Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Yuefeng Chu | -- | 4038 | 2024-02-19 18:09:09 | | | |
| 2 | Lindsay Dong | Meta information modification | 4038 | 2024-02-20 02:26:23 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Liu, S.; Li, Z.; Lan, S.; Hao, H.; Baz, A.A.; Yan, X.; Gao, P.; Chen, S.; Chu, Y. ATF3 in Inflammation, Apoptosis, Ferroptosis, and Pathogen Infection. Encyclopedia. Available online: https://encyclopedia.pub/entry/55192 (accessed on 06 June 2026).
Liu S, Li Z, Lan S, Hao H, Baz AA, Yan X, et al. ATF3 in Inflammation, Apoptosis, Ferroptosis, and Pathogen Infection. Encyclopedia. Available at: https://encyclopedia.pub/entry/55192. Accessed June 06, 2026.
Liu, Shuang, Zhangcheng Li, Shimei Lan, Huafang Hao, Ahmed Adel Baz, Xinmin Yan, Pengcheng Gao, Shengli Chen, Yuefeng Chu. "ATF3 in Inflammation, Apoptosis, Ferroptosis, and Pathogen Infection" Encyclopedia, https://encyclopedia.pub/entry/55192 (accessed June 06, 2026).
Liu, S., Li, Z., Lan, S., Hao, H., Baz, A.A., Yan, X., Gao, P., Chen, S., & Chu, Y. (2024, February 19). ATF3 in Inflammation, Apoptosis, Ferroptosis, and Pathogen Infection. In Encyclopedia. https://encyclopedia.pub/entry/55192
Liu, Shuang, et al. "ATF3 in Inflammation, Apoptosis, Ferroptosis, and Pathogen Infection." Encyclopedia. Web. 19 February, 2024.
Copy Citation
Transcription factors are pivotal regulators in the cellular life process. Activating transcription factor 3 (ATF3), a member of the ATF/CREB (cAMP response element-binding protein) family, plays a crucial role as cells respond to various stresses and damage. As a transcription factor, ATF3 significantly influences signal transduction regulation, orchestrating a variety of signaling pathways, including apoptosis, ferroptosis, and cellular differentiation. In addition, ATF3 serves as an essential link between inflammation, oxidative stress, and immune responses.
ATF3
inflammatory
apoptosis
ferroptosis
microorganism
1. Introduction
Transcription factors (TFs) play a crucial role in the regulation of cellular function and the development of diseases due to their direct influence on gene expression, thus becoming a focal point of research in recent years. TFs regulate a variety of cellular physiological activities and immune responses by initiating or inhibiting the transcription of specific genes. Serving as key regulatory factors in the coordination of immune responses, TFs are essential for the activation of immune cells and their direct involvement in the production of inflammatory cytokines [1][2][3]. However, in the prolonged struggle between pathogens and hosts, some pathogenic microorganisms have evolved strategies to manipulate host TFs, thereby promoting their survival and replication within the host. Studies have elucidated that specific pathogenic organisms manipulate the activation of the NF-κB pathway as a survival strategy, which can effectively protect the host cell from apoptosis and allow bacterial survival within host cells [2]. Therefore, TFs are not only central to the host immune response but are also a vital strategy for some pathogens to establish infection. Increasing evidence suggests that TFs, such as STAT1, STAT2, JunB, CHOP, ATF3, and NF-κB, are involved in the interaction between host and pathogen and the regulation of host immune responses [2][4][5][6][7]. Thus, understanding the specific roles of TF in different infectious environments can provide insight for the development of targeted therapies to enhance host defense capability.
Activating transcription factor 3 (ATF3) is a stress-induced transcription factor that belongs to the activating transcription factor/cAMP response element-binding protein (ATF/CREB) family [8][9], whose members include ATF1, CREB, CREM, ATF2, ATF3, ATF4, ATF5, ATF6, ATF7, and B-ATF [10]. ATF3 regulates gene transcription by forming homodimers or heterodimers through its basic-region leucine zipper (bZIP) domain, thus modulating the biological functions of genes. ATF3 expression is relatively stable under normal physiological conditions, but changes in its expression are associated with various pathophysiological responses such as inflammation, oxidative stress, stress of the endoplasmic reticulum, and cell death [8][9][11]. ATF3 expression is induced by a variety of signals, including those initiated by cytokines, genotoxic agents, or physiological stressors [10]. ATF3 is upregulated under multiple stress conditions, regulating the interaction between cellular metabolism and immune and inflammatory responses, thus maintaining cellular homeostasis.
2. Mechanisms of ATF3 Induction under Stressful Physiological Conditions
ATF3 shares the same binding site, 5′-TGACGTCA-3′, found in other transcription factors of the ATF/CREB family [10]. They interact with target DNA by binding to the entire region within the bZIP domain [8][10]. Several genes have been identified to possess this recognition sequence, including Nrf2, JunD, c-Jun, and IL-6 [12][13]. Intriguingly, some promoters of target genes regulated by ATF3 have binding sequences that differ from these common motifs (Table S1). ATF3 expression is relatively stable under normal physiological conditions; however, it rapidly changes in response to disturbances in the internal environment or external stimuli [14]. Changes in ATF3 expression are induced by inflammatory responses, cell death, cytokines, and cellular stresses (oxidative stress, DNA damage, or endoplasmic reticulum stress (ERS)). The specific induction mechanism may vary depending on the type of stress but usually involves the activation of stress-responsive kinases and upstream transcription factors, which then bind to the ATF3 promoter and stimulate its transcription [15][16]. Due to its induction in response to various stress signals, ATF3 is considered a stress-induced gene [17][18][19], which participates in cellular growth, tissue remodeling, cytoskeletal reorganization, and inflammation [15]. NF-E2-related factor 2 (Nrf2) transcriptionally upregulates ATF3 expression in astrocytes, thus promoting antioxidant and cytoprotective functions [20]. Naringin (Nar) activates ATF3 and inhibits PINK1 transcription by suppressing ERS and mitochondrial autophagy-related genes, as well as the expression of downstream ERS proteins [21].
ATF3 interacts with the cAMP response element (CRE) sequences through its basic region and forms homodimers or heterodimers with other CREB family members through its bZip domain [15][22]. Heterodimers, such as those formed by ATF3 with C-jun, ATF2, and JunB, have been demonstrated to possess transcriptional activation capabilities, enhancing the transcriptional activity of downstream target genes [23][24][25]. Currently, it is widely accepted that ATF3 might occur through stabilizing inhibitory cofactors on the promoter, thereby suppressing the transcriptional activity of downstream genes [13][22]. ATF3 recruits histone deacetylase 1 (HDAC1) to promoters containing ATF3 binding sites. Subsequently, HDAC1 causes histone deacetylation, leading to chromatin condensation and transcriptional repression [13][26]. Histone acetylation is a crucial physiological process that opens the chromatin structure, allowing transcription factors to bind to gene promoters and activate transcription [13][27].
3. Implications of PAMPs and PRRs Activation on the Expression of ATF3
The Toll-like receptor (TLR) family consists of pattern recognition receptors (PRRs) involved in the detection of pathogen-associated molecular patterns (PAMPs) [10][28]. TLR1, 2, 4, 5, and 6 are expressed on the cell surface membrane and recognize bacterial and fungal products, while TLR3, 7, 8, and 9 are located in intracellular endosomes, specializing in the detection of pathogen-associated nucleic acids [10][28]. Upon recognition of their ligands, TLRs initiate complex cell signaling pathways, conferring antiviral and antibacterial states to the cell and promoting the expression of inflammatory cytokines, chemokines, and co-stimulatory molecules, which are crucial for the activation of adaptive immune responses [10][28]. Studies have shown that murine bone marrow macrophages treated with multiple PAMPs recognized by TLRs such as LPS, pIC, CpG-ODN, pIC/CpG-ODN, and zymosan significantly increase ATF3 protein expression [29], indicating that ATF3 expression is induced by various TLR responses.
