
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Eric Dannaoui | + 7789 word(s) | 7789 | 2021-03-17 04:48:23 | | | |
| 2 | Eric Dannaoui | + 10 word(s) | 7799 | 2021-03-24 16:28:45 | | | | |
| 3 | Catherine Yang | -4252 word(s) | 3547 | 2021-03-25 03:27:31 | | |
Video Upload Options
The general approach of studying antifungal combinations is (i) to choose an experimental technique, (ii) to obtain raw numerical data, (iii) to analyze these data either graphically or numerically; (iv) to interpret the results, and (v) to conclude on the mode of interaction: synergy, indifference (no interaction), or antagonism.
1. Introduction
Fungal infections are serious pathologies that, despite adequate treatment, have high mortality rates [1][2]. In addition, besides natural resistance in some species, acquired resistance to antifungals is increasing [3][4]. Therefore, new therapeutic alternatives are needed. At present, only a few antifungals belonging to a limited number of antifungal classes with different mechanisms of action are on the market [3]. Despite the urgent need for new antifungals and antifungal classes [5], a promising therapeutic strategy would be to use antifungals in combination. Indeed, one of the main advantages of combining antifungals is to overcome resistance [6]. Moreover, antifungal combination can increase the efficacy of the combined molecules yielding to synergy. Combination therapy can also reduce toxicity by decreasing antifungal dosages, and improve the pharmacokinetics of one or both molecules [7]. Antifungal combinations are already used in clinical practice, such as 5-flucytosine combined with amphotericin B as first-line treatment for cryptococcal meningitis [8]. Moreover, it is also important to know if a combination exhibits antagonism.
Regarding the general approach of studying antifungal combinations, several steps are needed to perform and interpret antifungal combination tests (Figure 1). The first step is to choose an experimental technique: a liquid dilution method (e.g., checkerboard), a method of agar diffusion (e.g., gradient concentration strips such as Etest), or a study of fungicidal effect (e.g., time-kill). Using these methods, raw numerical data are obtained: minimal inhibitory concentrations (MIC), inhibition diameters, or number of colony-forming units (CFU) over time. The MIC data for example, are then analyzed, either using a graphical method (surface analysis), or by calculation of the inhibitory fractional concentration index (FIC index), and interpreted according to consensual thresholds or predetermined criteria. Finally, based on the results, a mode of interaction that is synergy, indifference (no interaction), or antagonism can be concluded. Currently, none of these steps are standardized, and therefore a large number of variables can influence the final results.
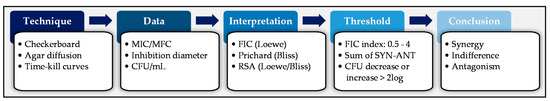
Figure 1. Summary of steps needed to perform and interpret antifungal combination tests. MIC, minimal inhibitory concentration; MFC, minimal fungicidal concentration; CFU, colony forming unit; FIC, fractional inhibitory concentration; RSA, response-surface analysis; SYN, synergy; ANT, antagonism.
2. In Vitro Techniques
To study antifungal combinations, several experimental techniques are possible. Each method has advantages, but also disadvantages (Table 1).
Table 1. Summary of the advantages and disadvantages of the different methods used to study antifungal combinations in vitro.
| Techniques | Advantages | Disadvantages |
|---|---|---|
| Checkerboard method | Quantitative | Discontinuous gradient of antifungal concentration |
| Automated reading of results | Lack of standardization in interpretation of results | |
| Agar diffusion assay (disks or gradient strips) | Continuous gradient of antifungal concentration | Qualitative for disks |
| Possible use of commercialized systems (gradient strips) | Difficult to assess at which concentrations interaction occurs | |
| Time-kill curves | Quantitative | Lack of standardization |
| Fungicidal exploration and rate of killing | Only a few concentrations studied at the same time |
2.1. Liquid Microdilution Technique: Checkerboard
The checkerboard method is generally based on the standardized EUCAST [9] or CLSI [10] broth micro-dilution techniques and performed in 96-well microplates [7][11]. Initially, each drug is diluted in series, usually using a dilution factor of two. These solutions are added to the culture medium (Roswell Park Memorial Institute Medium, RPMI), which is then distributed in a 96-well microplate (Figure 2). After preparation of the microplates, each well is inoculated with the fungal inoculum (yeast cells or conidia), and microplates are then incubated. To be able to interpret the results correctly, sufficient two-fold dilutions below and above the MIC have to be included for each antifungal. Reading can be performed either visually or spectrophotometrically. Nevertheless, for a more objective MIC determination and possible automation, the spectrophotometric method should be preferred [7][12]. After reading of the microplates, the quantitative data can be analyzed in different ways.
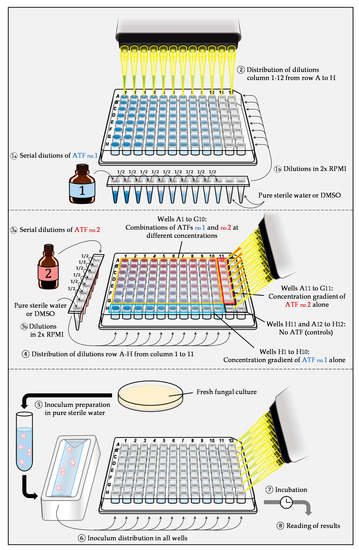
Figure 2. Example of preparation and inoculation of microplates using the checkerboard method based on the EUCAST methodology for antifungal susceptibility testing.
The checkerboard is the most often used in vitro technique [11][13], and is therefore considered to be the “reference” method, even though there is currently no consensus regarding the reference technique to be used for assessing antifungal combinations. Nevertheless, this technique has some drawbacks, in particular, the range of concentrations tested is discontinuous and the dilutions are performed in a geometric manner. This means that only certain combinations of concentrations can be evaluated on the microplate, and the error in determining the MICs is not the same over the entire concentration range. The checkerboard technique can also be used to test triple combinations [14]. It has been used to test triple combinations in the field of antivirals (e.g., against HIV) [15], antibiotics (e.g., against Mycobacteriacae and Enterobacteriacae) [16][17], and antifungal agents against Aspergillus spp. [18], Cryptococcus neoformans [19][20], Candida albicans [21][22][23], Mucorales [24], and Scedosporium spp. [25][26].
2.2. Agar-Medium Diffusion Techniques
Agar-medium diffusion techniques are widely used to determine antifungal susceptibilities. These methods can be adapted in different ways to study antifungal combinations.
2.2.1. Disk Diffusion Method
Disks impregnated with one of the two antifungal agents are placed face to face on an agar previously inoculated with the strain to be studied. The optimal distance between the discs to visualize the interaction should be determined in preliminary experiments. After growth of the microorganism, growth inhibition zones are obtained around each of the disks. In the zone were the diffusion of both antifungals is overlapping, special inhibition zones can be recognized. Depending on the growth characteristic of the strain on these zones, the interaction can be concluded.
Another technique is to use a disk impregnated with an antifungal agent, while the second antifungal is incorporated into the agar at a sub-inhibitory concentration. The inhibition zone obtained is compared to that of the control, i.e., agar without an antifungal agent. Compared to the control agar, an increase or decrease of the inhibition diameter around the disk will be obtained in cases of synergy or antagonism, respectively. Another way to detect antagonism is to incorporate the antifungal in the agar at a concentration higher than the MIC of the strain. In case of antagonism, growth of the microorganism will occur only around the disk [27].
This method has been used to evaluate antifungal combinations or combinations of antifungals with non-antifungal drugs against Candida spp. [28][29][30], Cryptococcus spp. [30], and dermatophytes [31].
