
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Mojtaba Golpich | -- | 3022 | 2024-03-05 17:09:56 | | | |
| 2 | Catherine Yang | + 1 word(s) | 3023 | 2024-03-06 02:06:19 | | |
Video Upload Options
Aptamers developed using in vitro Systematic Evolution of Ligands by Exponential Enrichment (SELEX) technology are single-stranded nucleic acids 10–100 nucleotides in length. Their targets, often with specificity and high affinity, range from ions and small molecules to proteins and other biological molecules as well as larger systems, including cells, tissues, and animals. Aptamers often rival conventional antibodies with improved performance, due to aptamers’ unique biophysical and biochemical properties, including small size, synthetic accessibility, facile modification, low production cost, and low immunogenicity. Therefore, there is sustained interest in engineering and adapting aptamers for many applications, including diagnostics and therapeutics.
1. Introduction
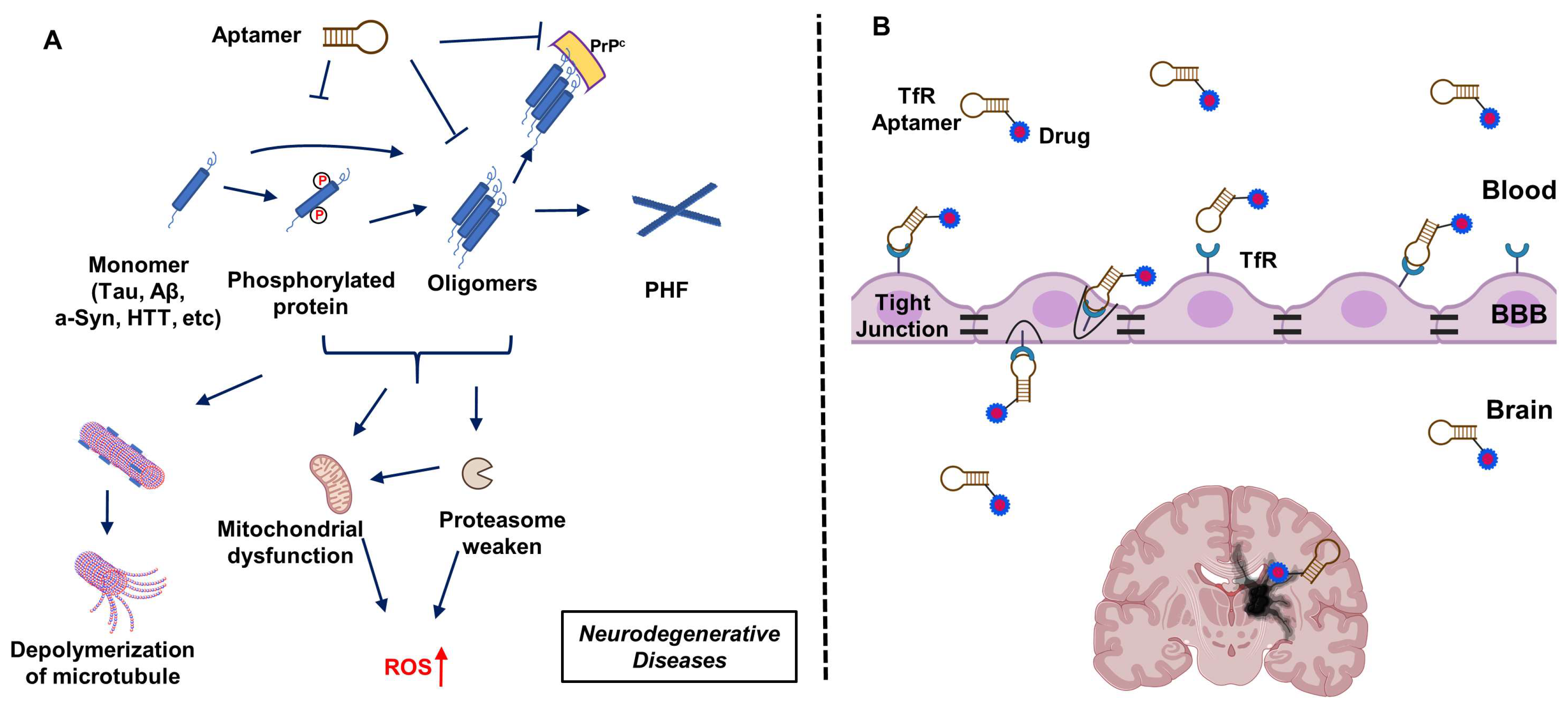
2. Aptamers Targeting Neuro-Medically Relevant Targets
| Target | Aptamers (Name) | Dissociation Constant |
Disease | Utility, Key Results | Ref. |
|---|---|---|---|---|---|
| GluR1 Ser845 | A1, A2, A3 (RNA) | 28–57 nM | Protein phosphorylation-related diseases | A2 binds GluR1 that inhibits GluR1/GluR1 containing AMPA receptor trafficking to the cell surface and abrogates forskolin-stimulated phosphorylation at GluR1 Ser845 | [28] |
| (MAPK) Erk1/2 | C5 (RNA) | 10 nM | CNS disorders, Alzheimer’s disease, Stroke, Epilepsy | C5 selective to inhibit the mitogen-activated kinase pathway in neurons | [29] |
| β2-adrenoceptor (β2-AR) (GPCR) | A1, A2, A13, and A16 (RNA) | 30.4–258.5 nM | - | RNA aptamers as allosteric GPCR modulators | [30] |
| Tau protein | IT(1–6), IT2a (DNA) | 5.5–68 nM | Traumatic brain injury, Alzheimer’s disease | Aptamers inhibit tau phosphorylation and oligomerization | [31] |
| Tau-1 (RNA) | ~200 nM | [32] | |||
| Aptamer 314 (DNA) | 13 ± 3 nM | Aptamer binds Tau protein | [33] | ||
| β-Amyloid protein |
β aptamers, e.g., β55 (RNA) | 29–48 nM | Alzheimer’s disease | β55 aptamer binds amyloid plaques in AD brain tissue | [34] |
| E1, E2, N1, N2, G2, etc. (RNA) | 10.9–21.6 μM | N2 aptamer is used to build a luminescent aptamer-ruthenium complex system for the detection of Aβ | [35] | ||
| Apt-GO (RNA) | 0.1–10 μM | Apt-GO selectively detects Aβ1–42 in an AD SH-SY5 cell model | [36] | ||
| Aβ42 | Aβ7-92-1H1 | 63.4 nM | Inhibits Aβ42 aggregation | [37] | |
| Aβ42 dimer | E22P–AbD43 (RNA) | 20 ± 6.0 nM | Aptamer inhibits the nucleation phase of the dimer and its associated neurotoxicity in SH-SY5Y human neuroblastoma cells. | [38] | |
| Aβ40 oligomers | KM (RNA) | - | Aptamers bind with Aβ40 fibrils that may serve as amyloid recognition tools | [39] | |
| α-synuclein protein | F5R1 (DNA) | 2.40 nM | Parkinson’s disease | Blocks the aberrant cellular effects of the overexpressed α-synuclein in cells | [40] |
| T-SO508 (DNA) | 68 nM | T-SO508 can bind to soluble α-synuclein oligomers | [41] | ||
| Apt11(DNA) | - | Parkinson’s disease and dementia with Lewy bodies | Apt11 aptamer binds to α-syn fibrils and inhibits α-syn aggregation in the in vitro model of PD and DLB. | [42] | |
