Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Nickolai A. Barlev | -- | 2977 | 2023-12-19 13:14:20 | | | |
| 2 | Lindsay Dong | Meta information modification | 2977 | 2023-12-22 01:59:07 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Szewczyk-Roszczenko, O.; Barlev, N.A. Role of p53 in Nanoparticle-Based Therapy for Cancer. Encyclopedia. Available online: https://encyclopedia.pub/entry/52927 (accessed on 15 May 2026).
Szewczyk-Roszczenko O, Barlev NA. Role of p53 in Nanoparticle-Based Therapy for Cancer. Encyclopedia. Available at: https://encyclopedia.pub/entry/52927. Accessed May 15, 2026.
Szewczyk-Roszczenko, Olga, Nikolai A. Barlev. "Role of p53 in Nanoparticle-Based Therapy for Cancer" Encyclopedia, https://encyclopedia.pub/entry/52927 (accessed May 15, 2026).
Szewczyk-Roszczenko, O., & Barlev, N.A. (2023, December 19). Role of p53 in Nanoparticle-Based Therapy for Cancer. In Encyclopedia. https://encyclopedia.pub/entry/52927
Szewczyk-Roszczenko, Olga and Nikolai A. Barlev. "Role of p53 in Nanoparticle-Based Therapy for Cancer." Encyclopedia. Web. 19 December, 2023.
Copy Citation
p53 is arguably one of the most important tumor suppressor genes in humans. Due to the paramount importance of p53 in the onset of cell cycle arrest and apoptosis, the p53 gene is found either silenced or mutated in the vast majority of cancers. Furthermore, activated wild-type p53 exhibits a strong bystander effect, thereby activating apoptosis in surrounding cells without being physically present there. For these reasons, p53-targeted therapy that is designed to restore the function of wild-type p53 in cancer cells seems to be a very appealing therapeutic approach. Systemic delivery of p53-coding DNA or RNA using nanoparticles proved to be feasible both in vitro and in vivo.
p53
gene therapy
nanoparticles
bystander effect
apoptosis
1. Introduction
To function properly, multicellular organisms require an exquisitely organized system of quality control. Both endocrine and paracrine regulatory mechanisms define the fate of cells thereby achieving their synchronous propagation or apoptosis. The pivotal element in the system of detection and elimination of defective cells from an organism is the product of the TP53 tumor suppressor gene [1]. The p53 protein belongs to the family of p53 proteins that involve two other members, p73 and p63. Although all members of the family are bona fide tumor suppressors, p73 and p63 play more important roles in the development of multicellular organisms than in oncogenesis [2].
Being a transcriptional factor, p53 promotes the expression of a number of genes involved in the activation of cell cycle arrest and apoptosis [3][4]. In addition, p53 is also able to suppress the transcription of certain genes by augmenting the expression of its target non-coding genes (lncRNAs and microRNAs) [5][6][7] (Figure 1).
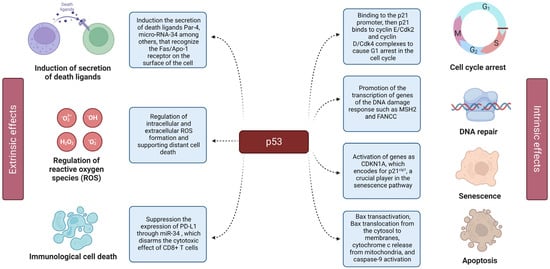
Figure 1. Extrinsic and intrinsic effects of p53. The left panel displays the extrinsic effects of p53, including its by-stander effect. The right panel shows the details of p53 action on the intracellular level (intrinsic effects), including cell cycle arrest, senescence, and apoptosis.
To avoid unnecessary activation of cell cycle arrest and cell death under normal conditions, the intracellular level of p53 is kept in check by post-translational modifications, among which ubiquitinoylation plays a critical role [8][9]. The principal p53-specific E3 ubiquitin ligase is Mdm2 (HDM2 in humans) [10][11], which targets the p53 lysine residues located in the C-terminus and targets the protein for degradation in proteasomes [12]. Importantly, Mdm2 can attenuate the activity of p53 both directly, via the binding and ubiquitylating of the latter and, indirectly, by affecting the interacting partners of p53 [11][13] or the degradation machinery [14][15]. In addition to ubiquitinoylation, other covalent post-translational modifications including the neddylation, sumoylation, and methylation of certain lysins promote p53 inactivation or its proteasomal degradation [16][17][18].
However, when cells experience virtually any type of stress, the signaling cues activate p53 via a sequential attachment of posttranscriptional modifications that include phosphorylation, methylation, and acetylation [19]. Contrary to ubiquitinoylation, acetylation and methylation generally promote p53 stabilization by outcompeting ubiquitinoylation of the same lysins [8]. Moreover, there is a crosstalk between post-translational modifications, and they can regulate each other’s functions in a positive or negative manner [20].