2.2.2. Right Angle Scattering Method
One of the common methods to assess interactions of antimicrobial drugs is the right angle scattering method [32]. It consists of placing two drug-impregnated paper strips at right angles on an agar plate. Depending on the growth characteristics of the microorganism in the area, where drug diffusion into the agar is overlapping, either synergy, indifference, or antagonism can be concluded. The technique is easy and fast to perform, but has only seldom been used for assessing antifungal combinations [33]. The fact that the technique is a diffusion method makes it possible to obtain a continuous gradient of the concentrations of the antifungals. However, the method has also several drawbacks. It is only qualitative, and the interpretation remains subjective as it depends on the growth of the microorganism on only a few millimeter-wide overlapping zone of the antifungals, which may vary between the experiments [7][27]. Additionally, the choice of concentrations of the antifungals on the paper strips makes preliminary experiments necessary.
2.2.3. Gradient Concentration Strip (Etest) method
Gradient concentration strips allow researchers to measure the MICs of antifungals. Strips are impregnated with concentration gradients of the molecules [34]. Even though this is not the reference method for antifungal susceptibility testing, it is a simple test to determine MICs. Gradient concentration strips can also be used to test interactions between drugs [7][27][35]. The endpoints used for MIC determination (complete or partial inhibition) for antifungal combination are the same as those used when drugs are tested alone. Due to the existence of registered trademarks (e.g., Etest), the reproducibility of the technique is good. Several methods are used to assess antifungal combinations.
The first method is used when strips are commercially available for both antifungals. After determination of the MICs alone, the MIC in combination can be evaluated in three different ways.
- (i)
-
The cross protocol
The strips of antifungal A and antifungal B are crossed at a 90° angle at the position of their MICs alone. This protocol has been used to test antibiotic combinations against gram-negative and gram-positive bacteria [36], but also to test voriconazole combined with either caspofungin or amphotericin B against Candida spp. [37], and to test various combinations against Candida glabrata [38][39][40].
- (ii)
-
The fixed ratio protocol
The strip of antifungal A is placed on the agar and is replaced after diffusion of the antifungal into the agar by the strip of antifungal B on exactly the same position as the first strip (Figure 3) [7][41]. This method has been used to test combinations against C. glabrata [38], C. neoformans [42], and Aspergillus spp. [43][44][45].
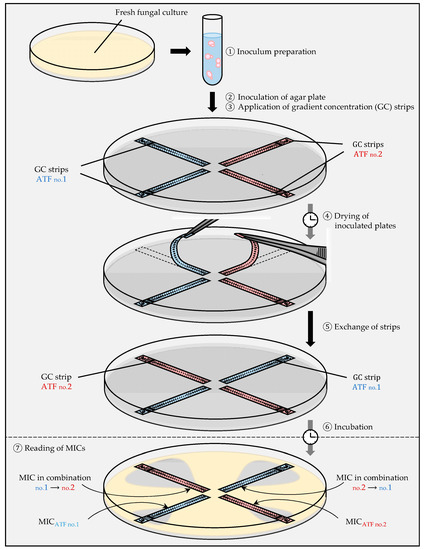
Figure 3. Gradient concentration strip method (Etest) for the determination of antifungal interactions: the fixed ratio protocol.
- (iii)
-
The MIC/MIC ratio protocol
The strip of antifungal A is applied onto the agar and is removed after 1 hour. After vertical transposition, the strip of antifungal B is applied on the agar surface, so the MIC of antifungal A meets the MIC of antifungal B, or a fraction of the MIC. Polymyxin B combined with fluconazole or caspofungin has been evaluated against C. glabrata by this method and showed synergistic interactions [46][47]. Synergy has also been found for combinations of doxycycline or tigecycline with fluconazole against C. glabrata [48].
The second method is used when no gradient strips are available for one of the two drugs. The MIC of antifungal A is determined by a gradient strip alone, and the MIC in combination after drug B has been incorporated in the agar at a fixed concentration [18][28][49][50][51][52][53][54][55]. A control plate with drug B alone is generally added to ensure that drug B is at a sub-inhibitory concentration. This method has been used to assess combinations of antifungals or combinations of antifungals with non-antifungal drugs against Candida spp. [42][49][56], Aspergillus spp. [18][51][54][57], and Mucorales [50].
2.3. Time-Kill Curves
Unlike the previous techniques which measure the inhibition of growth after a predetermined time point, time-kill curves measure the kinetics of fungicidal activity [7][41]. Fungal killing is calculated by measuring the colony forming units (CFU) at predetermined time points. The CFU are determined from tubes containing RPMI medium with the antifungals either alone or in combination. The concentrations of the antifungals are either fractions or multiples of the MICs. To interpret the results of this technique, it is necessary to compare the fungicidal activity of the combination to that obtained by the most active antifungal alone [6][7][27][41]. This method has been used to evaluate antifungal combinations or combinations of antifungals with non-antifungal drugs against Candida spp. [12][38][39][41][46][47][49][58][59][60][61][62][63][64][65][66][67][68][69][70][71][72][73][74][75][76][77][78][79][80][81][82][83][84][85][86], Cryptococcus spp. [42][59][68][70][83], and Aspergillus spp. [65][68][87][88][89][90][91]. Synergy or antagonism are defined by a decrease or an increase of ≥ 2 log10 CFU/mL of the combination compared to the most active drug [7][27]. The main advantage of this quantitative technique is the possibility to explore the fungicidal activity of combinations. The disadvantages are that technical parameters and the interpretation of the results are not standardized.
2.4. Analysis of Results and Interpretation
Several methods can be used to assess the combined effect of drugs that are tested in combination experiments (Figure 1).
There are several theoretical approaches to model the interaction between pharmacologically active molecules. In the field of antifungal drugs, two theories are mainly used.
The first is based on the Loewe additivity model. The model is based on the hypothesis that a drug does not interact with itself, which means the combination of a drug with itself, gives, by definition, an indifferent interaction. It is a dose-effect based strategy, meaning that concentrations that give a certain effect are compared [92]. Several methods can be applied to analyze the interactions of two drugs based on the Loewe theory, for example, intuitive graphical analysis such as the isobologram [93] or its algebraic counterpart based on the calculation of the FIC index [11]. Other approaches can also be used, such as the Greco model [94], the median-effect approach of Chou and Talabay [95], or response surface approaches [96].
The second is based on the Bliss independence model. The model is based on the hypothesis that two drugs act independently of each other. No interaction is obtained when the effect of the combination is equal to the product of the effects of the drugs alone. This approach compares the effects, instead of the concentrations, of drugs alone, or in combination. If the observed effect is better or worse than the expected indifferent interaction, the combination is defined as synergistic or antagonistic, respectively. Several methods of analysis, such as the Prichard model [97], have been developed based on the Bliss theory. Response-surface analysis can also be implemented based on the Bliss independence model [96]. Besides the Bliss independence model, other effect-based strategies can be used. These include the combination sub-thresholding, the highest single agent, and the response additivity approach [92].
2.4.1. FIC Index
The fractional inhibitory concentration index, or FIC index, can be used to determine the effect of a tested combination. To determine the FIC index, the fractional inhibitory concentrations (FIC) of both drugs are added. The FIC is calculated by division of the MIC in combination and the MIC alone of the tested drug. The FIC index is calculated according to the following formula:
MIC 1 alone and MIC 2 alone are the MICs of antifungals 1 and 2 when tested alone, and MIC combo 1 and MIC combo 2 are the MICs of antifungals 1 and 2 in combination.
In theory, a FIC index = 1 represents an additivity, while a FIC index < 1 is indicative of a synergy and a FIC index > 1 of an antagonism. Nevertheless, broth microdilution techniques have an intrinsic variability of at least one log2 dilution. Therefore, the FIC index threshold used to analyze the results should reflect this variability. Currently, the recommendation to interpret the FIC index is as follows: interaction is synergistic when the FIC index is ≤ 0.5, indifferent if the FIC index is > 0.5 to 4, and antagonistic if the FIC index > 4 [98] (Figure 4).