| TMG-79 (DNA) | - | TMG-79 aptamer detects α-syn in Lewy body and PD-associated dementia. | [43] | ||
| M5-15 (DNA) | 14.3 nM | M5-15 aptamer detects α-syn in Lewy body and PD-associated dementia. | [44] | ||
| α-synuclein & amyloid-β (Aβ) | AD-PAINT (DNA) | 500 nM–2 μM | Parkinson’s disease | AD-PAINT aptamer binds to fibrillar aggregates of α-syn and Aβ aggregates detected in both serum and CSF in PD | [45] |
| Dopamine | dopa2 (129 nucleotides); dopa2/c.1 |
2.8 µM 1.6 µM |
Parkinson’s disease | Dopa2 and dopa2/c.1 are characterized to bind a dopamine affinity column; the dopamine binding site is obtained by secondary selection | [46] |
| Toll-like receptor 4 (TLR4) | ApTLR#1R, ApTLR#4F (DNA) | - | Stroke disease | Aptamers have a TLR4 antagonistic effect | [24] |
| ApTOLL (DNA) | 20 nM | Acute ischemic stroke | ApTOLL aptamer binds and antagonizes TLR4 and improves functional outcomes in AIS patients | [25] | |
| Platelet-derived growth factor receptor β (PDGFRβ) | Gint4.T aptamer (RNA) | 9.6 nM | Glioma | Aptamer binds to human DGFRβ ectodomain, causing a strong inhibition of ligand-dependent receptor activation | [47][48] |
| Myelin basic protein | MBPcl3 MBPcl9 (DNA) |
- | Multiple Sclerosis | MBPc13 detects myelin-rich regions in paraffin-embedded mouse brain tissue; aptamer was found more sensitive than a commercial antibody. MBPcl3 blocks the binding of the antibody to MBP | [49] |
| Myelin basic protein (MBP) autoantibody | Apt2-9c (RNA) | 1.2 ± 0.1 nM | Multiple Sclerosis | Apt2-9c provides proof-of-principle for the detection of MS-specific autoantibodies | [50] |
| L1-CAM (Neurites) | yly12 (DNA) | 3.51 nM | Neurite-surface targeting and inhibitory effect on neurite outgrowth between cells | [51] | |
| Prion protein (PrP) | R12 (RNA) | ~10 nM | Creutzfeldt-Jakob disease; prion diseases | R12 binding to PrP results in the dissociation of PrP with Aβ. | [52] |
| clone 4–9 (DNA) | 113 nM | binds to PrP | [53] | ||
| DP7 (RNA) | 0.1–1.7 µM | Prion-protein-specific aptamer reduces PrPSc formation | [54] | ||
| A1 (DNA) | 232 nM | Aptamers modulate phase separation and promote PrP fibrillation | [55] | ||
| R24 (RNA) | 18 nM | R24 exhibited the lowest recorded IC50 and the highest anti-prion activity | [56] | ||
| Crossing BBB (target unknown) | A15 (RNA) | - | Neurological disorders or diseases. | In vivo SELEX (brain-penetrating aptamers) | [57] |
| CD20+B cells | TD05 (DNA) | 256 nM | Glioma | TD05-488 can diagnose CNS lymphoma within 11 min of biopsy from xenograft brain tumor models | [58] |
| U87 glioma cell line/EGFRvIII | QD-A32 (DNA) | - | Glioma | QD-apt can penetrate the BBB and then selectively accumulate in the tumors through binding to EGFRvIII | [59] |
| The Regulator of calcineurin 1 (RCAN1) | R1SR13 (RNA) | 0.3 µM | Down syndrome and Alzheimer’s disease | Inhibits the regulatory function of RCAN1 in NFAT and NF-kB signaling pathways | [60] |
| 0.23–30 nM | Acute ischemic stroke | R1SR13 aptamer alleviates the RCAN1.1 L-induced neuronal apoptosis in the human SHSY-5Y neuroblastoma cells and in the mouse model of AIS | [26] | ||