Another important feature of p53 is its promotion of bystander effects in neighboring cells, forcing the execution of a suicidal program in those cells despite the fact that they have not experienced the harmful consequences of stress themselves [21]. This feature of p53 is particularly important for successful elimination of tumor cells by p53-containing nanoparticles.
Thus, p53 acts as an integrator of signaling cascades directed against almost all forms of cell stress and contributes to the maintenance of higher-order structures—tissues and organs. In doing so, p53 employs several modes of action: as a transcriptional activator [22] and as a scaffold for protein–protein interactions [23]. The latter, however, poses the main threat to antitumor defense mechanisms: p53 mutations not only interfere with the normal functioning of defense mechanisms against malignant transformation, but, on the contrary, can lead to an imbalance in signaling cascades, resulting in the emergence of positive feedback loops and, ultimately, leading to cancer [24]. Based on this, many mutations that occur in the TP53 gene transform it from a tumor suppressor into an oncogene. Indeed, TP53 mutations are observed in more than 50% of human cancers [25]. It is important to note that transcriptionally incompetent mutant forms of p53 fail to induce the expression of Mdm2, thereby indirectly stabilizing mutant p53 at the protein level.
Given the fact that p53 can exist in cancer cells in two mutually opposite forms (wt vs. GOF, respectively), the task of designing an effective p53 therapy becomes very challenging. In general, the aim of successful p53-targeted anticancer therapy should be two-fold: (1) activating the p53 molecule in the event of its wild-type conformation and (2) either neutralizing the mutant form of p53 or restoring its wild-type conformation by specific compounds. Apparently, looked at from this perspective, it seems that degradation of p53 by E3 ligases, in particular Mdm2, seems to be the most promising axis for pharmacological intervention in wild-type p53-expressing tumors. In addition to a number of chemical inhibitors of Mdm2 that have already been developed over the years [26][27][28], several new inhibitors have entered clinical trials showing preliminarily promising results [29][30]. However, it should be noted that, so far, the success of Mdm2 inhibitors in clinical settings has been rather limited. There are a few reasons for this: in addition to robust side effects such as thrombocytopenia [31][32] and gastrointestinal toxicity associated with Mdm2 experimental drugs [30], cancer cells were able to adapt to the prolonged therapy by enhancing the expression of other p53-targeting E3 ligases (Pirh2, WWP1, etc.) [33], higher efflux of Mdm2 inhibitors, and their increased metabolic degradation [11].
2. p53 in Nanoparticle-Based Gene Therapy for Cancer
Delivering the p53 gene in its wild-type (WT) form to cancer cells via gene therapy is an intriguing approach to restoring p53 activity. Among the delivery approaches is the application of adenoviruses as carriers, which has been used effectively in a formulation called gendicine [34]. However, adenoviruses are not always suitable carriers, despite demonstrating encouraging outcomes in preclinical investigations, thus gene therapy may be limited by the absence of an effective delivery mechanism in the late stages of clinical trials [35]. The use of nanoparticles (NPs), which increase the stability of the given particles and exhibit higher absorption by cancer cells, might offer a method for more effectively delivering the p53 gene [36]. Multiple requirements must be accomplished for a p53 gene delivery system to be effective. The vector should not be immunogenic or toxic, permitting numerous injections if necessary. Because the p53 protein is effective but unstable, persistent gene expression within tumors is required for long-lasting therapeutic benefits. Since transfected cells may impact the tumor’s nontransfected areas, substantial levels of transfection may not be required to slow the growth of the tumor. This is because p53 exhibits strong bystander effects discussed in the previous chapter [37]. There are also attempts to deliver peptide activators of the p53 protein into cells, which may provide an alternative to gene therapy [38].
2.1. Liposomal Vectors
DDC is a delivery system that is based on DOTAP, DOPE, and Cholesterol. DOTAP (dioleoyltrimethylamino propane) is a cationic liposome, whereas DOPE (1,2-dioeoyl-3-phosphophatidylethanolamine) and cholesterol diminish fibrinogen, prothrombin, and vitamin K affinity for the lipid surface. DDC effectively transported plasmid DNA into ovarian cancer cells. High levels of p53 WT mRNA and protein expression were detected in OVCAR-3 cells because of transfection with the liposome-complexed p53 gene. Compared to control cells, cancer cells transfected with DDC/p53-EGFP complexes showed significant growth suppression. The apoptotic pathway was reinstated in ovarian cancer cells after wild-type p53 function was restored. The volumes of tumors in nude mice were considerably decreased by more than 60% in comparison to the control group after the inoculation of DDC/p53-EGFP complexes [39].
