
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Chunmei Li | -- | 2351 | 2024-02-16 13:59:09 | | | |
| 2 | Lindsay Dong | -3 word(s) | 2348 | 2024-02-18 02:14:51 | | |
Video Upload Options
As an important barrier between the cytoplasm and the microenvironment of the cell, the cell membrane is essential for the maintenance of normal cellular physiological activities. An abnormal cell membrane is a crucial symbol of body dysfunction and the occurrence of variant diseases; therefore, the visualization and monitoring of biomolecules associated with cell membranes and disease markers are of utmost importance in revealing the biological functions of cell membranes. Due to their biocompatibility, programmability, and modifiability, DNA nanomaterials have become increasingly popular in cell fluorescence imaging in recent years. In addition, DNA nanomaterials can be combined with the cell membrane in a specific manner to enable the real-time imaging of signal molecules on the cell membrane, allowing for the real-time monitoring of disease occurrence and progression.
1. Introduction
2. Application of Cell Membrane Imaging
2.1. Monitoring Imaging Triggered by the Tumor Cell Microenvironment
2.1.1. ATP
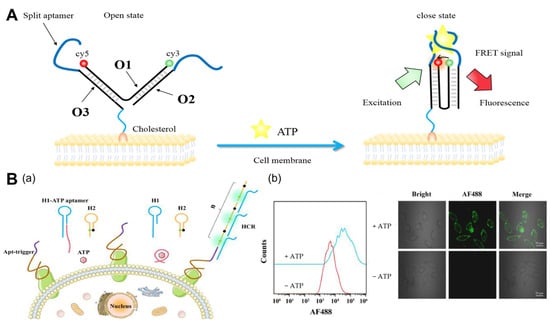
2.1.2. pH
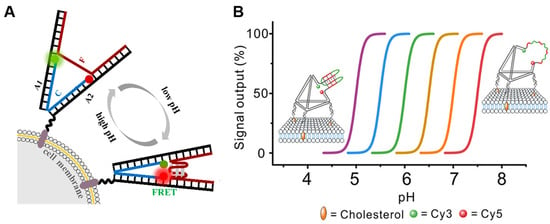
2.1.3. Metal Ions
2.2. Imaging of Cell Membrane Receptors
2.3. Imaging of Other Molecules
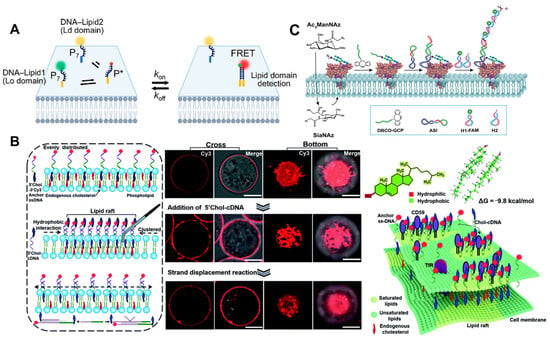
3. Conclusions
Although DNA nanomaterials are widely employed in cell surface fluorescence, the following obstacles must still be addressed. (1) Nowadays, most current imaging strategies involve imaging and analyzing only one or two targets, but using signal amplification techniques, such as HCR and CHA, to achieve the in situ accurate and highly sensitive imaging of multiple targets in the tumor microenvironment remains a challenge. (2) Aptamers must be designed specifically for each receptor. Currently, SELEX technology is used to screen all aptamers, and there are still receptors without aptamers. (3) The majority of DNA nanomaterials are attached to the cell membrane via covalent or noncovalent methods [57][58] but are susceptible to endocytosis. Even though some scientists have developed polymer molecular skeletons to increase the anchoring time of probes in cell membranes, long-term stability remains a problem. Monitoring intra- and extracellular signaling, cellular morphology, and structural changes requires prolonged in situ imaging, especially for slow-response events such as apoptosis. The creation of probes that can remain attached to the cell membrane for extended durations without being endocytosed remains an area of active research. (4) DNA nanomaterials are used not only for the fluorescence imaging of cell membranes but also for the functional regulation of cell membrane receptors. How to realize the integration of long-term imaging and the functional regulation of cell membranes should also be investigated. (5) FRET is dependent not only on the donor (D)–acceptor (A) separation distance and energetic resonance (i.e., D–A spectral overlap), but also on the orientation of the D emission and A absorption transition dipole moments [59][60].
References
- Yu, X.M.; Sha, L.J.; Liu, Q.; Zhao, Y.Y.; Fang, H.; Cao, Y.; Zhao, J. Recent Advances in Cell Membrane Camouflage-based Biosensing Application. Biosens. Bioelectron. 2021, 194, 113623.
