Aptamers developed using in vitro Systematic Evolution of Ligands by Exponential Enrichment (SELEX) technology are single-stranded nucleic acids 10–100 nucleotides in length. Their targets, often with specificity and high affinity, range from ions and small molecules to proteins and other biological molecules as well as larger systems, including cells, tissues, and animals. Aptamers often rival conventional antibodies with improved performance, due to aptamers’ unique biophysical and biochemical properties, including small size, synthetic accessibility, facile modification, low production cost, and low immunogenicity. Therefore, there is sustained interest in engineering and adapting aptamers for many applications, including diagnostics and therapeutics.
- aptamers
- neuro-diagnostics development
- neuro-medicine
- neuroscience
1. Introduction
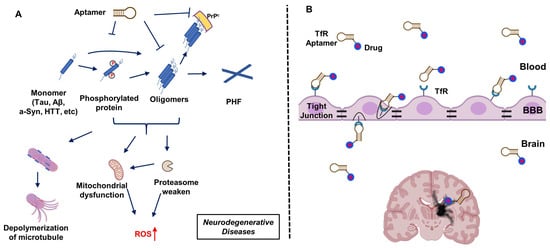
2. Aptamers Targeting Neuro-Medically Relevant Targets
| Target | Aptamers (Name) | Dissociation Constant |
Disease | Utility, Key Results | Ref. | ||
|---|---|---|---|---|---|---|---|
| GluR1 Ser845 | A1, A2, A3 (RNA) | 28–57 nM | Protein phosphorylation-related diseases | A2 binds GluR1 that inhibits GluR1/GluR1 containing AMPA receptor trafficking to the cell surface and abrogates forskolin-stimulated phosphorylation at GluR1 Ser845 | [28][78] | ||
| (MAPK) Erk1/2 | C5 (RNA) | 10 nM | [CNS disorders, Alzheimer’s disease, Stroke, Epilepsy | C5 selective to inhibit the mitogen-activated kinase pathway in neurons | 156][29][79] | ||
| β2-adrenoceptor (β2-AR) (GPCR) | A1, A2, A13, and A16 (RNA) | ||||||
| Human transferrin receptor specific C2. targets the apical domain of the human transferrin receptor (hTfR) Sequence (5′-3′): CAUCUCACAGAUCAAUCCAAGGCACCUCGUUAAAGGACGACUCCCUUACAUGCGAGAUG |
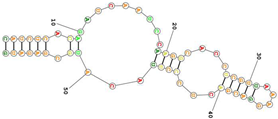 | 30.4–258.5 nM | [94- | ]RNA aptamers as allosteric GPCR modulators | [158][30][80] | ||
| Tau protein | IT(1–6), IT2a (DNA) | 5.5–68 nM | Traumatic brain injury, Alzheimer’s disease | Aptamers inhibit tau phosphorylation and oligomerization | |||
| Aptamer name: Min2/Waz (RNA) non-competitive for Transferrin. Targets the apical domain of the human transferrin receptor (hTfR) Sequence (5′-3′): GGGUUCUACGAUAAACGGUUAAUGACCAGCUUAUGGCUGGCAGUUCCC | [ | 31 | 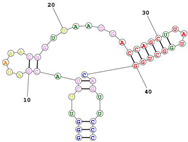 |
[77][141] | ][81] | ||
| Tau-1 (RNA) | ~200 nM | [ | |||||
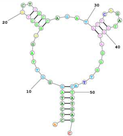
| Human TfR (cell-SELEX) (XQ-2d Shares a Similar Binding Site on CD71 with Transferrin) Aptamer name: XQ-2d (DNA) Sequence (5′-3′): ACTCATAGGGTTAGGGGCTGCTGGCCAGATACTCAGATGGTAGGGTTACTATGAGC | 32 | ][82] | |||||
 |
[ | 95][96][159,160] | Aptamer 314 (DNA) | 13 ± 3 nM | Aptamer binds Tau protein | [33][83] | |
| β-Amyloid protein |
β aptamers, e.g., β55 (RNA) | 29–48 nM | Alzheimer’s disease | β55 aptamer binds amyloid plaques in AD brain tissue | [34][84] | ||
| E1, E2, N1, N2, G2, etc. (RNA) | 10.9–21.6 μM | N2 aptamer is used to build a luminescent aptamer-ruthenium complex system for the detection of Aβ | [35][85] | ||||
| Apt-GO (RNA) | 0.1–10 μM | Apt-GO selectively detects Aβ1–42 in an AD SH-SY5 cell model | [36][86] | ||||
| Human TfR (cell-SELEX) Aptamer name: HG1-9 (DNA) HG1-9 aptamer binds human TfR with affinity (Kd < 20 nM) and completes a same bind site of human TfR with transferrin. Sequence (5′-3′): GGATAGGGATTCTGTTGGTCGGCTGGTTGGTATCC |
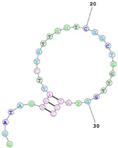 |
[97][161 | Aβ42 | Aβ7-92-1H1 | 63.4 nM | Inhibits Aβ42 aggregation | [37][87] |
| Aβ42 dimer | E22P–AbD43 (RNA) | 20 ± 6.0 nM | Aptamer inhibits the nucleation phase of the dimer and its associated neurotoxicity in SH-SY5Y human neuroblastoma cells. | [38][88] | |||
| Aβ40 oligomers | KM (RNA) | - | Aptamers bind with Aβ40 fibrils that may serve as amyloid recognition tools | [39][89] | |||
| α-synuclein protein | F5R1 (DNA) | 2.40 nM | Parkinson’s disease | Blocks the aberrant cellular effects of the overexpressed α-synuclein in cells | [40][90] | ||
| ] | T-SO508 (DNA) | 68 nM | T-SO508 can bind to soluble α-synuclein oligomers | [41][91] | |||
| Apt11(DNA) | - | Parkinson’s disease and dementia with Lewy bodies | Apt11 aptamer binds to α-syn fibrils and inhibits α-syn aggregation in the in vitro model of PD and DLB. | [42][92] | |||