2.2. Polymer NPs
A breast cancer cell line MDA-MB-435S treated with D, L-lactide-co-glycolide (PLGA (PLGA is an FDA-approved biocompatible and biodegradable polymer with a wide range of disintegration times and customizable mechanical properties)) nanoparticles containing p53 WT DNA experienced a persistent antiproliferative impact, whose strength increased with time. Plasmid DNA-containing nanoparticles were created using a multiple emulsion–solvent evaporation process. Researchers tracked the intracellular trafficking of the nanoparticles and the nanoparticle-entrapped DNA. They measured the amounts of p53 mRNA over time to comprehend the mechanism of sustained gene expression with nanoparticles. When compared to cells of the MDA-MB-435S transfected with bare p53 WT DNA or p53 WT DNA complexed with a commercially available transfecting agent (Lipofectamine), cells transfected with p53 WT DNA-loaded nanoparticles showed a persistent and much higher antiproliferative impact. The study’s findings point to the possibility that p53 WT DNA-loaded nanoparticles could be helpful in the treatment of breast cancer [40] and other malignancies linked to p53 gene mutations [41].
2.3. Metallic NPs
Because of their scalable design, functional variety, control over particle size and surface, and capacity to distinguish between different types of cells via surface coatings, gold nanoparticles are currently considered to be a highly perspective drug delivery technology. For the delivery of p53 WT to ovarian cancer cells, an EGFR (epidermal growth factor receptor)-targeted method based on gold nanoparticles was created. EGFR is overexpressed on the surface of many malignancies, including up to 90% of ovarian tumors. Thus, for specific targeting, cetuximab (C225), an FDA-approved monoclonal antibody that targets EGFR was used to deliver the p53 coding DNA to ovarian cancer cells. Targeting ovarian malignancies in vitro (SKOV-3 cell line) and in vivo (SKOV-3 xenograft mice) has shown encouraging results using a sophisticated gold nanoconjugate system (Au-C225-p53) including gold nanoparticles, cetuximab, and the pCMVp53 plasmid. Although xenograft mice in this study demonstrated its usefulness, it is still too early to say if this medication delivery system will advance to clinical trials [42].
2.4. Other NPs
An integrative method based on the production of nanoparticulated carriers in conjunction with the supercoiled (sc) isoform purification of a p53 tumor suppressor expressing plasmid was developed. Under mild conditions, the sc topoisoform is recovered with great purity and structural stability. Furthermore, naked sc pDNA was encased within chitosan nanoparticles by ionotropic gelation to improve protection and transfection efficiency. The technique’s gentle particle production conditions allowed for a high encapsulation efficiency for sc pDNA.
Short amphipathic peptides that combine with mRNA to generate stable, neutral nanoparticles are the foundation of ADGN technology. On 20 different cancer cell lines harboring various types of p53 mutations (null, deletion, nonsense, and missense), ADGN-531 nanoparticles containing full-length p53-mRNA were assessed. On colorectal SW403 (p53-deleted) and osteosarcoma SaOs2 (p53 null) mouse xenografts, the in vivo effectiveness of IV-administered ADGN-531 nanoparticles was assessed. On PARPi resistant SUM-149PT and OVCAR-8 cells as well as on PARPi-sensitive MDA-MB436 cells, sensitivity to veliparib (PARPi) was assessed in vitro after ADGN-531 treatment.
The ARF-mimicking MDM2-trapping peptide nanoparticles (Mtrap NPs) which can reassemble, were developed to treat p53-positive tumors. This approach is based on the fact that the alternative reading frame (ARF) protein sequesters away Mdm2 in cytoplasm, thereby protecting p53 from the Mdm2-mediated degradation. The findings on U2OS, A549, SK-BR-3, and H1299 cell lines revealed that Mtrap NPs respond to MDM2 and build a nanofiber structure which traps Mdm2. Thus, Mtra NPs suppress p53-wild-type cancers by stabilizing and activating p53 via inactivation of MDM2.
3. Impact of NPs on the p53 Protein
Although the use of nanoparticles has unquestionable benefits in terms of more effective medicine delivery, we must acknowledge the risks of using nanoparticles. Nanoparticles are not innocuous to the body on their own, which should inspire researchers to work towards developing a safer and more effective technology (summarized in Table 1).
Table 1. Interaction of NPs and p53.
