
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Danila Coradini | -- | 2938 | 2024-02-28 10:44:19 | | | |
| 2 | Fanny Huang | Meta information modification | 2938 | 2024-02-29 04:42:47 | | |
Video Upload Options
Cholesterol (CHOL) is a multifaceted lipid molecule. It is an essential structural component of cell membranes, where it cooperates in regulating the intracellular trafficking and signaling pathways. Additionally, it serves as a precursor for vital biomolecules, including steroid hormones, isoprenoids, vitamin D, and bile acids. Although CHOL is normally uptaken from the bloodstream, cells can synthesize it de novo in response to an increased requirement due to physiological tissue remodeling or abnormal proliferation, such as in cancer. Cumulating evidence indicated that increased CHOL biosynthesis is a common feature of breast cancer and is associated with the neoplastic transformation of normal mammary epithelial cells.
1. Introduction
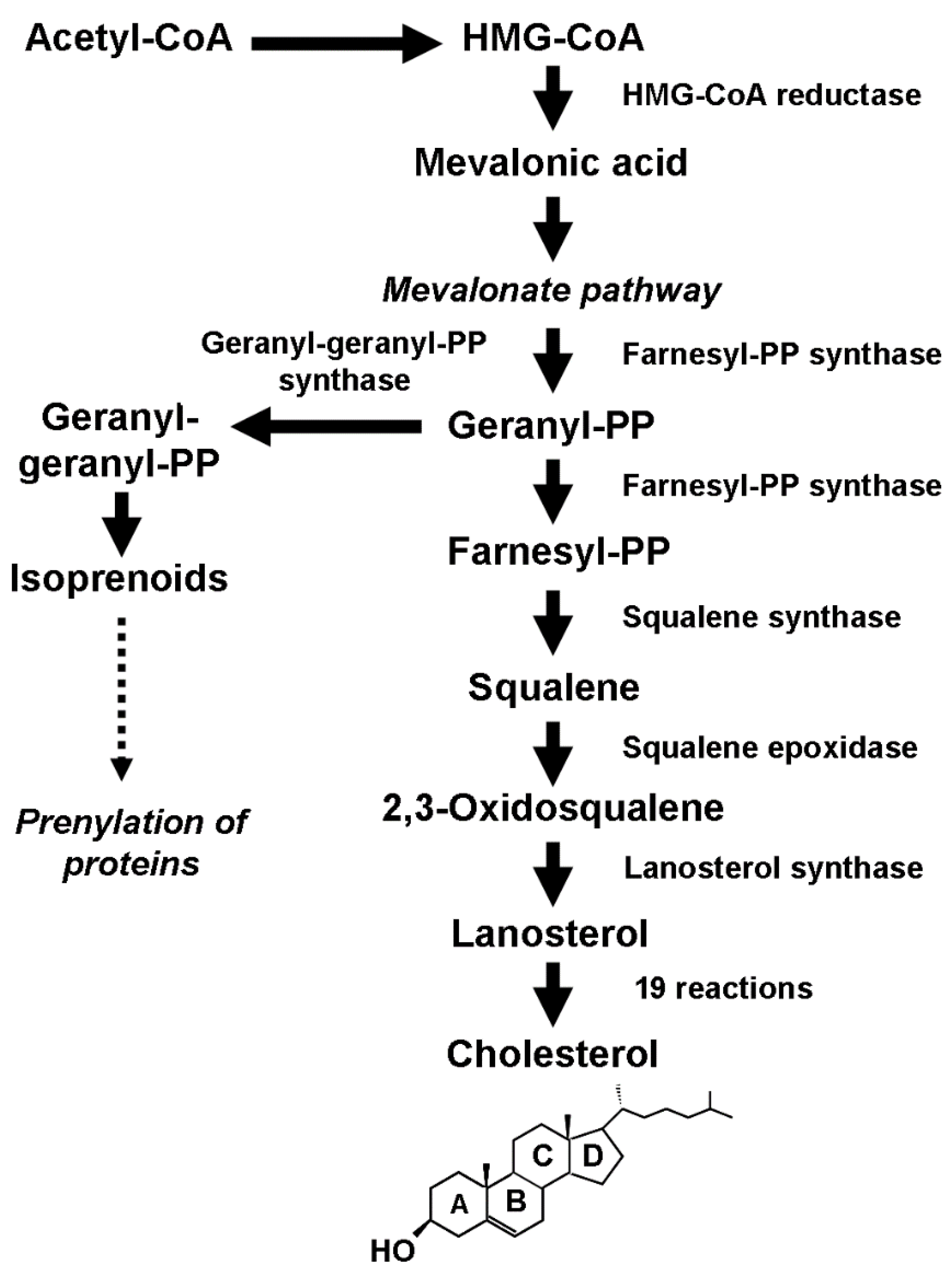