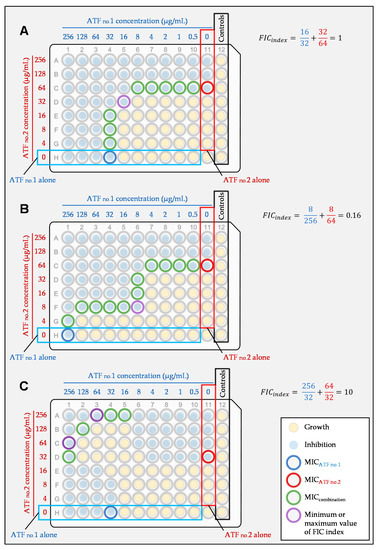
Figure 4. Example of a synergistic (A), indifferent (B), and antagonistic (C) interaction of two antifungals according to the checkerboard method and calculated by the FIC index. If there is no FIC index > 4, then the lowest FIC index is retained. If there is at least one FIC index > 4, then the highest FIC index is retained. Synergy is defined as a FIC index ≤ 0.5, indifference as a FIC index > 0.5 to 4, and antagonism as a FIC index > 4.
With the checkerboard method, different combinations of concentrations of the antifungals are tested at the same time. It is therefore possible to calculate several FIC indices for the tested combination. The minimum FIC index is reported in absence of antagonism, and the maximum FIC index in case of antagonism. Defining the threshold of the FIC index is one of the problems of this approach, but there are others, such as the evaluation of the MIC itself. Depending on the endpoint used for MIC determination (50% or 90% of growth inhibition compared to the growth control), one can come to completely different conclusions [11][27].
2.4.2. Surface Response Modeling
Response surface analysis is an alternative approach that does not require the determination of MICs. Unlike the FIC index, it is therefore independent of an inhibition endpoint. Moreover, it allows for the calculation and visualization of the combined effect of the two molecules for all tested concentrations, and not only for those corresponding to an MIC. This approach can be based on different theories (Loewe, Bliss, and other) and calculations are generally performed by dedicated software. In this approach, the inhibition curve of each antifungal agent is modeled on the basis of the growth rate obtained in each well containing the molecule alone [96]. From these dose-response curves, a theoretical growth inhibition matrix (represented by a theoretical dose-response surface) is modeled, corresponding to the inhibition rates expected in each well for the case where the interaction is purely indifferent, according to the chosen theory (Loewe, Bliss, or other model). The matrix of the experimental data (represented by an experimental dose-response surface) is then compared to the theoretical matrix. If the observed growth is weaker (stronger inhibition), synergy is concluded (Figure 5), whereas if the observed growth is stronger (weaker inhibition), antagonism is concluded. Apart of the graphical output, it is possible to generate metrics (for example the SUM-SYN-ANT metric in the Combenefit software), which can be used to quantitatively assess the drug interactions. Taking into account the intrinsic variability of the broth microdilution checkerboard technique, it is necessary to generate experimental data of the antifungals combined with themselves in order to define the threshold used for the interpretation of the metric [21][22].
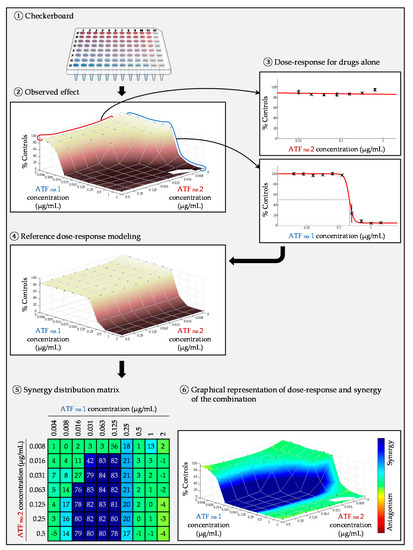
Figure 5. Example of a response-surface analysis for an in vitro antifungal combination experiment. Graphics were generated by the Combenefit software [96].
3. In Vivo Techniques
It is important to confirm the in vitro data by in vivo data. As the incidence of most fungal infections compared to bacterial infections is lower, it is very difficult to perform clinical trials in patients, although it has been done in some instances. Combination of amphotericin B with flucytosine has been tested for the treatment of cryptococcal meningitis [99][100], or combination of voriconazole with anidulafungin for the treatment of invasive aspergillosis [101]. Therefore, animal models are essential to evaluate antifungal drug combinations in vivo.
There are no standardized techniques for testing antifungal combinations in animal models. Mammalian models (e.g., mice) are most often used. At least three groups of animals are needed to study the combination of two antifungals: one receiving the combination (A + B), one with the molecule A alone, and one with the molecule B alone. A control group of infected but non-treated animals should also be included in the experiments. The most frequently used evaluation criteria are the mortality rate and the fungal load in the target organs (determination of the number of CFU per gram of tissue by culture). To evaluate the effectiveness of the combination (mortality or number of CFUs in the organs), the group receiving the combination therapy is compared to the groups receiving monotherapy. It has to be noted that the inoculum size used to study the CFU in the organs or mortality rate is not the same. To determine the most suitable inoculum size and antifungal dosages, preliminary experiments have to be performed. To assess whether the combination is more effective than the monotherapies, the drugs alone should not give a maximum response, i.e., either a survival of all animals or a sterilization of the organs. This may therefore imply that the dosages of antifungals could be lower than those usually used in humans. Several studies of antifungal combinations in animal models of invasive candidiasis [76][102][103][104][105][106], cryptococcosis [107][108][109][110], and aspergillosis [107][111][112][113] have been realized.
Mammalian animal models have several drawbacks. Indeed, they need dedicated infrastructures, time-consuming experiments, and ethical considerations limit their use. To avoid these limitations, alternative models have been developed [114]. The Galleria mellonella model has been one of the most often used models in recent years [115]. The G. mellonella model is interesting because it is inexpensive, easy to use, and does not require a dedicated infrastructure. The larvae of G. mellonella are small, making them easy to handle. Additionally, the larvae can survive at 37 °C, which makes them suitable to study human fungal pathogens. This model was first used for virulence studies, but is now also used for the evaluation of antifungal combinations [116][117][118]. Larval inoculation is performed by injecting a small volume (10 µl) into a proleg on the ventral face [119]. In general, 10 to 20 larvae per group are used. Preliminary experiments to determine the lethal dose that results in 90% mortality (LD90), or the sub-lethal dose that results in 10% mortality (LD10) have to be performed according to the main endpoint (mortality or fungal load in the larvae). Most often, the main endpoint is the mortality [120][121]. G. mellonella has been used to test antifungal combinations against different species of yeasts and filamentous fungi. In Candida spp. the combination of amphotericin B and flucytosine improved the survival of infected larvae [122]. Combinations of antifungals with antibiotics have also been tested and gave similar results [123][124][125][126]. Finally, other studies have used this model to demonstrate the synergistic interaction between fluconazole and other drugs against C. albicans [127][128][129][130][131]. Many studies have used G. mellonella as a model for the evaluation of antifungal combinations against Cryptococcus spp. [132]. One study used the conventional antifungal agents used for the treatment of Cryptococcus infection (combination of amphotericin B with flucytosine) [133], another study assessed drug repurposing using the compound astemizole (antihistaminic drug) [134]. Combination therapy decreased the mortality of the larvae compared to those receiving monotherapy. This model was also used to evaluate antifungal combinations against Aspergillus spp. [131][135]. Combination of amphotericin B with an Hsp70 inhibitor increased survival of larvae compared to monotherapies [135]. Another study demonstrated that combination of itraconazole with EGTA (ethylene glycol tetra-acetic acid), a calcium chelator, is synergistic [131].
4. Conclusions
In vitro and in vivo studies of antifungal combinations are important to evaluate new therapeutic strategies in difficult-to-treat fungal infections. There are robust in vitro methods based on reference techniques, although standardization has to be improved. Advances have been made in the process of interpretation of combination results. Alternative animal models in invertebrates, which are now commonly used for testing virulence and antifungal resistance, have proven to be useful in the field of antifungal combinations. Although standardization is not fully achieved, significant results can be obtained due to the possibility of concomitantly using several techniques and several form of analysis for the interpretation of the results.