- Feng, L.; Li, J.; Sun, J.; Wang, L.; Fan, C.; Shen, J. Recent Advances of DNA Nanostructure-Based Cell Membrane Engineering. Adv. Healthc. Mater. 2021, 10, 2001718.
- Qian, R.C.; Zhou, Z.R.; Guo, W.J.; Wu, Y.T.; Yang, Z.L.; Lu, Y. Cell Surface Engineering Using DNAzymes: Metal Ion Mediated Control of Cell-Cell Interactions. J. Am. Chem. Soc. 2021, 143, 5737–5744.
- Reading, E.; Hall, Z.; Martens, C.; Haghighi, T.; Findlay, H.; Ahdash, Z.; Politis, A.; Booth, P.J. Interrogating Membrane Protein Conformational Dynamics within Native Lipid Compositions. Angew. Chem. Int. Ed. 2017, 56, 15654–15657.
- Shi, P.; Wang, Y. Synthetic DNA for Cell-Surface Engineering. Angew. Chem. Int. Ed. 2021, 60, 11580–11591.
- Wang, H.; Wang, Y.S.; Wan, Y.Q.; Shang, J.H.; Wang, Q.; Jiang, Y.Q.; Liu, X.Q.; Wang, F. A Cellular Membrane-Confined Concatenate DNA Circuit for Non-Invasive Cell Modulation with High Accuracy and Efficiency. Adv. Funct. Mater. 2023, 33, 2302708.
- Zhang, S.Y.; Li, L.; Chen, J.Y.; Chen, Z.Q.; Zhang, W.L.; Lu, H.B. Quantitative Imaging of Gd Nanoparticles in Mice Using Benchtop Cone-Beam X-ray Fluorescence Computed Tomography System. Int. J. Mol. Sci. 2019, 20, 2315.
- Wang, Z.L.; Xue, X.D.; He, Y.X.; Lu, Z.W.; Jia, B.; Wu, H.; Yuan, Y.; Huang, Y.; Wang, H.; Lu, H.W.; et al. Novel Redox-Responsive Polymeric Magnetosomes with Tunable Magnetic Resonance Property for In Vivo Drug Release Visualization and Dual-Modal Cancer Therapy. Adv. Funct. Mater. 2018, 28, 1802158.
- Miao, X.Y.; Mao, R.; You, Y.J.; Zhou, H.C.; Qiu, C.; Li, X.H.; Chen, Z.H.; Ren, J.; Chen, M.H.; Wang, P.; et al. Intracolic Ultrasound Molecular Imaging: A Novel Method for Assessing Colonic Tumor Necrosis Factor-α Expression in Inflammatory Bowel Disease. Mol. Med. 2021, 27, 119.
- Wozniak, M.; Ploska, A.; Siekierzycka, A.; Dobrucki, L.W.; Kalinowski, L.; Dobrucki, I.T. Molecular Imaging and Nanotechnology-Emerging Tools in Diagnostics and Therapy. Int. J. Mol. Sci. 2022, 23, 2658.
- Li, F.; Li, J.; Dong, B.; Wang, F.; Fan, C.; Zuo, X. DNA Nanotechnology-empowered Nanoscopic Imaging of Biomolecules. Chem. Soc. Rev. 2021, 50, 5650–5667.
- Xu, Y.W.; Lv, Z.; Yao, C.; Yang, D.Y. Construction of Rolling Circle Amplification-based DNA Nanostructures for Biomedical Applications. Biomater. Sci. 2022, 10, 3054–3061.
- Meng, H.M.; Liu, H.; Kuai, H.L.; Peng, R.Z.; Mo, L.T.; Zhang, X.B. Aptamer-integrated DNA Nanostructures for Biosensing, Bioimaging and Cancer Therapy. Chem. Soc. Rev. 2016, 45, 2583–2602.
- Wang, K.; Gao, H.Q.; Zhang, Y.W.; Yan, H.Y.; Si, J.H.; Mi, X.Y.; Xia, S.A.; Feng, X.Q.; Liu, D.B.; Kong, D.L.; et al. Highly Bright AIE Nanoparticles by Regulating the Substituent of Rhodanine for Precise Early Detection of Atherosclerosis and Drug Screening. Adv. Mater. 2022, 34, 2106994.
- Ma, W.; Zhan, Y.; Zhang, Y.; Mao, C.; Xie, X.; Lin, Y. The Biological Applications of DNA Nanomaterials: Current Challenges and Future Directions. Signal Transduct. Target. Ther. 2021, 6, 351.