2. Cholesterol Biosynthesis and Breast Cancer Initiation and Development
2.1. Cholesterol Biosynthesis and the Hippo Signaling Pathway
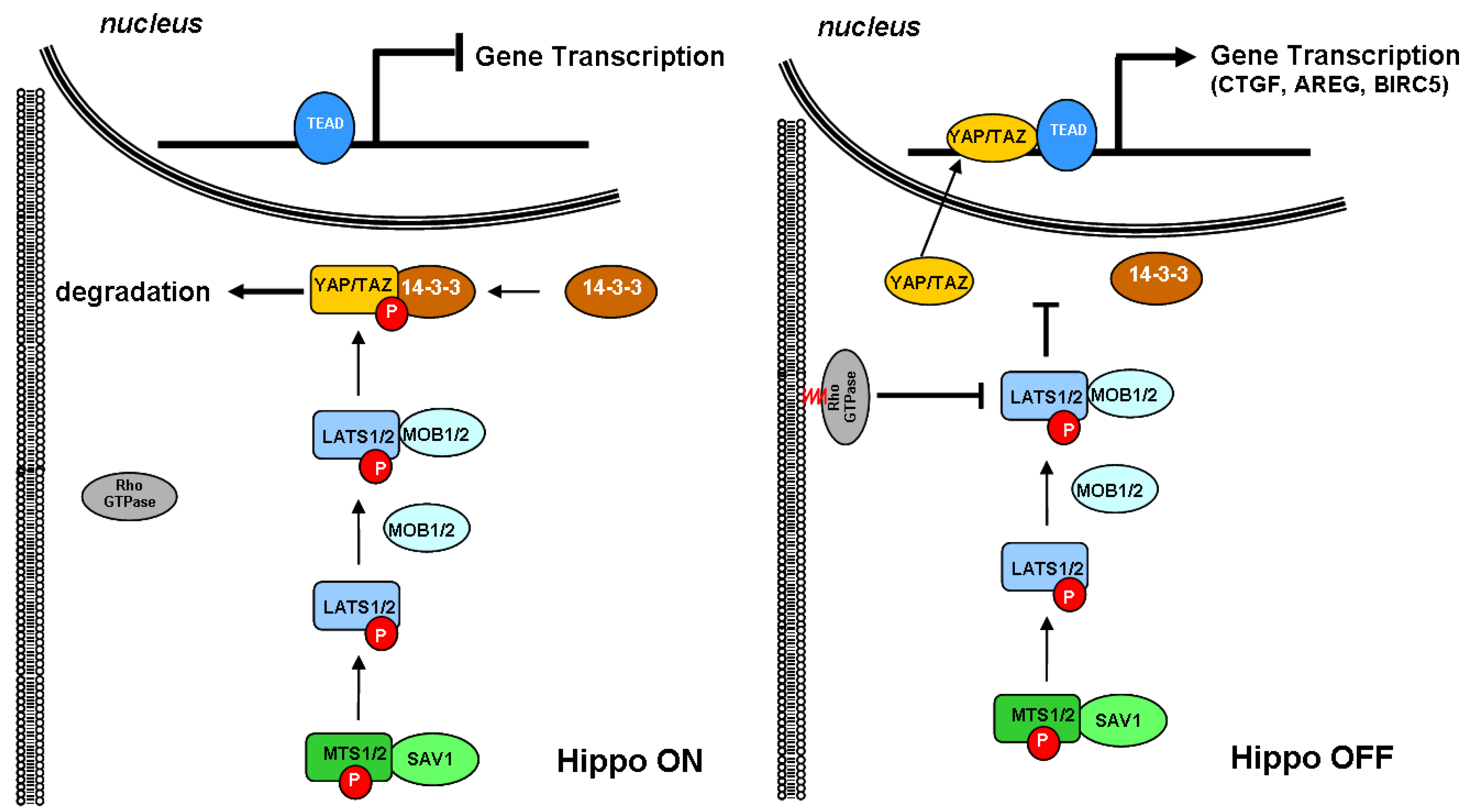
2.2. Cholesterol Biosynthesis, Mammary Stem Cells, and Breast Cancer Stem Cells
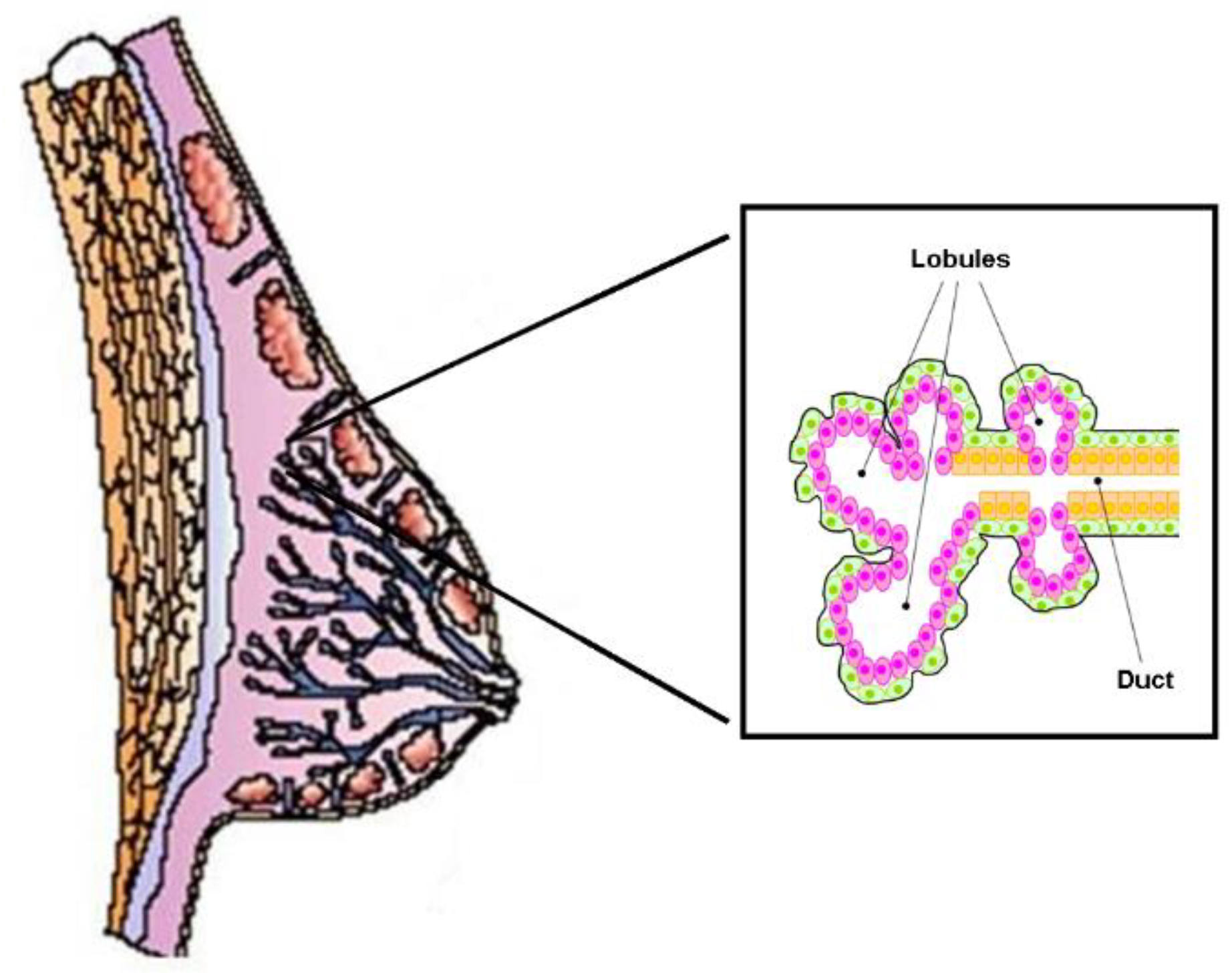
2.3. Cholesterol Biosynthesis and Ductal Carcinoma In Situ
2.4. Cholesterol Biosynthesis and Invasive Breast Cancer
References
- Luo, J.; Yang, H.; Song, B.L. Mechanisms and Regulation of Cholesterol Homeostasis. Nat. Rev. Mol. Cell Biol. 2020, 21, 225–245.
