Contraction, organization of the actin filaments, and protein components of the furrow. Proc. Natl. Acad. Sci. USA 1988, 85, 5966–5970.
- Henson, J.H.; Ditzler, C.E.; Germain, A.; Irwin, P.M.; Vogt, E.T.; Yang, S.; Wu, X.; Shuster, C.B. The ultrastructural organization of actin and myosin II filaments in the contractile ring: New support for an old model of cytokinesis. Mol. Biol. Cell 2017, 28, 613–623.
- Murthy, K.; Wadsworth, P. Myosin-II-dependent localization and dynamics of F-actin during cytokinesis. Curr. Biol. 2005, 15, 724–731.
- Yumura, S. Myosin II dynamics and cortical flow during contractile ring formation in Dictyostelium cells. J. Cell Biol. 2001, 154, 137–146.
- Frappaolo, A.; Sechi, S.; Kumagai, T.; Robinson, S.; Fraschini, R.; Karimpour-Ghahnavieh, A.; Belloni, G.; Piergentili, R.; Tiemeyer, K.H.; Tiemeyer, M.; et al. COG7 deficiency in Drosophila generates multifaceted developmental, behavioral and protein glycosylation phenotypes. J. Cell Sci. 2017, 130, 3637–3649.
- Schweitzer, J.K.; Burke, E.E.; Goodson, H.V.; D’Souza-Schorey, C. Endocytosis resumes during late mitosis and is required for cytokinesis. J. Biol. Chem. 2005, 280, 41628–41635.
- Sechi, S.; Frappaolo, A.; Fraschini, R.; Capalbo, L.; Gottardo, M.; Belloni, G.; Glover, D.M.; Wainman, A.; Giansanti, M.G. Rab1 interacts with GOLPH3 and controls Golgi structure and contractile ring constriction during cytokinesis in Drosophila melanogaster. Open Biol. 2017, 7, 160257.
- Shuster, C.B.; Burgess, D.R. Targeted new membrane addition in the cleavage furrow is a late, separate event in cytokinesis. Proc. Natl. Acad. Sci. USA 2002, 99, 3633–3638.
- Kanada, M.; Nagasaki, A.; Uyeda, T.Q. Adhesion-dependent and contractile ring-independent equatorial furrowing during cytokinesis in mammalian cells. Mol. Biol. Cell 2005, 16, 3865–3872.
- Mendes Pinto, I.; Rubinstein, B.; Kucharavy, A.; Unruh, J.R.; Li, R. Actin depolymerization drives actomyosin ring contraction during budding yeast cytokinesis. Dev. Cell 2012, 22, 1247–1260.
- Dix, C.L.; Matthews, H.K.; Uroz, M.; McLaren, S.; Wolf, L.; Heatley, N.; Win, Z.; Almada, P.; Henriques, R.; Boutros, M.; et al. The Role of Mitotic Cell-Substrate Adhesion Re-modeling in Animal Cell Division. Dev. Cell 2018, 45, 132–145.
- Taneja, N.; Rathbun, L.; Hehnly, H.; Burnette, D.T. The balance between adhesion and contraction during cell division. Curr. Opin. Cell Biol. 2019, 56, 45–52.
- Christ, L.; Raiborg, C.; Wenzel, E.M.; Campsteijn, C.; Stenmark, H. Cellular functions and molecular mechanisms of the ESCRT membrane-scission machinery. Trends Biochem. Sci. 2017, 42, 42–56.
- Gromley, A.; Yeaman, C.; Rosa, J.; Redick, S.; Chen, C.T.; Mirabelle, S.; Guha, M.; Sillibourne, J.; Doxsey, S.J. Centriolin anchoring of exocyst and SNARE complexes at the midbody is required for secretory-vesicle-mediated abscission. Cell 2005, 123, 75–87.
- Elia, N.; Sougrat, R.; Spurlin, T.A.; Hurley, J.H.; Lippincott-Schwartz, J. Dynamics of endosomal sorting complex required for transport (ESCRT) machinery during cytokinesis and its role in abscission. Proc. Natl. Acad. Sci. USA 2011, 108, 4846–4851.
- Pruyne, D.; Bretscher, A. Polarization of cell growth in yeast. I. Establishment and maintenance of polarity states. J. Cell Sci. 2000, 113, 365–375.
- Byers, B.; Goetsch, L. A highly ordered ring of membrane-associated filaments in budding yeast. J. Cell Biol. 1976, 69, 717–721.
- Glomb, O.; Gronemeyer, T. Septin Organization and Functions in Budding Yeast. Front. Cell Dev. Biol. 2016, 4, 123.
- Cassani, C.; Raspelli, E.; Chiroli, E.; Fraschini, R. Vhs2 is a novel regulator of septin dynamics in budding yeast. Cell Cycle 2014, 13, 1590–1601.