4. Modulatory Role of ATF3 in Inflammatory Cytokine
As a pivotal hub in the network of cellular adaptive responses and a transcription factor regulating immune response genes, ATF3 is a key regulator of both local and systemic inflammation, aiding cells in responding to disruptions in internal homeostasis. This is due to its ability to positively or negatively modulate the functional status or bioactivities of immune and nonimmune cells. The role of ATF3 has been studied in different contexts. In macrophages, ATF3 is a crucial regulator of innate immune responses. It is a product of TLR signaling and modulates inflammation in lipopolysaccharide (LPS)-stimulated cells by returning to this pathway [29][30][31][32]. Monocyte chemotactic protein-1-induced protein 1 (MCPIP1) inhibits macrophage polarization of M1 and promotes polarization of M2 by regulating the ATF3-AP1S2 signaling pathway, limiting the expression of pro-inflammatory cytokines in monocytes from patients with active inflammatory bowel disease (IBD). ATF3 can bind to the AP1S2 promoter and induce inflammation [33]. Postprandial triglyceride-rich lipoprotein (TGRL) lipolysis products (TL) induce the expression of pro-inflammatory factors in human brain microvascular endothelial cells by upregulating ATF3 through the activation of mitochondrial oxidative stress [34]. NF-κB regulates its expression by binding to the IL-6 promoter [35][36], while ATF3 has been shown to modulate NF-κB-dependent transcription by altering the phosphorylation of IκBα [22]. In osteoarthritis (OA), ATF3 modulates phosphorylation of p65 by altering IκB phosphorylation, which regulates the NF-κB signaling pathway, thus regulating the expression of IL-6 in chondrocytes [37]. ATF3 mediates particulate matter (PM)-induced inflammatory cytokine expression through the NF-κB and AP-1 pathways [38].
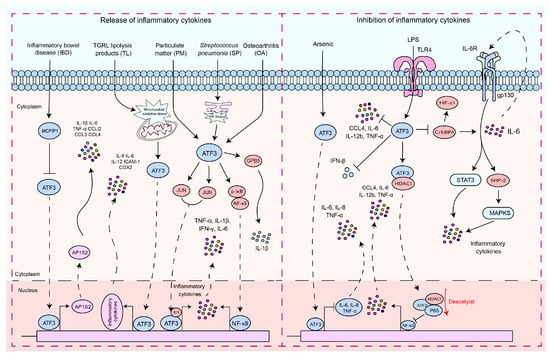
ATF3 plays a dual role in the regulation of inflammatory responses (Figure 1). In endotoxin-stimulated monocytes, the stress associated with reactive oxygen species (ROS) leads to the induction of ATF3 expression and inhibits IL-6 production, making mice highly susceptible to secondary bacterial and fungal infections [39]. Negative regulation of transcription by ATF3 may be achieved through the inhibition of CCAAT/enhancer binding protein δ (C/EBPδ), a positive regulator of cytokine gene induction [40][41]. C/EBPδ enhances the expression of IL-6 [40], and in turn, IL-6, through activation of IL-6R, can activate the STAT3, SHP-2, and MAPK pathways, thus promoting cytokine secretion [40][42][43]. It is reported that ATF3 recruits HDAC1 to the ATF3/p65 complex and promotes the deacetylation of p65 to inhibit the production of pro-inflammatory cytokines [44]. The role of ATF3 in macrophages is not limited to the regulation of pro-inflammatory cytokines. It is also essential for regulating the production of interferon (IFN)-β downstream of innate immune receptors. ATF3 acts as a transcriptional repressor by binding to regulatory sites on the Ifnb1 promoter [45]. The expression of macrophage inflammatory protein 1β (CCL4) leads to the onset of inflammatory diseases [30][46]. ATF3 inhibits the expression of CCL4 in mouse macrophages induced by LPS [30]. ATF3 negatively regulates the gene expression of IL-6 and IL-12 in macrophages by altering the structure of chromatin [13][29]. LPS activation of TLR4 induces ATF3 expression, which in turn suppresses the expression of various inflammatory genes induced by TLR4 signaling, including IL-6, IL-12b, and TNF-α [13][14].

Figure 1. The signaling pathways by which ATF3 regulates the inflammatory response. These signaling pathways by which ATF3 regulates the inflammatory response can be divided into two types: (1) pro-inflammatory response—ATF3 increased the production of pro-inflammatory cytokines and chemokines by enhancing AP1S2 expression by binding to AP1S2 promoters in inflammation of the intestine (IBD). S. pneumoniae (SP) stimulates the formation of an ATF3 complex with c-Jun, and this complex binds to cytokine promoters of cytokines (TNF-α, IL-1β, and IFN-γ), resulting in increased cytokine production. During an infection, lung macrophages quickly phagocytose invasive S. pneumoniae, resulting in ER stress and ATF3 activation. ATF3 then promotes GBP5 activation, triggering IL-1β secretion. TGRL lipolysis products (TL) potentiate ROS in mitochondria, activating mitochondrial oxidative stress and ATF3 signaling. Furthermore, ATF3 regulates TL-induced inflammation. ATF3 may positively regulate IL-6 expression in osteoarthritis (OA) chondrocytes through modulation of NF-κB-dependent transcription by modifying IκB phosphorylation. ATF3 may heterodimerize with c-JUN and activate IL-6 transcription in HBE cells induced by PM. (2) Anti-inflammatory response, including regulation of the ATF3/HDAC1/NF-κB axis and ATF3/C-EBPδ axis—ATF3 inhibits the production of inflammatory cytokines by suppressing C/EBPδ. ATF3 inhibits the production of pro-inflammatory cytokines by recruiting HDAC1 into the ATF3/p65 complex and facilitating the deacetylation of p65. ATF3 acted as a transcriptional repressor and regulated IFN-β. LPS activates ATF3 by stimulating TLRs, thus inhibiting the production of inflammatory cytokines. Solid arrows indicate promotion, dashed arrows denote translocation, horizontal arrows represent inhibition, and red arrows signify the processes of modification undergone.
5. The Role of ATF3 in the Regulation of Cell Death
5.1. The Regulation of Apoptosis
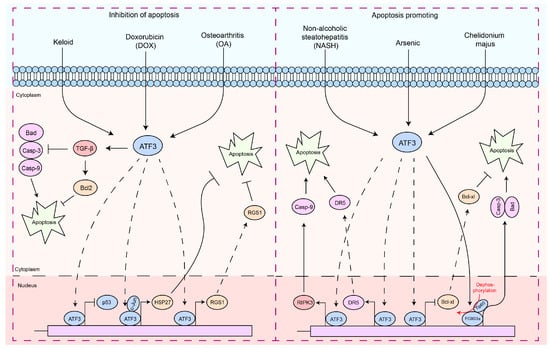
Apoptosis is a finely regulated process of programmed cell death that plays a crucial role in the maintenance of physiological functions in organisms, as well as in interactions between pathogens and hosts [47][48][49][50]. Apoptosis is a form of cell death by which the body maintains the homeostasis of the internal environment [5][51]. In addition, it plays an important role in the regulation of the immune system, especially in autoreactive immune cells [5][52][53][54]. In the interaction between pathogens and hosts, apoptosis can serve as a host defense mechanism to limit the replication and spread of pathogens [5][55][56]. In contrast, certain pathogens may exploit host defense mechanisms to evade host immune responses or induce excessive inflammation and tissue damage during infection [5][55][56]. Therefore, the diverse and complex functions of apoptosis are essential to understanding cellular behavior in physiological and pathological states. In particular, ATF3 induction appears to be consistently associated with cellular damage, as most of the signals that induce ATF3 also induce cellular injury [15][23][57]. Interestingly, ATF3 exhibits a dual role in the regulation of apoptosis (Figure 2). In cardiomyocytes, ATF3 effectively inhibits doxorubicin-induced apoptosis [58].

Figure 2. The functions of ATF3 in the response to apoptosis. These signaling pathways by which ATF3 regulates the apoptosis response can be divided into two types: (1) antiapoptotic response—ATF3 can regulate apoptosis of cells by upregulating the expression of the anti-apoptotic gene (HSP27, RGS1, and Bcl2) by binding to promoters, preventing p53 expression, which inhibits Caspases-3/9 activities. In addition, ATF3 inhibits Bad expression via TGF-β. (2) Pro-apoptosis response—ATF3 triggers the apoptotic pathway by upregulating RIPK3, DR5, and Caspase-9 by binding to their promoter, while concurrently inhibiting BCL-XL by binding to its promoter. Furthermore, ATF3 can also promote Caspase-3 and Bad transcription by activating FOXO3a, thereby regulating cell apoptosis. Solid arrows indicate promotion, dashed arrows denote translocation, horizontal arrows represent inhibition, and red arrows signify the processes of modification undergone.
