
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Smriti Verma | + 7103 word(s) | 7103 | 2020-04-08 11:46:08 | | | |
| 2 | Rita Xu | -2538 word(s) | 4565 | 2020-04-18 07:08:56 | | | | |
| 3 | Rita Xu | -20 word(s) | 4545 | 2020-10-29 09:22:09 | | |
Video Upload Options
Salmonella enterica serovars are important pathogens of humans and animals that are responsible for enormous morbidity, mortality and economic loss worldwide. Models used to study the disease pathology so far have provided valuable advancements, however, the molecular complexity of its pathogenesis remains poorly understood, particularly in humans. Therefore there remains a disconnect between what works at the bench versus at the bedside, especially in case of vaccines. The development of organoids/enteroids offers a tremendous opportunity to bridge this gap by bringing human-specific factors into the research models as well as elevate our understanding of the interactions and crosstalk between multiple cell types and the microbiota with Salmonella. Thus the use of organoids in studying Salmonella biology has the potential for improving clinical outcomes and future prophylactic and therapeutic intervention strategies.
1. Salmonella enterica
Several members of the genus Salmonella are major foodborne pathogens [1]. It comprises two species: S. bongori and S. enterica. Almost all Salmonella organisms that cause disease in humans and domestic animals are serovars belonging to S. enterica subspecies enterica. Broadly, the diseases caused by Salmonella in humans are of two kinds: 1) systemic febrile illness termed typhoid/enteric fever, and 2) acute self-limiting gastroenteritis. The serovars that cause typhoid are referred to as typhoidal Salmonella and include S. enterica subsp. enterica serovar Typhi and S. enterica subsp. enterica serovar Paratyphi A, B, and C. S. Typhi causes approximately 76.3% of global enteric fever cases [2]. S. Typhi and S. Paratyphi are restricted to humans and higher primates, and clinical manifestations of the infection include sustained high fever, abdominal pain, headache, weakness, malaise, and transient diarrhea/constipation. Without appropriate and effective antibiotic therapy, the infection may lead to gastrointestinal bleeding, intestinal perforation, septic shock, and death [3][4]. The Non-Typhoidal Salmonella (NTS) serovars cause self-limiting gastroenteritis and include S. enterica subsp. enterica serovar Typhimurium and S. enterica subsp. enterica serovar Enteridis, which are the most prevalent clinical isolates, according to the World Health Organization (WHO). These pathogens are broader in host range and infect humans and animals such as poultry, cattle, reptiles, and amphibians. Infections with NTS typically involve self-limiting diarrhea, stomach cramps, headache, vomiting, and fever that resolve on their own; however, the infection can be severe in children and the elderly and can sometimes be fatal [5]. NTS serovars have been reported to cause an invasive infection similar to typhoid, particularly in sub-Saharan Africa, predominantly in children and HIV-positive adults, with several co-morbidities such as ongoing or recent malaria infection and malnutrition contributing to higher mortality [6][7][8]. Salmonella infections represent a considerable economic burden and public health concern in both developing and developed countries. The Center for Disease Control and Prevention (CDC), estimates that 1.35 million infections by Salmonella occur in the United States. NTS causes more than 93 million global infections per year, with 155,000 deaths [9]. A study examining the global burden of diseases, injuries, and risk factors in 2017 reported 14.3 million cases of enteric fever globally, with a case fatality of 0.95% resulting in approximately 135,000 deaths [10]. Children, the elderly, and those residing in lower-income countries account for the greatest incidences [11][12].
Salmonella is acquired via contaminated food and water. Luminal bacteria invade M cells and absorptive enterocytes via a specialized apparatus called the type three secretion system (T3SS) [13] encoded on the Salmonella Pathogenicity Islands (SPIs) [14]. The T3SS injects bacterial proteins into host cells allowing the bacteria to essentially commandeer host cellular processes to induce cytoskeletal rearrangements that engulf the bacteria into specialized vesicles called the Salmonella containing vacuoles (SCVs) [15]. This invasion process, with subsequent translocation across the epithelium, is followed by the uptake of the bacteria by macrophages and dendritic cells in the intestinal submucosa, where bacterial proteins interfere with phagolysosomal maturation and allow the bacteria to survive inside the cells [16][17][18]. NTS serovars undergo prolific growth in the intestine, while T3SS effectors induce fluid secretion and promote inflammation. Immune signaling via recognition of pathogen-associated molecular patterns such as lipopolysaccharide (LPS) and flagella also induce a robust inflammatory response, which actually provides Salmonella with a growth advantage over resident microflora [5][19][20][21]. The immune response eventually limits Salmonella growth; nevertheless, the short-term proliferation is sufficient to ensure propagation. Typhoidal Salmonella strains elicit a more attenuated inflammatory response in the intestine, especially in terms of limited neutrophil recruitment [3][22][23]. Bacteria migrate to the mesenteric lymph nodes (MLN) and systemically within the reticuloendothelial cells, and as free bacteria in the blood or lymph, to establish new foci of infection in the liver, spleen, bone marrow, and gallbladder [24]. At these new sites, the bacteria replicate and re-enter the intestinal lumen via secretion in bile, promoting the shedding of the bacteria to continue the cycle of new infections by contaminated food and water. In 3%–5% of cases, the bacteria can persist for long durations in the gallbladder, which serves as a reservoir of chronic infection [25]. Chronic infection with Salmonella has been found to be a risk factor for the development of malignant neoplasms, including gallbladder cancer [26][27] and colorectal cancer [27][28].
The emergence of multi-drug resistance to conventional antibiotics complicates the treatment of Salmonella infection [29][30][31][32]. Antibiotic treatment destroys the resident microflora, provides a niche for Salmonella to proliferate, and may lead to increased levels of bacterial shedding [33]. Additionally, bacterial populations that express an antibiotic-tolerant phenotype can evade treatment and persist, causing relapses of the infection as well as the evolution of bacterial virulence [34][35]. Asymptomatic carriers act as reservoirs, contribute to the continued propagation of the pathogen, and are particularly important for food safety considerations. Currently, there are no effective vaccines against gastrointestinal Salmonella infections. Several typhoid vaccines are available and licensed in many countries; however, robust protection is limited and has been associated with injection site reactions. Furthermore, the vaccines have not been widely adopted by public health programs [36]. With the significant aging populations in both developed and developing countries [37][38], more people are at risk for severe consequences of Salmonella infections. Advances in the mechanistic understanding of Salmonella infections will facilitate the development of improved control strategies, particularly, safe and effective vaccines. The broad conservation of host responses as well as the molecular machinery used by Salmonella strains during infection of various hosts, namely T3SSs encoded by SPI1 and SPI2 that enable invasion of epithelial cells and subsequent intracellular survival, allows several model systems to be applied for Salmonella research [39]. Each non-human model has pros and cons and varies in the ability to recapitulate natural infection. Given the host-specific aspects of infection physiology, it is necessary to be cautious in applying the results from these model systems to human patients. Additionally, with Salmonella being an important tool to understand host physiology, metabolism, immune function, and interactions with microbes, human-specific investigations are valuable to the research community.