References
- Arendrup, M.C.; Boekhout, T.; Akova, M.; Meis, J.F.; Cornely, O.A.; Lortholary, O. European Society of Clinical Microbiology and Infectious Diseases Fungal Infection Study Group; European Confederation of Medical Mycology ESCMID and ECMM Joint Clinical Guidelines for the Diagnosis and Management of Rare Invasive Yeast Infections. Clin. Microbiol. Infect. 2014, 20 (Suppl. S3), 76–98.
- Cuenca-Estrella, M.; Verweij, P.E.; Arendrup, M.C.; Arikan-Akdagli, S.; Bille, J.; Donnelly, J.P.; Jensen, H.E.; Lass-Flörl, C.; Richardson, M.D.; Akova, M.; et al. ESCMID* Guideline for the Diagnosis and Management of Candida Diseases 2012: Diagnostic Procedures. Clin. Microbiol. Infect. 2012, 18 (Suppl. S7), 9–18.
- Arendrup, M.C.; Patterson, T.F. Multidrug-Resistant Candida: Epidemiology, Molecular Mechanisms, and Treatment. J. Infect. Dis. 2017, 216, S445–S451.
- Lestrade, P.P.A.; Meis, J.F.; Melchers, W.J.G.; Verweij, P.E. Triazole Resistance in Aspergillus fumigatus: Recent Insights and Challenges for Patient Management. Clin. Microbiol. Infect. 2019, 25, 799–806.
- Ball, B.; Bermas, A.; Carruthers-Lay, D.; Geddes-McAlister, J. Mass Spectrometry-Based Proteomics of Fungal Pathogenesis, Host–Fungal Interactions, and Antifungal Development. J. Fungi 2019, 5, 52.
- Kontoyiannis, D.P.; Lewis, R.E. Toward More Effective Antifungal Therapy: The Prospects of Combination Therapy. Br. J. Haematol. 2004, 126, 165–175.
- Vitale, R.G.; Afeltra, J.; Dannaoui, E. Antifungal Combinations. Methods Mol. Med. 2005, 118, 143–152.
- Perfect, J.R.; Dismukes, W.E.; Dromer, F.; Goldman, D.L.; Graybill, J.R.; Hamill, R.J.; Harrison, T.S.; Larsen, R.A.; Lortholary, O.; Nguyen, M.-H.; et al. Clinical Practice Guidelines for the Management of Cryptococcal Disease: 2010 Update by the Infectious Diseases Society of America. Clin. Infect. Dis. 2010, 50, 291–322.
- Arendrup, M.C.; Cuenca-Estrella, M.; Lass-Flörl, C.; Hope, W. EUCAST-AFST EUCAST Technical Note on the EUCAST Definitive Document EDef 7.2: Method for the Determination of Broth Dilution Minimum Inhibitory Concentrations of Antifungal Agents for Yeasts EDef 7.2 (EUCAST-AFST). Clin. Microbiol. Infect. 2012, 18, E246–E247.
- Clinical and laboratory Standards Institute (CLSI). Reference Method for Broth Dilution Antifungal Susceptibility Testing of Filamentous Fungi, 2nd ed.; Approved standard. Document M-38A2; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2008.
- Spitzer, M.; Robbins, N.; Wright, G.D. Combinatorial Strategies for Combating Invasive Fungal Infections. Virulence 2016, 8, 169–185.
- Katragkou, A.; McCarthy, M.; Meletiadis, J.; Hussain, K.; Moradi, P.W.; Strauss, G.E.; Myint, K.L.; Zaw, M.H.; Kovanda, L.L.; Petraitiene, R.; et al. In Vitro Combination Therapy with Isavuconazole against Candida spp. Med. Mycol. 2017, 55, 859–868.
- Mukherjee, P.K.; Sheehan, D.J.; Hitchcock, C.A.; Ghannoum, M.A. Combination Treatment of Invasive Fungal Infections. Clin. Microbiol. Rev. 2005, 18, 163–194.
- Berenbaum, M.C. A Method for Testing for Synergy with Any Number of Agents. J. Infect. Dis. 1978, 137, 122–130.
- Prichard, M.N.; Prichard, L.E.; Shipman, C. Strategic Design and Three-Dimensional Analysis of Antiviral Drug Combinations. Antimicrob. Agents Chemother. 1993, 37, 540–545.
- Pryjma, M.; Burian, J.; Thompson, C.J. Rifabutin Acts in Synergy and Is Bactericidal with Frontline Mycobacterium abscessus Antibiotics Clarithromycin and Tigecycline, Suggesting a Potent Treatment Combination. Antimicrob. Agents Chemother. 2018, 62.
- Stein, C.; Makarewicz, O.; Bohnert, J.A.; Pfeifer, Y.; Kesselmeier, M.; Hagel, S.; Pletz, M.W. Three Dimensional Checkerboard Synergy Analysis of Colistin, Meropenem, Tigecycline against Multidrug-Resistant Clinical Klebsiella pneumonia Isolates. PLoS ONE 2015, 10, e0126479.
- Dannaoui, E.; Lortholary, O.; Dromer, F. In Vitro Evaluation of Double and Triple Combinations of Antifungal Drugs against Aspergillus fumigatus and Aspergillus terreus. Antimicrob. Agents Chemother. 2004, 48, 970–978.
- Ghannoum, M.A.; Fu, Y.; Ibrahim, A.S.; Mortara, L.A.; Shafiq, M.C.; Edwards, J.E.; Criddle, R.S. In Vitro Determination of Optimal Antifungal Combinations against Cryptococcus neoformans and Candida albicans. Antimicrob. Agents Chemother. 1995, 39, 2459–2465.
- Schwarz, P.; Dromer, F.; Lortholary, O.; Dannaoui, E. In Vitro Interaction of Flucytosine with Conventional and New Antifungals against Cryptococcus neoformans Clinical Isolates. Antimicrob. Agents Chemother. 2003, 47, 3361–3364.
- Bidaud, A.L.; Botterel, F.; Chowdhary, A.; Dannaoui, E. In Vitro Antifungal Combination of Flucytosine with Amphotericin B, Voriconazole, or Micafungin against Candida auris Shows No Antagonism. Antimicrob. Agents Chemother. 2019, 63.
- Bidaud, A.L.; Djenontin, E.; Botterel, F.; Chowdhary, A.; Dannaoui, E. Colistin Interacts Synergistically with Echinocandins against Candida auris. Int. J. Antimicrob. Agents 2020, 55, 105901.
- Imbert, S.; Palous, M.; Meyer, I.; Dannaoui, E.; Mazier, D.; Datry, A.; Fekkar, A. In Vitro Combination of Voriconazole and Miltefosine against Clinically Relevant Molds. Antimicrob. Agents Chemother. 2014, 58, 6996–6998.
- Schwarz, P.; Schwarz, P.V.; Felske-Zech, H.; Dannaoui, E. In Vitro Interactions between Isavuconazole and Tacrolimus, Cyclosporin A or Sirolimus against Mucorales. J. Antimicrob. Chemother. 2019, 74, 1921–1927.
- Afeltra, J.; Dannaoui, E.; Meis, J.F.G.M.; Rodriguez-Tudela, J.L.; Verweij, P.E. In Vitro Synergistic Interaction between Amphotericin B and Pentamidine against Scedosporium prolificans. Antimicrob. Agents Chemother. 2002, 46, 3323–3326.
- Compain, F.; Botterel, F.; Sitterlé, E.; Paugam, A.; Bougnoux, M.-E.; Dannaoui, E. In Vitro Activity of Miltefosine in Combination with Voriconazole or Amphotericin B against Clinical Isolates of Scedosporium spp. J. Med. Microbiol. 2015, 64, 309–311.
- Dannaoui, E.; Lortholary, O.; Dromer, F. Methods for antifungal combination studies in vitro and in vivo in animal models. J. Med. Microbiol. 2003, 13, 73–85.