These findings suggest a role for ATF3 in inhibiting apoptosis. However, ectopic expression of ATF3 improves the apoptotic capacity of topotecan-induced HeLa cells or camptothecin-induced HeLa cells [59]. ATF3 may act as a downstream target of the NF-κB and JNK/SAPK signaling pathways, promoting β-cell apoptosis [57]. ATF3 expression intensifies t-butyl hydroperoxide (TBHP)-induced apoptosis in nucleus pulposus cells (NPC) [60]. ATF3-dependent induction of RIPK3 causes a shift from apoptosis to necroptosis in hepatocytes [61]. The opposing regulation of DR5 and Bcl-xL expression by ATF3 promotes arsenic-induced apoptosis [62]. Forkhead transcription factors (FOXO3a) are a key molecule that promotes apoptosis, primarily functioning by facilitating the transcription of apoptosis-related factors, thereby mediating cell apoptosis [63][64][65][66]. The PI3K/Akt pathway inhibits apoptosis by phosphorylating FOXO3a, which prevents its nuclear translocation [65][66][67]. Chelidonium majus induces apoptosis in SKOV-3 cells by increasing the expression levels of ATF3 and its downstream protein, Tip60 [68].
5.2. The Regulation of Ferroptosis
Ferroptosis is a nonapoptotic form of cell death that can be induced by metabolic stress, such as glutathione (GSH) depletion [69][70][71]. Recently defined as a newly discovered form of cell death, ferroptosis is different from apoptosis in that it does not involve caspase activation [70]. Ferroptosis leads to an increase in ROS and malondialdehyde (MDA), which ultimately causes overwhelming lipid peroxidation and results in cell death [70][72][73]. ATF3 is often involved in vital cellular activities such as metabolism. Currently, a large amount of research data indicates that ATF3 plays a significant role in the regulation of ferroptosis [69][73][74][75][76][77][78][79]. Nuclear factor erythroid 2 related factor 2 (Nrf2) can promote the expression of SLC7A11 and GPX4 under oxidative stress, which is crucial to mediate the onset of ferroptosis [80][81][82]. As an endogenous inhibitor of SLC7A11, ATF3 promotes erastin-induced ferroptosis by inhibiting the cystine/glutamate antiporter (system Xc-) [69].
6. The Functions of ATF3 in the Pathogenic Microbial Infection Process
6.1. The Functions of ATF3 in Viral Infection
6.1.1. DNA Virus
The HBx protein, one of the seven proteins made by the Hepatitis B virus (HBV), is very toxic and can activate various genes in cells [83][84]. This affects many cell processes, such as the regulation of intracellular gene transcription, signal transduction, protein degradation, the cell cycle, and apoptosis [83][84]. Furthermore, HBx is a protein with a short half-life, primarily degraded in the cell via the ubiquitin-dependent proteasome pathway. Inhibiting HBx degradation and increasing the level of intracellular expression of HBx is one way HBV can cause liver cancer [84]. IL-1β/ATF3 promotes HBx mRNA degradation by mediating the expression of Ski2, which helps prevent complications mediated by HBx [83]. In particular, the level of Ski2 is also regulated by the HBx protein, forming a significant negative feedback loop to suppress HBx levels [83]. This mechanism is likely crucial for the virus to maintain an optimal concentration of HBx to support viral replication without triggering apoptosis [83][85]. Interestingly, HBx appears to induce Ski2 promoter activity by interacting with ATF3 [83][86].
Murine cytomegalovirus (MCMV) is a virus particularly susceptible to control by IFN-γ, produced by NK cells. The viral load of MCMV in the liver is regulated by IFN-γ, which is secreted by NK cells in response to IL-12 [87][88][89]. Previous studies have shown that depletion of IFN-γ exacerbates infection, leading to an increase in liver MCMV viral load and the appearance of hepatitis [88][90][91].
A key characteristic of the Herpes Simplex Virus (HSV) is its ability to establish latent infection in the autonomic ganglia and reactivate under physical, hormonal, or emotional stress [92][93]. ATF3 is currently recognized as a significant stress-induced factor associated with the suppression of latent viral activation [93]. ATF3 has been reported to play a role in maintaining the latent state of HSV [93]. ATF3 is specifically expressed after neuronal injury, and it acts in synergy with STAT3 to induce the expression of downstream genes [94]. The main function of ATF3 in cells infected with HSV-1 has been reported to be to maintain neuronal integrity [93].
Human Papillomavirus (HPV) infection is a primary risk factor for cervical cancer [95][96]. HPV facilitates the proteolytic degradation and inactivation of p53 through the expression of the E6 protein [97][98]. Therefore, inhibiting the E6-promoted degradation of p53 appears to be an effective intervention against HPV-induced cervical cancer [98][99][100].
6.1.2. RNA Virus
The binding of type I interferons (IFN) to their receptors leads to receptor dimerization, subsequently activating the IRF and STAT families of TF [101][102]. STAT1 and STAT2 undergo dimerization and interact with IRF9, resulting in the formation of the interferon-stimulated gene factor 3 (ISGF3) complex [101][102]. This complex then translocates to the nucleus and binds to the conserved interferon-stimulated response element, thus inducing a range of interferon-sensitive genes (ISGs) that inhibit the replication of the Japanese Encephalitis Virus (JEV) [120
Interestingly, compared to its response to JEV infection, ATF3 exhibits a different function in hosts infected with the Zika virus (ZIKV). Research found that ATF3 inhibits ZIKV infection by differentially regulating the transcription of specific innate immune response and autophagy genes [103]. During the infection with ZIKV of A549 cells, ATF3 promotes the transcription of the RIG-I, STAT1, IRF9, and ISG15 genes while simultaneously inhibiting the transcription levels of IFNβ and IFIT2 [103].
Viruses often need to inhibit host cell death in the early stages of infection to allow sufficient replication time for the production of adequate viral progeny [104]. Later in the infection, promoting host cell death or utilizing budding mechanisms can facilitate viral dissemination [104].
Infection with the Dengue virus (DENV) can induce a robust cytokine storm in the brain, leading to neurological symptoms or death in the host [105]. An upregulation of ATF3 expression has been observed in blood samples of patients infected with DENV [106]. Research has discovered that monocytes infected with DENV secrete extracellular vesicles (EV), which are internalized by microglia [107]. The miR-148a carried within these EVs inhibits the expression level of the ubiquitin-specific peptidase 33 (USP33) protein. The reduction in USP33, in turn, decreases the stability of cellular ATF3 protein through deubiquitination, thereby promoting the expression of pro-inflammatory genes such as TNF-α, NF-κB, and IFN-β [107]. This indicates that DENV manipulates the EV pathway to transfer miR-148a, thereby regulating the levels of USP33 and downstream ATF3 in human microglia and leading to neuroinflammation within the central nervous system.
In the context of the pathogenesis of integrated viruses, a pivotal aspect involves the exploitation of host cellular machinery for the expression of the viral genome during host infection. Specifically, the viral genome’s integration into the host’s chromatin architecture necessitates a strategic utilization of the host’s gene regulatory systems [108][109]. This phenomenon is exemplified in the case of Human Immunodeficiency Virus Type 1 (HIV-1), where, post-infection, the viral genome becomes assimilated into the host genome as a component of chromatin [108][109]. Central to this process is Nuc-1, a nucleosome situated immediately downstream of the HIV-1 transcription initiation site, which inherently inhibits the activity of the long-terminal repeat (LTR) [108][109][110]. The initiation of LTR-driven transcription and consequent viral expression are contingent upon both epigenetic modifications and the disruption of nuc-1 [109]. Within this nucleosome, the presence of three AP1 sites is critical for the facilitation of viral transcription and replication [111][112][113].
6.2. The Functions of ATF3 in Bacterial Infection
6.2.1. Gram-Positive Bacteria
During the infection process of Staphylococcus aureus (S. aureus), the host mediates the secretion of immune factors such as cytokines and chemokines through TLR-2, subsequently promoting the production of IL-17 to coordinate the host immune response [114][115]. IL-17, together with IL-22/IL-23, modulates macrophage function, thus inducing the expression of antimicrobial peptides (AMP) that kill or inactivate the pathogen [116][117]. Inactivation of IL-17, IL-22, and IL-23 leads to an increased S. aureus load and exacerbates the disease [116][118]. ATF3 has been reported to promote bacterial clearance by regulating the production of inflammatory cytokines, thus alleviating lethal S. aureus pneumonia [114]. ATF3 positively regulates the host’s resistance to S. aureus infection by modulating macrophage Reg3 expression and AMPs gene-mediated bacterial clearance, as well as the recruitment of macrophages, thus playing a significant role in the early stages of S. aureus infection [114].
IL-17A is critical in the early defense against Streptococcus pneumoniae (S. pneumoniae), as mice lacking IL-17A and IL-17RA show increased vulnerability to bacterial pathogens that incite lung diseases [119][120][121]. ATF3 has been reported to facilitate the generation of IL-17A in γδ T cells through macrophage-mediated secretion of IL-1β, thus modulating the response to infection [4]. ATF3 regulates the immune response by maintaining the intracellular balance of ROS and calcium ions (Ca2+), influencing macrophage production of IL-1β and IL-23p19 [4]. This process is essential for stimulating the secretion of IL-17A, which is necessary for early defense against infections and crucial to eradicating S. pneumoniae [4].