- Nishikawa, M.; Tan, M.M.; Liao, W.Q.; Kusamori, K. Nanostructured DNA for the Delivery of Therapeutic Agents. Adv. Drug Deliv. Rev. 2019, 147, 29–36.
- Zhang, J.J.; Li, F.; Yang, D.Y. DNA: From Carrier of Genetic Information to Polymeric Materials. Trans. Tianjin Univ. 2019, 25, 301–311.
- Nicolson, F.; Ali, A.; Kircher, M.F.; Pal, S. DNA Nanostructures and DNA-Functionalized Nanoparticles for Cancer Theranostics. Adv. Sci. 2020, 7, 2001669.
- Wang, D.X.; Wang, J.; Wang, Y.X.; Du, Y.C.; Huang, Y.; Tang, A.N.; Cui, Y.X.; Kong, D.M. DNA Nanostructure-based Nucleic Acid Probes: Construction and Biological Applications. Chem. Sci. 2021, 12, 7602–7622.
- Song, L.; Zhuge, Y.; Zuo, X.L.; Li, M.; Wang, F. DNA Walkers for Biosensing Development. Adv. Sci. 2022, 9, 2200327.
- Wu, X.H.; Wu, T.T.; Liu, J.B.; Ding, B.Q. Gene Therapy Based on Nucleic Acid Nanostructure. Adv. Healthc. Mater. 2020, 9, 2001046.
- Wang, H.M.; Luo, D.; Wang, H.; Wang, F.; Liu, X.Q. Construction of Smart Stimuli-Responsive DNA Nanostructures for Biomedical Applications. Chem. Eur. J. 2021, 27, 3929–3943.
- Li, C.; Tebo, A.G.; Gautier, A. Fluorogenic Labeling Strategies for Biological Imaging. Int. J. Mol. Sci. 2017, 18, 1473.
- Zhou, Z.X.; Lu, Z.R. Molecular Imaging of the Tumor Microenvironment. Adv. Drug Deliv. Rev. 2017, 113, 24–48.
- Zhang, Y.; Cheng, F.; Liang, R.; Shuai, X.J.; Nie, K.H.; Li, J.; Zeng, Q.L.; Huang, C.Z.; Li, C.M. In Situ Activation of the Receptor Aggregation for Cell Apoptosis by an AI-Au Intelligent Nanomachine via Tumor Extracellular Acidity. ACS Appl. Mater. Interfaces 2023, 15, 32262–32271.
- Park, H.; Saravanakumar, G.; Kim, J.; Lim, J.; Kim, W.J. Tumor Microenvironment Sensitive Nanocarriers for Bioimaging and Therapeutics. Adv. Healthc. Mater. 2021, 10, 2000834.
- Wang, Y.; Tang, L.H.; Li, Z.H.; Lin, Y.H.; Li, J.H. In Situ Simultaneous Monitoring of ATP and GTP using a Graphene Oxide Nanosheet-based Sensing Platform in Living Cells. Nat. Protoc. 2014, 9, 1944–1955.
- Yuan, J.; Deng, Z.W.; Liu, H.; Li, X.F.; Li, J.B.; He, Y.; Qing, Z.H.; Yang, Y.J.; Zhong, S.A. Cell-Surface-Anchored Ratiometric DNA Nanoswitch for Extracellular ATP Imaging. ACS Sens. 2019, 4, 1648–1653.
- Zong, H.; Wang, X.X.; Mu, X.J.; Wang, J.G.; Sun, M.T. Plasmon-Enhanced Fluorescence Resonance Energy Transfer. Chem. Rec. 2019, 19, 818–842.
- Yang, X.J.; Cui, M.R.; Li, X.L.; Chen, H.Y.; Xu, J.J. A Self-Powered 3D DNA Walker with Programmability and Signal-amplification for Illuminating MicroRNA in Living Cells. Chem. Commun. 2020, 56, 2135–2138.
- Bi, S.; Yue, S.Z.; Zhang, S.S. Hybridization Chain Reaction: A Versatile Molecular Tool for Biosensing, Bioimaging, and Biomedicine. Chem. Soc. Rev. 2017, 46, 4281–4298.
- Li, L.; Li, S.P.; Wang, J.; Wen, X.H.; Yang, M.; Chen, H.; Guo, Q.; Wang, K. Extracellular ATP-activated Hybridization Chain Reaction for Accurate and Sensitive Detection of Cancer Cells. Chin. Chem. Lett. 2023, 34, 108399.