| TAR-DNA-Binding Protein 43 (TDP-43) | Apt-1 (DNA) | 1 μM | Amyotrophic Lateral Sclerosis | Apt-1 aptamer binds to TDP-43 in the ALS model. | [27] |
3. Applying Experience from Non-CNS Therapeutic Aptamers towards Neuro-Aptamer Development
4. Aptamers Targeting Transferrin Receptor 1 to Facilitate Drug Transport across the BBB
| Transferrin Aptamer Name, Nucleotide Sequence |
2-D Structure | Ref. |
|---|---|---|
| Mouse transferrin receptor-specific GS24, reduced to 50 nucleotides. Sequence (5′-3′): GCGTGTGCACACGGTCACAGTTAGTATCGCTACGTTCTTTGGTAGTCCGTTCGG |
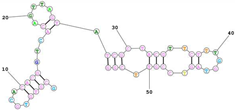 |
[91] |
| Target Mouse TfR Aptamer name: TfRA1 Truncated GS24; 14 nucleotides Sequence (5′-3′): GCGTGTGCACACGC |
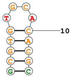 |
[92] |
| Human transferrin receptor specific C2. targets the apical domain of the human transferrin receptor (hTfR) Sequence (5′-3′): CAUCUCACAGAUCAAUCCAAGGCACCUCGUUAAAGGACGACUCCCUUACAUGCGAGAUG |
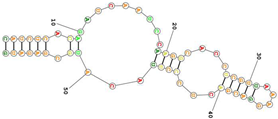 |
[94] |
| Aptamer name: Min2/Waz (RNA) non-competitive for Transferrin. Targets the apical domain of the human transferrin receptor (hTfR) Sequence (5′-3′): GGGUUCUACGAUAAACGGUUAAUGACCAGCUUAUGGCUGGCAGUUCCC |
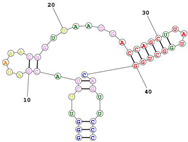 |
[77] |
| Human TfR (cell-SELEX) (XQ-2d Shares a Similar Binding Site on CD71 with Transferrin) Aptamer name: XQ-2d (DNA) Sequence (5′-3′): ACTCATAGGGTTAGGGGCTGCTGGCCAGATACTCAGATGGTAGGGTTACTATGAGC |
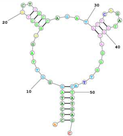 |
[95][96] |
| Human TfR (cell-SELEX) Aptamer name: HG1-9 (DNA) HG1-9 aptamer binds human TfR with affinity (Kd < 20 nM) and completes a same bind site of human TfR with transferrin. Sequence (5′-3′): GGATAGGGATTCTGTTGGTCGGCTGGTTGGTATCC |
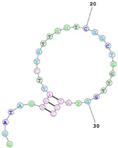 |
[97] |
References
- Ellington, A.D.; Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 1990, 346, 818–822.
- Tuerk, C.; Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 1990, 249, 505–510.
- Gold, L.; Ayers, D.; Bertino, J.; Bock, C.; Bock, A.; Brody, E.N.; Carter, J.; Dalby, A.B.; Eaton, B.E.; Fitzwater, T.; et al. Aptamer-Based Multiplexed Proteomic Technology for Biomarker Discovery. PLoS ONE 2010, 5, e15004.
- Gawande, B.N.; Rohloff, J.C.; Carter, J.D.; von Carlowitz, I.; Zhang, C.; Schneider, D.J.; Janjic, N. Selection of DNA aptamers with two modified bases. Proc. Natl. Acad. Sci. USA 2017, 114, 2898–2903.
- Kimoto, M.; Yamashige, R.; Matsunaga, K.-i.; Yokoyama, S.; Hirao, I. Generation of high-affinity DNA aptamers using an expanded genetic alphabet. Nat. Biotechnol. 2013, 31, 453–457.
- Sefah, K.; Yang, Z.; Bradley, K.M.; Hoshika, S.; Jiménez, E.; Zhang, L.; Zhu, G.; Shanker, S.; Yu, F.; Turek, D. In vitro selection with artificial expanded genetic information systems. Proc. Natl. Acad. Sci. USA 2014, 111, 1449–1454.
- Chen, Z.; Lichtor, P.A.; Berliner, A.P.; Chen, J.C.; Liu, D.R. Evolution of sequence-defined highly functionalized nucleic acid polymers. Nat. Chem. 2018, 10, 420–427.
- Berezhnoy, A.; Stewart, C.A.; Mcnamara II, J.O.; Thiel, W.; Giangrande, P.; Trinchieri, G.; Gilboa, E. Isolation and optimization of murine IL-10 receptor blocking oligonucleotide aptamers using high-throughput sequencing. Mol. Ther. 2012, 20, 1242–1250.