Wild-type mice demonstrate more effective bacterial clearance than ATF3-null mice during Listeria monocytogenes (L. monocytogenes) infection [122]. This suggests that ATF3 plays a critical role in resisting L. monocytogenes infection. PLY induces ATF3 expression through the TLR4/MAPK pathway [123]. Listeriolysin O (LLO), a member of the cytolysins released by L. monocytogenes, is known to stimulate TLR4-dependent cytokine expression and acts as a TLR4 agonist [124]. Furthermore, ATF3 significantly improves the expression levels of TNF-α, IL-1β, and IFN-γ during L. monocytogenes infection [122].
6.2.2. Gram-Negative Bacteria
During episodes of bacterial sepsis, the host modulates its response to infection by upregulating or suppressing cytokines through ATF3 [39][125]. LPS significantly induces ATF3, which functions as a negative regulator in the production of inflammatory cytokines [13][39].
E. coli sepsis is currently one of the most important types of sepsis [126]. ATF3 facilitates the progression of E. coli sepsis by suppressing IL-6 transcription [39]. Due to immunosuppression associated with ATF3-mediated sepsis, ATF3 knockout mice exhibit longer survival than wild-type mice after infection with E. coli [39]. Post-infection with uropathogenic Escherichia coli (UPEC), cytokines such as IL-1β, IL-6, and IFN-γ are significantly suppressed, while the bacterial load in the lungs and spleens of wild-type mice is substantially higher than in ATF3 knockout mice [122]. This indicates that ATF3 exhibits a divergent mechanism in Gram-positive and Gram-negative bacterial infections. This contradictory result may be caused by the following reasons: (1) LPS-induced ATF3 competes with NF-κB for binding to the promoters of target cytokines, thus inhibiting the production of inflammatory cytokines [13][122]. (2) LPS-induced ATF3 binds to cytokine promoters, and its interaction with HDAC leads to histone deacetylation. This process results in chromatin condensation, which suppresses cytokine gene transcription [13][122]. (3) The regulation of TLR4 differs; in Gram-positive bacterial infections, ATF3 may positively regulate TLR4 expression and stimulate cytokine production [122][123][125].
6.2.3. Other Bacteria
ATF3 expression is significantly upregulated during Mycobacterium tuberculosis (Mtb) infection [127]. ATF3 cooperates with BRG1 to activate the expression of the inflammatory cytokines IL-6, TNF-α, and IL-12p40 and increase the production of nitric oxide [127][128]. Foamy macrophages, a subpopulation of macrophages, play a pivotal role in the pathogenesis of tuberculosis [128]. They are characterized by an abundance of liposomes (LB), which may provide a survival environment for mycobacteria in granulomas [128][129]. Mtb stimulates the formation of LB-rich foamy macrophages [129][130]. ATF3 has been reported to inhibit liposome formation by regulating the expression of genes related to lipid metabolism [128].
6.3. The Functions of ATF3 in Fungal and Parasite Infections
Patulin is a fungal toxin primarily released by Aspergillus and Penicillium species [131]. Patulin exerts its toxic effect by covalently binding to reactive sulfhydryl groups in cellular proteins and by depleting glutathione, resulting in oxidative damage and the generation of reactive oxygen species (ROS) [132][133]. Research has found that Patulin induces transcription factor EGR-1 phosphorylation through increased oxidative stress, thereby enhancing ATF3 expression and promoting apoptosis in colorectal cancer cells [134]. Deoxynivalenol (DON), commonly known as vomitoxin, is a type B trichothecene mycotoxin predominantly produced by Fusarium species, such as F. culmorum and F. graminearum [135].
7. Prospects for Clinical Applications
Considering the multifaceted role of ATF3 in the regulation of physiological functions, it presents a broad prospect as a clinical pharmacological target: (1) Treatments that target ATF3 to control inflammation, useful in autoimmune and other inflammatory diseases. (2) Exploiting ATF3 to develop treatments for neurological disorders and nerve damage. (3) Utilizing ATF3 expression levels as a diagnostic and prognostic tool, especially in cancer. However, we need to carefully consider ATF3’s dual functions to avoid adverse effects in treatment. Targeting ATF3 presents significant opportunities for novel therapeutic developments and improved disease management in clinical medicine. Its potential as a drug target and a biomarker can lead to advances in personalized medicine. However, the complexity of ATF3’s roles necessitates careful research and development to ensure the efficacy and safety of these new approaches.
References
- Wang, W.; Chang, X.; Yao, W.; Wei, N.; Huo, N.; Wang, Y.; Wei, Q.; Liu, H.; Wang, X.; Zhang, S.; et al. Host CARD11 Inhibits Newcastle Disease Virus Replication by Suppressing Viral Polymerase Activity in Neurons. J. Virol. 2019, 93, e01499-19.
- Caamano, J.; Hunter, C.A. NF-kappaB family of transcription factors: Central regulators of innate and adaptive immune functions. Clin. Microbiol. Rev. 2002, 15, 414–429.
- Le Negrate, G. Subversion of innate immune responses by bacterial hindrance of NF-kappaB pathway. Cell Microbiol. 2012, 14, 155–167.
- Lee, S.; Kim, G.L.; Kim, N.Y.; Kim, S.J.; Ghosh, P.; Rhee, D.K. ATF3 Stimulates IL-17A by Regulating Intracellular Ca2+/ROS-Dependent IL-1beta Activation During Streptococcus pneumoniae Infection. Front. Immunol. 2018, 9, 1954.
- Hu, H.; Tian, M.; Ding, C.; Yu, S. The C/EBP Homologous Protein (CHOP) Transcription Factor Functions in Endoplasmic Reticulum Stress-Induced Apoptosis and Microbial Infection. Front. Immunol. 2018, 9, 3083.
- Zhang, K.; Yang, B.; Shen, C.; Zhang, T.; Hao, Y.; Zhang, D.; Liu, H.; Shi, X.; Li, G.; Yang, J.; et al. MGF360-9L Is a Major Virulence Factor Associated with the African Swine Fever Virus by Antagonizing the JAK/STAT Signaling Pathway. mBio 2022, 13, e0233021.
- Ariffianto, A.; Deng, L.; Harada, S.; Liang, Y.; Matsui, C.; Abe, T.; Shoji, I. Transcription Factor JunB Suppresses Hepatitis C Virus Replication. Kobe J. Med. Sci. 2023, 69, E86–E95.
- Li, D.; Jing, J.; Dong, X.; Zhang, C.; Wang, J.; Wan, X. Activating transcription factor 3: A potential therapeutic target for inflammatory pulmonary diseases. Immun. Inflamm. Dis. 2023, 11, e1028.
- Rohini, M.; Haritha Menon, A.; Selvamurugan, N. Role of activating transcription factor 3 and its interacting proteins under physiological and pathological conditions. Int. J. Biol. Macromol. 2018, 120, 310–317.
- Thompson, M.R.; Xu, D.; Williams, B.R. ATF3 transcription factor and its emerging roles in immunity and cancer. J. Mol. Med. 2009, 87, 1053–1060.
- Zhou, H.; Li, N.; Yuan, Y.; Jin, Y.G.; Guo, H.; Deng, W.; Tang, Q.Z. Activating transcription factor 3 in cardiovascular diseases: A potential therapeutic target. Basic. Res. Cardiol. 2018, 113, 37.
- Zhao, J.; Li, X.; Guo, M.; Yu, J.; Yan, C. The common stress responsive transcription factor ATF3 binds genomic sites enriched with p300 and H3K27ac for transcriptional regulation. BMC Genom. 2016, 17, 335.
- Gilchrist, M.; Thorsson, V.; Li, B.; Rust, A.G.; Korb, M.; Roach, J.C.; Kennedy, K.; Hai, T.; Bolouri, H.; Aderem, A. Systems biology approaches identify ATF3 as a negative regulator of Toll-like receptor 4. Nature 2006, 441, 173–178.
- Hai, T.; Wolford, C.C.; Chang, Y.S. ATF3, a hub of the cellular adaptive-response network, in the pathogenesis of diseases: Is modulation of inflammation a unifying component? Gene Expr. 2010, 15, 1–11.
- Hai, T.; Wolfgang, C.D.; Marsee, D.K.; Allen, A.E.; Sivaprasad, U. ATF3 and stress responses. Gene Expr. 1999, 7, 321–335.
- Chen, H.; Luo, S.; Chen, H.; Zhang, C. ATF3 regulates SPHK1 in cardiomyocyte injury via endoplasmic reticulum stress. Immun. Inflamm. Dis. 2023, 11, e998.