- Sugii, S.; Reid, P.C.; Ohgami, N.; Du, H.; Chang, T.Y. Distinct Endosomal Compartments in Early Trafficking of Low Density Lipoprotein-Derived Cholesterol. J. Biol. Chem. 2003, 278, 27180–27189.
- Chang, T.Y.; Chang, C.C.; Ohgami, N.; Yamauchi, Y. Cholesterol Sensing, Trafficking, and Esterification. Annu. Rev. Cell Dev. Biol. 2006, 22, 129–157.
- Gao, Y.; Zhou, Y.; Goldstein, J.L.; Brown, M.S.; Radhakrishnan, A. Cholesterol-Induced Conformational Changes in the Sterol-Sensing Domain of the Scap Protein Suggest Feedback Mechanism to Control Cholesterol Synthesis. J. Biol. Chem. 2017, 292, 8729–8737.
- Sun, L.P.; Li, L.; Goldstein, J.L.; Brown, M.S. Insig Required for Sterol-Mediated Inhibition of Scap/SREBP Binding to COPII Proteins in vitro. J. Biol. Chem. 2005, 280, 26483–26490.
- Sever, N.; Yang, T.; Brown, M.S.; Goldstein, J.L.; DeBose-Boyd, R.A. Accelerated Degradation of HMG CoA Reductase Mediated by Binding of Insig-1 to Its Sterol-Sensing Domain. Mol. Cell. 2003, 11, 25–33.
- DeBose-Boyd, R.A. Feedback Regulation of Cholesterol Synthesis: Sterol-Accelerated Ubiquitination and Degradation of HMG CoA Reductase. Cell Res. 2008, 18, 609–621.
- Clark, L.T. Cholesterol and Heart Disease: Current Concepts in Pathogenesis and Treatment. J. Natl. Med. Assoc. 1986, 78, 743–751.
- Brown, A.J. Cholesterol, Statins and Cancer. Clin. Exp. Pharmacol. Physiol. 2007, 34, 135–141.
- Kuzu, O.F.; Noory, M.A.; Robertson, G.P. The Role of Cholesterol in Cancer. Cancer Res. 2016, 76, 2063–2070.
- Murdock, D.J.; Sanchez, R.J.; Mohammadi, K.A.; Fazio, S.; Geba, G.P. Serum Cholesterol and the Risk of Developing Hormonally Driven Cancers: A Narrative Review. Cancer Med. 2023, 12, 6722–6767.
- Li, C.; Yang, L.; Zhang, D.; Jiang, W. Systematic Review and Meta-Analysis Suggest that Dietary Cholesterol Intake Increases Risk of Breast Cancer. Nutr. Res. 2016, 36, 627–635.
- Johnson, K.E.; Siewert, K.M.; Klarin, D.; Damrauer, S.M.; Chang, K.M.; Tsao, P.S.; Assimes, T.L.; Maxwell, K.N.; Voight, B.F. The Relationship Between Circulating Lipids and Breast Cancer Risk: A Mendelian randomization study. PLoS Med. 2020, 17, e1003302.
- Touvier, M.; Fassier, P.; His, M.; Norat, T.; Chan, D.S.; Blacher, J.; Hercberg, S.; Galan, P.; Druesne-Pecollo, N.; Latino-Martel, P. Cholesterol and Breast Cancer Risk: A Systematic Review and Meta-Analysis of Prospective Studies. Br. J. Nutr. 2015, 114, 347–357.
- Baek, A.E.; Nelson, E.R. The Contribution of Cholesterol and Its Metabolites to the Pathophysiology of Breast Cancer. Horm. Cancer 2016, 7, 219–228.
- Ben Hassen, C.; Goupille, C.; Vigor, C.; Durand, T.; Guéraud, F.; Silvente-Poirot, S.; Poirot, M.; Frank, P.G. Is Cholesterol a Risk Factor for Breast Cancer Incidence and Outcome? J. Steroid Biochem. Mol. Biol. 2023, 232, 106346.