- Kacherovsky, N.; Cardle, I.I.; Cheng, E.L.; Yu, J.L.; Baldwin, M.L.; Salipante, S.J.; Jensen, M.C.; Pun, S.H. Traceless aptamer-mediated isolation of CD8+ T cells for chimeric antigen receptor T-cell therapy. Nat. Biomed. Eng. 2019, 3, 783–795.
- Famulok, M.; Blind, M.; Mayer, G. Intramers as promising new tools in functional proteomics. Chem. Biol. 2001, 8, 931–939.
- Kobeissy, F.H.; Guingab-Cagmat, J.D.; Zhang, Z.; Moghieb, A.; Glushakova, O.Y.; Mondello, S.; Boutté, A.M.; Anagli, J.; Rubenstein, R.; Bahmad, H. Neuroproteomics and systems biology approach to identify temporal biomarker changes post experimental traumatic brain injury in rats. Front. Neurol. 2016, 7, 198.
- Jaber, Z.; Aouad, P.; Al Medawar, M.; Bahmad, H.; Abou-Abbass, H.; Ghandour, H.; Mondello, S.; Kobeissy, F. Role of systems biology in brain injury biomarker discovery: Neuroproteomics application. Methods Mol. Biol. 2016, 1462, 157–174.
- Razafsha, M.; Khaku, A.; Azari, H.; Alawieh, A.; Behforuzi, H.; Fadlallah, B.; Kobeissy, F.H.; Wang, K.K.; Gold, M.S. Biomarker identification in psychiatric disorders: From neuroscience to clinical practice. J. Psychiatr. Pract. 2015, 21, 37–48.
- Kobeissy, F.H.; Guingab-Cagmat, J.D.; Razafsha, M.; O’Steen, L.; Zhang, Z.; Hayes, R.L.; Chiu, W.-T.; Wang, K.K. Leveraging biomarker platforms and systems biology for rehabilomics and biologics effectiveness research. PMR 2011, 3, S139–S147.
- Feigin, V.L.; Nichols, E.; Alam, T.; Bannick, M.S.; Beghi, E.; Blake, N.; Culpepper, W.J.; Dorsey, E.R.; Elbaz, A.; Ellenbogen, R.G. Global, regional, and national burden of neurological disorders, 1990–2016: A systematic analysis for the Global Burden of Disease Study 2016. Lancet Neurol. 2019, 18, 459–480.
- Martier, R.; Konstantinova, P. Gene therapy for neurodegenerative diseases: Slowing down the ticking clock. Front. Neurosci. 2020, 14, 580179.
- Khan, I.; Preeti, K.; Fernandes, V.; Khatri, D.K.; Singh, S.B. Role of MicroRNAs, Aptamers in Neuroinflammation and Neurodegenerative Disorders. Cell. Mol. Neurobiol. 2022, 42, 2075–2095.
- Hauser, S.L.; Cree, B.A. Treatment of multiple sclerosis: A review. Am. J. Med. 2020, 133, 1380–1390.e2.
- Selkoe, D.J. Alzheimer disease and aducanumab: Adjusting our approach. Nat. Rev. Neurol. 2019, 15, 365–366.
- Gandhi, J.; Antonelli, A.C.; Afridi, A.; Vatsia, S.; Joshi, G.; Romanov, V.; Murray, I.V.; Khan, S.A. Protein misfolding and aggregation in neurodegenerative diseases: A review of pathogeneses, novel detection strategies, and potential therapeutics. Rev. Neurosci. 2019, 30, 339–358.
- Zhou, G.; Wilson, G.; Hebbard, L.; Duan, W.; Liddle, C.; George, J.; Qiao, L. Aptamers: A promising chemical antibody for cancer therapy. Oncotarget 2016, 7, 13446.
- Roesler, T.W.; Marvian, A.T.; Brendel, M.; Nykaenen, N.-P.; Hoellerhage, M.; Schwarz, S.C.; Hopfner, F.; Koeglsperger, T.; Respondek, G.; Schweyer, K. Four-repeat tauopathies. Prog. Neurobiol. 2019, 180, 101644.
- Ma, J.; Gao, J.; Wang, J.; Xie, A. Prion-like mechanisms in Parkinson’s disease. Front. Neurosci. 2019, 13, 552.
- Hernández-Jiménez, M.; Abad-Santos, F.; Cotgreave, I.; Gallego, J.; Jilma, B.; Flores, A.; Jovin, T.G.; Vivancos, J.; Molina, C.A.; Montaner, J. APRIL: A double-blind, placebo-controlled, randomized, Phase Ib/IIa clinical study of ApTOLL for the treatment of acute ischemic stroke. Front. Neurol. 2023, 14, 1127585.
- Yun, Y.; Yang, X.; Tan, S.; Wang, P.; Ji, Y.; Sun, X. Targeting upregulated RNA binding protein RCAN1. 1: A promising strategy for neuroprotection in acute ischemic stroke. CNS Neurosci. Ther. 2022, 28, 1814–1828.
- Zacco, E.; Kantelberg, O.; Milanetti, E.; Armaos, A.; Panei, F.P.; Gregory, J.; Jeacock, K.; Clarke, D.J.; Chandran, S.; Ruocco, G. Probing TDP-43 condensation using an in silico designed aptamer. Nat. Commun. 2022, 13, 3306.