- Yu, R.; Shtil, A.A.; Tan, T.H.; Roninson, I.B.; Kong, A.N. Adriamycin activates c-jun N-terminal kinase in human leukemia cells: A relevance to apoptosis. Cancer Lett. 1996, 107, 73–81.
- Drysdale, B.E.; Howard, D.L.; Johnson, R.J. Identification of a lipopolysaccharide inducible transcription factor in murine macrophages. Mol. Immunol. 1996, 33, 989–998.
- Weir, E.; Chen, Q.; DeFrances, M.C.; Bell, A.; Taub, R.; Zarnegar, R. Rapid induction of mRNAs for liver regeneration factor and insulin-like growth factor binding protein-1 in primary cultures of rat hepatocytes by hepatocyte growth factor and epidermal growth factor. Hepatology 1994, 20, 955–960.
- Kim, K.H.; Jeong, J.Y.; Surh, Y.J.; Kim, K.W. Expression of stress-response ATF3 is mediated by Nrf2 in astrocytes. Nucleic Acids Res. 2010, 38, 48–59.
- Wei, Y.; Sun, L.; Liu, C.; Li, L. Naringin regulates endoplasmic reticulum stress and mitophagy through the ATF3/PINK1 signaling axis to alleviate pulmonary fibrosis. Naunyn Schmiedebergs Arch. Pharmacol. 2023, 396, 1155–1169.
- Kim, E.Y.; Shin, H.Y.; Kim, J.Y.; Kim, D.G.; Choi, Y.M.; Kwon, H.K.; Rhee, D.K.; Kim, Y.S.; Choi, S. ATF3 plays a key role in Kdo2-lipid A-induced TLR4-dependent gene expression via NF-kappaB activation. PLoS ONE 2010, 5, e14181.
- Hai, T.; Hartman, M.G. The molecular biology and nomenclature of the activating transcription factor/cAMP responsive element binding family of transcription factors: Activating transcription factor proteins and homeostasis. Gene 2001, 273, 1–11.
- Nilsson, M.; Toftgard, R.; Bohm, S. Activated Ha-Ras but not TPA induces transcription through binding sites for activating transcription factor 3/Jun and a novel nuclear factor. J. Biol. Chem. 1995, 270, 12210–12218.
- Nakagomi, S.; Suzuki, Y.; Namikawa, K.; Kiryu-Seo, S.; Kiyama, H. Expression of the activating transcription factor 3 prevents c-Jun N-terminal kinase-induced neuronal death by promoting heat shock protein 27 expression and Akt activation. J. Neurosci. 2003, 23, 5187–5196.
- Chen, B.P.; Liang, G.; Whelan, J.; Hai, T. ATF3 and ATF3 delta Zip. Transcriptional repression versus activation by alternatively spliced isoforms. J. Biol. Chem. 1994, 269, 15819–15826.
- Shvedunova, M.; Akhtar, A. Modulation of cellular processes by histone and non-histone protein acetylation. Nat. Rev. Mol. Cell Biol. 2022, 23, 329–349.
- Kawai, T.; Akira, S. TLR signaling. Cell Death Differ. 2006, 13, 816–825.
- Whitmore, M.M.; Iparraguirre, A.; Kubelka, L.; Weninger, W.; Hai, T.; Williams, B.R. Negative regulation of TLR-signaling pathways by activating transcription factor-3. J. Immunol. 2007, 179, 3622–3630.
- Khuu, C.H.; Barrozo, R.M.; Hai, T.; Weinstein, S.L. Activating transcription factor 3 (ATF3) represses the expression of CCL4 in murine macrophages. Mol. Immunol. 2007, 44, 1598–1605.
- Suganami, T.; Yuan, X.; Shimoda, Y.; Uchio-Yamada, K.; Nakagawa, N.; Shirakawa, I.; Usami, T.; Tsukahara, T.; Nakayama, K.; Miyamoto, Y.; et al. Activating transcription factor 3 constitutes a negative feedback mechanism that attenuates saturated Fatty acid/toll-like receptor 4 signaling and macrophage activation in obese adipose tissue. Circ. Res. 2009, 105, 25–32.
- Yuan, X.; Yu, L.; Li, J.; Xie, G.; Rong, T.; Zhang, L.; Chen, J.; Meng, Q.; Irving, A.T.; Wang, D.; et al. ATF3 suppresses metastasis of bladder cancer by regulating gelsolin-mediated remodeling of the actin cytoskeleton. Cancer Res. 2013, 73, 3625–3637.
- Lu, H.; Zhang, C.; Wu, W.; Chen, H.; Lin, R.; Sun, R.; Gao, X.; Li, G.; He, Q.; Gao, H.; et al. MCPIP1 restrains mucosal inflammation by orchestrating the intestinal monocyte to macrophage maturation via an ATF3-AP1S2 axis. Gut 2023, 72, 882–895.
- Nyunt, T.; Britton, M.; Wanichthanarak, K.; Budamagunta, M.; Voss, J.C.; Wilson, D.W.; Rutledge, J.C.; Aung, H.H. Mitochondrial oxidative stress-induced transcript variants of ATF3 mediate lipotoxic brain microvascular injury. Free Radic. Biol. Med. 2019, 143, 25–46.
- Dendorfer, U.; Oettgen, P.; Libermann, T.A. Multiple regulatory elements in the interleukin-6 gene mediate induction by prostaglandins, cyclic AMP, and lipopolysaccharide. Mol. Cell Biol. 1994, 14, 4443–4454.
- de Haij, S.; Bakker, A.C.; van der Geest, R.N.; Haegeman, G.; Vanden Berghe, W.; Aarbiou, J.; Daha, M.R.; van Kooten, C. NF-kappaB mediated IL-6 production by renal epithelial cells is regulated by c-jun NH2-terminal kinase. J. Am. Soc. Nephrol. 2005, 16, 1603–1611.
- Iezaki, T.; Ozaki, K.; Fukasawa, K.; Inoue, M.; Kitajima, S.; Muneta, T.; Takeda, S.; Fujita, H.; Onishi, Y.; Horie, T.; et al. ATF3 deficiency in chondrocytes alleviates osteoarthritis development. J. Pathol. 2016, 239, 426–437.
- Yan, F.; Wu, Y.; Liu, H.; Wu, Y.; Shen, H.; Li, W. ATF3 is positively involved in particulate matter-induced airway inflammation in vitro and in vivo. Toxicol. Lett. 2018, 287, 113–121.
- Hoetzenecker, W.; Echtenacher, B.; Guenova, E.; Hoetzenecker, K.; Woelbing, F.; Bruck, J.; Teske, A.; Valtcheva, N.; Fuchs, K.; Kneilling, M.; et al. ROS-induced ATF3 causes susceptibility to secondary infections during sepsis-associated immunosuppression. Nat. Med. 2011, 18, 128–134.
- Balamurugan, K.; Mendoza-Villanueva, D.; Sharan, S.; Summers, G.H.; Dobrolecki, L.E.; Lewis, M.T.; Sterneck, E. C/EBPdelta links IL-6 and HIF-1 signaling to promote breast cancer stem cell-associated phenotypes. Oncogene 2019, 38, 3765–3780.
- Litvak, V.; Ramsey, S.A.; Rust, A.G.; Zak, D.E.; Kennedy, K.A.; Lampano, A.E.; Nykter, M.; Shmulevich, I.; Aderem, A. Function of C/EBPdelta in a regulatory circuit that discriminates between transient and persistent TLR4-induced signals. Nat. Immunol. 2009, 10, 437–443.
- Wu, X.; Wei, S.; Chen, M.; Li, J.; Wei, Y.; Zhang, J.; Dong, W. P2RY13 Exacerbates Intestinal Inflammation by Damaging the Intestinal Mucosal Barrier via Activating IL-6/STAT3 Pathway. Int. J. Biol. Sci. 2022, 18, 5056–5069.
- Tanaka, T.; Narazaki, M.; Kishimoto, T. IL-6 in inflammation, immunity, and disease. Cold Spring Harb. Perspect. Biol. 2014, 6, a016295.
- Kwon, J.W.; Kwon, H.K.; Shin, H.J.; Choi, Y.M.; Anwar, M.A.; Choi, S. Activating transcription factor 3 represses inflammatory responses by binding to the p65 subunit of NF-kappaB. Sci. Rep. 2015, 5, 14470.
- Labzin, L.I.; Schmidt, S.V.; Masters, S.L.; Beyer, M.; Krebs, W.; Klee, K.; Stahl, R.; Lutjohann, D.; Schultze, J.L.; Latz, E.; et al. ATF3 Is a Key Regulator of Macrophage IFN Responses. J. Immunol. 2015, 195, 4446–4455.