- Zeng, S.; Liu, D.; Li, C.; Yu, F.; Fan, L.; Lei, C.; Huang, Y.; Nie, Z.; Yao, S. Cell-Surface-Anchored Ratiometric DNA Tweezer for Real-Time Monitoring of Extracellular and Apoplastic pH. Anal. Chem. 2018, 90, 13459–13466.
- Li, Z.Z.; Wang, J.B.; Zhou, Z.X.; O’Hagan, M.P.; Willner, I. Gated Transient Dissipative Dimerization of DNA Tetrahedra Nanostructures for Programmed DNAzymes Catalysis. ACS Nano 2022, 16, 3625–3636.
- Zhao, C.; Zhang, L.; Hu, Y.; Nie, C.; Chen, T.T.; Chu, X. Simultaneous Imaging and Visualizing the Association of Survivin mRNA and Telomerase in Living Cells by Using a Dual-Color Encoded DNA Nanomachine. Anal. Chem. 2023, 95, 1498–1504.
- Liu, J.X.; Li, W.W.; Li, R.S.; Yin, X.Z.; He, S.L.; Hu, J.Q.; Ruan, S.C. Programmable DNA Framework Sensors for In Situ Cell-Surface pH Analysis. Anal. Chem. 2021, 93, 12170–12174.
- Aksentijevic, D.; Karlstaedt, A.; Basalay, M.V.; O’Brien, B.A.; Sanchez-Tatay, D.; Eminaga, S.; Thakker, A.; Tennant, D.A.; Fuller, W.; Eykyn, T.R.; et al. Intracellular Sodium Elevation Reprograms Cardiac Metabolism. Nat. Commun. 2020, 11, 4337.
- Deng, Z.; Gao, P.; Liu, H.; He, Y.; Zhong, S.; Yang, Y. Cell-Surface-Anchored DNA Sensors for Simultaneously Monitoring Extracellular Sodium and Potassium Levels. Anal. Chem. 2021, 93, 16432–16438.
- Anees, P.; Saminathan, A.; Rozmus, E.R.; Di, A.K.; Malik, A.B.; Delisle, B.P.; Krishnan, Y. Detecting Organelle-specific Activity of Potassium Channels with A DNA Nanodevice. Nat. Biotechnol. 2023, 1546–1696.
- Vodnala, S.K.; Eil, R.; Kishton, R.J.; Sukumar, M.; Yamamoto, T.N.; Ha, N.H.; Lee, P.H.; Shin, M.; Patel, S.J.; Yu, Z.Y.; et al. T Cell Stemness and Dysfunction in Tumors are Triggered by a Common Mechanism. Science 2019, 363, eaau0135.
- Eil, R.; Vodnala, S.K.; Clever, D.; Klebanoff, C.A.; Sukumar, M.; Pan, J.H.; Palmer, D.C.; Gros, A.; Yamamoto, T.N.; Patel, S.J.; et al. Ionic Immune Suppression within the Tumour Microenvironment Limits T Cell Effector Function. Nature 2016, 537, 539–543.
- Cheng, F.; Jiang, Y.J.; Kong, B.; Lin, H.R.; Shuai, X.J.; Hu, P.P.; Gao, P.F.; Zhan, L.; Huang, C.Z.; Li, C.M. Multi-Catcher Polymers Regulate the Nucleolin Cluster on the Cell Surface for Cancer Therapy. Adv. Healthc. Mater. 2023, 12, 2300102.
- Wang, L.P.; Liang, H.; Sun, J.; Liu, Y.C.; Li, J.Y.; Li, J.Y.; Li, J.; Yang, H.H. Bispecific Aptamer Induced Artificial Protein-Pairing: A Strategy for Selective Inhibition of Receptor Function. J. Am. Chem. Soc. 2019, 141, 12673–12681.
- Nishida, N.; Osawa, M.; Takeuchi, K.; Imai, S.; Stampoulis, P.; Kofuku, Y.; Ueda, T.; Shimada, I. Functional Dynamics of Cell Surface Membrane Proteins. J. Magn. Reson. 2014, 241, 86–96.
- He, F.; Wang, M.X.; Wang, J.Y.; Wang, H.H.; Nie, Z. An Extracellular miRNA-Responsive Artificial Receptor via Dynamic DNA Nano-assembly for Biomarker-Driven Therapy. Angew. Chem. Int. Ed. 2023, 135, e202305227.