- Llaverias, G.; Danilo, C.; Mercier, I.; Daumer, K.; Capozza, F.; Williams, T.M.; Sotgia, F.; Lisanti, M.P.; Frank, P.G. Role of Cholesterol in the Development and Progression of Breast Cancer. Am. J. Pathol. 2011, 178, 402–412.
- Clendening, J.W.; Pandyra, A.; Boutros, P.C.; El Ghamrasni, S.; Khosravi, F.; Trentin, G.A.; Martirosyan, A.; Hakem, A.; Hakem, R.; Jurisica, I.; et al. Dysregulation of the Mevalonate Pathway Promotes Transformation. Proc. Natl. Acad. Sci. USA 2010, 107, 15051–15056.
- Zhao, B.; Tumaneng, K.; Guan, K.L. The Hippo Pathway in Organ Size Control, Tissue Regeneration and Stem Cell Self-Renewal. Nat. Cell Biol. 2011, 13, 877–883.
- Zhao, B.; Wei, X.; Li, W.; Udan, R.S.; Yang, Q.; Kim, J.; Xie, J.; Ikenoue, T.; Yu, J.; Li, L.; et al. Inactivation of YAP Oncoprotein by the Hippo Pathway Is Involved in Cell Contact Inhibition and Tissue Growth Control. Genes Dev. 2007, 21, 2747–2761.
- Yu, F.X.; Guan, K.L. The Hippo Pathway: Regulators and Regulations. Genes Dev. 2013, 27, 355–371.
- Etienne-Manneville, S.; Hall, A. Rho GTPases in Cell Biology. Nature 2002, 420, 629–635.
- Ridley, A.J.; Hall, A. The Small GTP-Binding Protein Rho Regulates the Assembly of Focal Adhesions and Actin Stress Fibers in Response to Growth Factors. Cell 1992, 70, 389–399.
- Jang, J.W.; Kim, M.K.; Bae, S.C. Reciprocal Regulation of YAP/TAZ by the Hippo Pathway and the Small GTPase Pathway. Small GTPases 2020, 11, 280–288.
- Zhao, B.; Li, L.; Lei, Q.; and Guan, K.L. The Hippo-YAP Pathway in Organ Size Control and Tumorigenesis: An Updated Version. Genes Dev. 2010, 24, 862–874.
- Steinhardt, A.A.; Gayyed, M.F.; Klein, A.P.; Dong, J.; Maitra, A.; Pan, D.; Montgomery, E.A.; Anders, R.A. Expression of Yes-associated Protein in Common Solid Tumors. Hum. Pathol. 2008, 39, 1582–1589.
- Stingl, J.; Eaves, C.J.; Kuusk, U.; Emerman, J.T. Phenotypic and Functional Characterization In Vitro of a Multipotent Epithelial Cell Present in the Normal Adult Human Breast. Differentiation 1998, 63, 201–213.
- Gudjonsson, T.; Villadsen, R.; Nielsen, H.L.; Rønnov-Jessen, L.; Bissell, M.J.; Petersen, O.W. Isolation, Immortalization, and Characterization of a Human Breast Epithelial Cell Line with Stem Cell Properties. Genes Dev. 2002, 16, 693–706.
- Fu, N.Y.; Nolan, E.; Lindeman, G.J.; Visvader, J.E. Stem Cells and the Differentiation Hierarchy in Mammary Gland Development. Physiol. Rev. 2020, 100, 489–523.
- Joshi, P.A.; Jackson, H.W.; Beristain, A.G.; Di Grappa, M.A.; Mote, P.A.; Clarke, C.L.; Stingl, J.; Waterhouse, P.D.; Khokha, R. Progesterone Induces Adult Mammary Stem Cell Expansion. Nature 2010, 465, 803–807.
- Honeth, G.; Schiavinotto, T.; Vaggi, F.; Marlow, R.; Kanno, T.; Shinomiya, I.; Lombardi, S.; Buchupalli, B.; Graham, R.; Gazinska, P.; et al. Models of Breast Morphogenesis Based on Localization of Stem Cells in the Developing Mammary Lobule. Stem Cell Rep. 2015, 4, 699–711.
- Dontu, G.; Abdallah, W.M.; Foley, J.M.; Jackson, K.W.; Clarke, M.F.; Kawamura, M.J.; Wicha, M.S. In Vitro Propagation and Transcriptional Profiling of Human Mammary Stem/Progenitor Cells. Genes Dev. 2003, 17, 1253–1270.