- Liu, Y.; Sun, Q.-A.; Chen, Q.; Lee, T.H.; Huang, Y.; Wetsel, W.C.; Michelotti, G.A.; Sullenger, B.A.; Zhang, X. Targeting inhibition of GluR1 Ser845 phosphorylation with an RNA aptamer that blocks AMPA receptor trafficking. J. Neurochem. 2009, 108, 147–157.
- Lennarz, S.; Alich, T.C.; Kelly, T.; Blind, M.; Beck, H.; Mayer, G. Selective Aptamer-Based Control of Intraneuronal Signaling. Angew. Chem. 2015, 127, 5459–5463.
- Kahsai, A.W.; Wisler, J.W.; Lee, J.; Ahn, S.; Cahill III, T.J.; Dennison, S.M.; Staus, D.P.; Thomsen, A.R.; Anasti, K.M.; Pani, B. Conformationally selective RNA aptamers allosterically modulate the β2-adrenoceptor. Nat. Chem. Biol. 2016, 12, 709–716.
- Teng, I.-T.; Li, X.; Yadikar, H.A.; Yang, Z.; Li, L.; Lyu, Y.; Pan, X.; Wang, K.K.; Tan, W. Identification and characterization of DNA aptamers specific for phosphorylation epitopes of tau protein. J. Am. Chem. Soc. 2018, 140, 14314–14323.
- Kim, J.H.; Kim, E.; Choi, W.H.; Lee, J.; Lee, J.H.; Lee, H.; Kim, D.-E.; Suh, Y.H.; Lee, M.J. Inhibitory RNA aptamers of tau oligomerization and their neuroprotective roles against proteotoxic stress. Mol. Pharm. 2016, 13, 2039–2048.
- Lisi, S.; Fiore, E.; Scarano, S.; Pascale, E.; Boehman, Y.; Ducongé, F.; Chierici, S.; Minunni, M.; Peyrin, E.; Ravelet, C. Non-SELEX isolation of DNA aptamers for the homogeneous-phase fluorescence anisotropy sensing of tau Proteins. Anal. Chim. Acta 2018, 1038, 173–181.
- Ylera, F.; Lurz, R.; Erdmann, V.A.; Fürste, J.P. Selection of RNA aptamers to the Alzheimer’s disease amyloid peptide. Biochem. Biophys. Res. Commun. 2002, 290, 1583–1588.
- Takahashi, T.; Tada, K.; Mihara, H. RNA aptamers selected against amyloid β-peptide (Aβ) inhibit the aggregation of Aβ. Mol. Biosyst. 2009, 5, 986–991.
- Song, G.; Shui, R.; Wang, D.; Fang, R.; Yuan, T.; Li, L.; Feng, J.; Gao, F.; Shen, Q.; Gong, J. Aptamer-conjugated graphene oxide-based surface assisted laser desorption ionization mass spectrometry for selective extraction and detection of Aβ1–42 in an Alzheimer’s disease SH-SY5 cell model. Front. Aging Neurosci. 2022, 14, 993281.
- Zheng, Y.; Wang, P.; Li, S.; Geng, X.; Zou, L.; Jin, M.; Zou, Q.; Wang, Q.; Yang, X.; Wang, K. Development of DNA aptamer as a β-amyloid aggregation inhibitor. ACS Appl. Bio Mater. 2020, 3, 8611–8618.
- Murakami, K.; Obata, Y.; Sekikawa, A.; Ueda, H.; Izuo, N.; Awano, T.; Takabe, K.; Shimizu, T.; Irie, K. An RNA aptamer with potent affinity for a toxic dimer of amyloid β42 has potential utility for histochemical studies of Alzheimer’s disease. J. Biol. Chem. 2020, 295, 4870–4880.
- Rahimi, F.; Murakami, K.; Summers, J.L.; Chen, C.-H.B.; Bitan, G. RNA aptamers generated against oligomeric Aβ40 recognize common amyloid aptatopes with low specificity but high sensitivity. PLoS ONE 2009, 4, e7694.
- Zheng, Y.; Qu, J.; Xue, F.; Zheng, Y.; Yang, B.; Chang, Y.; Yang, H.; Zhang, J. Novel DNA aptamers for Parkinson’s disease treatment inhibit α-synuclein aggregation and facilitate its degradation. Mol. Ther. Nucleic Acids 2018, 11, 228–242.
- Tsukakoshi, K.; Abe, K.; Sode, K.; Ikebukuro, K. Selection of DNA aptamers that recognize α-synuclein oligomers using a competitive screening method. Anal. Chem. 2012, 84, 5542–5547.
- Hmila, I.; Sudhakaran, I.P.; Ghanem, S.S.; Vaikath, N.N.; Poggiolini, I.; Abdesselem, H.; El-Agnaf, O.M. Inhibition of α-Synuclein Seeding-Dependent Aggregation by ssDNA Aptamers Specific to C-Terminally Truncated α-Synuclein Fibrils. ACS Chem. Neurosci. 2022, 13, 3330–3341.
- Rock, M.; Zouganelis, G.D.; de Andrade, A.F.B.; Drake, S.J.; Alexiou, A.; Albrakati, A.; Batiha, G.E.-S.; Illingworth, T.A. Development and validation of anti-human Alpha synuclein DNA aptamer using computer modelling techniques—An in silico study. J. Integr. Neurosci. 2022, 21, 5.
- Tsukakoshi, K.; Harada, R.; Sode, K.; Ikebukuro, K. Screening of DNA aptamer which binds to α-synuclein. Biotechnol. Lett. 2010, 32, 643–648.