| TMG-79 (DNA) | - | TMG-79 aptamer detects α-syn in Lewy body and PD-associated dementia. | [43][93] | ||||
| M5-15 (DNA) | 14.3 nM | M5-15 aptamer detects α-syn in Lewy body and PD-associated dementia. | [44][94] | ||||
| α-synuclein & amyloid-β (Aβ) | AD-PAINT (DNA) | 500 nM–2 μM | Parkinson’s disease | AD-PAINT aptamer binds to fibrillar aggregates of α-syn and Aβ aggregates detected in both serum and CSF in PD | [45][95] | ||
| Dopamine | dopa2 (129 nucleotides); dopa2/c.1 |
2.8 µM 1.6 µM |
Parkinson’s disease | Dopa2 and dopa2/c.1 are characterized to bind a dopamine affinity column; the dopamine binding site is obtained by secondary selection | [46][96] | ||
| Toll-like receptor 4 (TLR4) | ApTLR#1R, ApTLR#4F (DNA) | - | Stroke disease | Aptamers have a TLR4 antagonistic effect | [24][74] | ||
| ApTOLL (DNA) | 20 nM | Acute ischemic stroke | ApTOLL aptamer binds and antagonizes TLR4 and improves functional outcomes in AIS patients | [25][75] | |||
| Platelet-derived growth factor receptor β (PDGFRβ) | Gint4.T aptamer (RNA) | 9.6 nM | Glioma | Aptamer binds to human DGFRβ ectodomain, causing a strong inhibition of ligand-dependent receptor activation | [47][48][97,98] | ||
| Myelin basic protein | MBPcl3 MBPcl9 (DNA) |
- | Multiple Sclerosis | MBPc13 detects myelin-rich regions in paraffin-embedded mouse brain tissue; aptamer was found more sensitive than a commercial antibody. MBPcl3 blocks the binding of the antibody to MBP | [49][99] | ||
| Myelin basic protein (MBP) autoantibody | Apt2-9c (RNA) | 1.2 ± 0.1 nM | Multiple Sclerosis | Apt2-9c provides proof-of-principle for the detection of MS-specific autoantibodies | [50][100] | ||
| L1-CAM (Neurites) | yly12 (DNA) | 3.51 nM | Neurite-surface targeting and inhibitory effect on neurite outgrowth between cells | [51][101] | |||
| Prion protein (PrP) | R12 (RNA) | ~10 nM | Creutzfeldt-Jakob disease; prion diseases | R12 binding to PrP results in the dissociation of PrP with Aβ. | [52][102] | ||
| clone 4–9 (DNA) | 113 nM | binds to PrP | [53][103] | ||||
| DP7 (RNA) | 0.1–1.7 µM | Prion-protein-specific aptamer reduces PrPSc formation | [54][104] | ||||
| A1 (DNA) | 232 nM | Aptamers modulate phase separation and promote PrP fibrillation | [55][105] | ||||
| R24 (RNA) | 18 nM | R24 exhibited the lowest recorded IC50 and the highest anti-prion activity | [56][106] | ||||
| Crossing BBB (target unknown) | A15 (RNA) | - | Neurological disorders or diseases. | In vivo SELEX (brain-penetrating aptamers) | [57][31] | ||
| CD20+B cells | TD05 (DNA) | 256 nM | Glioma | TD05-488 can diagnose CNS lymphoma within 11 min of biopsy from xenograft brain tumor models | [58][107] | ||
| U87 glioma cell line/EGFRvIII | QD-A32 (DNA) | - | Glioma | QD-apt can penetrate the BBB and then selectively accumulate in the tumors through binding to EGFRvIII | [59][108] | ||
| The Regulator of calcineurin 1 (RCAN1) | R1SR13 (RNA) | 0.3 µM | Down syndrome and Alzheimer’s disease | Inhibits the regulatory function of RCAN1 in NFAT and NF-kB signaling pathways | [60][109] | ||
| 0.23–30 nM | Acute ischemic stroke | R1SR13 aptamer alleviates the RCAN1.1 L-induced neuronal apoptosis in the human SHSY-5Y neuroblastoma cells and in the mouse model of AIS | [26][76] | ||||
| TAR-DNA-Binding Protein 43 (TDP-43) | Apt-1 (DNA) | 1 μM | Amyotrophic Lateral Sclerosis | Apt-1 aptamer binds to TDP-43 in the ALS model. | [27][77] |
3. Applying Experience from Non-CNS Therapeutic Aptamers towards Neuro-Aptamer Development