- Li, Y.; Wan, Z.; Liu, W.; Li, R. Synergistic Activity of Chloroquine with Fluconazole against Fluconazole-Resistant Isolates of Candida Species. Antimicrob. Agents Chemother. 2015, 59, 1365–1369.
- Yang, J.; Gao, L.; Yu, P.; Kosgey, J.C.; Jia, L.; Fang, Y.; Xiong, J.; Zhang, F. In Vitro Synergy of Azole Antifungals and Methotrexate against Candida albicans. Life Sci. 2019, 235, 116827.
- Liu, W.; Li, L.P.; Zhang, J.D.; Li, Q.; Shen, H.; Chen, S.M.; He, L.J.; Yan, L.; Xu, G.T.; An, M.M.; et al. Synergistic Antifungal Effect of Glabridin and Fluconazole. PLoS ONE 2014, 9, e103442.
- Aneke, C.I.; Rhimi, W.; Otranto, D.; Cafarchia, C. Synergistic Effects of Efflux Pump Modulators on the Azole Antifungal Susceptibility of Microsporum canis. Mycopathologia 2020, 185, 279–288.
- Pillai, S.K.; Moellerig, R.C., Jr.; Eliopoulos, G.M. Antimicrobial combinations. In Antibiotics in Laboratory Medecine, 5th ed.; Lorian, V., Ed.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2005; pp. 537–595.
- Siau, H.; Kerridge, D. The Effect of Antifungal Drugs in Combination on the Growth of Candida glabrata in Solid and Liquid Media. J. Antimicrob. Chemother. 1998, 41, 357–366.
- Dannaoui, E.; Espinel-Ingroff, A. Antifungal Susceptibly Testing by Concentration Gradient Strip Etest Method for Fungal Isolates: A Review. J. Fungi 2019, 5, 108.
- Doern, C.D. When Does 2 Plus 2 Equal 5? A Review of Antimicrobial Synergy Testing. J. Clin. Microbiol. 2014, 52, 4124–4128.
- White, R.L.; Burgess, D.S.; Manduru, M.; Bosso, J.A. Comparison of Three Different in Vitro Methods of Detecting Synergy: Time-Kill, Checkerboard, and E Test. Antimicrob. Agents Chemother. 1996, 40, 1914–1918.
- Siopi, M.; Siafakas, N.; Zerva, L.; Meletiadis, J. Evaluation of Paper Gradient Concentration Strips for Antifungal Combination Testing of Candida spp. Mycoses 2015, 58, 679–687.
- Kiraz, N.; Dag, I.; Yamac, M.; Kiremitci, A.; Kasifoglu, N.; Akgun, Y. Antifungal Activity of Caspofungin in Combination with Amphotericin B against Candida glabrata: Comparison of Disk Diffusion, Etest, and Time-Kill Methods. Antimicrob. Agents Chemother. 2009, 53, 788–790.
- Kiraz, N.; Dag, I.; Yamac, M.; Kiremitci, A.; Kasifoglu, N.; Oz, Y. Synergistic Activities of Three Triazoles with Caspofungin against Candida glabrata Isolates Determined by Time-Kill, Etest, and Disk Diffusion Methods. Antimicrob. Agents Chemother. 2010, 54, 2244–2247.
- Cantón, E.; Pemán, J.; Romero, M.; Gobernado, M. Usefulness of the E-test and its assay conditions in the study of the interaction of antifungal agents. A pilot study. Rev. Espanola Quimioter. 2004, 17, 48–56.
- Lewis, R.E.; Diekema, D.J.; Messer, S.A.; Pfaller, M.A.; Klepser, M.E. Comparison of Etest, Chequerboard Dilution and Time-Kill Studies for the Detection of Synergy or Antagonism between Antifungal Agents Tested against Candida Species. J. Antimicrob. Chemother. 2002, 49, 345–351.
- Schwarz, P.; Janbon, G.; Dromer, F.; Lortholary, O.; Dannaoui, E. Combination of Amphotericin B with Flucytosine Is Active in Vitro against Flucytosine-Resistant Isolates of Cryptococcus neoformans. Antimicrob. Agents Chemother. 2007, 51, 383–385.
- Planche, V.; Ducroz, S.; Alanio, A.; Bougnoux, M.-E.; Lortholary, O.; Dannaoui, E. In Vitro Combination of Anidulafungin and Voriconazole against Intrinsically Azole-Susceptible and -Resistant Aspergillus spp. Antimicrob. Agents Chemother. 2012, 56, 4500–4503.
- Raffetin, A.; Courbin, V.; Jullien, V.; Dannaoui, E. In Vitro Combination of Isavuconazole with Echinocandins against Azole-Susceptible and -Resistant Aspergillus spp. Antimicrob. Agents Chemother. 2018, 62.
- Denardi, L.B.; Keller, J.T.; de Azevedo, M.I.; Oliveira, V.; Piasentin, F.B.; Severo, C.B.; Santurio, J.M.; Alves, S.H. Comparison between Etest and Broth Microdilution Methods for Testing Itraconazole-Resistant Aspergillus fumigatus Susceptibility to Antifungal Combinations. Mycopathologia 2018, 183, 359–370.
- Pankey, G.; Ashcraft, D.; Kahn, H.; Ismail, A. Time-Kill Assay and Etest Evaluation for Synergy with Polymyxin B and Fluconazole against Candida glabrata. Antimicrob. Agents Chemother. 2014, 58, 5795–5800.
- Adams, E.K.; Ashcraft, D.S.; Pankey, G.A. In Vitro Synergistic Activity of Caspofungin Plus Polymyxin B Against Fluconazole-Resistant Candida glabrata. Am. J. Med. Sci. 2016, 351, 265–270.
- Hooper, R.W.; Ashcraft, D.S.; Pankey, G.A. In Vitro Synergy with Fluconazole plus Doxycycline or Tigecycline against Clinical Candida glabrata Isolates. Med. Mycol. 2019, 57, 122–126.
- Cantón, E.; Pemán, J.; Gobernado, M.; Viudes, A.; Espinel-Ingroff, A. Synergistic Activities of Fluconazole and Voriconazole with Terbinafine against Four Candida Species Determined by Checkerboard, Time-Kill, and Etest Methods. Antimicrob. Agents Chemother. 2005, 49, 1593–1596.
- Dannaoui, E.; Schwarz, P.; Lortholary, O. In Vitro Interactions between Antifungals and Immunosuppressive Drugs against Zygomycetes. Antimicrob. Agents Chemother. 2009, 53, 3549–3551.
- Ben-Ami, R.; Lewis, R.E.; Kontoyiannis, D.P. In Vitro Interactions among Echinocandins against Aspergillus fumigatus: Lack of Concordance among Methods. Med. Mycol. 2011, 49, 285–288.
- Cowen, L.E.; Lindquist, S. Hsp90 Potentiates the Rapid Evolution of New Traits: Drug Resistance in Diverse Fungi. Science 2005, 309, 2185–2189.
- Cowen, L.E.; Singh, S.D.; Köhler, J.R.; Collins, C.; Zaas, A.K.; Schell, W.A.; Aziz, H.; Mylonakis, E.; Perfect, J.R.; Whitesell, L.; et al. Harnessing Hsp90 Function as a Powerful, Broadly Effective Therapeutic Strategy for Fungal Infectious Disease. Proc. Natl. Acad. Sci. USA 2009, 106, 2818–2823.
- Kontoyiannis, D.P.; Lewis, R.E.; Sagar, N.; May, G.; Prince, R.A.; Rolston, K.V. Itraconazole-Amphotericin B Antagonism in Aspergillus fumigatus: An E-Test-Based Strategy. Antimicrob. Agents Chemother. 2000, 44, 2915–2918.
- Lafleur, M.D.; Sun, L.; Lister, I.; Keating, J.; Nantel, A.; Long, L.; Ghannoum, M.; North, J.; Lee, R.E.; Coleman, K.; et al. Potentiation of Azole Antifungals by 2-Adamantanamine. Antimicrob. Agents Chemother. 2013, 57, 3585–3592.