2. Model Systems to Study Salmonella Biology
Transformed cell lines such as Cos-1, MDCK, HeLa, HepG2, CaCo2, and T-84 have been used to carry out several fundamental studies on Salmonella pathogenicity, such as the identification of the T3SS and the SPIs [13][40]. However, these cells have drawbacks in that the cells do not adequately represent the physiological characteristics of normal human tissue [41][42]. Explant tissue cultures have organotypic properties that can be vital for studies on development and physiology but are limited by culturing difficulties and short life span [43][44]. Classically, animal models have offered solutions for several of these limitations, and have been used to corroborate data obtained from other model systems as well as to investigate the deeper molecular mechanism of infection. For example, S. Typhimurium is a natural pathogen of calves and causes gastroenteritis with clinical and pathological manifestations similar to humans, namely diarrhea, anorexia, fever, localized infection, and neutrophil infiltration [45][46]. Meanwhile, the bovine ligated loop was instrumental in characterizing fluid accumulation and host inflammatory responses following S. Typhimurium infection. For example, mutants lacking the invasion proteins SipA, SopA, and SopD were shown to have little to no effect on the ability of S. Typhimurium to invade epithelial cells, but were shown to reduce the fluid accumulation and neutrophil immigration in bovine loops [47][48].
More importantly, the vast majority of studies on Salmonella pathogenesis have been conducted in the murine model, including studies for the development of new vaccines. This model has been useful in clarifying various aspects of in vivo Salmonella pathogenesis; however, it does have limitations with respect to its ability to faithfully reproduce all aspects of Salmonella infection in humans. S. Typhimurium causes two very different types of diseases in human and mice. In humans, the infection is localized, dominated by the infiltration of neutrophils and self-limiting. In mice, S. Typhimurium spreads systemically, with slow infiltration of mononuclear inflammatory cells and little or no localized intestinal tissue injury. The host responses involved are also dramatically different. Since the clinical manifestations and pathology of mice infected with S. Typhimurium resemble those observed in humans with S. Typhi infection, the model has been used as a surrogate to study typhoid pathogenesis [49]. A mouse model of S. Typhimurium-induced enterocolitis has been developed and involves pre-treatment of mice with a single dose of streptomycin. This procedure diminishes the colonization resistance by the commensal microbiota, allowing Salmonella (that carry the gene for resistance to streptomycin) to grow to high densities in the cecum and large intestine and trigger acute gastroenteritis [50]. Mouse models have also been developed to mimic chronic infections of Salmonella observed in certain carrier individuals, which typically involve infecting susceptible mice with avirulent strains, or sub-lethal doses of S. Typhimurium in resistant mice. Experimental interpretations from the mouse model may not translate to human disease. S. Typhi and S. Typhimurium share about 89% of their genes, with approximately 500 genes unique to S. Typhimurium and 600 genes unique to S. Typhi, including the genes encoding typhoid toxin and the immunoprotective capsule [51][52]. S. Typhi and S. Paratyphi also have pseudogenes as well as small sequence differences in genes encoding the T3SS apparatuses and related effectors that may have important implications in pathogenesis. It is possible that virulence factors that may have no role in S. Typhi-mediated infection in humans may be important for mouse infection by S. Typhimurium and vice versa. Finally, S. Typhi genes required for causing typhoid but absent in S. Typhimurium cannot be studied easily using the latter. Typhoid fever can be induced experimentally by oral infection in higher primates or human volunteers, but these studies come with their own set of difficulties and ethical objections. Researchers have attempted to compensate for the discrepancies by developing humanized mice, i.e., immunodeficient mice engrafted with human hematopoietic cells [53][54], but these models are cumbersome, expensive, and still do not guarantee that mouse-specific factors will not add complexities or variations to the data generated. In addition, host genetic background has been found to play a role in susceptibility to invasive NTS infections, a concept that borders on the realm of personalized medicine [55]. Thus, organoids offer a promising model to mechanistically study host-specific aspects of infection.
3. Organoids/Enteroids in Salmonella Biology
Organoids consist of a three-dimensional (3D) organ-like structure that is made up of organ-specific differentiated cells of multiple lineages and that recapitulates the unique organizational and functional characteristics of the corresponding organ in vivo. The 3D structures are generated in vitro from cells with a whole range of origins, such as tissue segments [56][57] and their derived adult stem cells (ASCs) [58][59], transformed cell lines [60], and pluripotent stem cells (PSCs). These models bridge the gap between simple 2D cell culture and complex in vivo experiments. Organoids have the following characteristics: 1) self-organization: individual cells arrange in vitro into a 3D structure that mimics the in vivo organ or tissue, 2) multicellularity: organoids are composed of multiple cell types typically found in the organ or tissue in equivalent proportions, 3) functionality: the organoid structure should be able to execute at least some of the organ- or tissue-specific functions, and 4) sustainability: organoids can propagate indefinitely without requiring transformation by maintaining a pool of progenitor cells. In 2012, the International Stem Cell Consortium [61] set guidelines for the nomenclature to be used to define the 3D structures generated in vitro depending on their origin and cellular composition. When referring to intestinal organotypic models, 3D structures that are composed of just epithelial cell types are generally known as “enteroids” if derived from the small intestinal epithelium, or “colonoids” if derived from colonic origin. The term “organoid” is usually reserved for 3D structures containing more than one cell lineage. However, it should be noted that these guidelines have not been uniformly adopted by the field. Many researchers commonly use “organoids” as a blanket term for 3D structures derived from ASCs, PSCs, or comprising transformed cell lines that resemble in vivo 3D architecture and physiology.
One of the earliest studies that utilized organotypic 3D structures to investigate Salmonella pathology was carried out by Nickerson, CA et al., in 2001 [62]. The authors generated 3D organotypic cultures by growing human embryonic intestinal cell line Int-407 in rotating wall vessel (RWV) bioreactors and subsequently infected the cells with S. Typhimurium. The resulting infection was quite different from what had previously been observed in monolayer cultures. There was minimal loss of structural integrity, lower ability of the bacteria to adhere to and invade epithelia, and lowered expression of cytokine in 3D Int-407 aggregates as compared to infected Int-407 monolayers. Since the authors observed that the 3D Int-407 aggregates more closely resembled in vivo characteristics (tissue organization, tight junctions, apical-to-basal polarity, microvilli development, expression of extracellular and basement membrane proteins, and greater M cell glycosylation pattern), the authors concluded that the infection phenotypes observed in the 3D aggregates were likely representative of an in vivo infection. This study laid the groundwork for the use of 3D organotypic cultures to study Salmonella biology. The following section will highlight research performed in both mouse and human organotypic models that have improved our understanding of Salmonella pathogenesis (Figure 1).
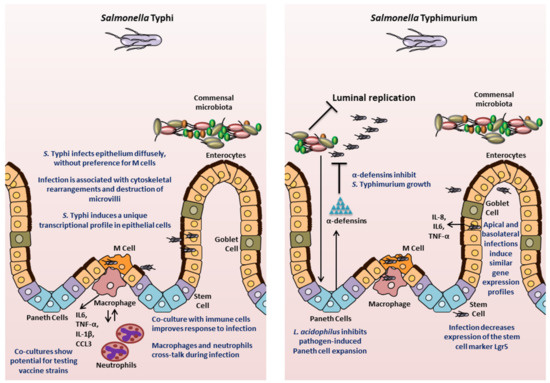
Figure 1. Insights into Salmonella pathogenesis from intestinal organoids/enteroids. The key findings for S. Typhi and S. Typhimurium are highlighted.