| Type of NPs | In Vitro/ In Vivo |
Tissue/Cell Line | Effect | Reference |
|---|---|---|---|---|
| Al2O3 | In vivo | Sub-brain regions of rats | Decreased expression of cyclin D1, bcl-2, Mdm2, and phospho-Rb and increased expression of p53, p21, Bax, and Rb | [43] |
| Ag | In vitro | GC1415, NCI-N87, and MKN45 | Increased p53 expression, inhibition of STAT3 | [44] |
| In vitro | HCT116 | Increased transcription of p53, p21, and caspases (3,8,9), decreased amount of AKT and NF-κB | [45] | |
| CuO | In vitro/ Ex vivo |
K562 and peripheral blood mononuclear cell | Increase in Bax/Bcl-2 ratio, upregulation of p53, and ROS production | [46] |
| Fe3O4 | In vitro | HepG2, A549, IMR-90 | Induction of ROS, upregulation of p53, and caspases 3 and 9 | [47] |
| Pt | In vitro | IMR-90, U251 | Upregulation of p53 and p21, DNA damage | [48] |
| Si | In vitro | HUVECs | Activation of c-Jun, p53, caspase-3, and NF-κB, increased Bax expression and suppression Bcl-2 | [49] |
| TiO2 | Ex vivo | peripheral blood lymphocytes | Accumulation of p53 and activation of DNA damage checkpoint kinases | [50] |
| In vitro | PC12 | ROS and JNK/p53 mediated apoptosis and causing. G2/M arrest by the activation of p53/p21 pathway |
[51] | |
| V2O5 | In vitro | B16F10, A549, and PANC1 | Impaired angiogenesis, increased ROS, overexpression of p53 | [52] |
| Zn | In vitro | HepG2 | ROS generation, DNA damage, activation of p53 and p38 | [53] |
4. Toxicity of NPs
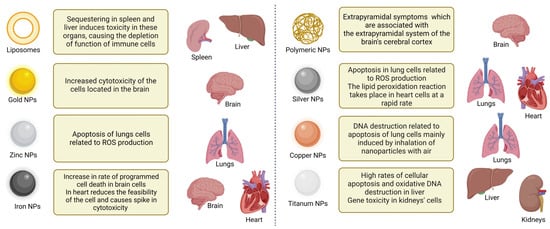
Metal, lipid, and protein NPs, the three types of NPs commonly utilized in medical delivery, have all been subjected to thorough evaluation of their toxicity profiles (Figure 2). Protein-based NPs, for example, have been linked to hepatotoxicity and nephrotoxicity, whereas metal-based NPs have been linked to increased oxidative stress and may penetrate the cell nucleus [54]. Given the expanding use of NPs in drug delivery, regulatory authorities are rightfully raising concerns about the toxicity of nanocarriers in organisms. Thus, it is critical to identify the gaps in such toxicity studies, including the toxicity of polymeric NPs [55].

Figure 2. Specific organ toxicity of NPs. Shown are various types of NPs and their effects on different organs.
In line with this, Adams and colleagues conducted one of the first studies investigating liposomal toxicity in vivo in 1977. Five different types of liposomes with various net surface charges were produced and injected into mouse brains in this investigation. Liposomes containing 45 mol% lecithin or dipalmitoyl lecithin were the least harmful, while others, e.g., containing 9% stearylamine, caused serious cognitive impairment and respiratory failure. These results are important to take into account since liposomes are generally thought to be pharmacologically inactive with low toxicity, yet their safety is highly dependent on the model, exposure period, dose, and/or surface features [56].
Silver deposition in the skin and eyes is one of the toxic post-exposure effects in humans. Silver-doped prosthetic restorations color the tissues surrounding the prosthesis blackish blue. After cessation of exposure, these effects are reversible; elimination primarily occurs through the liver and kidney. According to findings generated using the rat liver cell line BRL 3A, the ability of cells cultured with silver nanoparticles was significantly reduced. The researchers discovered a change in mitochondrial membrane potential, a drop in glutathione concentration, and a considerable increase in reactive oxygen species concentration.
The mechanisms that regulate copper metabolism are easily disturbed, resulting in oxidative damage. The most common is a cascade effect that impairs liver function or disrupts mitochondrial respiration. Copper particle accumulation has been reported in plants and animals. The respiratory system is the primary route of toxicity caused by copper nanoparticles. Long-term exposure can result in dose-dependent pneumonia and lung cell damage. CuO NPs induce high toxicity in human lung epithelial cell A549, leukemia cell HL60, human breast cancer cell MCF-7, and hepatocellular carcinoma cell HepG2. Nanoparticles that enter the body through the lungs can also affect the cardiovascular system. One hypothesis is that inhaled particles promote inflammation and release inflammatory cytokines into the bloodstream. Another theory holds that nanoparticles are freely discharged into the circulatory system via lung blood veins.
5. Limitations
There are several significant obstacles that must be overcome for p53-based gene therapy to gain clinical relevance. The first problem stems from the nature of mutations in the p53 gene: the most frequent mutations in p53 found in cancer cells exhibit dominant negative effects towards p53 WT, thereby reducing or even completely blunting its tumor suppressive activity. Since GOF mutants of p53 are not degraded by Mdm2 due to their structural features and their inability to induce the Mdm2 expression, they incorporate in cells at high levels and often outcompete the exogenously delivered p53 [57]. Perhaps not surprisingly, it was found that in a population of cancer cells with mixed p53 genotypes, cells bearing p53 WT responded better to p53 therapy compared to those cells that contained mutant p53 [58][59].