- Schwarz, P.; Bidaud, A.L.; Dannaoui, E. In Vitro Synergy of Isavuconazole in Combination with Colistin against Candida auris. Sci. Rep. 2020, 10, 21448.
- Schwarz, P.; Djenontin, E.; Dannaoui, E. Colistin and Isavuconazole Interact Synergistically in Vitro against Aspergillus nidulans and Aspergillus niger. Microorganisms 2020, 8, 1447.
- Ernst, E.J.; Yodoi, K.; Roling, E.E.; Klepser, M.E. Rates and Extents of Antifungal Activities of Amphotericin B, Flucytosine, Fluconazole, and Voriconazole against Candida lusitaniae Determined by Microdilution, Etest, and Time-Kill Methods. Antimicrob. Agents Chemother. 2002, 46, 578–581.
- Keele, D.J.; DeLallo, V.C.; Lewis, R.E.; Ernst, E.J.; Klepser, M.E. Evaluation of Amphotericin B and Flucytosine in Combination against Candida albicans and Cryptococcus neoformans Using Time-Kill Methodology. Diagn. Microbiol. Infect. Dis. 2001, 41, 121–126.
- Roling, E.E.; Klepser, M.E.; Wasson, A.; Lewis, R.E.; Ernst, E.J.; Pfaller, M.A. Antifungal Activities of Fluconazole, Caspofungin (MK0991), and Anidulafungin (LY 303366) Alone and in Combination against Candida spp. and Crytococcus neoformans via Time-Kill Methods. Diagn. Microbiol. Infect. Dis. 2002, 43, 13–17.
- Mashaly, G.; Shrief, R. Candida glabrata Complex from Patients with Healthcare-Associated Infections in Mansoura University Hospitals, Egypt: Distribution, Antifungal Susceptibility and Effect of Fluconazole and Polymyxin B Combination. Germs 2019, 9, 125–132.
- Oz, Y.; Dag, I.; Kiraz, N. Broth Microdilution and Time-Kill Testing of Caspofungin, Voriconazole, Amphotericin B and Their Combinations against Clinical Isolates of Candida krusei. Mycopathologia 2012, 173, 27–34.
- Li, Y.; Sun, S.; Guo, Q.; Ma, L.; Shi, C.; Su, L.; Li, H. In Vitro Interaction between Azoles and Cyclosporin A against Clinical Isolates of Candida albicans Determined by the Chequerboard Method and Time-Kill Curves. J. Antimicrob. Chemother. 2008, 61, 577–585.
- Khodavandi, A.; Alizadeh, F.; Vanda, N.A.; Karimi, G.; Chong, P.P. Possible Mechanisms of the Antifungal Activity of Fluconazole in Combination with Terbinafine against Candida albicans. Pharm. Biol. 2014, 52, 1505–1509.
- Li, H.; Chen, Z.; Zhang, C.; Gao, Y.; Zhang, X.; Sun, S. Resistance Reversal Induced by a Combination of Fluconazole and Tacrolimus (FK506) in Candida glabrata. J. Med. Microbiol. 2015, 64, 44–52.
- Kumar, S.N.; Nambisan, B.; Mohandas, C.; Sundaresan, A. In Vitro Synergistic Activity of Diketopiperazines Alone and in Combination with Amphotericin B or Clotrimazole against Candida albicans. Folia Microbiol. (Praha) 2013, 58, 475–482.
- Girmenia, C.; Venditti, M.; Martino, P. Fluconazole in Combination with Flucytosine in the Treatment of Fluconazole-Resistant Candida Infections. Diagn. Microbiol. Infect. Dis. 2003, 46, 227–231.
- Nakajima, R.; Kitamura, A.; Someya, K.; Tanaka, M.; Sato, K. In Vitro and in Vivo Antifungal Activities of DU-6859a, a Fluoroquinolone, in Combination with Amphotericin B and Fluconazole against Pathogenic Fungi. Antimicrob. Agents Chemother. 1995, 39, 1517–1521.
- Rukayadi, Y.; Lee, K.; Lee, M.-S.; Yong, D.; Hwang, J.-K. Synergistic Anticandidal Activity of Xanthorrhizol in Combination with Ketoconazole or Amphotericin B. FEMS Yeast Res. 2009, 9, 1302–1311.
- Yousfi, H.; Ranque, S.; Rolain, J.-M.; Bittar, F. In Vitro Polymyxin Activity against Clinical Multidrug-Resistant Fungi. Antimicrob. Resist. Infect. Control 2019, 8, 66.
- Shin, S.; Pyun, M.-S. Anti-Candida Effects of Estragole in Combination with Ketoconazole or Amphotericin B. Phytother. Res. 2004, 18, 827–830.
- Alnajjar, L.M.; Bulatova, N.R.; Darwish, R.M. Evaluation of Four Calcium Channel Blockers as Fluconazole Resistance Inhibitors in Candida glabrata. J. Glob. Antimicrob. Resist. 2018, 14, 185–189.
- Sun, L.; Liao, K.; Hang, C. Caffeic Acid Phenethyl Ester Synergistically Enhances the Antifungal Activity of Fluconazole against Resistant Candida albicans. Phytomedicine Int. J. Phytother. Phytopharm. 2018, 40, 55–58.
- Marchetti, O.; Moreillon, P.; Glauser, M.P.; Bille, J.; Sanglard, D. Potent Synergism of the Combination of Fluconazole and Cyclosporine in Candida albicans. Antimicrob. Agents Chemother. 2000, 44, 2373–2381.
- Dennis, E.K.; Garneau-Tsodikova, S. Synergistic Combinations of Azoles and Antihistamines against Candida Species in Vitro. Med. Mycol. 2019, 57, 874–884.
- Louie, A.; Kaw, P.; Banerjee, P.; Liu, W.; Chen, G.; Miller, M.H. Impact of the Order of Initiation of Fluconazole and Amphotericin B in Sequential or Combination Therapy on Killing of Candida albicans in Vitro and in a Rabbit Model of Endocarditis and Pyelonephritis. Antimicrob. Agents Chemother. 2001, 45, 485–494.
- Yang, D.L.; Hu, Y.L.; Yin, Z.X.; Zeng, G.S.; Li, D.; Zhang, Y.Q.; Xu, Z.H.; Guan, X.M.; Weng, L.X.; Wang, L.H. Cis-2-Dodecenoic Acid Mediates Its Synergistic Effect with Triazoles by Interfering with Efflux Pumps in Fluconazole-Resistant Candida albicans. Biomed. Environ. Sci. 2019, 32, 199–209.
- Li, L.P.; Liu, W.; Liu, H.; Zhu, F.; Zhang, D.Z.; Shen, H.; Xu, Z.; Qi, Y.P.; Zhang, S.Q.; Chen, S.M.; et al. Synergistic Antifungal Activity of Berberine Derivative B-7b and Fluconazole. PLoS ONE 2015, 10, e0126393.
- Yousfi, H.; Cassagne, C.; Ranque, S.; Rolain, J.-M.; Bittar, F. Repurposing of Ribavirin as an Adjunct Therapy against Invasive Candida Strains in an in Vitro Study. Antimicrob. Agents Chemother. 2019, 63.
- Shrestha, S.K.; Fosso, M.Y.; Garneau-Tsodikova, S. A Combination Approach to Treating Fungal Infections. Sci. Rep. 2015, 5, 17070.
- Li, H.; Zhang, C.; Chen, Z.; Shi, W.; Sun, S. A Promising Approach of Overcoming the Intrinsic Resistance of Candida krusei to Fluconazole (FLC)—Combining Tacrolimus with FLC. FEMS Yeast Res. 2014, 14, 808–811.
- Makarasen, A.; Reukngam, N.; Khlaychan, P.; Chuysinuan, P.; Isobe, M.; Techasakul, S. Mode of Action and Synergistic Effect of Valinomycin and Cereulide with Amphotericin B against Candida albicans and Cryptococcus albidus. J. Mycol. Med. 2018, 28, 112–121.
