Circular RNAs (circRNAs) are single-stranded closed non-coding RNA molecules that are aberrantly expressed and produce tumor-specific gene signatures in human cancers. They exert biological functions by acting as transcriptional regulators, microRNA sponges, and protein scaffolds, regulating the formation of protein–RNA complexes and, ultimately, regulating gene expression. Triple-negative breast cancer (TNBC) is one of the most aggressive cancers of the mammary gland and has a poor prognosis. Studies of circRNAs in TNBC are limited but have demonstrated these molecules’ pivotal roles in cell proliferation, invasion, metastasis, and resistance to chemo/radiotherapy, suggesting that they could be potential prognostic biomarkers and novel therapeutic targets.
1. Metastasis in Breast Cancer
Metastasis is the most devastating feature of malignant neoplasia and is responsible for a large number of cancer-related deaths. Disseminated disease occurs when highly aggressive cancer cells detach from a primary tumor, migrating to distant tissues and organs and achieving complete adaptation to a new niche to initiate the growth of a secondary malignant tumor. Breast cancer metastasis occurs mainly in the bones, lungs, liver, and brain, which drastically shortens the overall survival of patients
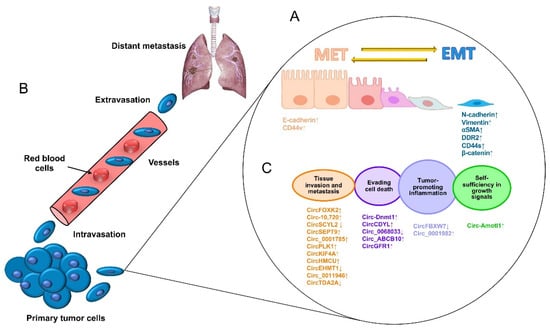
[1]. The escape of tumor cells from a primary aggressive tumor occurs as a series of steps beginning with local invasion and followed by the entry of tumor cells into blood vessels (intravasation), the survival of cancer cells in blood circulation, and the outflow of blood vessels (extravasation) in a specialized microenvironment, which serves as a previously prepared niche for secondary tumor growth (
Figure 1A)
[2]. Therefore, the metastatic process is not a disorderly process; on the contrary, it takes place as a series of well-coordinated cellular and molecular events driven by dynamic, genetic, and epigenetic programs. At the cellular level, alterations in the tight contacts between cancer cells and extracellular matrix (ECM) proteins occur, characterized by a morphology change known as an epithelial–mesenchymal transition (EMT), which allows for cancer cells to acquire enhanced migratory abilities, to detach from the ECM, and to initiate the invasion of proximal tissues. Diverse specialized transmembrane proteins drive the EMT, and alterations in their functions may result in chemo/radiotherapy resistance
[3]. For instance, E-cadherin is a cell adhesion protein that keeps cells together and whose low expression is related to EMT activation, whereas high expression of its family member N-cadherin is associated with the loss of cell adhesions mediated by integrins, which are mechano-transduction proteins that sense the ECM (
Figure 1B). During the invasion of nearby tissues, the ECM is degraded by the actions of metalloproteinases (MMPs) and the urokinase plasminogen activator (uPA) system
[4]. Then, coordinated tumor cell migration is carried out with additional cellular processes, including the activation of the “angiogenic switch”, characterized by a change in the local balance of proangiogenic factors, such as the vascular endothelial growth factor (VEGF), and antiangiogenic factors, including thrombospondins and endostatins, among others, which indicates that angiogenesis is a key mechanism for tumor growth and invasive behavior
[4]. Interestingly, more recent studies have been focused on the identification of potential regulatory non-coding RNAs such as microRNAs, long non-coding RNAs, and circular RNAs (circRNAs) as molecular drivers of metastasis and therapy resistance, which may lead to the identification of potential biomarkers and novel therapeutic targets
[5][6].
Figure 1. The process of the metastasis of tumor cells. (
A) Metastasis begins with paracrine signals that activate genes to initiate the switch from the mesenchymal–epithelial transition (MET) to the EMT status, changing the interactions of cancer cell junctions and extracellular matrix proteins, including cadherins, vimentin, CD44, and beta-catenin, among others. (
B) Next, cancer cells break off from the primary tumor and move through the wall of blood vessels (intravasation) to travel through the bloodstream in search of a new site to grow (extravasation). The cells leave the bloodstream to settle in tissues surrounding the blood vessels from which they emerged. (
C) Diverse circular RNAs have been identified as regulators of genes that participate in cell proliferation, apoptosis inflammation, apoptosis, tissue invasion, and metastasis. ↓ Downregulated. ↑ Upregulated.