3.1. Mouse-Derived Models
Following the establishment of protocols to generate crypt-derived mouse intestinal enteroids (referred to as organoids by the authors) by Sato et al., Zhang and colleagues [63] in 2014 utilized the system to analyze the interaction of S. Typhimurium with epithelial cells. The authors visualized bacterial infection, while also observing bacterial-induced disruption of tight junctions, activation of the nuclear factor kappa-light-chain-enhancer of activated B cells (NF-kB)-mediated inflammatory response, and a decrease in the stem cell marker Lgr5. The authors noted that these observations were similar to findings in animal models. The caveat to this study is that the Salmonella were not delivered into the lumen of the enteroids, the location of the initial contact of bacteria with epithelial cells in vivo; instead, the bacteria were added to the medium and came in contact with the enteroids basolaterally. Nevertheless, this study established mouse-derived enteroids as a model system for studying Salmonella infection biology.
Since this initial study, enteroids have been used to interrogate various aspects of Salmonella pathology, including investigating cell types that were previously not accessible to study in vitro. Farin and colleagues [64], in a 2014 study, used mouse intestinal enteroids to study the control of Paneth cell (PC) degranulation in response to bacteria or bacterial molecules such as LPS. The authors found that PC degranulation did not occur upon stimulation with microbial molecules or Salmonella, but was induced by a novel mechanism requiring only the presence of recombinant interferon gamma (IFN-γ) [64]. In another study, Wilson and colleagues [65] interrogated the antimicrobial role of Paneth cell α-defensin peptides. The authors developed small intestinal epithelial enteroids from both wild-type mice or mice mutated for α-defensin production (Mmp7-/- mice, MMP7 is a matrix metalloprotease that is required to generate bactericidally active α-defensins in mice [66]), and infected the enteroids with S. Typhimurium by microinjecting the bacteria directly into the lumen. The absence of mature α-defensins reduced the intra-luminal bacterial killing, which could be partially restored by the expression of the human defensin HD5 [65]. This study demonstrated the contribution of α-defensins to the innate immune response to Salmonella, which previously had been a challenge to examine since most of the earlier experimental systems inadequately recapitulated in vivo cellular processes [67].
Salmonella has been suggested to contribute to the development of cancer by epidemiological studies [27]. Scanu and colleague [68] probed this phenomenon in a 2015 study using the case of gallbladder cancer (GBC). The authors derived gallbladder enteroids from mice carrying mutations that inactivate p53 and are known to be found in GBC patients in India, where the disease is prevalent. When exposed to wild-type S. Typhimurium, single cells derived from the gallbladder enteroids carrying these predisposing mutations generated new enteroids that exhibited growth factor independence, which is one of the hallmarks of transformation, and had histopathological features consistent with neoplastic transformation, thus establishing a direct association between Salmonella and cancer. To delve into the mechanism of this transformation, the authors looked at the Salmonella T3SS-mediated activation of AKT or mitogen-activated protein (MAP) kinase pathways, which have been shown to be elevated in human cancers. The signals activated by AKT and MAPK were found to be key in driving the cellular transformation and were sustained even after the eradication of the Salmonella infection. Studies have shown that AKT and MAPK pathways are activated by other bacteria and viruses that have been associated with various cancers [69][70][71][72]. Although the authors employed a murine gallbladder enteroid model and S. Typhimurium to study GBC, the authors proposed that the AKT and MAPK pathways are activated by both S. Typhimurium and S. Typhi serovars, and contribute to development of cancer that is associated with chronic S. Typhi infection in humans. Chronic infection by Salmonella has also been found to be a risk factor for developing cancers in the ascending and transverse colon [28]. However, detailed mechanistic studies of Salmonella-associated colon carcinogenesis need to be carried out and organoid model systems may prove to be extremely useful for this purpose.
Finally, another important aspect of Salmonella is the interaction of the pathogen with the host microbiome. Lu and colleagues [73] recently demonstrated that Lactobacillus acidophilus, a well-established probiotic bacterium, can alleviate damage caused by S. Typhimurium. Earlier studies had shown that L. acidophilus can inhibit adhesion of Salmonella to CaCo2 cells [74]. In this study, the authors extended the mechanism of protection to include the effects on the host. L. acidophilus altered the differentiation of epithelial cells in crypt-derived enteroids by impeding the Salmonella-mediated expansion of Paneth cells [64], thus maintaining homeostasis and appropriate epithelial composition during the infection. This study not only improved our understanding of the role of L. acidophilus in protecting the epithelial lining, but also demonstrated the ability to include microbiota-specific analyses to study Salmonella infection with enteroids/organoids.
3.2. Human-Derived Models
The potential to gain significant insight into Salmonella pathogenesis is particularly relevant in relation to the use of human-specific organoids, especially since these models possess human genetic specificities absent in mice. Studies have interrogated the usefulness of intestinal enteroids/organoids derived from human ASCs or PSCs to understand complex interactions between the epithelium and Salmonella. In 2015 Forbester and colleagues [75] used RNA sequencing to examine the epithelial transcriptional signature following injection of S. Typhimurium into the lumen of organoids derived from human induced PSCs (hiPSCs). The analysis showed significant up-regulation of genes for cytokine-mediated signaling, NF-κB activation, angiogenesis, and chemotaxis. Enhanced release of pro-inflammatory cytokines IL-8, IL-6, and TNF-α was also confirmed. The findings were consistent with prior studies in animal and mouse organoid models, thus establishing the human organoids as a viable infection model for Salmonella. The study also demonstrated that a noninvasive mutant strain (deficient in invA gene) could be used in the model to examine Salmonella pathogenicity and the functionality of defined mutants [75]. Furthermore, the authors generated an RNA sequencing data set following basolateral administration of Salmonella to the organoids. Interestingly, 49 of the 100 most highly upregulated genes were also significantly induced in the data set obtained by microinjecting the bacteria for apical infection. The data provide credence to the results of the Zhang and colleagues study [63], which documented similar patterns of gene expression upon basolateral administration of Salmonella to mouse organoids as had been observed earlier in literature with other model systems. Thus, the hiPSC organoids maintain a conserved response to Salmonella infection and provide a human-specific model for pathogenesis studies.
A subsequent study further demonstrated the validity of hiPSCs as a model to study human-specific responses to Salmonella infection. Using the same model system, the role of the cytokine IL-22 in priming intestinal epithelial cells towards a more effective response to S. Typhimurium was also explored [76]. The study showed that IL-22 pre-treated hiPSC-derived organoids increased phagolysosomal fusion leading to enhanced antimicrobial activity. Thus, this study confirmed earlier observations made in mouse organoids [77].