- Kumar, M.; Chaturvedi, A.K.; Kavishwar, A.; Shukla, P.K.; Kesarwani, A.P.; Kundu, B. Identification of a Novel Antifungal Nonapeptide Generated by Combinatorial Approach. Int. J. Antimicrob. Agents 2005, 25, 313–320.
- Nash, J.D.; Burgess, D.S.; Talbert, R.L. Effect of Fluvastatin and Pravastatin, HMG-CoA Reductase Inhibitors, on Fluconazole Activity against Candida albicans. J. Med. Microbiol. 2002, 51, 105–109.
- Serena, C.; Mariné, M.; Quindós, G.; Carrillo, A.J.; Cano, J.F.; Pastor, F.J.; Guarro, J. In Vitro Interactions of Micafungin with Amphotericin B against Clinical Isolates of Candida spp. Antimicrob. Agents Chemother. 2008, 52, 1529–1532.
- Huang, S.; Cao, Y.Y.; Dai, B.D.; Sun, X.R.; Zhu, Z.Y.; Cao, Y.B.; Wang, Y.; Gao, P.H.; Jiang, Y.Y. In Vitro Synergism of Fluconazole and Baicalein against Clinical Isolates of Candida albicans Resistant to Fluconazole. Biol. Pharm. Bull. 2008, 31, 2234–2236.
- Cafini, F.; Sevillano, D.; Alou, L.; Gómez-Aguado, F.; Corcuera, M.T.; González, N.; Guinea, J.; Prieto, J. Effect of Protein Binding on the Activity of Voriconazole Alone or Combined with Anidulafungin against Aspergillus spp. Using a Time-Kill Methodology. Rev. Espanola Quimioter. 2012, 25, 47–55.
- Ruhil, S.; Kumar, V.; Balhara, M.; Malik, M.; Dhankhar, S.; Kumar, M.; Kumar Chhillar, A. In Vitro Evaluation of Combination of Polyenes with EDTA against Aspergillus spp. by Different Methods (FICI and CI Model). J. Appl. Microbiol. 2014, 117, 643–653.
- Ganesan, L.T.; Manavathu, E.K.; Cutright, J.L.; Alangaden, G.J.; Chandrasekar, P.H. In-Vitro Activity of Nikkomycin Z Alone and in Combination with Polyenes, Triazoles or Echinocandins against Aspergillus fumigatus. Clin. Microbiol. Infect. 2004, 10, 961–966.
- Natesan, S.K.; Chandrasekar, P.H.; Alangaden, G.J.; Manavathu, E.K. Fluvastatin Potentiates the Activity of Caspofungin against Aspergillus fumigatus in Vitro. Diagn. Microbiol. Infect. Dis. 2008, 60, 369–373.
- Hughes, C.E.; Harris, C.; Peterson, L.R.; Gerding, D.N. Enhancement of the in Vitro Activity of Amphotericin B against Aspergillus spp. by Tetracycline Analogs. Antimicrob. Agents Chemother. 1984, 26, 837–840.
- Foucquier, J.; Guedj, M. Analysis of Drug Combinations: Current Methodological Landscape. Pharmacol. Res. Perspect. 2015, 3, e00149.
- Huang, R.-Y.; Pei, L.; Liu, Q.; Chen, S.; Dou, H.; Shu, G.; Yuan, Z.-X.; Lin, J.; Peng, G.; Zhang, W.; et al. Isobologram Analysis: A Comprehensive Review of Methodology and Current Research. Front. Pharmacol. 2019, 10, 1222.
- Greco, W.R.; Bravo, G.; Parsons, J.C. The Search for Synergy: A Critical Review from a Response Surface Perspective. Pharmacol. Rev. 1995, 47, 331–385.
- Chou, T.-C. Drug Combination Studies and Their Synergy Quantification Using the Chou-Talalay Method. Cancer Res. 2010, 70, 440–446.
- Di Veroli, G.Y.; Fornari, C.; Wang, D.; Mollard, S.; Bramhall, J.L.; Richards, F.M.; Jodrell, D.I. Combenefit: An Interactive Platform for the Analysis and Visualization of Drug Combinations. Bioinformatics 2016, 32, 2866–2868.
- Prichard, M.N.; Shipman, C. A Three-Dimensional Model to Analyze Drug-Drug Interactions. Antiviral Res. 1990, 14, 181–205.
- Odds, F.C. Synergy, Antagonism, and What the Chequerboard Puts between Them. J. Antimicrob. Chemother. 2003, 52, 1.
- Brouwer, A.E.; Rajanuwong, A.; Chierakul, W.; Griffin, G.E.; Larsen, R.A.; White, N.J.; Harrison, T.S. Combination Antifungal Therapies for HIV-Associated Cryptococcal Meningitis: A Randomised Trial. Lancet 2004, 363, 1764–1767.
- Bennett, J.E.; Dismukes, W.E.; Duma, R.J.; Medoff, G.; Sande, M.A.; Gallis, H.; Leonard, J.; Fields, B.T.; Bradshaw, M.; Haywood, H.; et al. A Comparison of Amphotericin B Alone and Combined with Flucytosine in the Treatment of Cryptoccal Meningitis. N. Engl. J. Med. 1979, 301, 126–131.
- Marr, K.A.; Schlamm, H.T.; Herbrecht, R.; Rottinghaus, S.T.; Bow, E.J.; Cornely, O.A.; Heinz, W.J.; Jagannatha, S.; Koh, L.P.; Kontoyiannis, D.P.; et al. Combination Antifungal Therapy for Invasive Aspergillosis: A Randomized Trial. Ann. Intern. Med. 2015, 162, 81–89.
- Sugar, A.M.; Liu, X.P. Interactions of Itraconazole with Amphotericin B in the Treatment of Murine Invasive Candidiasis. J. Infect. Dis. 1998, 177, 1660–1663.
- Louie, A.; Banerjee, P.; Drusano, G.L.; Shayegani, M.; Miller, M.H. Interaction between Fluconazole and Amphotericin B in Mice with Systemic Infection Due to Fluconazole-Susceptible or -Resistant Strains of Candida albicans. Antimicrob. Agents Chemother. 1999, 43, 2841–2847.
- Flattery, A.M.; Hickey, E.; Gill, C.J.; Powles, M.A.; Misura, A.S.; Galgoci, A.M.; Ellis, J.D.; Zhang, R.; Sandhu, P.; Ronan, J.; et al. Efficacy of Caspofungin in a Juvenile Mouse Model of Central Nervous System Candidiasis. Antimicrob. Agents Chemother. 2011, 55, 3491–3497.
- Sanati, H.; Ramos, C.F.; Bayer, A.S.; Ghannoum, M.A. Combination Therapy with Amphotericin B and Fluconazole against Invasive Candidiasis in Neutropenic-Mouse and Infective-Endocarditis Rabbit Models. Antimicrob. Agents Chemother. 1997, 41, 1345–1348.
- Johnson, M.D.; Perfect, J.R. Use of Antifungal Combination Therapy: Agents, Order, and Timing. Curr. Fungal Infect. Rep. 2010, 4, 87–95.
- Barchiesi, F.; Schimizzi, A.M.; Caselli, F.; Novelli, A.; Fallani, S.; Giannini, D.; Arzeni, D.; Di Cesare, S.; Di Francesco, L.F.; Fortuna, M.; et al. Interactions between Triazoles and Amphotericin B against Cryptococcus neoformans. Antimicrob. Agents Chemother. 2000, 44, 2435–2441.
- Diamond, D.M.; Bauer, M.; Daniel, B.E.; Leal, M.A.; Johnson, D.; Williams, B.K.; Thomas, A.M.; Ding, J.C.; Najvar, L.; Graybill, J.R.; et al. Amphotericin B Colloidal Dispersion Combined with Flucytosine with or without Fluconazole for Treatment of Murine Cryptococcal Meningitis. Antimicrob. Agents Chemother. 1998, 42, 528–533.