2. Circular RNAs
In 1976, circRNAs were observed in the light of confocal microscopy, by Sanger et al., for the first time. At first, these sequences were thought to be viroids due to their circular shape. Over time and with technological advances, they went from being classified as rare types of RNA that resulted from splicing artifacts or genetic rearrangements to real regulatory elements in eukaryotic cells
[7]. CircRNAs are derived from precursor-messenger RNAs (pre-mRNAs), which are transcribed by RNA polymerase II and characterized by circular shapes that result from continuous, covalently closed single-stranded loops. These unique molecular structures make circRNAs resistant to RNAse-R and exonuclease digestion; hence, they exhibit high RNA stability
[8]. CircRNAs are classified into three types, as follows, according to their structures and mechanisms of circularization: (i) exon circRNAs (ecircRNAs), (ii) circular intron RNAs (ciRNAs), and (iii) exon–intron circRNAs (ElciRNAs)
[8]. Consisting of a single exon or multiple numbers of exons and formed through a shearing process called “head-to-tail” or “backsplicing,” ecircRNAs make up more than 80% of circRNAs and mostly exist in the cytoplasm. EIciRNAs are predominantly located in the nucleus and circularized in the form of retention introns between exons
[9]. There are currently three models of circular RNAs production, namely, (i) intron-pairing-driven circularization, (ii) RNA-binding protein (RBP)-dependent circularization, and (iii) loop-driven circularization, which have been recognized to produce EcircRNAs and EIciRNAs (
Figure 2A)
[6]. Recent studies have found that circRNAs are exported to cytoplasm through a mechanism that is dependent on the length of a closed RNA molecule. The UAP56/DDX39B complex is required for the export of long circRNAs, whereas the URH49/DDX39A proteins are required for the transport of short circRNAs (<400 nt). Interestingly, the depletion of the UAP56 or URH49 factors can affect stationary levels of cytoplasmic circRNAs (
Figure 2B).
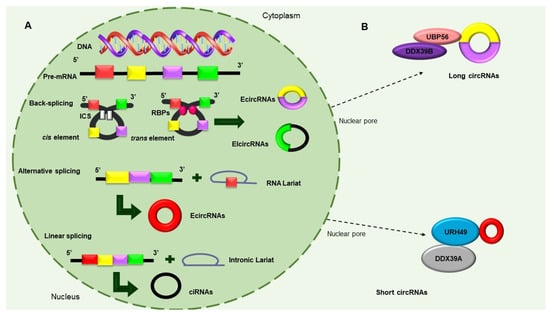
Figure 2. Biosynthesis of circular RNAs. (
A) Precursor mRNA (pre-mRNA) consensus sequences, such as the cis elements known as intronic complementary sequences (ICSs) and flanking circRNA-forming exons, in addition to trans elements, including some RBPs, participate in the formation of circRNAs. The backsplicing process originates EcircRNAs and circular RNAs with introns (EIcircRNAs) via the donor splicer and the receptor-splicing site through base-pairing interactions between the ICSs or the dimerization of the RBPs. Alternative splicing circularization allows the covalent attachment of an upstream exon and a downstream exon that generates a lariat RNA precursor. When an intronic lariat is formed via the canonical splicing pathway, it generates a linear lariat intron that contains skipped exons that eventually undergo reverse splicing. (
B) The export of circRNAs from the nucleus to the cytoplasm is dependent on their size. The UBP56 protein is responsible for the export of long circRNAs (>1300 nucleotides), while URH49 exports short circRNAs (<400 nt)
[10][11][12].