- Slepicka, P.F.; Somasundara, A.V.H.; Dos Santos, C.O. The Molecular Basis of Mammary Gland Development and Epithelial Differentiation. Semin. Cell Dev. Biol. 2021, 114, 93–112.
- Dontu, G.; El-Ashry, D.; Wicha, M.S. Breast Cancer, Stem/Progenitor Cells and the Estrogen Receptor. Trends Endocrinol. Metab. 2004, 15, 193–197.
- Ercan, C.; van Diest, P.J.; Vooijs, M. Mammary Development and Breast Cancer: The Role of Stem Cells. Curr. Mol. Med. 2011, 11, 270–285.
- Economopoulou, P.; Kaklamani, V.G.; Siziopikou, K. The Role of Cancer Stem Cells in Breast Cancer Initiation and Progression: Potential Cancer Stem Cell-directed Therapies. Oncologist 2012, 17, 1394–1401.
- Charafe-Jauffret, E.; Ginestier, C.; Bertucci, F.; Cabaud, O.; Wicinski, J.; Finetti, P.; Josselin, E.; Adelaide, J.; Nguyen, T.T.; Monville, F.; et al. ALDH1-positive Cancer Stem Cells Predict Engraftment of Primary Breast Tumors and Are Governed by a Common Stem Cell Program. Cancer Res. 2013, 73, 7290–7300.
- Ehmsen, S.; Pedersen, M.H.; Wang, G.; Terp, M.G.; Arslanagic, A.; Hood, B.L.; Conrads, T.P.; Leth-Larsen, R.; Ditzel, H.J. Increased Cholesterol Biosynthesis Is a Key Characteristic of Breast Cancer Stem Cells Influencing Patient Outcome. Cell Rep. 2019, 27, 3927–3938.e6.
- Ginestier, C.; Monville, F.; Wicinski, J.; Cabaud, O.; Cervera, N.; Josselin, E.; Finetti, P.; Guille, A.; Larderet, G.; Viens, P.; et al. Mevalonate Metabolism Regulates Basal Breast Cancer Stem Cells and Is a Potential Therapeutic Target. Stem Cells 2012, 30, 1327–1337.
- Walsh, C.A.; Akrap, N.; Garre, E.; Magnusson, Y.; Harrison, H.; Andersson, D.; Jonasson, E.; Rafnsdottir, S.; Choudhry, H.; Buffa, F.; et al. The Mevalonate Precursor Enzyme HMGCS1 Is a Novel Marker and Key Mediator of Cancer Stem Cell Enrichment in Luminal and Basal Models of Breast Cancer. PLoS ONE 2020, 15, e0236187.
- Liu, S.; Clouthier, S.G.; Wicha, M.S. Role of microRNAs in the Regulation of Breast Cancer Stem Cells. J. Mammary Gland Biol. Neoplasia 2012, 17, 15–21.
- Monchusi, B.; Kaur, M. miRNAs as Modulators of Cholesterol in Breast Cancer Stem Cells: An Approach to Overcome Drug Resistance in Cancer. Curr. Drug Targets 2022, 23, 656–677.
- Giaquinto, A.N.; Sung, H.; Miller, K.D.; Kramer, J.L.; Newman, L.A.; Minihan, A.; Jemal, A.; Siegel, R.L. Breast Cancer Statistics, 2022. CA Cancer J. Clin. 2022, 72, 524–541.
- Sanders, M.E.; Schuyler, P.A.; Simpson, J.F.; Page, D.L.; Dupont, W.D. Continued Observation of the Natural History of Low-grade Ductal Carcinoma in situ Reaffirms Proclivity for Local Recurrence Even after More than 30 Years of Follow-Up. Mod. Pathol. 2015, 28, 662–669.
- Schuetz, C.S.; Bonin, M.; Clare, S.E.; Nieselt, K.; Sotlar, K.; Walter, M.; Fehm, T.; Solomayer, E.; Riess, O.; Wallwiener, D.; et al. Progression-specific Genes Identified by Expression Profiling of Matched Ductal Carcinomas in situ and Invasive Breast Tumors, Combining Laser Capture Microdissection and Oligonucleotide Microarray Analysis. Cancer Res. 2006, 66, 5278–5286.