- Lobanova, E.; Whiten, D.; Ruggeri, F.S.; Taylor, C.G.; Kouli, A.; Xia, Z.; Emin, D.; Zhang, Y.P.; Lam, J.Y.; Williams-Gray, C.H. Imaging protein aggregates in the serum and cerebrospinal fluid in Parkinson’s disease. Brain 2022, 145, 632–643.
- Mannironi, C.; Di Nardo, A.; Fruscoloni, P.; Tocchini-Valentini, G. In vitro selection of dopamine RNA ligands. Biochemistry 1997, 36, 9726–9734.
- Fernández, G.; Moraga, A.; Cuartero, M.I.; García-Culebras, A.; Peña-Martínez, C.; Pradillo, J.M.; Hernández-Jiménez, M.; Sacristán, S.; Ayuso, M.I.; Gonzalo-Gobernado, R. TLR4-binding DNA aptamers show a protective effect against acute stroke in animal models. Mol. Ther. 2018, 26, 2047–2059.
- Camorani, S.; Esposito, C.L.; Rienzo, A.; Catuogno, S.; Iaboni, M.; Condorelli, G.; De Franciscis, V.; Cerchia, L. Inhibition of receptor signaling and of glioblastoma-derived tumor growth by a novel PDGFRβ aptamer. Mol. Ther. 2014, 22, 828–841.
- Monaco, I.; Camorani, S.; Colecchia, D.; Locatelli, E.; Calandro, P.; Oudin, A.; Niclou, S.; Arra, C.; Chiariello, M.; Cerchia, L. Aptamer functionalization of nanosystems for glioblastoma targeting through the blood–brain barrier. J. Med. Chem. 2017, 60, 4510–4516.
- Rozenblum, G.T.; Kaufman, T.; Vitullo, A.D. Myelin basic protein and a multiple sclerosis-related MBP-peptide bind to oligonucleotides. Mol. Ther. Nucleic Acids 2014, 3, e192.
- Vorobjeva, M.A.; Krasitskaya, V.V.; Fokina, A.A.; Timoshenko, V.V.; Nevinsky, G.A.; Venyaminova, A.G.; Frank, L.A. RNA aptamer against autoantibodies associated with multiple sclerosis and bioluminescent detection probe on its basis. Anal. Chem. 2014, 86, 2590–2594.
- Wang, L.; Bing, T.; Liu, Y.; Zhang, N.; Shen, L.; Liu, X.; Wang, J.; Shangguan, D. Imaging of neurite network with an anti-L1CAM aptamer generated by neurite-SELEX. J. Am. Chem. Soc. 2018, 140, 18066–18073.
- Iida, M.; Mashima, T.; Yamaoki, Y.; So, M.; Nagata, T.; Katahira, M. The anti-prion RNA aptamer R12 disrupts the Alzheimer’s disease-related complex between prion and amyloid β. FEBS J. 2019, 286, 2355–2365.
- Ogasawara, D.; Hasegawa, H.; Kaneko, K.; Sode, K.; Ikebukuro, K. Screening of DNA aptamer against mouse prion protein by competitive selection. Prion 2007, 1, 248–254.
- Proske, D.; Gilch, S.; Wopfner, F.; Schätzl, H.M.; Winnacker, E.L.; Famulok, M. Prion-protein-specific aptamer reduces PrPSc formation. Chembiochem 2002, 3, 717–725.
- Matos, C.O.; Passos, Y.M.; do Amaral, M.J.; Macedo, B.; Tempone, M.H.; Bezerra, O.C.; Moraes, M.O.; Almeida, M.S.; Weber, G.; Missailidis, S. Liquid-liquid phase separation and fibrillation of the prion protein modulated by a high-affinity DNA aptamer. FASEB J. 2020, 34, 365–385.
- Mashima, T.; Lee, J.-H.; Kamatari, Y.O.; Hayashi, T.; Nagata, T.; Nishikawa, F.; Nishikawa, S.; Kinoshita, M.; Kuwata, K.; Katahira, M. Development and structural determination of an anti-PrPC aptamer that blocks pathological conformational conversion of prion protein. Sci. Rep. 2020, 10, 4934.
- Cheng, C.; Chen, Y.H.; Lennox, K.A.; Behlke, M.A.; Davidson, B.L. In vivo SELEX for Identification of Brain-penetrating Aptamers. Mol. Ther. Nucleic Acids 2013, 2, e67.
- Georges, J.; Qi, X.; Liu, X.; Zhou, Y.; Woolf, E.C.; Valeri, A.; Al-Atrache, Z.; Belykh, E.; Feuerstein, B.G.; Preul, M. Provision of rapid and specific ex vivo diagnosis of central nervous system lymphoma from rodent xenograft biopsies by a fluorescent aptamer. J. Neurosurg. 2020, 134, 1783–1790.
- Tang, J.; Huang, N.; Zhang, X.; Zhou, T.; Tan, Y.; Pi, J.; Pi, L.; Cheng, S.; Zheng, H.; Cheng, Y. Aptamer-conjugated PEGylated quantum dots targeting epidermal growth factor receptor variant III for fluorescence imaging of glioma. Int. J. Nanomed. 2017, 12, 3899.
- Yun, Y.; Zhang, Y.; Zhang, C.; Huang, L.; Tan, S.; Wang, P.; Vilariño-Gúell, C.; Song, W.; Sun, X. Regulator of calcineurin 1 is a novel RNA-binding protein to regulate neuronal apoptosis. Mol. Psychiatry 2021, 26, 1361–1375.