3. Roles of Circular RNAs in Cancer Cells
Functions in RNA Polymerase II Transcription and Splicing
A better-described function of circRNAs is that of being microRNA sponges because they can bind to those tiny regulatory ncRNAs; thus, they are also dubbed endogenous competitors
[13]. CircRNAs have both oncogenic and tumor-suppressing properties in human cancer depending on the functions of the kidnapped microRNA (
Figure 3A). Interestingly, some circRNAs have been discovered to also exert functions in transcriptional regulation through interaction with specific DNA loci to form RNA–DNA hybrids, which will cause a transcriptional pause and the recruitment of the splicing factors, as in the cases of circSEP3 and circSMARCA5
[14][15]. RBPs are important to the regulation of gene expression and splicing. Notably, circEIF3J and circPAIP2 have been detected to associate with the RNA polymerase II protein and enhance gene expression. Another example is intronic ci-ankrd52, which upregulates transcriptions driven by RNA polymerase II
[16]. Likewise, the EIF3J and PAIP2 circRNAs interact with U1 snRNP, which, in turn, interacts with RNA polymerase II in the promoter regions of their parental genes, increasing transcriptions (
Figure 3C)
[17]. Another example is circ-UBR5, which regulates RNA splicing when it binds to regulators like QKI, NOVA1, and the U1 proteins (
Figure 3B)
[18]. CircRNAs also may contain small open-reading frames (ORFs) to encode polypeptides that mediate diverse cellular activities. An example is circ-ZNF609, which binds to polysomes in murine and human myoblasts to produce a 30 kDa protein in a splicing-dependent and 5´cap-independent fashion (
Figure 3D)
[19].
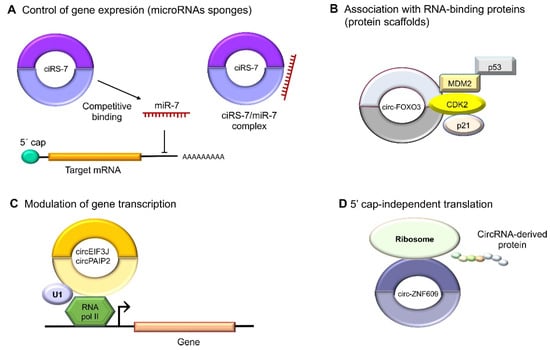
Figure 3. Molecular functions of circular RNAs. (
A) CircRNAs act indirectly on gene expression as microRNA sponges: for example, oncogenic ciRS-7 (CDR1as) competitively binds to tumor suppressor miR-7, since it contains miRNA response elements (MREs). (
B) CircRNAs can also bind to RBPs such as circ-FOXO3, which interacts with MDM2 (murine double-minute 2) to degrade p53 by ubiquitination, forming a ternary complex with CDK2 and p21. (
C) The nuclear circRNAs that conserve the intronic sequences of their parental genes, circEIF3J and circPAIP2, to improve those genes’ cis expressions could associate with RNA polymerase II, which would improve their functions in a U1 snRNP-dependent manner. (
D) Multiple circRNAs contain internal ribosome entry sites (IRESs), which suggests that they can be translated into peptides or proteins in a 5′ cap-independent manner, as in the case of circ-ZNF609, the translation of which plays an important role in myogenesis.
4. Circular RNAs Functions in Triple-Negative Breast Cancer
CircRNAs Sponge Relevant MicroRNAs Involved in Tumor Progression and Metastasis
At least 60% of the human transcriptome belongs to ncRNAs that participate in biological processes as differential regulators with pathological impacts
[20]. The role of circRNAs in breast cancer, especially in the triple-negative subtype, has recently been demonstrated. Some studies have shown that circRNAs act mainly through the kidnapping of microRNAs, which regulates important oncogenic and tumor-suppressor genes involved in the regulation of cancer hallmarks. Moreover, breast cancer metastasis has been associated with alterations in the expressions of multiple circRNAs and microRNAs. On the other hand, circBCBM1 was detected in the brain as a promoter of metastasis through the modulation of the miR-125a/BRD4 axis
[21]. Another relevant circular RNA is circZEB1, which has been markedly overexpressed in TNBC tissues and cell lines, promoting proliferation, and reducing apoptosis through the miR-448/eEF2K axis
[22].
Similarly, the overexpression of circBACH2 has been reported to facilitate the EMT, activating cancer cell invasion and migration as well as proliferation through the binding of both tumor suppressors miR-186-5p and miR-548c-3p, thus promoting the expression of the oncogenic chemokine receptor CXC type 4, or CXCR4. Moreover, circBACH2 was found expressed on the surfaces of various types of cancer cells; thus, its depletion has resulted in the suppression of the malignant progression of TNBC cells
[23].
Intriguingly, Li et al. discovered that a circRNA derived from the HER2 gene, circ-HER2, was expressed in TNBC even when its classification indicated the absence of the HER2 receptor. The expression of the circ-HER2 contributed to homo/hetero EGFR/HER3 dimerization, which originated the sustained phosphorylation of AKT that promotes proliferation, invasion, and metastasis. The circ-HER2 expression correlated with poor prognosis in patients
[24].