- Ma, J.Q.; Li, Z.; Xie, W.R.; Liu, C.M.; Liu, S.S. Quercetin protects mouse liver against CCl4-induced inflammation by the TLR2/4 and MAPK/NF-kappaB pathway. Int. Immunopharmacol. 2015, 28, 531–539.
- Yue, J.; Lopez, J.M. Understanding MAPK Signaling Pathways in Apoptosis. Int. J. Mol. Sci. 2020, 21, 2346.
- Morana, O.; Wood, W.; Gregory, C.D. The Apoptosis Paradox in Cancer. Int. J. Mol. Sci. 2022, 23, 1328.
- D’Arcy, M.S. Cell death: A review of the major forms of apoptosis, necrosis and autophagy. Cell Biol. Int. 2019, 43, 582–592.
- Bertheloot, D.; Latz, E.; Franklin, B.S. Necroptosis, pyroptosis and apoptosis: An intricate game of cell death. Cell Mol. Immunol. 2021, 18, 1106–1121.
- Kitakata, H.; Endo, J.; Ikura, H.; Moriyama, H.; Shirakawa, K.; Katsumata, Y.; Sano, M. Therapeutic Targets for DOX-Induced Cardiomyopathy: Role of Apoptosis vs. Ferroptosis. Int. J. Mol. Sci. 2022, 23, 1414.
- Williams, G.T. Role of apoptosis in the immune system. Biochem. Cell Biol. 1994, 72, 447–450.
- Nagata, S.; Tanaka, M. Programmed cell death and the immune system. Nat. Rev. Immunol. 2017, 17, 333–340.
- Ekert, P.G.; Vaux, D.L. Apoptosis and the immune system. Br. Med. Bull. 1997, 53, 591–603.
- Place, D.E.; Lee, S.; Kanneganti, T.D. PANoptosis in microbial infection. Curr. Opin. Microbiol. 2021, 59, 42–49.
- Chen, Y.; Zychlinsky, A. Apoptosis induced by bacterial pathogens. Microb. Pathog. 1994, 17, 203–212.
- Hartman, M.G.; Lu, D.; Kim, M.L.; Kociba, G.J.; Shukri, T.; Buteau, J.; Wang, X.; Frankel, W.L.; Guttridge, D.; Prentki, M.; et al. Role for activating transcription factor 3 in stress-induced beta-cell apoptosis. Mol. Cell Biol. 2004, 24, 5721–5732.
- Nobori, K.; Ito, H.; Tamamori-Adachi, M.; Adachi, S.; Ono, Y.; Kawauchi, J.; Kitajima, S.; Marumo, F.; Isobe, M. ATF3 inhibits doxorubicin-induced apoptosis in cardiac myocytes: A novel cardioprotective role of ATF3. J. Mol. Cell Cardiol. 2002, 34, 1387–1397.
- Mashima, T.; Udagawa, S.; Tsuruo, T. Involvement of transcriptional repressor ATF3 in acceleration of caspase protease activation during DNA damaging agent-induced apoptosis. J. Cell Physiol. 2001, 188, 352–358.
- Li, Y.; Pan, D.; Wang, X.; Huo, Z.; Wu, X.; Li, J.; Cao, J.; Xu, H.; Du, L.; Xu, B. Silencing ATF3 Might Delay TBHP-Induced Intervertebral Disc Degeneration by Repressing NPC Ferroptosis, Apoptosis, and ECM Degradation. Oxid. Med. Cell Longev. 2022, 2022, 4235126.
- Inaba, Y.; Hashiuchi, E.; Watanabe, H.; Kimura, K.; Oshima, Y.; Tsuchiya, K.; Murai, S.; Takahashi, C.; Matsumoto, M.; Kitajima, S.; et al. The transcription factor ATF3 switches cell death from apoptosis to necroptosis in hepatic steatosis in male mice. Nat. Commun. 2023, 14, 167.
- Shi, Q.; Hu, B.; Yang, C.; Zhao, L.; Wu, J.; Qi, N. ATF3 Promotes Arsenic-Induced Apoptosis and Oppositely Regulates DR5 and Bcl-xL Expression in Human Bronchial Epithelial Cells. Int. J. Mol. Sci. 2021, 22, 4223.
- Yang, J.Y.; Zong, C.S.; Xia, W.; Yamaguchi, H.; Ding, Q.; Xie, X.; Lang, J.Y.; Lai, C.C.; Chang, C.J.; Huang, W.C.; et al. ERK promotes tumorigenesis by inhibiting FOXO3a via MDM2-mediated degradation. Nat. Cell Biol. 2008, 10, 138–148.
- Maiese, A.; Spina, F.; Visi, G.; Del Duca, F.; De Matteis, A.; La Russa, R.; Di Paolo, M.; Frati, P.; Fineschi, V. The Expression of FOXO3a as a Forensic Diagnostic Tool in Cases of Traumatic Brain Injury: An Immunohistochemical Study. Int. J. Mol. Sci. 2023, 24, 2584.
- Maiese, A.; Manetti, A.C.; Santoro, P.; Del Duca, F.; De Matteis, A.; Turillazzi, E.; Frati, P.; Fineschi, V. FOXO3 Depletion as a Marker of Compression-Induced Apoptosis in the Ligature Mark: An Immunohistochemical Study. Int. J. Mol. Sci. 2023, 24, 1396.
- Li, D.; Qu, Y.; Mao, M.; Zhang, X.; Li, J.; Ferriero, D.; Mu, D. Involvement of the PTEN-AKT-FOXO3a pathway in neuronal apoptosis in developing rat brain after hypoxia-ischemia. J. Cereb. Blood Flow. Metab. 2009, 29, 1903–1913.
- Van Der Heide, L.P.; Hoekman, M.F.; Smidt, M.P. The ins and outs of FoxO shuttling: Mechanisms of FoxO translocation and transcriptional regulation. Biochem. J. 2004, 380, 297–309.
- Shen, L.; Lee, S.; Joo, J.C.; Hong, E.; Cui, Z.Y.; Jo, E.; Park, S.J.; Jang, H.J. Chelidonium majus Induces Apoptosis of Human Ovarian Cancer Cells via ATF3-Mediated Regulation of Foxo3a by Tip60. J. Microbiol. Biotechnol. 2022, 32, 493–503.
- Wang, L.; Liu, Y.; Du, T.; Yang, H.; Lei, L.; Guo, M.; Ding, H.F.; Zhang, J.; Wang, H.; Chen, X.; et al. ATF3 promotes erastin-induced ferroptosis by suppressing system Xc−. Cell Death Differ. 2020, 27, 662–675.
- Stockwell, B.R.; Friedmann Angeli, J.P.; Bayir, H.; Bush, A.I.; Conrad, M.; Dixon, S.J.; Fulda, S.; Gascon, S.; Hatzios, S.K.; Kagan, V.E.; et al. Ferroptosis: A Regulated Cell Death Nexus Linking Metabolism, Redox Biology, and Disease. Cell 2017, 171, 273–285.
- Shimada, K.; Skouta, R.; Kaplan, A.; Yang, W.S.; Hayano, M.; Dixon, S.J.; Brown, L.M.; Valenzuela, C.A.; Wolpaw, A.J.; Stockwell, B.R. Global survey of cell death mechanisms reveals metabolic regulation of ferroptosis. Nat. Chem. Biol. 2016, 12, 497–503.
- Xie, Y.; Hou, W.; Song, X.; Yu, Y.; Huang, J.; Sun, X.; Kang, R.; Tang, D. Ferroptosis: Process and function. Cell Death Differ. 2016, 23, 369–379.
- Fu, D.; Wang, C.; Yu, L.; Yu, R. Induction of ferroptosis by ATF3 elevation alleviates cisplatin resistance in gastric cancer by restraining Nrf2/Keap1/xCT signaling. Cell Mol. Biol. Lett. 2021, 26, 26.
- American Association of Neurological Surgeons (AANS); American Society of Neuroradiology (ASNR); Cardiovascular and Interventional Radiology Society of Europe (CIRSE); Canadian Interventional Radiology Association (CIRA); Sacks, D.; Baxter, B.; Campbell, B.C.V.; Carpenter, J.S.; Cognard, C.; Dippel, D.; et al. Multisociety Consensus Quality Improvement Revised Consensus Statement for Endovascular Therapy of Acute Ischemic Stroke. Int. J. Stroke 2018, 13, 612–632.