The potential for using aptamers in therapy was recognized in 1990 when Tuerk and Gold selected an RNA aptamer to target bacteriophage T4 DNA polymerase in the first SELEX experiment [2]. In the same year, Sullenger et al. reported the inhibitory effect of transactivation response (TAR)-containing sequences (TAR decoys) on HIV-1 viral infection in host cells. These TAR decoys (which perform as if they were aptamers) prevent the Tar protein from binding to the endogenous TAR RNA, inhibiting HIV gene expression and replication. TAR decoy RNA-mediated HIV inhibition was also suggested to be effective against natural HIV isolates despite their hypervariable nature because the replication of SIVmac was also inhibited in cells expressing HIV-1 TAR decoys [69][133]. These two independent discoveries suggested that specific nucleic acid sequences have potential for therapies in general. Since then, aptamers have been studied in preclinical and clinical tests with different strategies employed for therapeutic applications, including aptamers as antagonists or agonists, and targeting ligands conjugated onto the drug carriers. In 2004, the first aptamer therapeutic, Macugen (Pegaptanib) was approved by the FDA, which is presently the only aptamer approved by the US FDA. It is effective as an anti-angiogenic medicine to treat neovascular (wet) age-related macular degeneration (AMD) [70][134]. Therapeutic targets for aptamers to date include thrombin [71][135] and nucleolin [72][136]. In addition, aptamers have been used to treat aging-related disorders [73][137], obesity and diabetes mellitus [74][75][138,139], cardiovascular diseases [76][140], infectious diseases [77][78][141,142], blood coagulation [79][143], bone diseases [80][144], immunological diseases [81][145] and cancers [82][146]. Experience from developing therapeutic aptamers for this range of targets, and knowledge regarding aptamer technology derived from this experience [83][147], will likely accelerate the development of aptamers toward managing neurological or neurodegenerative disorders.4. Aptamers Targeting Transferrin Receptor 1 to Facilitate Drug Transport across the BBB
The BBB impedes the entry of blood-borne molecules and is necessary to maintain brain homeostasis. The BBB comprises endothelial cells joined by highly polarized tight junctions and supported by astrocytes and pericytes responsible for the isolation of the brain from peripheral circulation [84][85][148,149]. The highly restrictive nature of the BBB limits access to most bio-therapeutics, including antibodies, aptamers, and most (~98%) small-molecule drugs to the brain microenvironment [86][150]. Aptamers targeting CNS proteins may not cross the BBB unless they are combined with BBB-penetrating agents or unique loading structures, such as neuronal cell-derived exosomes, exploiting the TfR-based transcytosis, or some efficient nano-systems (such as tetrahedra or circular DNA structures and nanoparticles). Structures implementing soft or solid nanomaterials improve the serum stability brain retention times of aptamers compared with the aptamers alone. This may mitigate the inherent drawbacks of rapid nuclease degradation and rapid renal clearance. This strategy may improve therapeutic efficacies [87][151]. Transferrin receptor 1, which is abundant in endothelial cells on the neurovascular material that forms the BBB, is considered a promising target for CNS delivery across the BBB. This has generated interest in developing antibodies [88][152] that bind TfR for CNS delivery [89][90][153,154]. Yu et al. reported the development of a low-affinity monoclonal antibody that binds TfR and therefore can cross the BBB to enhance the delivery of conjugated therapeutic antibodies that bind enzyme β-secretase (BACE1), a target for Alzheimer’s drugs. Several TfR DNA and RNA aptamers are now known. Thus, Neufeld et al. first developed the GS24 DNA aptamer (50 nucleotides long [91][155]) for its ability to bind mouse transferrin receptor 1 (TfR1) as shown in Table 23. GS24 was subsequentially truncated and mutated by MacDonald et al. to give a short aptamer, TfRA1, only 14 nucleotides in length [92][156]. This team then further exploited a bispecific strategy that conjugated TfRA1 with the epithelial cell adhesion molecule (EpCAM) to treat brain cancer metastases [93][157].| Transferrin Aptamer Name, Nucleotide Sequence |
2-D Structure | Ref. |
|---|---|---|
| Mouse transferrin receptor-specific GS24, reduced to 50 nucleotides. Sequence (5′-3′): GCGTGTGCACACGGTCACAGTTAGTATCGCTACGTTCTTTGGTAGTCCGTTCGG |
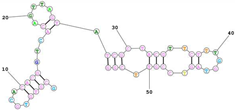 |
[91][155] |
| Target Mouse TfR Aptamer name: TfRA1 Truncated GS24; 14 nucleotides Sequence (5′-3′): GCGTGTGCACACGC |
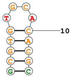 |
[92] |