The fidelity of organoid-derived data in representing human disease was further demonstrated in 2018 by Nickerson, KP and colleagues [78], who compared infection of human tissue biopsies and human intestinal enteroid-derived monolayers seeded on a 2D Transwell system, and observed that the enteroid-derived epithelial monolayers recapitulated S. Typhi infection observations made in the tissue biopsy model. The authors also carried out transcriptional profiling of both the host tissue and the bacteria in order to determine early critical interactions. Infection with S. Typhi significantly down-regulated several host genes, including those involved in activation of the mucosal immune response, bacterial clearance, and cytoskeletal rearrangement. Interestingly in this model, a down-regulation of SPI1 genes in S. Typhi was observed. This work demonstrated that S. Typhi reduces intestinal inflammation by limiting the induction of pathogen-induced processes through the regulation of virulence gene expression, which is a characteristic feature of human infection with S. Typhi. Transmission electron microscopic comparisons of the tissues and human organoid-derived epithelial monolayers showed that the monolayers reproduced the cytoskeletal arrangements, microvilli destruction, and vesicle-bound bacteria observed in tissues. There were no changes observed in paracellular permeability, increased death of host cells, or bacterial association with M cells, suggesting divergence from S. Typhimurium infection in mice. This study highlights the ability of organoids to compare human-specific responses to each Salmonella serovar, which is important in the context of translational capacity for developing prophylactic or therapeutic intervention strategies against S. Typhimurium versus S. Typhi infections.
Despite the multiple advantages of the organoids as an experimental system, the technology is still in its infancy and has certain limitations. The complex structure of the organoids poses a practical limitation in accessing the internal luminal compartment. Researchers have used microinjections to access the apical epithelium. This approach may preserve the internal microenvironment, but is resource-intensive, may not allow synchronous exposure and suffers from variability in the volume that can be injected due to heterogeneity of the organoid/enteroid sizes. In addition, the lumen of 3D organoid/enteroid accumulates cellular debris, which may bind bacteria or hamper interactions with the apical membranes. Researchers have turned to organoid/enteroid-derived 2D monolayers to better access the apical side of the model and enable more efficient, user-friendly analyses in a multiple-well plate format. However, this modification can limit the number of processes that can be interrogated, especially when considering the lack of 3D structure. Interestingly, Co and colleagues [79] demonstrated in a recent study that the polarity of human enteroids could be reversed such that the apical surface faced the medium and was readily accessible. The enteroids released mucus and extruded cells outwards into the culture medium rather than having the cells embedded in the basement membrane. Using enteroids with reversed polarity, the authors showed that S. Typhimurium invades and induces actin ruffles more efficiently at the apical surface compared to the basolateral surface. The authors observed a more diffuse process of epithelial invasion rather than invasion only or predominantly at the M cells [79], which confirmed the S. Typhi observations by Nickerson, KP et al. [78].
Current organoid/enteroid models are devoid of muscles, innervation, vascularization, and immune cells. There are a couple of approaches being carried out to increase the complexity of organoid models, including co-culturing techniques. In 2011, Salerno-Goncalves and colleagues [60] generated an organotypic model using the human ileococal adenocarcinoma cell line HCT-8 and adding primary endothelial cells, fibroblasts, and peripheral blood mononuclear cells (PBMCs), which they used in a 2019 study to probe the crosstalk between these cell types during infection with S. Typhi, S. Paratyphi A, or S. Paratyphi B [80]. An ECM composed of collagen-I enriched with other gastrointestinal basement proteins was embedded with the fibroblasts and epithelial cells, and transferred to an RWV bioreactor containing epithelial cells. Under low microgravity and low shear conditions, the HCT-8 cells behaved as multipotent progenitor cells and gave rise to multiple cell types, including absorptive enterocytes, goblet cells, and M cells. After one to two weeks, PBMCs were added to the system. The co-culture model was then infected with the various Salmonella serovars to compare responses to the three strains. The authors found that the presence of the immune cells in the model resulted in secretion of the cytokines IL-1β and CCL3, while secretion of cytokines IL-6 and TNF-α was enhanced. Using depletion experiments, the authors showed that macrophages were the PBMC cell type responsible for the enhanced secretion of IL-6 and TNF-α. The authors further used the Transwell system to show that supernatants from organotypic models built with whole or macrophage-depleted PBMCs infected with the three Salmonella strains varied in their ability to elicit transmigration of macrophages and neutrophils [80]. Interestingly, the two immune cells displayed crosstalk during infections with S. Paratyphi A and S. Paratyphi B, such that the presence of macrophages in the co-culture reduced neutrophil migration as compared to the system built without macrophages [80]. This study illustrates that co-cultures can aid in probing the contribution of immune cells to Salmonella infection at the mucosal surface. Finally, this model has also been used to assess the inflammatory response to several candidate S. Typhi vaccine strains in comparison to the response elicited by the oral vaccine strain Ty21a strain and its parent wild-type Ty2 strain [81]. Salerno-Goncalves and colleagues [81] found that specific changes to the genetic makeup of the candidate vaccine strains (in the form of deletions of specific metabolic genes) elicited host changes in intestinal permeability, inflammatory cytokine secretion, as well as activation of innate immunity pathways. Higgins and colleagues [82] also used the model to test the inflammatory response of an S. Typhimurium vaccine strain that they generated. These studies highlight the usefulness of co-cultured organoid/enteroid models in assessing important factors to be considered while designing vaccines.
Schulte and colleagues [83] generated a co-culture system of human intestinal epithelial cell line (Caco-2), primary human microvascular endothelial cells, primary intestinal collagen scaffold, and PBMCs in a Transwell set up. Using GFP-labeled S. Typhimurium, microscopy, and flow cytometry, the authors demonstrated that the bacteria can be found in epithelial but not endothelial cells, thus modeling the epithelium-restricted infection of humans with S. Typhimurium. These findings are in contrast to those of Spadoni and colleagues [24] in the mouse model of S. Typhimurium infections where a breach of the gut-vascular barrier by the bacteria was observed. The endothelial cells respond to the infection process by bringing about changes in the transcription of various genes and releasing the phagocyte chemoattractant IL-8. Such models, ideally with enteroid/organoid-derived cells replacing cell lines where used, should prove to be extremely useful and versatile in interrogating the role of different immune cells, vasculature, and the related crosstalk with epithelial cells during infection with Salmonella, especially for S. Typhi, where the bacteria spread systemically both as free bacteria and within reticuloendothelial cells [84].
References
- Raquel M. Callejón; M. Isabel Rodríguez-Naranjo; Cristina Ubeda; Ruth Hornedo-Ortega; M. Carmen García-Parrilla; Ana M. Troncoso; Reported Foodborne Outbreaks Due to Fresh Produce in the United States and European Union: Trends and Causes. Foodborne Pathogens and Disease 2015, 12, 32-38, 10.1089/fpd.2014.1821.
- Stanaway, J.D.; Reiner, R.C.; Blacker, B.F.; Goldberg, E.M.; Khalil, I.A.; Troeger, C.E.; Andrews, J.R.; Bhutta, Z.A.; Crump, J.A.; Im, J; et al. The global burden of typhoid and paratyphoid fevers: a systematic analysis for the Global Burden of Disease Study 2017. The Lancet Infectious Diseases 2019, 19, 369-381, 10.1016/s1473-3099(18)30685-6.