- McCann, H.; Stevens, C.H.; Cartwright, H.; Halliday, G.M. α-Synucleinopathy phenotypes. Park. Relat. Disord. 2014, 20, S62–S67.
- Ren, X.; Zhao, Y.; Xue, F.; Zheng, Y.; Huang, H.; Wang, W.; Chang, Y.; Yang, H.; Zhang, J. Exosomal DNA aptamer targeting α-synuclein aggregates reduced neuropathological deficits in a mouse Parkinson’s disease model. Mol. Ther. Nucleic Acids 2019, 17, 726–740.
- Shamili, F.H.; Alibolandi, M.; Rafatpanah, H.; Abnous, K.; Mahmoudi, M.; Kalantari, M.; Taghdisi, S.M.; Ramezani, M. Immunomodulatory properties of MSC-derived exosomes armed with high affinity aptamer toward mylein as a platform for reducing multiple sclerosis clinical score. J. Control Release 2019, 299, 149–164.
- Goldenberg, M.M. Multiple sclerosis review. Pharm. Ther. 2012, 37, 175.
- Qu, W.; Yuan, B.; Liu, J.; Liu, Q.; Zhang, X.; Cui, R.; Yang, W.; Li, B. Emerging role of AMPA receptor subunit GluA1 in synaptic plasticity: Implications for Alzheimer’s disease. Cell Prolif. 2021, 54, e12959.
- Kiefel, H.; Bondong, S.; Hazin, J.; Ridinger, J.; Schirmer, U.; Riedle, S.; Altevogt, P. L1CAM: A major driver for tumor cell invasion and motility. Cell Adh Migr. 2012, 6, 374–384.
- Shi, S.; Fu, W.; Lin, S.; Tian, T.; Li, S.; Shao, X.; Zhang, Y.; Zhang, T.; Tang, Z.; Zhou, Y. Targeted and effective glioblastoma therapy via aptamer-modified tetrahedral framework nucleic acid-paclitaxel nanoconjugates that can pass the blood brain barrier. Nanomed. Nanotechnol. Biol. Med. 2019, 21, 102061.
- Wang, F.; Zhou, Y.; Cheng, S.; Lou, J.; Zhang, X.; He, Q.; Huang, N.; Cheng, Y. Gint4.T-modified DNA tetrahedrons loaded with doxorubicin inhibits glioma cell proliferation by targeting PDGFRβ. Nanoscale Res. Lett. 2020, 15, 150.
- Sullenger, B.A.; Gallardo, H.F.; Ungers, G.E.; Gilboa, E. Overexpression of TAR sequences renders cells resistant to human immunodeficiency virus replication. Cell 1990, 63, 601–608.
- Ng, E.W.M.; Shima, D.T.; Calias, P.; Cunningham, E.T.; Guyer, D.R.; Adamis, A.P. Pegaptanib, a targeted anti-VEGF aptamer for ocular vascular disease. Nat. Rev. Drug Discov. 2006, 5, 123–132.
- Hughes, Q.W.; Le, B.T.; Gilmore, G.; Baker, R.I.; Veedu, R.N. Construction of a Bivalent Thrombin Binding Aptamer and Its Antidote with Improved Properties. Molecules 2017, 22, 1770.
- Yazdian-Robati, R.; Bayat, P.; Oroojalian, F.; Zargari, M.; Ramezani, M.; Taghdisi, S.M.; Abnous, K. Therapeutic applications of AS1411 aptamer, an update review. Int. J. Biol. Macromol. 2020, 155, 1420–1431.
- Yamagishi, S.-i.; Taguchi, K.; Fukami, K. DNA-aptamers raised against AGEs as a blocker of various aging-related disorders. Glycoconj. J. 2016, 33, 683–690.
- Helmling, S.; Maasch, C.; Eulberg, D.; Buchner, K.; Schröder, W.; Lange, C.; Vonhoff, S.; Wlotzka, B.; Tschöp, M.H.; Rosewicz, S. Inhibition of ghrelin action in vitro and in vivo by an RNA-Spiegelmer. Proc. Natl. Acad. Sci. USA 2004, 101, 13174–13179.
- Vater, A.; Sell, S.; Kaczmarek, P.; Maasch, C.; Buchner, K.; Pruszynska-Oszmalek, E.; Kolodziejski, P.; Purschke, W.G.; Nowak, K.W.; Strowski, M.Z. A mixed mirror-image DNA/RNA aptamer inhibits glucagon and acutely improves glucose tolerance in models of type 1 and type 2 diabetes. J. Biol. Chem. 2013, 288, 21136–21147.
- Wang, P.; Yang, Y.; Hong, H.; Zhang, Y.; Cai, W.; Fang, D. Aptamers as therapeutics in cardiovascular diseases. Curr. Med. Chem. 2011, 18, 4169–4174.
- Maier, K.E.; Jangra, R.K.; Shieh, K.R.; Cureton, D.K.; Xiao, H.; Snapp, E.L.; Whelan, S.P.; Chandran, K.; Levy, M. A new transferrin receptor aptamer inhibits new world hemorrhagic fever mammarenavirus entry. Mol. Ther. Nucleic Acids 2016, 5, e321.
- Li, W.; Feng, X.; Yan, X.; Liu, K.; Deng, L. A DNA Aptamer Against Influenza A Virus: An Effective Inhibitor to the Hemagglutinin–Glycan Interactions. Nucleic Acid Ther. 2016, 26, 166–172.