One of the solutions to this problem is to re-engineer the p53 molecule by either increasing its expression levels and/or alter its structure to avoid the interaction with mutant p53 [60][61]. For example, chimeric proteins resulting from the fusion of mitochondrial targeting signals (MTS) and functional domains of p53 (e.g., the DNA binding domain) are capable of rapidly (but transiently) inducing apoptosis through their action inside the mitochondria. In this way, the MTS-p53 fusion proteins can escape the interaction with endogenous p53 located in the nucleus and cytoplasm of cancer cells [62]. This is a new mechanism of apoptosis that is not associated with transcriptional activation of p53 target genes in the nucleus. Another variant of the same approach is represented by the p53-Bad chimeric protein, which is a fusion product of monomeric p53 with the proapoptotic factor Bad and contains an integrated MTS [63]. Both chimeric proteins (p53-MTS and p53-Bad) were shown to successfully overcome the dominant negative effect of mutant p53. This new gene therapy approach showed the ability to trigger apoptosis in cancer cell lines with various p53 mutations, indicating that it could be used as a therapy regardless of p53 status.
Other enhanced p53 gene therapies include creation of “super-p53,” in which the p53 tetramerization domain (TD) is replaced with alternative TD. The tetramerization domain is required for effective transcriptional activity of p53. However, at the same time this domain allows wild-type p53 to hetero-oligomerize with dominant negative p53 mutants. To circumvent this undesirable phenomenon, the TD of wild-type p53 was replaced with an engineered leucine zipper that assembles into a four-stranded coiled coil. The ability of the engineered zipper to drive tetramerization was critical to p53 function, since p53 molecules engineered only to dimerize have been shown to be poor tumor suppressors [64].
References
- Lane, D.P. P53, Guardian of the Genome. Nature 1992, 358, 15–16.
- Rozenberg, J.M.; Zvereva, S.; Dalina, A.; Blatov, I.; Zubarev, I.; Luppov, D.; Bessmertnyi, A.; Romanishin, A.; Alsoulaiman, L.; Kumeiko, V.; et al. The P53 Family Member P73 in the Regulation of Cell Stress Response. Biol. Direct 2021, 16, 23.
- Thomas, A.F.; Kelly, G.L.; Strasser, A. Of the Many Cellular Responses Activated by TP53, which Ones Are Critical for Tumour Suppression? Cell Death Differ. 2022, 29, 961–971.
- Riley, T.; Sontag, E.D.; Chen, P.; Levine, A.J. Transcriptional Control of Human P53-Regulated Genes. Nat. Rev. Mol. Cell Biol. 2008, 9, 402–412.
- Barlev, N.A.; Sayan, B.S.; Candi, E.; Okorokov, A.L. The microRNA and P53 Families Join Forces against Cancer. Cell Death Differ. 2010, 17, 373–375.
- Parfenyev, S.; Singh, A.; Fedorova, O.A.; Daks, A.; Kulshreshtha, R.; Barlev, N.A. Interplay between P53 and Non-Coding RNAs in the Regulation of EMT in Breast Cancer. Cell Death Dis. 2021, 12, 17.
- Hermeking, H. MicroRNAs in the P53 Network: Micromanagement of Tumour Suppression. Nat. Rev. Cancer 2012, 12, 613–626.
- Marouco, D.; Garabadgiu, A.V.; Melino, G.; Barlev, N.A.; Barlev, N.A. Lysine-Specific Modifications of P53: A Matter of Life and Death? Oncotarget 2013, 4, 1556–1571.
- Liu, Y.; Tavana, O.; Gu, W. P53 Modifications: Exquisite Decorations of the Powerful Guardian. J. Mol. Cell Biol. 2019, 11, 564–577.
- Konopleva, M.; Martinelli, G.; Daver, N.; Papayannidis, C.; Wei, A.; Higgins, B.; Ott, M.; Mascarenhas, J.; Andreeff, M. MDM2 Inhibition: An Important Step Forward in Cancer Therapy. Leukemia 2020, 34, 2858–2874.
- Klein, A.M.; de Queiroz, R.M.; Venkatesh, D.; Prives, C. The Roles and Regulation of MDM2 and MDMX: It Is Not Just about P53. Genes Dev. 2021, 35, 575–601.
- Morgunkova, A.; Barlev, N.A. Lysine Methylation Goes Global. Cell Cycle 2006, 5, 1308–1312.
- Lezina, L.; Aksenova, V.; Fedorova, O.; Malikova, D.; Shuvalov, O.; Antonov, A.V.; Tentler, D.; Garabadgiu, A.V.; Melino, G.; Barlev, N.A. KMT Set7/9 Affects Genotoxic Stress Response via the Mdm2 Axis. Oncotarget 2015, 6, 25843–25855.
- Petukhov, A.; Ag, M.; Moiseeva, T.N.; Tn, M.; Barlev, N.A. Role of Proteasomes in Transcription and Their Regulation by Covalent Modifications. Front. Biosci. 2008, 13, 7184–7192.
- Sdek, P.; Ying, H.; Chang, D.L.F.; Qiu, W.; Zheng, H.; Touitou, R.; Allday, M.J.; Allday, M.J.; Xiao, Z.-X.J. MDM2 Promotes Proteasome-Dependent Ubiquitin-Independent Degradation of Retinoblastoma Protein. Mol. Cell 2005, 20, 699–708.