- Renée M. Tsolis; Glenn M. Young; Jay V. Solnick; Andreas J. Bäumler; From bench to bedside: stealth of enteroinvasive pathogens. Nature Reviews Genetics 2008, 6, 883-892, 10.1038/nrmicro2012.
- Parry, C.M.; Hein, T.T.; Dougan, G.; White, N.J.; Farrar, J.J; Typhoid fever. N. Engl. J. Med. 2002, 347, 1770–1782.
- Anderson, C.; Kendall, M.M; Salmonella enterica Serovar Typhimurium strategies for host adaptation. Front. Microbiol. 2017, 8, 1983.
- Nicholas Feasey; Gordon Dougan; Robert A Kingsley; Robert S Heyderman; Melita A Gordon; Invasive non-typhoidal salmonella disease: an emerging and neglected tropical disease in Africa. The Lancet 2012, 379, 2489-2499, 10.1016/S0140-6736(11)61752-2.
- Amreeta Dhanoa; Quek Kia Fatt; Non-typhoidal Salmonella bacteraemia: Epidemiology, clinical characteristics and its' association with severe immunosuppression. Annals of Clinical Microbiology and Antimicrobials 2009, 8, 15, 10.1186/1476-0711-8-15.
- Aj Brent; Joe O. Oundo; Isaiah Mwangi; Lucy Ochola; Brett Lowe; James A. Berkley; Salmonella Bacteremia in Kenyan Children. The Pediatric Infectious Disease Journal 2006, 25, 230-236, 10.1097/01.inf.0000202066.02212.ff.
- Shannon E. Majowicz; Jennie Musto; Elaine Scallan; Frederick J. Angulo; Martyn D. Kirk; Sarah J. O'brien; Timothy F. Jones; Aamir Fazil; Robert M. Hoekstra; The Global Burden of NontyphoidalSalmonellaGastroenteritis. Clinical Infectious Diseases 2010, 50, 882-889, 10.1086/650733.
- Stanaway, J.D.; Reiner, R.C.; Blacker, B.F.; Goldberg, E.M.; Khalil, I.A.; Troeger, C.E.; Andrews, J.R.; Bhutta, Z.A.; Crump, J.A.; Im, J; et al. The global burden of typhoid and paratyphoid fevers: a systematic analysis for the Global Burden of Disease Study 2017. The Lancet Infectious Diseases 2019, 19, 369-381, 10.1016/s1473-3099(18)30685-6.
- Marina Antillon; Joshua L. Warren; Forrest W. Crawford; Daniel M. Weinberger; Esra KÜRÜM; Gi Deok Pak; Florian Marks; Virginia E. Pitzer; The burden of typhoid fever in low- and middle-income countries: A meta-regression approach. PLOS Neglected Tropical Diseases 2017, 11, e0005376, 10.1371/journal.pntd.0005376.
- Denise DeRoeck; Luis Jódar; John Clemens; Putting Typhoid Vaccination on the Global Health Agenda. New England Journal of Medicine 2007, 357, 1069-1071, 10.1056/nejmp078144.
- J. E. Galan; R. Curtiss; Cloning and molecular characterization of genes whose products allow Salmonella typhimurium to penetrate tissue culture cells.. Proceedings of the National Academy of Sciences 1989, 86, 6383-6387, 10.1073/pnas.86.16.6383.
- Renée M. Tsolis; L. Garry Adams; Thomas A. Ficht; Andreas J. Bäumler; Contribution of Salmonella typhimuriumVirulence Factors to Diarrheal Disease in Calves. Infection and Immunity 1999, 67, 4879-4885, 10.1128/iai.67.9.4879-4885.1999.
- Jose Antonio Ibarra; Olivia Steele-Mortimer; Salmonella – the ultimate insider. Salmonella virulence factors that modulate intracellular survival. Cellular Microbiology 2009, 11, 1579-1586, 10.1111/j.1462-5822.2009.01368.x.
- B. B. Finiay; S. Falkow; Salmonella as an intracellular parasite. Molecular Microbiology 1989, 3, 1833-1841, 10.1111/j.1365-2958.1989.tb00170.x.
- Agneta Richter-Dahlfors; A.M.J. Buchan; B B Finlay; Murine Salmonellosis Studied by Confocal Microscopy: Salmonella typhimurium Resides Intracellularly Inside Macrophages and Exerts a Cytotoxic Effect on Phagocytes In Vivo. Journal of Experimental Medicine 1997, 186, 569-580, 10.1084/jem.186.4.569.
- Swart, A.L.; Hensel, M; Interactions of Salmonella enterica with dendritic cells. Virulence 2012, 3, 660–667.
- Wu, J.; Sabag-Daigle, A.; Borton, M.A.; Kop, L.F.M.; Szkoda, B.E.; Kaiser, B.L.D.; Lindemann, S.R.; Renslow, R.S.; Wei, S.; Nicora, C.D; et al. Salmonella-Mediated Inflammation Eliminates Competitors for Fructose-Asparagine in the Gut. Infection and Immunity 2018, 86, e00945-17, 10.1128/IAI.00945-17.
- Thiennimitr, P.; Winter, S.E.; Winter, M.G.; Xavier, M.N.; Tolstikov, V.; Huseby, U.L.; Sterzenbach, T.; Tsolis, R.M.; Roth, J.R.; Bäumler, A.J; et al. Intestinal inflammation allows Salmonella to use ethanolamine to compete with the microbiota. Proceedings of the National Academy of Sciences 2011, 108, 17480-17485, 10.1073/pnas.1107857108.
- Winter, S.E.; Thiennimitr, P.; Winter, M.G.; Butler, B.P.; Huseby, U.L.; Crawford, R.W.; Russell, J.M.; Bevins, C.L.; Adams, L.G.; Tsolis, R.M; et al. Gut inflammation provides a respiratory electron acceptor for Salmonella. Nature 2010, 467, 426-9, 10.1038/nature09415.
- Winter, S.E.; Winter, M.G.; Poon, V.; Keestra, A.M.; Sterzenbach, T.; Faber, F.; Costa, L.F.; Cassou, F.; Costa, E.A.; Alves, G.E.S; et al. Salmonella enterica Serovar Typhi Conceals the Invasion-Associated Type Three Secretion System from the Innate Immune System by Gene Regulation. PLOS Pathogens 2014, 10, e1004207, 10.1371/journal.ppat.1004207.
- Wangdi, T.; Lee, C.-Y.; Spees, A.M.; Yu, C.; Kingsbury, D.D.; Winter, S.E.; Hastey, C.J.; Wilson, R.P.; Heinrich, V.; Bäumler, A.J; et al. The Vi Capsular Polysaccharide Enables Salmonella enterica Serovar Typhi to Evade Microbe-Guided Neutrophil Chemotaxis. PLOS Pathogens 2014, 10, e1004306, 10.1371/journal.ppat.1004306.
- Ilaria Spadoni; Elena Zagato; Alice Bertocchi; Roberta Paolinelli; Edina Hot; Antonio Di Sabatino; Flavio Caprioli; Luca Bottiglieri; Amanda Oldani; Giuseppe Viale; G. Penna; E. Dejana; Maria Rescigno; A gut-vascular barrier controls the systemic dissemination of bacteria. Science 2015, 350, 830-834, 10.1126/science.aad0135.