- Soule, E.E.; Bompiani, K.M.; Woodruff, R.S.; Sullenger, B.A. Targeting two coagulation cascade proteases with a bivalent aptamer yields a potent and antidote-controllable anticoagulant. Nucleic Acid Ther. 2016, 26, 1–9.
- Jin, L.; Nonaka, Y.; Miyakawa, S.; Fujiwara, M.; Nakamura, Y. Dual therapeutic action of a neutralizing anti-FGF2 aptamer in bone disease and bone cancer pain. Mol. Ther. 2016, 24, 1974–1986.
- Li, W.; Lan, X. Aptamer oligonucleotides: Novel potential therapeutic agents in autoimmune disease. Nucleic Acid Ther. 2015, 25, 173–179.
- Zhu, G.; Ye, M.; Donovan, M.J.; Song, E.; Zhao, Z.; Tan, W. Nucleic acid aptamers: An emerging frontier in cancer therapy. Chem. Commun. 2012, 48, 10472–10480.
- Xing, H.; Hwang, K.; Li, J.; Torabi, S.-F.; Lu, Y. DNA aptamer technology for personalized medicine. Curr. Opin. Chem. Eng. 2014, 4, 79–87.
- Ballabh, P.; Braun, A.; Nedergaard, M. The blood–brain barrier: An overview: Structure, regulation, and clinical implications. Neurobiol. Dis. 2004, 16, 1–13.
- Abbott, N.J.; Patabendige, A.A.; Dolman, D.E.; Yusof, S.R.; Begley, D.J. Structure and function of the blood–brain barrier. Neurobiol. Dis. 2010, 37, 13–25.
- Gabathuler, R. Approaches to transport therapeutic drugs across the blood–brain barrier to treat brain diseases. Neurobiol. Dis. 2010, 37, 48–57.
- Wang, R.E.; Wu, H.; Niu, Y.; Cai, J. Improving the stability of aptamers by chemical modification. Curr. Med. Chem. 2011, 18, 4126–4138.
- Kariolis, M.S.; Wells, R.C.; Getz, J.A.; Kwan, W.; Mahon, C.S.; Tong, R.; Kim, D.J.; Srivastava, A.; Bedard, C.; Henne, K.R. Brain delivery of therapeutic proteins using an Fc fragment blood-brain barrier transport vehicle in mice and monkeys. Sci. Transl. Med. 2020, 12, eaay1359.
- Yu, Y.J.; Zhang, Y.; Kenrick, M.; Hoyte, K.; Luk, W.; Lu, Y.; Atwal, J.; Elliott, J.M.; Prabhu, S.; Watts, R.J. Boosting brain uptake of a therapeutic antibody by reducing its affinity for a transcytosis target. Sci. Transl. Med. 2011, 3, ra44–ra84.
- Johnsen, K.B.; Burkhart, A.; Thomsen, L.B.; Andresen, T.L.; Moos, T. Targeting the transferrin receptor for brain drug delivery. Prog. Neurobiol. 2019, 181, 101665.
- Chen, C.-h.B.; Dellamaggiore, K.R.; Ouellette, C.P.; Sedano, C.D.; Lizadjohry, M.; Chernis, G.A.; Gonzales, M.; Baltasar, F.E.; Fan, A.L.; Myerowitz, R. Aptamer-based endocytosis of a lysosomal enzyme. Proc. Natl. Acad. Sci. USA 2008, 105, 15908–15913.
- Macdonald, J.; Houghton, P.; Xiang, D.; Duan, W.; Shigdar, S. Truncation and mutation of a transferrin receptor aptamer enhances binding affinity. Nucleic Acid. Ther. 2016, 26, 348–354.
- Macdonald, J.; Denoyer, D.; Henri, J.; Jamieson, A.; Burvenich, I.J.; Pouliot, N.; Shigdar, S. Bifunctional aptamer–doxorubicin conjugate crosses the blood–brain barrier and selectively delivers its payload to EpCAM-positive tumor cells. Nucleic Acid. Ther. 2020, 30, 117–128.
- Wilner, S.E.; Wengerter, B.; Maier, K.; Magalhães, M.d.L.B.; Del Amo, D.S.; Pai, S.; Opazo, F.; Rizzoli, S.O.; Yan, A.; Levy, M. An RNA alternative to human transferrin: A new tool for targeting human cells. Mol. Ther. Nucleic Acids 2012, 1, e21.
- Wu, X.; Zhao, Z.; Bai, H.; Fu, T.; Yang, C.; Hu, X.; Liu, Q.; Champanhac, C.; Teng, I.-T.; Ye, M. DNA aptamer selected against pancreatic ductal adenocarcinoma for in vivo imaging and clinical tissue recognition. Theranostics 2015, 5, 985.
- Wu, X.; Liu, H.; Han, D.; Peng, B.; Zhang, H.; Zhang, L.; Li, J.; Liu, J.; Cui, C.; Fang, S. Elucidation and structural modeling of CD71 as a molecular target for cell-specific aptamer binding. J. Am. Chem. Soc. 2019, 141, 10760–10769.
- Zhang, N.; Wang, J.; Bing, T.; Liu, X.; Shangguan, D. Transferrin receptor-mediated internalization and intracellular fate of conjugates of a DNA aptamer. Mol. Ther. Nucleic Acids 2022, 27, 1249–1259.