- Stindt, M.H.; Carter, S.A.; Vigneron, A.M.; Ryan, K.M.; Vousden, K.H. MDM2 Promotes SUMO-2/3 Modification of P53 to Modulate Transcriptional Activity. Cell Cycle 2011, 10, 3176–3188.
- Abida, W.M.; Nikolaev, A.; Zhao, W.; Zhang, W.; Gu, W.; Abida, W.M.; Nikolaev, A.; Zhao, W.; Zhang, W.; Gu, W. FBXO11 Promotes the Neddylation of P53 and Inhibits Its Transcriptional Activity. J. Biol. Chem. 2007, 282, 1797–1804.
- Rada, M.; Vasileva, E.; Lezina, L.; Marouco, D.; Antonov, A.V.; Macip, S.; Melino, G.; Barlev, N.A. Human EHMT2/G9a Activates P53 through Methylation-Independent Mechanism. Oncogene 2017, 36, 922–932.
- Daks, A.; Shuvalov, O.; Fedorova, O.; Parfenyev, S.; Simon, H.-U.; Barlev, N.A. Barlev Methyltransferase Set7/9 as a Multifaceted Regulator of ROS Response. Int. J. Biol. Sci. 2023, 19, 2304–2318.
- Ivanov, G.S.; Ivanova, T.; Kurash, J.; Ivanov, A.; Chuikov, S.; Gizatullin, F.; Herrera-Medina, E.M.; Rauscher, F.; Reinberg, D.; Barlev, N.A. Methylation-Acetylation Interplay Activates P53 in Response to DNA Damage. Mol. Cell. Biol. 2007, 27, 6756–6769.
- Xu, M.; Kumar, D.; Kumar, D.; Kumar, D.; Srinivas, S.; DeTolla, L.J.; Yu, S.F.; Stass, S.A.; Mixson, A.J. Parenteral Gene Therapy with P53 Inhibits Human Breast Tumors in Vivo through a Bystander Mechanism without Evidence of Toxicity. Hum. Gene Ther. 1997, 8, 177–185.
- Pfister, N.T.; Prives, C. Transcriptional Regulation by Wild-Type and Cancer-Related Mutant Forms of P53. Cold Spring Harb. Perspect. Med. 2017, 7, a026054.
- Brady, C.A.; Attardi, L.D. P53 at a Glance. J. Cell Sci. 2010, 123, 2527–2532.
- Bellazzo, A.; Sicari, D.; Valentino, E.; Del Sal, G.; Collavin, L. Complexes Formed by Mutant P53 and Their Roles in Breast Cancer. Breast Cancer 2018, 10, 101–112.
- Bykov, V.J.N.; Eriksson, S.E.; Bianchi, J.; Wiman, K.G. Targeting Mutant P53 for Efficient Cancer Therapy. Nat. Rev. Cancer 2018, 18, 89–102.
- Vassilev, L.T. MDM2 Inhibitors for Cancer Therapy. Trends Mol. Med. 2007, 13, 23–31.
- Davidovich, P.; Aksenova, V.; Petrova, V.; Tentler, D.; Orlova, D.; Smirnov, S.; Gurzhiy, V.; Okorokov, A.L.; Garabadzhiu, A.; Melino, G.; et al. Discovery of Novel Isatin-Based P53 Inducers. ACS Med. Chem. Lett. 2015, 6, 856–860.
- Fallatah, M.M.J.; Law, F.V.; Chow, W.A.; Kaiser, P. Small-Molecule Correctors and Stabilizers to Target P53. Trends Pharmacol. Sci. 2023, 44, 274–289.
- Dumbrava, E.E.; Johnson, M.L.; Tolcher, A.W.; Shapiro, G.I.; Thompson, J.A.; El-Khoueiry, A.B.; Vandross, A.L.; Kummar, S.; Parikh, A.R.; Munster, P.N.; et al. First-in-Human Study of PC14586, a Small Molecule Structural Corrector of Y220C Mutant P53, in Patients with Advanced Solid Tumors Harboring a TP53 Y220C Mutation. J. Clin. Oncol. 2022, 40 (Suppl. S16), 3003.
- Gounder, M.; Bauer, T.; Schwartz, G.; Weise, A.; LoRusso, P.; Kumar, P.; Tao, B.; Hong, Y.; Patel, P.; Lu, Y.; et al. A First-in-Human Phase I Study of Milademetan, an MDM2 Inhibitor, in Patients With Advanced Liposarcoma, Solid Tumors, or Lymphomas. J. Clin. Oncol. 2023, 41, 1714–1724.
- Mahfoudhi, E.; Lordier, L.; Marty, C.; Pan, J.; Roy, A.; Roy, L.; Rameau, P.; Abbes, S.; Debili, N.; Raslova, H.; et al. P53 Activation Inhibits All Types of Hematopoietic Progenitors and All Stages of Megakaryopoiesis. Oncotarget 2016, 7, 31980–31992.
- Khurana, A.; Shafer, D.A. MDM2 Antagonists as a Novel Treatment Option for Acute Myeloid Leukemia: Perspectives on the Therapeutic Potential of Idasanutlin (RG7388). OncoTargets Ther. 2019, 12, 2903–2910.