- R. B. Hornick; S. E. Greisman; T. E. Woodward; H. L. Dupont; A. T. Dawkins; M. J. Snyder; Typhoid Fever: Pathogenesis and Immunologic Control. New England Journal of Medicine 1970, 283, 686-691, 10.1056/nejm197009242831306.
- Shukla, V.; Singh, H.; Pandey, R.; Upadhyay, S.; Nath, G; Carcinoma of the Gallbladder—Is it a sequel of typhoid. Dig. Dis. Sci. 2000, 45, 900–903.
- C.P.J. Caygill; M.J. Hill; M. Braddick; J.C.M. Sharp; Cancer mortality in chronic typhoid and paratyphoid carriers. The Lancet 1994, 343, 83-84, 10.1016/s0140-6736(94)90816-8.
- Lapo Mughini-Gras; Michael Schaapveld; Jolanda Kramers; Sofie Mooij; E. Andra Neefjes-Borst; Wilfrid Van Pelt; Jacques Neefjes; Increased colon cancer risk after severe Salmonella infection. PLOS ONE 2018, 13, e0189721, 10.1371/journal.pone.0189721.
- Ian F. Connerton; John Wain; Tran T. Hien; Tahir Ali; Christopher Parry; Nguyen T. Chinh; Ha Vinh; Vo A. Ho; To S. Diep; Nicholas P. J. Day; Nicholas J. White; Gordon Dougan; Jeremy Farrar; Epidemic Typhoid in Vietnam: Molecular Typing of Multiple-Antibiotic-Resistant Salmonella enterica Serotype Typhi from Four Outbreaks. Journal of Clinical Microbiology 2000, 38, 895-897, 10.1128/jcm.38.2.895-897.2000.
- Aoife Molloy; Satheesh Nair; Fiona J. Cooke; John Wain; Mark Farrington; Paul J. Lehner; Estee Torok; First Report of Salmonella enterica Serotype Paratyphi A Azithromycin Resistance Leading to Treatment Failure. Journal of Clinical Microbiology 2010, 48, 4655-4657, 10.1128/JCM.00648-10.
- Bernard Rowe; Linda R. Ward; E. John Threlfall; Multidrug-Resistant Salmonella typhi: A Worldwide Epidemic. Clinical Infectious Diseases 1997, 24, S106-S109, 10.1093/clinids/24.Supplement_1.S106.
- Xuchu Wang; Silpak Biswas; Narayan Paudyal; Hang Pan; Xiaoliang Li; Weihuan Fang; Min Yue; Antibiotic Resistance in Salmonella Typhimurium Isolates Recovered From the Food Chain Through National Antimicrobial Resistance Monitoring System Between 1996 and 2016.. Frontiers in Microbiology 2019, 10, 985, 10.3389/fmicb.2019.00985.
- Smita Gopinath; Joshua Lichtman; nna M. Bouley; Joshua E. Elias; Denise M Monack; Role of disease-associated tolerance in infectious superspreaders. Proceedings of the National Academy of Sciences 2014, 111, 15780-15785, 10.1073/pnas.1409968111.
- Médéric Diard; Mikael E. Sellin; Tamas Dolowschiak; Markus Arnoldini; Martin Ackermann; Wolf-Dietrich Hardt; Antibiotic Treatment Selects for Cooperative Virulence of Salmonella Typhimurium. Current Biology 2014, 24, 2000-2005, 10.1016/j.cub.2014.07.028.
- Sandra Y. Wotzka; Bidong D. Nguyen; Wolf-Dietrich Hardt; Salmonella Typhimurium Diarrhea Reveals Basic Principles of Enteropathogen Infection and Disease-Promoted DNA Exchange. Cell Host & Microbe 2017, 21, 443-454, 10.1016/j.chom.2017.03.009.
- Camilo J. Acosta; Cm Galindo; Jl Deen; Rl Ochiai; HJ Lee; L Von Seidlein; R Carbis; Jd Clemens; Leon Ochiai; Vaccines against cholera, typhoid fever and shigellosis for developing countries. Expert Opinion on Biological Therapy 2004, 4, 1939-1951, 10.1517/14712598.4.12.1939.
- Ageing United Nations. Available online: https://www.un.org/en/sections/issues-depth/ageing/ (accessed on 9 February 2020).
- Featured Graphic: Many Countries’ Populations Are Aging—Population Reference Bureau. Available online: https://www.prb.org/insight/featured-graphic-many-countries-populations-are-aging/ (accessed on 9 February 2020).
- Smriti Verma; C. V. Srikanth; Understanding the complexities of Salmonella-host crosstalk as revealed byin vivomodel organisms. IUBMB Life 2015, 67, 482-497, 10.1002/iub.1393.
- A K Criss; D M Ahlgren; T S Jou; B A McCormick; J E Casanova; The GTPase Rac1 selectively regulates Salmonella invasion at the apical plasma membrane of polarized epithelial cells.. Journal of Cell Science 2001, 114, 1331–1341.
- Huadong Sun; Edwin Cy Chow; Shanjun Liu; Yimin Du; K. Pang; The Caco-2 cell monolayer: usefulness and limitations. Expert Opinion on Drug Metabolism & Toxicology 2008, 4, 395-411, 10.1517/17425255.4.4.395.
- Y. Sambuy; Isabella De Angelis; Giulia Ranaldi; M. L. Scarino; A. Stammati; F. Zucco; The Caco-2 cell line as a model of the intestinal barrier: influence of cell and culture-related factors on Caco-2 cell functional characteristics. Cell Biology and Toxicology 2005, 21, 1-26, 10.1007/s10565-005-0085-6.
- Grossmann, J. Progress on isolation and short-term ex-vivo culture of highly purified non-apoptotic human intestinal epithelial cells (IEC). Eur. J. Cell Boil. 2003, 82, 262–270.
- M. C. Aldhous; A. N. Shmakov; J. Bode; S. Ghosh; Characterization of conditions for the primary culture of human small intestinal epithelial cells. Clinical and Experimental Immunology 2001, 125, 32-40, 10.1046/j.1365-2249.2001.01522.x.
- Renato De Lima Santos; Renée M. Tsolis; Shuping Zhang; Thomas A. Ficht; Andreas J. Bäumler; L. Garry Adams; Salmonella-Induced Cell Death Is Not Required for Enteritis in Calves. Infection and Immunity 2001, 69, 4610-4617, 10.1128/iai.69.7.4610-4617.2001.
- Renato De Lima Santos; S. Zhang; Renée M. Tsolis; Andreas J. Bäumler; Leslie Garry Adams; Morphologic and Molecular Characterization ofSalmonella typhimuriumInfection in Neonatal Calves. Veterinary Pathology 2002, 39, 200-215, 10.1354/vp.39-2-200.
- Jepson, M.A.; Kenny, B.; Leard, A.D. Role of sipA in the early stages of Salmonella Typhimurium entry into epithelial cells. Cell Microbiol. 2001, 3, 417–426.