- Lu, S.; Wang, X.Z.; He, C.; Wang, L.; Liang, S.P.; Wang, C.C.; Li, C.; Luo, T.F.; Feng, C.S.; Wang, Z.C.; et al. ATF3 contributes to brucine-triggered glioma cell ferroptosis via promotion of hydrogen peroxide and iron. Acta Pharmacol. Sin. 2021, 42, 1690–1702.
- Li, Y.; Yan, J.; Zhao, Q.; Zhang, Y.; Zhang, Y. ATF3 promotes ferroptosis in sorafenib-induced cardiotoxicity by suppressing Slc7a11 expression. Front. Pharmacol. 2022, 13, 904314.
- Zhao, Y.; Du, Y.; Gao, Y.; Xu, Z.; Zhao, D.; Yang, M. ATF3 Regulates Osteogenic Function by Mediating Osteoblast Ferroptosis in Type 2 Diabetic Osteoporosis. Dis. Markers 2022, 2022, 9872243.
- Wang, Y.; Chen, D.; Xie, H.; Jia, M.; Sun, X.; Peng, F.; Guo, F.; Tang, D. AUF1 protects against ferroptosis to alleviate sepsis-induced acute lung injury by regulating NRF2 and ATF3. Cell Mol. Life Sci. 2022, 79, 228.
- Wang, Y.; Quan, F.; Cao, Q.; Lin, Y.; Yue, C.; Bi, R.; Cui, X.; Yang, H.; Yang, Y.; Birnbaumer, L.; et al. Quercetin alleviates acute kidney injury by inhibiting ferroptosis. J. Adv. Res. 2021, 28, 231–243.
- Zhu, B.; Ni, Y.; Gong, Y.; Kang, X.; Guo, H.; Liu, X.; Li, J.; Wang, L. Formononetin ameliorates ferroptosis-associated fibrosis in renal tubular epithelial cells and in mice with chronic kidney disease by suppressing the Smad3/ATF3/SLC7A11 signaling. Life Sci. 2023, 315, 121331.
- Dong, H.; Qiang, Z.; Chai, D.; Peng, J.; Xia, Y.; Hu, R.; Jiang, H. Nrf2 inhibits ferroptosis and protects against acute lung injury due to intestinal ischemia reperfusion via regulating SLC7A11 and HO-1. Aging 2020, 12, 12943–12959.
- Ge, M.H.; Tian, H.; Mao, L.; Li, D.Y.; Lin, J.Q.; Hu, H.S.; Huang, S.C.; Zhang, C.J.; Mei, X.F. Zinc attenuates ferroptosis and promotes functional recovery in contusion spinal cord injury by activating Nrf2/GPX4 defense pathway. CNS Neurosci. Ther. 2021, 27, 1023–1040.
- Shiromoto, F.; Aly, H.H.; Kudo, H.; Watashi, K.; Murayama, A.; Watanabe, N.; Zheng, X.; Kato, T.; Chayama, K.; Muramatsu, M.; et al. IL-1beta/ATF3-mediated induction of Ski2 expression enhances hepatitis B virus x mRNA degradation. Biochem. Biophys. Res. Commun. 2018, 503, 1854–1860.
- Chen, H.; Zhang, Y.; Ye, S.; Wu, Q.; Lin, Y.; Sheng, K.; Chen, W.; Lin, X.; Lin, X. Chromatin remodelling factor BAF155 protects hepatitis B virus X protein (HBx) from ubiquitin-independent proteasomal degradation. Emerg. Microbes Infect. 2019, 8, 1393–1405.
- Kim, K.H.; Seong, B.L. Pro-apoptotic function of HBV X protein is mediated by interaction with c-FLIP and enhancement of death-inducing signal. EMBO J. 2003, 22, 2104–2116.
- Barnabas, S.; Hai, T.; Andrisani, O.M. The hepatitis B virus X protein enhances the DNA binding potential and transcription efficacy of bZip transcription factors. J. Biol. Chem. 1997, 272, 20684–20690.
- Orange, J.S.; Biron, C.A. An absolute and restricted requirement for IL-12 in natural killer cell IFN-gamma production and antiviral defense. Studies of natural killer and T cell responses in contrasting viral infections. J. Immunol. 1996, 156, 1138–1142.
- Rosenberger, C.M.; Clark, A.E.; Treuting, P.M.; Johnson, C.D.; Aderem, A. ATF3 regulates MCMV infection in mice by modulating IFN-gamma expression in natural killer cells. Proc. Natl. Acad. Sci. USA 2008, 105, 2544–2549.
- Pien, G.C.; Satoskar, A.R.; Takeda, K.; Akira, S.; Biron, C.A. Cutting edge: Selective IL-18 requirements for induction of compartmental IFN-gamma responses during viral infection. J. Immunol. 2000, 165, 4787–4791.
- Orange, J.S.; Salazar-Mather, T.P.; Opal, S.M.; Biron, C.A. Mechanisms for virus-induced liver disease: Tumor necrosis factor-mediated pathology independent of natural killer and T cells during murine cytomegalovirus infection. J. Virol. 1997, 71, 9248–9258.
- Orange, J.S.; Wang, B.; Terhorst, C.; Biron, C.A. Requirement for natural killer cell-produced interferon gamma in defense against murine cytomegalovirus infection and enhancement of this defense pathway by interleukin 12 administration. J. Exp. Med. 1995, 182, 1045–1056.
- Kang, W.; Mukerjee, R.; Fraser, N.W. Establishment and maintenance of HSV latent infection is mediated through correct splicing of the LAT primary transcript. Virology 2003, 312, 233–244.
- Shu, M.; Du, T.; Zhou, G.; Roizman, B. Role of activating transcription factor 3 in the synthesis of latency-associated transcript and maintenance of herpes simplex virus 1 in latent state in ganglia. Proc. Natl. Acad. Sci. USA 2015, 112, E5420–E5426.
- Kiryu-Seo, S.; Kato, R.; Ogawa, T.; Nakagomi, S.; Nagata, K.; Kiyama, H. Neuronal injury-inducible gene is synergistically regulated by ATF3, c-Jun, and STAT3 through the interaction with Sp1 in damaged neurons. J. Biol. Chem. 2008, 283, 6988–6996.
- Schiffman, M.; Castle, P.E.; Jeronimo, J.; Rodriguez, A.C.; Wacholder, S. Human papillomavirus and cervical cancer. Lancet 2007, 370, 890–907.
- zur Hausen, H. Papillomaviruses causing cancer: Evasion from host-cell control in early events in carcinogenesis. J. Natl. Cancer Inst. 2000, 92, 690–698.
- Scheffner, M.; Werness, B.A.; Huibregtse, J.M.; Levine, A.J.; Howley, P.M. The E6 oncoprotein encoded by human papillomavirus types 16 and 18 promotes the degradation of p53. Cell 1990, 63, 1129–1136.
- Martinez-Zapien, D.; Ruiz, F.X.; Poirson, J.; Mitschler, A.; Ramirez, J.; Forster, A.; Cousido-Siah, A.; Masson, M.; Vande Pol, S.; Podjarny, A.; et al. Structure of the E6/E6AP/p53 complex required for HPV-mediated degradation of p53. Nature 2016, 529, 541–545.
- Wang, H.; Mo, P.; Ren, S.; Yan, C. Activating transcription factor 3 activates p53 by preventing E6-associated protein from binding to E6. J. Biol. Chem. 2010, 285, 13201–13210.
- Kooti, A.; Abuei, H.; Farhadi, A.; Behzad-Behbahani, A.; Zarrabi, M. Activating transcription factor 3 mediates apoptotic functions through a p53-independent pathway in human papillomavirus 18 infected HeLa cells. Virus Genes. 2022, 58, 88–97.
- Sood, V.; Sharma, K.B.; Gupta, V.; Saha, D.; Dhapola, P.; Sharma, M.; Sen, U.; Kitajima, S.; Chowdhury, S.; Kalia, M.; et al. ATF3 negatively regulates cellular antiviral signaling and autophagy in the absence of type I interferons. Sci. Rep. 2017, 7, 8789.
- Schindler, C.; Levy, D.E.; Decker, T. JAK-STAT signaling: From interferons to cytokines. J. Biol. Chem. 2007, 282, 20059–20063.
- Badu, P.; Pager, C.T. Activation of ATF3 via the Integrated Stress Response Pathway Regulates Innate Immune and Autophagy Processes to Restrict Zika Virus. bioRxiv 2023.
- Carthy, C.M.; Granville, D.J.; Watson, K.A.; Anderson, D.R.; Wilson, J.E.; Yang, D.; Hunt, D.W.; McManus, B.M. Caspase activation and specific cleavage of substrates after coxsackievirus B3-induced cytopathic effect in HeLa cells. J. Virol. 1998, 72, 7669–7675.