- Daks, A.; Petukhov, A.; Fedorova, O.; Shuvalov, O.; Merkulov, V.; Vasileva, E.; Antonov, A.; Barlev, N.A. E3 Ubiquitin Ligase Pirh2 Enhances Tumorigenic Properties of Human Non-Small Cell Lung Carcinoma Cells. Genes Cancer 2016, 7, 383–393.
- Lundstrom, K. Viral Vectors in Gene Therapy: Where Do We Stand in 2023? Viruses 2023, 15, 698.
- Zeimet, A.G.; Marth, C. Why Did P53 Gene Therapy Fail in Ovarian Cancer. Lancet Oncol. 2003, 4, 415–422.
- Szewczyk, O.K.; Roszczenko, P.; Czarnomysy, R.; Bielawska, A.; Bielawski, K. An Overview of the Importance of Transition-Metal Nanoparticles in Cancer Research. Int. J. Mol. Sci. 2022, 23, 6688.
- Rezaei, M.; Esmailzadeh, A.; Shanei, A. Bystander Effect of Therapeutic Ultrasound in the Presence of Cisplatin: An in Vitro Study on Human Melanoma Cells. J. Biomed. Phys. Eng. 2023, 13, 433–442.
- He, W.; Yan, J.; Li, Y.; Yan, S.; Wang, S.; Hou, P.; Lu, W. Resurrecting a P53 Peptide Activator—An Enabling Nanoengineering Strategy for Peptide Therapeutics. J. Control. Release 2020, 325, 293–303.
- Kim, C.-K.; Choi, E.J.; Choi, E.-J.; Choi, S.-H.; Park, J.-S.; Haider, K.H.; Ahn, W.S. Enhanced P53 Gene Transfer to Human Ovarian Cancer Cells Using the Cationic Nonviral Vector, DDC. Gynecol. Oncol. 2003, 90, 265–272.
- Marvalim, C.; Datta, A.; Lee, S.C. Role of P53 in Breast Cancer Progression: An Insight into P53 Targeted Therapy. Theranostics 2023, 13, 1421–1442.
- Prabha, S.; Labhasetwar, V. Nanoparticle-Mediated Wild-Type P53 Gene Delivery Results in Sustained Antiproliferative Activity in Breast Cancer Cells. Mol. Pharm. 2004, 1, 211–219.
- Kotcherlakota, R.; Vydiam, K.; Srinivasan, D.J.; Mukherjee, S.; Roy, A.; Kuncha, M.; Rao, T.N.; Sistla, R.; Gopal, V.; Patra, C.R. Restoration of P53 Function in Ovarian Cancer Mediated by Gold Nanoparticle-Based EGFR Targeted Gene Delivery System. ACS Biomater. Sci. Eng. 2019, 5, 3631–3644.
- Liu, H.; Zhang, W.; Fang, Y.; Yang, H.; Tian, L.; Li, K.; Lai, W.; Bian, L.; Lin, B.; Liu, X.; et al. Neurotoxicity of Aluminum Oxide Nanoparticles and Their Mechanistic Role in Dopaminergic Neuron Injury Involving P53-Related Pathways. J. Hazard. Mater. 2020, 392, 122312.
- Huang, D.; Wang, J.; Zhou, S.; Zhang, T.; Cai, J.; Liu, Y. Ag Nanoparticles Green-Mediated by Scrophularia Striata Aqueous Extract Induce Apoptosis via P53 and Signal Transducer and Activator of Transcription 3 Signaling Pathways in Gastric Cancer Cells. Inorg. Chem. Commun. 2023, 155, 110942.
- Satapathy, S.R.; Mohapatra, P.; Preet, R.; Das, D.; Sarkar, B.; Choudhuri, T.; Wyatt, M.D.; Kundu, C.N. Silver-Based Nanoparticles Induce Apoptosis in Human Colon Cancer Cells Mediated through P53. Nanomed. Nanotechnol. Biol. Med. 2013, 8, 1307–1322.
- Shafagh, M.; Rahmani, F.; Delirezh, N. CuO nanoparticles induce cytotoxicity and apoptosis in human K562 cancer cell line via mitochondrial pathway, through reactive oxygen species and P53. Iran. J. Basic Med. Sci. 2015, 18, 993–1000.
- Ahamed, M.; Alhadlaq, H.A.; Khan, M.A.M.; Akhtar, M.J. Selective Killing of Cancer Cells by Iron Oxide Nanoparticles Mediated through Reactive Oxygen Species via P53 Pathway. J. Nanopart. Res. 2013, 15, 1225.
- Asharani, P.V.; Xinyi, N.; Hande, M.P.; Valiyaveettil, S. DNA Damage and P53-Mediated Growth Arrest in Human Cells Treated with Platinum Nanoparticles. Nanomed. Nanotechnol. Biol. Med. 2010, 5, 51–64.