- Shuping Zhang; Renato De Lima Santos; Renée M. Tsolis; Silke Stender; Wolf-Dietrich Hardt; Andreas J. Bäumler; L. Garry Adams; The Salmonella enterica Serotype Typhimurium Effector Proteins SipA, SopA, SopB, SopD, and SopE2 Act in Concert To Induce Diarrhea in Calves. Infection and Immunity 2002, 70, 3843-3855, 10.1128/iai.70.7.3843-3855.2002.
- Renato De Lima Santos; Shuping Zhang; Renée M. Tsolis; Robert A. Kingsley; L. Garry Adams; Andreas J. Bäumler; Animal models of Salmonella infections: enteritis versus typhoid fever. Microbes and Infection 2001, 3, 1335-1344, 10.1016/s1286-4579(01)01495-2.
- Barthel, M.; Hapfelmeier, S.; Quintanilla-Martinez, L.; Kremer, M.; Rohde, M.; Hogardt, M.; Pfeffer, K.; Rüssmann, H.; Hardt, W.-D; Pretreatment of Mice with Streptomycin Provides a Salmonella enterica Serovar Typhimurium Colitis Model That Allows Analysis of Both Pathogen and Host. Infection and Immunity 2003, 71, 2839-2858, 10.1128/iai.71.5.2839-2858.2003.
- Isabelle Virlogeux-Payant; H. Waxin; C. Ecobichon; M. Y. Popoff; Role of the viaB locus in synthesis, transport and expression of Salmonella typhi Vi antigen. Microbiology 1995, 141, 3039-3047, 10.1099/13500872-141-12-3039.
- Sébastien C. Sabbagh; Chantal G. Forest; Christine Lepage; Jean-Mathieu Leclerc; France Daigle; So similar, yet so different: uncovering distinctive features in the genomes of Salmonella enterica serovars Typhimurium and Typhi. FEMS Microbiology Letters 2010, 305, 1-13, 10.1111/j.1574-6968.2010.01904.x.
- M. Firoz Mian; Elisabeth A. Pek; Meghan Chenoweth; Brian K. Coombes; Ali A Ashkar; Humanized mice for Salmonella typhi infection: new tools for an old problem. Virulence 2011, 2, 248-252, 10.4161/viru.2.3.16133.
- Libby, S.J.; Brehm, M.A.; Greiner, D.L.; Shultz, L.D.; McClelland, M.; Smith, K.D.; Cookson, B.T.; Karlinsey, J.E.; Kinkel, T.L.; Porwollik, S; et al. Humanized nonobese diabetic-scid IL2rγnull mice are susceptible to lethal Salmonella Typhi infection. Proceedings of the National Academy of Sciences 2010, 107, 15589-15594, 10.1073/pnas.1005566107.
- Aparna Bhaduri; Madeline G. Andrews; Walter Mancia Leon; Diane Jung; David Shin; Denise Allen; Dana Jung; Galina Schmunk; Maximilian Haeussler; Jahan Salma; Alex A. Pollen; Tomasz J. Nowakowski; Arnold R. Kriegstein; Cell stress in cortical organoids impairs molecular subtype specification. Nature 2020, 578, 142-148, 10.1038/s41586-020-1962-0.
- Y Sambuy; I De Angelis; Formation of organoid structures and extracellular matrix production in an intestinal epithelial cell line during long-term in vitro culture.. Cell Differentiation 1986, 19, 139–147.
- Emile Levy; Edgard E. Delvin; Daniel Ménard; Jean-François Beaulieu; Functional Development of Human Fetal Gastrointestinal Tract. Advanced Structural Safety Studies 2009, 550, 205-224, 10.1007/978-1-60327-009-0_13.
- Toshiro Sato; Robert G. Vries; Hugo J. Snippert; Marc Van De Wetering; Nick Barker; Daniel Stange; Johan H. Van Es; Arie Abo; Pekka Kujala; Peter J. Peters; Hans Clevers; Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 2009, 459, 262-265, 10.1038/nature07935.
- Toshiro Sato; Daniel Stange; Marc Ferrante; Robert G. Vries; Johan H. Van Es; Stieneke Van Den Brink; Winan J. Van Houdt; Apollo Pronk; Joost M Van Gorp; Peter D. Siersema; Hans Clevers; Long-term Expansion of Epithelial Organoids From Human Colon, Adenoma, Adenocarcinoma, and Barrett's Epithelium. Gastroenterology 2011, 141, 1762-1772, 10.1053/j.gastro.2011.07.050.
- Rosângela Salerno-Gonçalves; Alessio Fasano; Marcelo B. Sztein; Engineering of a multicellular organotypic model of the human intestinal mucosa.. Gastroenterology 2011, 141, e18-20, 10.1053/j.gastro.2011.04.062.
- Stelzner, M.G.; Helmrath, M.; Dunn, J.C.Y.; Henning, S.J.; Houchen, C.W.; Kuo, C.; Lynch, J.; Li, L.; Magness, S.T.; Martin, M.G; et al. A nomenclature for intestinal in vitro cultures. American Journal of Physiology-Gastrointestinal and Liver Physiology 2012, 302, G1359–G1363, 10.1152/ajpgi.00493.2011.
- Cheryl A. Nickerson; Thomas J. Goodwin; Jacqueline Terlonge; C. Mark Ott; Kent L. Buchanan; William C. Uicker; Kamal Emami; Carly L. Leblanc; Rajee Ramamurthy; Mark S. Clarke; Charles R. Vanderburg; Timothy Hammond; Duane L. Pierson; Three-Dimensional Tissue Assemblies: Novel Models for the Study of Salmonella enterica Serovar Typhimurium Pathogenesis. Infection and Immunity 2001, 69, 7106-7120, 10.1128/iai.69.11.7106-7120.2001.
- Yong‐Guo Zhang; Shaoping Wu; Yinglin Xia; Jun Sun; Salmonella‐infected crypt‐derived intestinal organoid culture system for host–bacterial interactions. Physiological Reports 2014, 2, e12147, 10.14814/phy2.12147.
- Henner F. Farin; Wouter R. Karthaus; Pekka Kujala; Maryam Rakhshandehroo; Gerald Schwank; Robert G.J. Vries; Eric Kalkhoven; Edward E.S. Nieuwenhuis; Hans Clevers; Paneth cell extrusion and release of antimicrobial products is directly controlled by immune cell–derived IFN-γ. Journal of Experimental Medicine 2014, 211, 1393-1405, 10.1084/jem.20130753.
- Sarah S Wilson; Autumn Tocchi; Mayumi K Holly; William C Parks; Jason G. Smith; A small intestinal organoid model of non-invasive enteric pathogen-epithelial cell interactions.. Mucosal Immunology 2014, 8, 352-61, 10.1038/mi.2014.72.
- Wilson, C.L.; Ouellette, A.J.; Satchell, D.P.; Ayabe, T.; López-Boado, Y.S.; Stratman, J.L.; Hultgren, S.J.; Matrisian, L.M.; Parks, W.C; Regulation of intestinal alpha-defensin activation by the metalloproteinase matrilysin in innate host defense. Science 1999, 286, 113–117.
- Charles L. Bevins; Nita Salzman; Paneth cells, antimicrobial peptides and maintenance of intestinal homeostasis. Nature Reviews Genetics 2011, 9, 356-368, 10.1038/nrmicro2546.