- Longtine, M.S.; Fares, H.; Pringle, J.R. Role of the yeast Gin4p protein kinase in septin assembly and the relationship between septin assembly and septin function. J. Cell Biol. 1998, 3, 719–736.
- Caudron, F.; Barral, Y. Septins and the lateral compartmentalization of eukaryotic membranes. Dev. Cell 2009, 16, 493–506.
- Lippincott, J.; Li, R. Involvement of PCH family proteins in cytokinesis and actin distribution. Microsc. Res. Tech. 2000, 49, 168–172.
- Bi, E. Cytokinesis in budding yeast: The relationship between actomyosin ring function and septum formation. Cell Struct. Funct. 2001, 26, 529–537.
- Bi, E. Involvement of an actomyosin contractile ring in Saccharomyces cerevisiae cytokinesis. J. Cell Biol. 1998, 142, 1301–1312.
- Shaw, J.A.; Mol, P.C.; Bowers, B.; Silverman, S.J.; Valdivieso, M.H.; Durán, A.; Cabib, E. The function of chitin synthases 2 and 3 in the Saccharomyces cerevisiae cell cycle. J. Cell Biol. 1991, 114, 111–123.
- VerPlank, L.; Li, R. Cell cycle-regulated trafficking of Chs2 controls actomyosin ring stability during cytokinesis. Mol. Biol. Cell 2005, 16, 2529–2543.
- Chin, C.F.; Bennett, A.M.; Ma, W.K.; Hall, M.C.; Yeong, F.M. Dependence of Chs2 ER export on dephosphorylation by cytoplasmic Cdc14 ensures that septum formation follows mitosis. Mol. Biol. Cell 2012, 23, 45–58.
- Devrekanli, A.; Foltman, M.; Roncero, C.; Sanchez-Diaz, A.; Labib, K. Inn1 and Cyk3 regulate chitin synthase during cytokinesis in budding yeasts. J. Cell Sci. 2012, 125, 5453–5466.
- Lesage, G.; Bussey, H. Cell wall assembly in Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 2006, 70, 317–343.
- Chin, C.F.; Tan, K.; Onishi, M.; Chew, Y.; Augustine, B.; Lee, W.R.; Yeong, F.M. Timely Endocytosis of Cytokinetic Enzymes Prevents Premature Spindle Breakage during Mitotic Exit. PLoS Genet. 2016, 12, e1006195.
- Yeong, F.M. Severing all ties between mother and daughter: Cell separation in budding yeast. Mol. Microbiol. 2005, 55, 1325–1331.
- Colman-Lerner, A.; Chin, T.E.; Brent, R. Yeast Cbk1 and Mob2 activate daughter-specific genetic programs to induce asymmetric cell fates. Cell 2001, 107, 739–750.
- Weiss, E.L.; Kurischko, C.; Zhang, C.; Shokat, K.; Drubin, D.G.; Luca, F.C. The Saccharomyces cerevisiae Mob2p–Cbk1p kinase complex promotes polarized growth and acts with the mitotic exit network to facilitate daughter cell-specific localization of Ace2p transcription factor. J. Cell Biol. 2002, 158, 885–900.
- Kuranda, M.J.; Robbins, P.W. Chitinase is required for cell separation during growth of Saccharomyces cerevisiae. J. Biol. Chem. 1991, 266, 19758–19767.
- Brace, J.L.; Doerfler, M.D.; Weiss, E.L. A cell separation checkpoint that enforces the proper order of late cytokinetic events. J. Cell Biol. 2019, 218, 150–170.
- Cabib, E.; Sburlati, A.; Bowers, B.; Silverman, S.J. Chitin synthase 1, an auxiliary enzyme for chitin synthesis in Saccharomyces cerevisiae. J. Cell Biol. 1989, 108, 1665–1672.
- Caydasi, A.K.; Pereira, G. SPOC alert—when chromosomes get the wrong direction. Exp. Cell Res. 2012, 318, 1421–1427.
- Baro, B.; Queralt, E.; Monje-Casas, F. Regulation of Mitotic Exit in Saccharomyces cerevisiae. Methods Mol. Biol. 2017, 1505, 3–17.
- Meitinger, F.; Petrova, B.; Lombardi, I.M.; Bertazzi, D.T.; Hub, B.; Zentgraf, H.; Pereira, G. Targeted localization of Inn1, Cyk3 and Chs2 by the mitotic-exit network regulates cytokinesis in budding yeast. J. Cell Sci. 2010, 123, 1851–1861.
- Bloom, J.; Cristea, I.M.; Procko, A.L.; Lubkov, V.; Chait, B.T.; Snyder, M.; Cross, F.R. Global analysis of Cdc14 phosphatase reveals diverse roles in mitotic processes. J. Biol. Chem. 2011, 286, 5434–5445.