- Tamimi, R.M.; Baer, H.J.; Marotti, J.; Galan, M.; Galaburda, L.; Fu, Y.; Deitz, A.C.; Connolly, J.L.; Schnitt, S.J.; Colditz, G.A.; et al. Comparison of Molecular Phenotypes of Ductal Carcinoma in situ and Invasive Breast Cancer. Breast Cancer Res. 2008, 10, R67.
- Wiechmann, L.; Kuerer, H.M. The Molecular Journey from Ductal Carcinoma in situ to Invasive Breast Cancer. Cancer 2008, 112, 2130–2142.
- Coradini, D. Interaction of de novo Cholesterol Biosynthesis and Hippo Signaling Pathway in Ductal Carcinoma in situ (DCIS)—Comparison with the Corresponding Normal Breast Epithelium. Transl. Breast. Cancer Res. 2023, 4, 26.
- Butt, S.; Butt, T.; Jirström, K.; Hartman, L.; Amini, R.M.; Zhou, W.; Wärnberg, F.; Borgquist, S. The Target for Statins, HMG-CoA Reductase, is Expressed in Ductal Carcinoma in situ and May Predict Patient Response to Radiotherapy. Ann. Surg. Oncol. 2014, 21, 2911–2919.
- Gill, S.; Stevenson, J.; Kristiana, I.; Brown, A.J. Cholesterol-dependent Degradation of Squalene Monooxygenase, a Control Point in Cholesterol Synthesis Beyond HMG-CoA Reductase. Cell Metab. 2011, 13, 260–273.
- Göbel, A.; Riffel, R.M.; Hofbauer, L.C.; Rachner, T.D. The Mevalonate Pathway in Breast Cancer Biology. Cancer Lett. 2022, 542, 215761.
- Kim, H.Y.; Bae, S.J.; Choi, J.W.; Han, S.; Bae, S.H.; Cheong, J.H.; Jang, H. Cholesterol Synthesis Is Important for Breast Cancer Cell Tumor Sphere Formation and Invasion. Biomedicines 2022, 10, 1908.
- Maja, M.; Mohammed, D.; Dumitru, A.C.; Verstraeten, S.; Lingurski, M.; Mingeot-Leclercq, M.P.; Alsteens, D.; Tyteca, D. Surface Cholesterol-Enriched Domains Specifically Promote Invasion of Breast Cancer Cell Lines by Controlling Invadopodia and Extracellular Matrix Degradation. Cell Mol. Life Sci. 2022, 79, 417.
- Tang, Q.; Liang, B.; Zhang, L.; Li, X.; Li, H.; Jing, W.; Jiang, Y.; Zhou, F.; Zhang, J.; Meng, Y.; et al. Enhanced Cholesterol Biosynthesis Promotes Breast Cancer Metastasis Via Modulating CCDC25 Expression and Neutrophil Extracellular Traps Formation. Sci. Rep. 2022, 12, 17350.
- Coradini, D.; Ambrogi, F.; Infante, G. Cholesterol de novo Biosynthesis in Paired Samples of Breast Cancer and Adjacent Histologically Normal Tissue: Association with Proliferation Index, Tumor Grade, and Recurrence-Free Survival. Arch. Breast Cancer 2023, 10, 187–199.
- Yoon, S.H.; Kim, H.S.; Kim, R.N.; Jung, S.Y.; Hong, B.S.; Kang, E.J.; Moon, H.G.; Noh, D.Y.; Han, W. NAD(P)-dependent Steroid Dehydrogenase-like is Involved in Breast Cancer Cell Growth and Metastasis. BMC Cancer 2020, 20, 375.
- Sotiriou, C.; Wirapati, P.; Loi, S.; Harris, A.; Fox, S.; Smeds, J.; Nordgren, H.; Farmer, P.; Praz, V.; Haibe-Kains, B.; et al. Gene Expression Profiling in Breast Cancer: Understanding the Molecular Basis of Histologic Grade to Improve Prognosis. J. Natl. Cancer Inst. 2006, 98, 262–272.
- Matikas, A.; Foukakis, T.; Swain, S.; Bergh, J. Avoiding over- and Undertreatment in Patients with Resected Node-Positive Breast Cancer with the Use of Gene Expression Signatures: Are We There Yet? Ann. Oncol. 2019, 30, 1044–1050.