- Rothman, A.L. Immunity to dengue virus: A tale of original antigenic sin and tropical cytokine storms. Nat. Rev. Immunol. 2011, 11, 532–543.
- Li, J.; Yan, X.; Li, B.; Huang, L.; Wang, X.; He, B.; Xie, H.; Wu, Q.; Chen, L. Identification and validation of ferroptosis-related genes in patients infected with dengue virus: Implication in the pathogenesis of DENV. Virus Genes. 2023, 59, 377–390.
- Mishra, R.; Lahon, A.; Banerjea, A.C. Dengue Virus Degrades USP33-ATF3 Axis via Extracellular Vesicles to Activate Human Microglial Cells. J. Immunol. 2020, 205, 1787–1798.
- Henderson, A.; Holloway, A.; Reeves, R.; Tremethick, D.J. Recruitment of SWI/SNF to the human immunodeficiency virus type 1 promoter. Mol. Cell Biol. 2004, 24, 389–397.
- Tripathy, M.K.; Abbas, W.; Herbein, G. Epigenetic regulation of HIV-1 transcription. Epigenomics 2011, 3, 487–502.
- Verdin, E.; Paras, P., Jr.; Van Lint, C. Chromatin disruption in the promoter of human immunodeficiency virus type 1 during transcriptional activation. EMBO J. 1993, 12, 3249–3259.
- el Kharroubi, A.; Martin, M.A. cis-acting sequences located downstream of the human immunodeficiency virus type 1 promoter affect its chromatin structure and transcriptional activity. Mol. Cell Biol. 1996, 16, 2958–2966.
- Rabbi, M.F.; Saifuddin, M.; Gu, D.S.; Kagnoff, M.F.; Roebuck, K.A. U5 region of the human immunodeficiency virus type 1 long terminal repeat contains TRE-like cAMP-responsive elements that bind both AP-1 and CREB/ATF proteins. Virology 1997, 233, 235–245.
- Van Lint, C.; Amella, C.A.; Emiliani, S.; John, M.; Jie, T.; Verdin, E. Transcription factor binding sites downstream of the human immunodeficiency virus type 1 transcription start site are important for virus infectivity. J. Virol. 1997, 71, 6113–6127.
- Du, Y.; Ma, Z.; Zheng, J.; Huang, S.; Yang, X.; Song, Y.; Dong, D.; Shi, L.; Xu, D. ATF3 Positively Regulates Antibacterial Immunity by Modulating Macrophage Killing and Migration Functions. Front. Immunol. 2022, 13, 839502.
- Bekeredjian-Ding, I.; Stein, C.; Uebele, J. The Innate Immune Response Against Staphylococcus aureus. Curr. Top. Microbiol. Immunol. 2017, 409, 385–418.
- De Luca, A.; Pariano, M.; Cellini, B.; Costantini, C.; Villella, V.R.; Jose, S.S.; Palmieri, M.; Borghi, M.; Galosi, C.; Paolicelli, G.; et al. The IL-17F/IL-17RC Axis Promotes Respiratory Allergy in the Proximal Airways. Cell Rep. 2017, 20, 1667–1680.
- Choi, S.M.; McAleer, J.P.; Zheng, M.; Pociask, D.A.; Kaplan, M.H.; Qin, S.; Reinhart, T.A.; Kolls, J.K. Innate Stat3-mediated induction of the antimicrobial protein Reg3gamma is required for host defense against MRSA pneumonia. J. Exp. Med. 2013, 210, 551–561.
- Robinson, K.M.; Choi, S.M.; McHugh, K.J.; Mandalapu, S.; Enelow, R.I.; Kolls, J.K.; Alcorn, J.F. Influenza A exacerbates Staphylococcus aureus pneumonia by attenuating IL-1beta production in mice. J. Immunol. 2013, 191, 5153–5159.
- Ritchie, N.D.; Ritchie, R.; Bayes, H.K.; Mitchell, T.J.; Evans, T.J. IL-17 can be protective or deleterious in murine pneumococcal pneumonia. PLoS Pathog. 2018, 14, e1007099.
- Wonnenberg, B.; Jungnickel, C.; Honecker, A.; Wolf, L.; Voss, M.; Bischoff, M.; Tschernig, T.; Herr, C.; Bals, R.; Beisswenger, C. IL-17A attracts inflammatory cells in murine lung infection with P. aeruginosa. Innate Immun. 2016, 22, 620–625.
- Lu, Y.J.; Gross, J.; Bogaert, D.; Finn, A.; Bagrade, L.; Zhang, Q.; Kolls, J.K.; Srivastava, A.; Lundgren, A.; Forte, S.; et al. Interleukin-17A mediates acquired immunity to pneumococcal colonization. PLoS Pathog. 2008, 4, e1000159.
- Nguyen, C.T.; Luong, T.T.; Lee, S.; Kim, G.L.; Pyo, S.; Rhee, D.K. ATF3 provides protection from Staphylococcus aureus and Listeria monocytogenes infections. FEMS Microbiol. Lett. 2016, 363, fnw062.
- Nguyen, C.T.; Kim, E.H.; Luong, T.T.; Pyo, S.; Rhee, D.K. TLR4 mediates pneumolysin-induced ATF3 expression through the JNK/p38 pathway in Streptococcus pneumoniae-infected RAW 264.7 cells. Mol. Cells 2015, 38, 58–64.
- Park, J.M.; Ng, V.H.; Maeda, S.; Rest, R.F.; Karin, M. Anthrolysin O and other gram-positive cytolysins are toll-like receptor 4 agonists. J. Exp. Med. 2004, 200, 1647–1655.
- Nguyen, C.T.; Kim, E.H.; Luong, T.T.; Pyo, S.; Rhee, D.K. ATF3 confers resistance to pneumococcal infection through positive regulation of cytokine production. J. Infect. Dis. 2014, 210, 1745–1754.
- Annane, D.; Bellissant, E.; Cavaillon, J.M. Septic shock. Lancet 2005, 365, 63–78.
- Zhang, B.; Li, H.; Zhang, J.; Hang, Y.; Xu, Y. Activating transcription factor 3 protects alveolar epithelial type II cells from Mycobacterium tuberculosis infection-induced inflammation. Tuberculosis 2022, 135, 102227.
- Kumar, M.; Majumder, D.; Mal, S.; Chakraborty, S.; Gupta, P.; Jana, K.; Gupta, U.D.; Ghosh, Z.; Kundu, M.; Basu, J. Activating transcription factor 3 modulates the macrophage immune response to Mycobacterium tuberculosis infection via reciprocal regulation of inflammatory genes and lipid body formation. Cell Microbiol. 2020, 22, e13142.
- Peyron, P.; Vaubourgeix, J.; Poquet, Y.; Levillain, F.; Botanch, C.; Bardou, F.; Daffe, M.; Emile, J.F.; Marchou, B.; Cardona, P.J.; et al. Foamy macrophages from tuberculous patients’ granulomas constitute a nutrient-rich reservoir for M. tuberculosis persistence. PLoS Pathog. 2008, 4, e1000204.
- Russell, D.G.; Cardona, P.J.; Kim, M.J.; Allain, S.; Altare, F. Foamy macrophages and the progression of the human tuberculosis granuloma. Nat. Immunol. 2009, 10, 943–948.
- Glaser, N.; Stopper, H. Patulin: Mechanism of genotoxicity. Food Chem. Toxicol. 2012, 50, 1796–1801.
- Heussner, A.H.; Dietrich, D.R.; O’Brien, E. In vitro investigation of individual and combined cytotoxic effects of ochratoxin A and other selected mycotoxins on renal cells. Toxicol Vitr. 2006, 20, 332–341.
- Wu, T.S.; Liao, Y.C.; Yu, F.Y.; Chang, C.H.; Liu, B.H. Mechanism of patulin-induced apoptosis in human leukemia cells (HL-60). Toxicol. Lett. 2008, 183, 105–111.
- Kwon, O.; Soung, N.K.; Thimmegowda, N.R.; Jeong, S.J.; Jang, J.H.; Moon, D.O.; Chung, J.K.; Lee, K.S.; Kwon, Y.T.; Erikson, R.L.; et al. Patulin induces colorectal cancer cells apoptosis through EGR-1 dependent ATF3 up-regulation. Cell Signal 2012, 24, 943–950.
- Wu, Q.; Dohnal, V.; Kuca, K.; Yuan, Z. Trichothecenes: Structure-toxic activity relationships. Curr. Drug Metab. 2013, 14, 641–660.
More
Information
Subjects:
Immunology
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
805
Revisions:
2 times
(View History)
Update Date:
20 Feb 2024
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