- Scanu, T.; Spaapen, R.M.; Bakker, J.M.; Pratap, C.B.; Wu, L.-E.; Hofland, I.; Broeks, A.; Shukla, V.K.; Kumar, M.; Janssen, H; et al. Salmonella Manipulation of Host Signaling Pathways Provokes Cellular Transformation Associated with Gallbladder Carcinoma. Cell Host & Microbe 2015, 17, 763-774, 10.1016/j.chom.2015.05.002.
- S-C Kuo; Yu-Wen Hu; C-J Liu; Y-T Lee; Y-T Chen; T-L Chen; T-J Chen; C-P Fung; Association between tuberculosis infections and non-pulmonary malignancies: a nationwide population-based study. British Journal of Cancer 2013, 109, 229-234, 10.1038/bjc.2013.220.
- Zhan, P.; Suo, L.-J.; Qian, Q.; Shen, X.-K.; Qiu, L.-X.; Yu, L.; Song, Y; Chlamydia pneumoniae infection and lung cancer risk: A meta-analysis. European Journal of Cancer 2011, 47, 742-747, 10.1016/j.ejca.2010.11.003.
- Arnheim-Dahlström, L.; Andersson, K.; Luostarinen, T.; Thoresen, S.; Ögmundsdottír, H.; Tryggvadottir, L.; Wiklund, F.; Skare, G.B.; Eklund, C.; Sjölin, K.; et al. Prospective seroepidemiologic study of uman Papillomavirus and other risk factors in cervical cancer. Cancer Epidemiol. Biomark. Prev. 2011, 20, 2541–2550.
- Aurélie Gagnaire; Bertrand Nadel; Didier Raoult; Jacques Neefjes; Jean-Pierre E. Gorvel; Collateral damage: insights into bacterial mechanisms that predispose host cells to cancer. Nature Reviews Genetics 2017, 15, 109-128, 10.1038/nrmicro.2016.171.
- Lu, X.; Xie, S.; Ye, L.; Zhu, L.; Yu, Q; Lu, X.; Xie, S.; Ye, L.; Zhu, L.; Yu, Q. Lactobacillus protects against S. Typhimurium –induced intestinal inflammation by determining the fate of epithelial proliferation and differentiation. Mol. Nutr. Food Res. 2020, 64, e1900655.. Mol. Nutr. Food Res. 2020, 64, e1900655.
- Pengcheng Li; Qinghua Yu; Xiaolan Ye; Zhisheng Wang; Qian Yang; Lactobacillus S-layer protein inhibition of Salmonella-induced reorganization of the cytoskeleton and activation of MAPK signalling pathways in Caco-2 cells. Microbiology 2011, 157, 2639-2646, 10.1099/mic.0.049148-0.
- Jessica L. Forbester; David Goulding; Ludovic Vallier; Nicholas R. F. Hannan; Christine Hale; Derek Pickard; Subhankar Mukhopadhyay; Gordon Dougan; Interaction of Salmonella enterica Serovar Typhimurium with Intestinal Organoids Derived from Human Induced Pluripotent Stem Cells.. Infection and Immunity 2015, 83, 2926-2934, 10.1128/IAI.00161-15.
- Jessica L. Forbester; Emily A. Lees; David Goulding; Sally Forrest; Amy Yeung; Anneliese O. Speak; Simon Clare; Eve L. Coomber; Subhankar Mukhopadhyay; Judith Kraiczy; Fernanda Schreiber; Trevor D. Lawley; Robert E.W. Hancock; Holm H. Uhlig; Matthias Zilbauer; Fiona Powrie; Gordon Dougan; Interleukin-22 promotes phagolysosomal fusion to induce protection against Salmonella enterica Typhimurium in human epithelial cells.. Proceedings of the National Academy of Sciences 2018, 115, 10118-10123, 10.1073/pnas.1811866115.
- Tu Anh N. Pham; Simon Clare; David Goulding; Julia M. Arasteh; Mark D. Stares; Hilary P. Browne; Jacqueline A. Keane; Andrew Page; Natsuhiko Kumasaka; Leanne Kane; Lynda Mottram; Katherine Harcourt; Christine Hale; Mark J. Arends; Daniel J. Gaffney; Gordon Dougan; T. D. Lawley; Epithelial IL-22RA1-Mediated Fucosylation Promotes Intestinal Colonization Resistance to an Opportunistic Pathogen. Cell Host & Microbe 2014, 16, 504-516, 10.1016/j.chom.2014.08.017.
- K.P. Nickerson; S. Senger; Y. Zhang; R. Lima; S. Patel; L. Ingano; W.A. Flavahan; D.K.V. Kumar; C.M. Fraser; Christina S. Faherty; M.B. Sztein; M. Fiorentino; Alessio Fasano; Salmonella Typhi Colonization Provokes Extensive Transcriptional Changes Aimed at Evading Host Mucosal Immune Defense During Early Infection of Human Intestinal Tissue. EBioMedicine 2018, 31, 92-109, 10.1016/j.ebiom.2018.04.005.
- Co, J.Y.; Margalef-Català, M.; Li, X.; Mah, A.T.; Kuo, C.J.; Monack, D.M.; Amieva, M; Controlling Epithelial Polarity: A Human Enteroid Model for Host-Pathogen Interactions. Cell Reports 2019, 26, 2509-2520, 10.1016/j.celrep.2019.01.108.
- Rosângela Salerno-Gonçalves; Darpan Kayastha; Alessio Fasano; Myron M. Levine; Marcelo B. Sztein; Crosstalk between leukocytes triggers differential immune responses against Salmonella enterica serovars Typhi and Paratyphi. PLOS Neglected Tropical Diseases 2019, 13, e0007650, 10.1371/journal.pntd.0007650.
- Salerno-Gonçalves, R.; Galen, J.E.; Levine, M.M.; Fasano, A.; Sztein, M.B; Manipulation of Salmonella Typhi Gene Expression Impacts Innate Cell Responses in the Human Intestinal Mucosa. Frontiers in Immunology 2018, 9, 2543, 10.3389/fimmu.2018.02543.
- Higginson, E.E.; Ramachandran, G.; Panda, A.; Shipley, S.T.; Kriel, E.H.; DeTolla, L.J.; Lipsky, M.; Perkins, D.J.; Salerno-Gonçalves, R.; Sztein, M.B; et al. Improved Tolerability of a Salmonella enterica Serovar Typhimurium Live-Attenuated Vaccine Strain Achieved by Balancing Inflammatory Potential with Immunogenicity. Infection and Immunity 2018, 86, e00440-18, 10.1128/iai.00440-18.
- Schulte, L.N.; Schweinlin, M.; Westermann, A.J.; Janga, H.; Santos, S.C.; Appenzeller, S.; Walles, H.; Vogel, J.; Metzger, M; An Advanced Human Intestinal Coculture Model Reveals Compartmentalized Host and Pathogen Strategies during Salmonella Infection. mBio 2020, 11, e03348-19, 10.1128/mBio.03348-19.
- Dougan, G.; Baker, S; Salmonella enterica serovar Typhi and the pathogenesis of typhoid fever. Annu. Rev. Microbiol. 2014, 68, 317–336.