- Schwarz, P.; Dromer, F.; Lortholary, O.; Dannaoui, E. Efficacy of Amphotericin B in Combination with Flucytosine against Flucytosine-Susceptible or Flucytosine-Resistant Isolates of Cryptococcus neoformans during Disseminated Murine Cryptococcosis. Antimicrob. Agents Chemother. 2006, 50, 113–120.
- George, D.; Kordick, D.; Miniter, P.; Patterson, T.F.; Andriole, V.T. Combination Therapy in Experimental Invasive Aspergillosis. J. Infect. Dis. 1993, 168, 692–698.
- Petraitis, V.; Petraitiene, R.; McCarthy, M.W.; Kovanda, L.L.; Zaw, M.H.; Hussain, K.; Shaikh, N.; Maung, B.B.W.; Sekhon, N.K.; Hope, W.W.; et al. Combination Therapy with Isavuconazole and Micafungin for Treatment of Experimental Invasive Pulmonary Aspergillosis. Antimicrob. Agents Chemother. 2017, 61.
- Seyedmousavi, S.; Brüggemann, R.J.M.; Melchers, W.J.G.; Rijs, A.J.M.M.; Verweij, P.E.; Mouton, J.W. Efficacy and Pharmacodynamics of Voriconazole Combined with Anidulafungin in Azole-Resistant Invasive Aspergillosis. J. Antimicrob. Chemother. 2013, 68, 385–393.
- Chamilos, G.; Lionakis, M.S.; Lewis, R.E.; Kontoyiannis, D.P. Role of Mini-Host Models in the Study of Medically Important Fungi. Lancet Infect. Dis. 2007, 7, 42–55.
- Jemel, S.; Guillot, J.; Kallel, K.; Botterel, F.; Dannaoui, E. Galleria mellonella for the Evaluation of Antifungal Efficacy against Medically Important Fungi, a Narrative Review. Microorganisms 2020, 8, 390.
- Arvanitis, M.; Glavis-Bloom, J.; Mylonakis, E. Invertebrate Models of Fungal Infection. Biochim. Biophys. Acta 2013, 1832, 1378–1383.
- Kavanagh, K.; Sheehan, G. The Use of Galleria mellonella Larvae to Identify Novel Antimicrobial Agents against Fungal Species of Medical Interest. J. Fungi 2018, 4, 113.
- Lionakis, M.S. Drosophila and Galleria Insect Model Hosts: New Tools for the Study of Fungal Virulence, Pharmacology and Immunology. Virulence 2011, 2, 521–527.
- Champion, O.L.; Titball, R.W.; Bates, S. Standardization of G. mellonella Larvae to Provide Reliable and Reproducible Results in the Study of Fungal Pathogens. J. Fungi 2018, 4, 108.
- Andrea, A.; Krogfelt, K.A.; Jenssen, H. Methods and Challenges of Using the Greater Wax Moth (Galleria mellonella) as a Model Organism in Antimicrobial Compound Discovery. Microorganisms 2019, 7, 85.
- Tsai, C.J.-Y.; Loh, J.M.S.; Proft, T. Galleria mellonella Infection Models for the Study of Bacterial Diseases and for Antimicrobial Drug Testing. Virulence 2016, 7, 214–229.
- Li, D.-D.; Deng, L.; Hu, G.-H.; Zhao, L.-X.; Hu, D.-D.; Jiang, Y.-Y.; Wang, Y. Using Galleria mellonella-Candida albicans Infection Model to Evaluate Antifungal Agents. Biol. Pharm. Bull. 2013, 36, 1482–1487.
- MacCallum, D.M.; Desbois, A.P.; Coote, P.J. Enhanced Efficacy of Synergistic Combinations of Antimicrobial Peptides with Caspofungin versus Candida albicans in Insect and Murine Models of Systemic Infection. Eur. J. Clin. Microbiol. Infect. Dis. 2013, 32, 1055–1062.
- Gu, W.; Yu, Q.; Yu, C.; Sun, S. In Vivo Activity of Fluconazole/Tetracycline Combinations in Galleria mellonella with Resistant Candida albicans Infection. J. Glob. Antimicrob. Resist. 2018, 13, 74–80.
- Lu, M.; Yang, X.; Yu, C.; Gong, Y.; Yuan, L.; Hao, L.; Sun, S. Linezolid in Combination with Azoles Induced Synergistic Effects against Candida albicans and Protected Galleria mellonella against Experimental Candidiasis. Front. Microbiol. 2018, 9, 3142.
- Lu, M.; Yu, C.; Cui, X.; Shi, J.; Yuan, L.; Sun, S. Gentamicin Synergises with Azoles against Drug-Resistant Candida albicans. Int. J. Antimicrob. Agents 2018, 51, 107–114.
- Li, X.; Wu, X.; Gao, Y.; Hao, L. Synergistic Effects and Mechanisms of Combined Treatment with Harmine Hydrochloride and Azoles for Resistant Candida albicans. Front. Microbiol. 2019, 10, 2295.
- Li, X.; Zhao, Y.; Huang, X.; Yu, C.; Yang, Y.; Sun, S. Ambroxol Hydrochloride Combined with Fluconazole Reverses the Resistance of Candida albicans to Fluconazole. Front. Cell. Infect. Microbiol. 2017, 7, 124.
- Sun, W.; Wang, D.; Yu, C.; Huang, X.; Li, X.; Sun, S. Strong Synergism of Dexamethasone in Combination with Fluconazole against Resistant Candida albicans Mediated by Inhibiting Drug Efflux and Reducing Virulence. Int. J. Antimicrob. Agents 2017, 50, 399–405.
- Zhang, M.; Yan, H.; Lu, M.; Wang, D.; Sun, S. Antifungal Activity of Ribavirin Used Alone or in Combination with Fluconazole against Candida albicans Is Mediated by Reduced Virulence. Int. J. Antimicrob. Agents 2020, 55, 105804.
- Liu, F.; Pu, L.; Zheng, Q.; Zhang, Y.; Gao, R.; Xu, X.; Zhang, S.; Lu, L. Calcium Signaling Mediates Antifungal Activity of Triazole Drugs in the Aspergilli. Fungal Genet. Biol. 2015, 81, 182–190.
- Lu, M.; Yan, H.; Yu, C.; Yuan, L.; Sun, S. Proton Pump Inhibitors Act Synergistically with Fluconazole against Resistant Candida albicans. Sci. Rep. 2020, 10, 498.
- Sangalli-Leite, F.; Scorzoni, L.; Alves de Paula e Silva, A.C.; da Silva, J.d.F.; de Oliveira, H.C.; de Lacorte Singulani, J.; Gullo, F.P.; Moraes da Silva, R.; Regasini, L.O.; Siqueira da Silva, D.H.; et al. Synergistic Effect of Pedalitin and Amphotericin B against Cryptococcus neoformans by in Vitro and in Vivo Evaluation. Int. J. Antimicrob. Agents 2016, 48, 504–511.
- Mylonakis, E.; Moreno, R.; El Khoury, J.B.; Idnurm, A.; Heitman, J.; Calderwood, S.B.; Ausubel, F.M.; Diener, A. Galleria mellonella as a Model System to Study Cryptococcus neoformans Pathogenesis. Infect. Immun. 2005, 73, 3842–3850.
- Vu, K.; Gelli, A. Astemizole and an Analogue Promote Fungicidal Activity of Fluconazole against Cryptococcus neoformans var. grubii and Cryptococcus gattii. Med. Mycol. 2010, 48, 255–262.
- Blatzer, M.; Blum, G.; Jukic, E.; Posch, W.; Gruber, P.; Nagl, M.; Binder, U.; Maurer, E.; Sarg, B.; Lindner, H.; et al. Blocking Hsp70 Enhances the Efficiency of Amphotericin B Treatment against Resistant Aspergillus terreus Strains. Antimicrob. Agents Chemother. 2015, 59, 3778–3788.