Remarkably, circRNAs have been described to not only promote the metastasis of cancer cells but possibly also play the opposite role: strong tumor suppression. For example, the overexpression of circCDYL has promoted apoptosis and inhibited cell proliferation through regulation of the miR-190a-3p/TP53INP1 axis, therefore upregulating the tumor-suppressing TP53INP1 protein in TNBC
[25].
Finally, circRNA molecules could be derived from diverse genomic regions, as in the case of hsa_circ_0091074, which is generated from a genomic region of the X-specific inactive transcript (XIST) and binds to miR-1297, partially blocking the antitumor action of its target microRNA and resulting in the overexpression of the TAZ protein and positive regulation of the cell cycle in TNBC cells
[26].
5. Circular RNAs Exported in Exosomes in TNBC
In recent decades, exosomes have become an important topic of study given their important roles in the regulation of cancer hallmarks. Exosomes are nanometric vesicles (40–100 nm in diameter) of endocytic origin that share a similar topology and lipid composition with the plasma membrane. From inside, a wide variety of cargo molecules, such as nucleic acids, proteins, and enzymes capable of modulating cellular activities in recipient cells through the transfer of functional genetic information, can be found
[27]. Exosome biogenesis is mediated by two pathways that classify the proteins that they contain. The endosomal sorting complex transport (ESCRT)-dependent pathway is the best-characterized mechanism, involving four large protein complexes and more than 30 proteins. The ESCRT-independent pathway involves the inhibition of a neutral sphingomyelinase (nSMase) that is necessary to hydrolyze sphingomyelin and originate ceramide
[28]. Exosomes are formed by budding from the membranes of multivesicular bodies (MVBs) from late endosomes: invaginations, called intraluminal vesicles (ILVs), that contain cytosolic components. MVBs can fuse with lysosomes for their degradation or may follow the endocytic pathway for the generation of exosomes
[29].
Exosomal circRNAs are known to play an important role in cancer biology, as they can be taken up by neighboring or distant cells and affect many aspects of the physiological and pathological conditions of recipient cells. Recently, exosomes were discovered to be enriched with circRNA molecules, as demonstrated by Li et al. More than 1000 circRNAs have been identified in human serum exosomes. Notably, the circRNA content in exosomes is regulated by changes in the levels of the associated microRNAs in the producer cells
[30]. On the other hand, Yang et al. found that circPSMA1 was overexpressed in serum exosomes from TNBC patients.
6. Circular RNAs Regulation of Resistance to Chemo/Radiotherapy
The expression of circRNAs related to the doxorubicin (DOX) response has been reported in TNBC. DOX is a cytotoxic anthracycline antibiotic that commonly generates resistance
[31]. The high expression of the circular RNA dubbed circUBE2D2 has been associated with a poor prognosis of TNBC patients in the advanced stages of the disease, as well as with lymph node metastases and resistance to DOX. CircUBE2D2 acts as a molecular sponge for miR-512-3p, which has a role as a suppressor of various tumors by acting on cell division cycle associated protein 3 (CDCA3)
[32]. On the other hand, the hsa_circ_0092276/miR-384/ATG7 axis promotes autophagy and DOX resistance through the overexpression of hsa_circ_0092276
[33].
Remarkably, circRNAs that perform the reverse function have been studied. For instance, the exogenous overexpression of circKDM4C has inhibited DOX resistance as well as proliferation and metastasis in breast cancer. CircKDM4C is a sponge for miR-548p, and the overexpression of this miRNA reverses the attenuation of malignant phenotypes
[34].
Several studies have corroborated the participation of circRNAs in the modulation of the proapoptotic and antiapoptotic proteins involved in resistance to chemotherapy. The overexpression of circAMOTL1 in TNBC has been related to significant increases in paclitaxel resistance
[35]. The role of circRNAs in apoptosis reduction, that is, modulating the expressions of AKT and proapoptotic factors BAX and BAK as well as the antiapoptotic BCL-2 protein, has been well-documented
[36].
7. Conclusions
CircRNAs are a type of ncRNA that has various regulatory roles in aggressive TNBC. Evidence has shown that they may greatly influence cell proliferation, migration, metastasis, apoptosis, and chemoresistance. CircRNAs have a dual role in metastasis, as they may act as oncogenes and tumor-suppressing genes through the regulation of microRNAs. Furthermore, circRNAs have been detected in exosomes in the serums of TNBC patients, representing potential biomarkers of disease, and participate in the response to anticancer drugs in TNBC.