- Liu, X.; Sun, J. Endothelial Cells Dysfunction Induced by Silica Nanoparticles through Oxidative Stress via JNK/P53 and NF-κB Pathways. Biomaterials 2010, 31, 8198–8209.
- Kang, S.J.; Kim, B.M.; Lee, Y.-J.; Chung, H.W. Titanium Dioxide Nanoparticles Trigger P53-Mediated Damage Response in Peripheral Blood Lymphocytes. Environ. Mol. Mutagen. 2008, 49, 399–405.
- Wu, J.; Sun, J.; Xue, Y. Involvement of JNK and P53 Activation in G2/M Cell Cycle Arrest and Apoptosis Induced by Titanium Dioxide Nanoparticles in Neuron Cells. Toxicol. Lett. 2010, 199, 269–276.
- Xi, W.; Tang, H.; Liu, Y.; Liu, C.; Gao, Y.; Cao, A.; Liu, Y.; Chen, Z.; Wang, H. Cytotoxicity of Vanadium Oxide Nanoparticles and Titanium Dioxide-coated Vanadium Oxide Nanoparticles to Human Lung Cells. J. Appl. Toxicol. 2020, 40, 567–577.
- Sharma, V.; Anderson, D.; Kumar, A.; Dhawan, A. Zinc Oxide Nanoparticles Induce Oxidative DNA Damage and ROS-Triggered Mitochondria Mediated Apoptosis in Human Liver Cells (HepG2). Apoptosis 2012, 17, 852–870.
- Roszczenko, P.; Szewczyk, O.K.; Czarnomysy, R.; Bielawski, K.; Bielawska, A. Biosynthesized Gold, Silver, Palladium, Platinum, Copper, and Other Transition Metal Nanoparticles. Pharmaceutics 2022, 14, 2286.
- Yao, Y.; Zang, Y.; Qu, J.; Tang, M.; Zhang, T. The Toxicity Of Metallic Nanoparticles On Liver: The Subcellular Damages, Mechanisms, And Outcomes. Int. J. Nanomed. 2019, 14, 8787–8804.
- Inglut, C.T.; Sorrin, A.J.; Kuruppu, T.; Vig, S.; Cicalo, J.; Ahmad, H.; Huang, H.-C. Immunological and Toxicological Considerations for the Design of Liposomes. Nanomaterials 2020, 10, 190.
- Dolma, L.; Muller, P.A.J. GOF Mutant P53 in Cancers: A Therapeutic Challenge. Cancers 2022, 14, 5091.
- Munisamy, M.; Mukherjee, N.; Thomas, L.; Pham, A.T.; Shakeri, A.; Zhao, Y.; Kolesar, J.; Rao, P.P.N.; Rangnekar, V.M.; Rao, M. Therapeutic Opportunities in Cancer Therapy: Targeting the P53-MDM2/MDMX Interactions. Am. J. Cancer Res. 2021, 11, 5762–5781.
- Zhang, C.; Liu, J.; Xu, D.; Zhang, T.; Hu, W.; Feng, Z. Gain-of-Function Mutant P53 in Cancer Progression and Therapy. J. Mol. Cell Biol. 2020, 12, 674–687.
- Matissek, K.J.; Mossalam, M.; Okal, A.; Lim, C.S. The DNA Binding Domain of P53 Is Sufficient To Trigger a Potent Apoptotic Response at the Mitochondria. Mol. Pharm. 2013, 10, 3592–3602.
- Okal, A.; Matissek, K.J.; Matissek, S.J.; Price, R.; Salama, M.E.; Janát-Amsbury, M.M.; Lim, C.S. Re-Engineered P53 Activates Apoptosis in Vivo and Causes Primary Tumor Regression in a Dominant Negative Breast Cancer Xenograft Model. Gene Ther. 2014, 21, 903–912.
- Matissek, K.J.; Okal, A.; Mossalam, M.; Lim, C.S. Delivery of a Monomeric P53 Subdomain with Mitochondrial Targeting Signals from Pro-Apoptotic Bak or Bax. Pharm. Res. 2014, 31, 2503–2515.
- Lu, P.; Redd Bowman, K.E.; Brown, S.M.; Joklik-Mcleod, M.; Vander Mause, E.R.; Nguyen, H.T.N.; Lim, C.S. P53-Bad: A Novel Tumor Suppressor/Proapoptotic Factor Hybrid Directed to the Mitochondria for Ovarian Cancer Gene Therapy. Mol. Pharm. 2019, 16, 3386–3398.
- Waterman, M.J.; Waterman, J.L.; Halazonetis, T.D. An Engineered Four-Stranded Coiled Coil Substitutes for the Tetramerization Domain of Wild-Type P53 and Alleviates Transdominant Inhibition by Tumor-Derived P53 Mutants. Cancer Res. 1996, 56, 158–163.
More
Information
Subjects:
Cell Biology
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
599
Revisions:
2 times
(View History)
Update Date:
22 Dec 2023
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