- Li, L.L.; Lv, W.Y.; Xu, Y.T.; Li, Y.F.; Li, C.M.; Huang, C.Z. DNA Logic Nanodevices for the Sequential Imaging of Cancer Markers through Localized Catalytic Hairpin Assembly Reaction. Anal. Chem. 2022, 94, 4399–4406.
- Fang, Y.Y.; Li, Y.T.; Li, Y.Y.; He, R.; Zhang, Y.; Zhang, X.B.; Liu, Y.; Ju, H.X. In Situ Protease Secretion Visualization and Metastatic Lymph Nodes Imaging via a Cell Membrane-Anchored Upconversion Nanoprobe. Anal. Chem. 2021, 93, 7258–7265.
- Yang, L.L.; Meng, L.Y.; Song, J.Y.; Xiao, Y.; Wang, R.W.; Kang, H.Z.; Han, D. Dynamic Colloidal Nanoparticle Assembly Triggered by Aptamer-receptor Interactions on Live Cell Membranes. Chem. Sci. 2019, 10, 7466–7471.
- Jia, H.R.; Zhu, Y.X.; Duan, Q.Y.; Wu, F.G. Cell Surface-localized Imaging and Sensing. Chem. Soc. Rev. 2021, 50, 6240–6277.
- Ma, W.J.; Sun, H.H.; Chen, B.A.; Jia, R.C.; Huang, J.; Cheng, H.; He, X.X.; Huang, M.M.; Wang, K.M. Engineering a Facile Aptamer “Molecule-Doctor” with Hairpin- Contained I-Motif Enables Accurate Imaging and Killing of Cancer Cells. Anal. Chem. 2021, 93, 14552–14559.
- Sezgin, E.; Levental, I.; Mayor, S.; Eggeling, C. The Mystery of Membrane Organization: Composition, Regulation and Roles of Lipid Rafts. Nat. Rev. Mol. Cell Biol. 2017, 18, 361–374.
- Cheng, X.L.; Smith, J.C. Biological Membrane Organization and Cellular Signaling. Chem. Rev. 2019, 119, 5849–5880.
- Ali, A.A.; Bagheri, Y.; Tian, Q.; You, M.X. Advanced DNA Zipper Probes for Detecting Cell Membrane Lipid Domains. Nano Lett. 2022, 22, 7579–7587.
- Lingwood, D.; Simons, K. Lipid Rafts As a Membrane-Organizing Principle. Science 2010, 327, 46–50.
- Sun, L.L.; Su, Y.Y.; Wang, J.G.; Xia, F.; Xu, Y.; Li, D. DNA Nanotweezers for Stabilizing and Dynamically Lighting up a Lipid Raft on Living Cell Membranes and The Activation of T Cells. Chem. Sci. 2020, 11, 1581–1586.
- Li, J.Y.; Liu, S.Y.; Sun, L.Q.; Li, W.; Zhang, S.Y.; Yang, S.; Li, J.; Yang, H.H. Amplified Visualization of Protein-Specific Glycosylation in Zebrafish via Proximity-Induced Hybridization Chain Reaction. J. Am. Chem. Soc. 2018, 140, 16589–16595.
- Xun, K.Y.; Pei, K.; Liu, X.J.; Peng, X.Y.; Du, Y.L.; Qiu, L.P.; Tan, W.H. Cell-Membrane-Anchored DNA Nanoplatform for Programming Cellular Interactions. J. Am. Chem. Soc. 2019, 141, 18013–18020.
- Chandra, R.A.; Douglas, E.S.; Mathies, R.A.; Bertozzi, C.R.; Francis, M.B. Programmable Cell Adhesion Encoded by DNA Hybridization. Angew. Chem. Int. Ed. 2006, 45, 896–901.
- Mathur, D.; Díaz, S.A.; Hildebrandt, N.; Pensack, R.D.; Yurke, B.; Biaggne, A.; Li, L.; Melinger, J.S.; Ancona, M.G.; Knowlton, W.B.; et al. Pursuing Excitonic Energy Transfer with Programmable DNA-based Optical Breadboards. Chem. Soc. Rev. 2023, 52, 7848–7948.
- Cervantes-Salguero, K.; Biaggne, A.; Youngsman, J.M.; Ward, B.M.; Kim, Y.C.; Li, L.; Hall, J.A.; Knowlton, W.B.; Graugnard, E.; Kuang, W. Strategies for Controlling the Spatial Orientation of Single Molecules Tethered on DNA Origami Templates Physisorbed on Glass Substrates: Intercalation and Stretching. Int. J. Mol. Sci. 2022, 23, 7690.