- Cassani, C.; Raspelli, E.; Santo, N.; Chiroli, E.; Lucchini, G.; Fraschini, R. Saccharomyces cerevisiae Dma proteins participate in cytokinesis by controlling two different pathways. Cell Cycle 2013, 12, 2794–2808.
- Simanis, V. SIN review Pombe’s thirteen-control of fission yeast cell division by the septation initiation network. J. Cell Sci. 2015, 128, 1465–1474.
- Mocciaro, A.; Schiebel, E. Cdc14: A highly conserved family of phosphatases with non-conserved functions? J. Cell Sci. 2010, 123, 2867–2876.
- Mardin, B.R.; Lange, C.; Baxter, J.E.; Hardy, T.; Scholz, S.R.; Fry, A.M.; Schiebel, E. Components of the Hippo pathway cooperate with Nek2 kinase to regulate centrosome disjunction. Nat. Cell Biol. 2010, 12, 1166–1176.
- Zhao, B.; Tumaneng, K.; Guan, K.L. The Hippo pathway in organ size control, tissue regeneration and stem cell self-renewal. Nat. Cell Biol. 2011, 13, 877–883.
- Ganem, N.J.; Godinho, S.A.; Pellman, D. A mechanism linking extra centrosomes to chromosomal instability. Nature 2009, 460, 278–282.
- Silkworth, W.T.; Nardi, I.K.; Scholl, L.M.; Cimini, D. Multipolar spindle pole coalescence is a major source of kinetochore mis-attachment and chromosome mis-segregation in cancer cells. PLoS ONE 2009, 4, e6564.
- Andreassen, P.R.; Lohez, O.D.; Lacroix, F.B.; Margolis, R.L. Tetraploid state induces p53-dependent arrest of nontransformed mammalian cells in G1. Mol. Biol. Cell 2001, 12, 1315–1328.
- Aylon, Y.; Michael, D.; Shmueli, A.; Yabuta, N.; Nojima, H.; Oren, M. A positive feedback loop between the p53 and Lats2 tumor suppressors prevents tetraploidization. Genes Dev. 2006, 20, 2687–2700.
- Ganem, N.J.; Cornils, H.; Chiu, S.Y.; O’Rourke, K.P.; Arnaud, J.; Yimlamai, D.; Théry, M.; Camargo, F.D.; Pellman, D. Cytokinesis failure triggers hippo tumor suppressor pathway activation. Cell 2014, 158, 833–848.
- Godinho, S.A.; Picone, R.; Burute, M.; Dagher, R.; Su, Y.; Leung, C.T.; Polyak, K.; Brugge, J.S.; Théry, M.; Pellman, D. Oncogene-like induction of cellular invasion from centrosome amplification. Nature 2014, 510, 167–171.
- Petsalaki, E.; Zachos, G. Building bridges between chromosomes: Novel insights into the abscission checkpoint. Cell. Mol. Life Sci. 2019, 76, 4291–4307.
- Daniels, M.J.; Wang, Y.; Lee, M.; Venkitaraman, A.R. Abnormal cytokinesis in cells deficient in the breast cancer susceptibility protein BRCA2. Science 2004, 306, 876–879.
- Takaki, T.; Trenz, K.; Costanzo, V.; Petronczki, M. Polo-like kinase 1 reaches beyond mitosis--cytokinesis, DNA damage response, and development. Curr. Opin. Cell Biol. 2008, 20, 650–660.
- Smolka, M.B.; Chen, S.H.; Maddox, P.S.; Enserink, J.M.; Albuquerque, C.P.; Wei, X.X.; Desai, A.; Kolodner, R.D.; Zhou, H. An FHA domain-mediated protein interaction network of Rad53 reveals its role in polarized cell growth. J. Cell Biol. 2006, 175, 743–753.
- Jackson, M.W.; Agarwal, M.K.; Yang, J.; Bruss, P.; Uchiumi, T.; Agarwal, M.L.; Stark, G.R.; Taylor, W.R. p130/p107/p105Rb-dependent transcriptional repression during DNA-damage-induced cell-cycle exit at G2. J. Cell Sci. 2005, 118, 1821–1832.
- Date, D.A.; Jacob, C.J.; Bekier, M.E.; Stiff, A.C.; Jackson, M.W.; Taylor, W.R. Borealin is repressed in response to p53/Rb signaling. Cell Biol. Int. 2007, 31, 1470–1481.
- Monaco, L.; Kolthur-Seetharam, U.; Loury, R.; Murcia, J.M.; de Murcia, G.; Sassone-Corsi, P. Inhibition of Aurora-B kinase activity by poly(ADP-ribosyl)ation in response to DNA damage. Proc. Natl. Acad. Sci. USA 2005, 102, 14244–14248.
More
Information
Subjects:
Cell Biology
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.1K
Revisions:
2 times
(View History)
Update Date:
20 May 2021
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





