| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Sajida Maryam | -- | 3193 | 2023-05-09 15:33:40 | | | |
| 2 | Peter Tang | Meta information modification | 3193 | 2023-05-10 03:16:54 | | | | |
| 3 | Peter Tang | Meta information modification | 3193 | 2023-05-10 03:59:41 | | |
Video Upload Options
Colorectal cancer (CRC) is the third most common cause of cancer-related deaths worldwide. Risk factors for CRC include obesity, a diet low in fruits and vegetables, physical inactivity, and smoking. CRC has a poor prognosis, and there is a critical need for new diagnostic and prognostic biomarkers to reduce related deaths. Studies have focused more on molecular testing to guide targeted treatments for CRC patients. The most crucial feature of activated immune cells is the production and release of growth factors and cytokines that modulate the inflammatory conditions in tumor tissues. The cytokine network is valuable for the prognosis and pathogenesis of colorectal cancer as they can aid in the cost-effective and non-invasive detection of cancer. A large number of interleukins (IL) released by the immune system at various stages of CRC can act as “biomarkers”. They play diverse functions in colorectal cancer, and include IL-4, IL-6, IL-8, IL-11, IL-17A, IL-22, IL-23, IL-33, TNF, TGF-β, and vascular endothelial growth factor (VEGF), which are pro-tumorigenic genes.
1. Introduction
2. Molecular Pathways and Cytokine Role in CRC

|
TME Cells |
Soluble Factors |
Target Cells |
Signaling Pathway |
Biological Effects |
Reference |
|---|---|---|---|---|---|
|
CD4+ |
IL-22 |
CRC |
STAT3/DOT1L (Signal transducer and activator of transcription)/(Disruptor of telomeric silencing 1-like) |
Stemness gene regulation |
[59] |
|
CAF (Cancer-associated fibroblasts) |
HGF/SDF1 (Hepatocyte growth factor)/Stromal cell-derived factor-1 |
Cancer stem cells (CSC) |
Wnt/β-catenin |
Clonogenic activity and expression of CD44v6 |
[60] |
|
Endothelial cells |
JAG1 (Jagged) |
CRC |
Notch |
CD133 expression, tumorigenicity and chemoresistance |
[61] |
|
MSC (Mesenchymal stem cells) |
PGE2 (Prostaglandin E2) |
CRC |
Wnt/β-catenin |
EMT (Epithelial-to-mesenchymal transition) and invasion |
[62] |
|
Myofibroblasts |
HGF |
CSC |
Wnt/β-catenin |
Clonogenicity |
[63] |
3. Advancements in Cytokine Detection and Monitoring Clinically
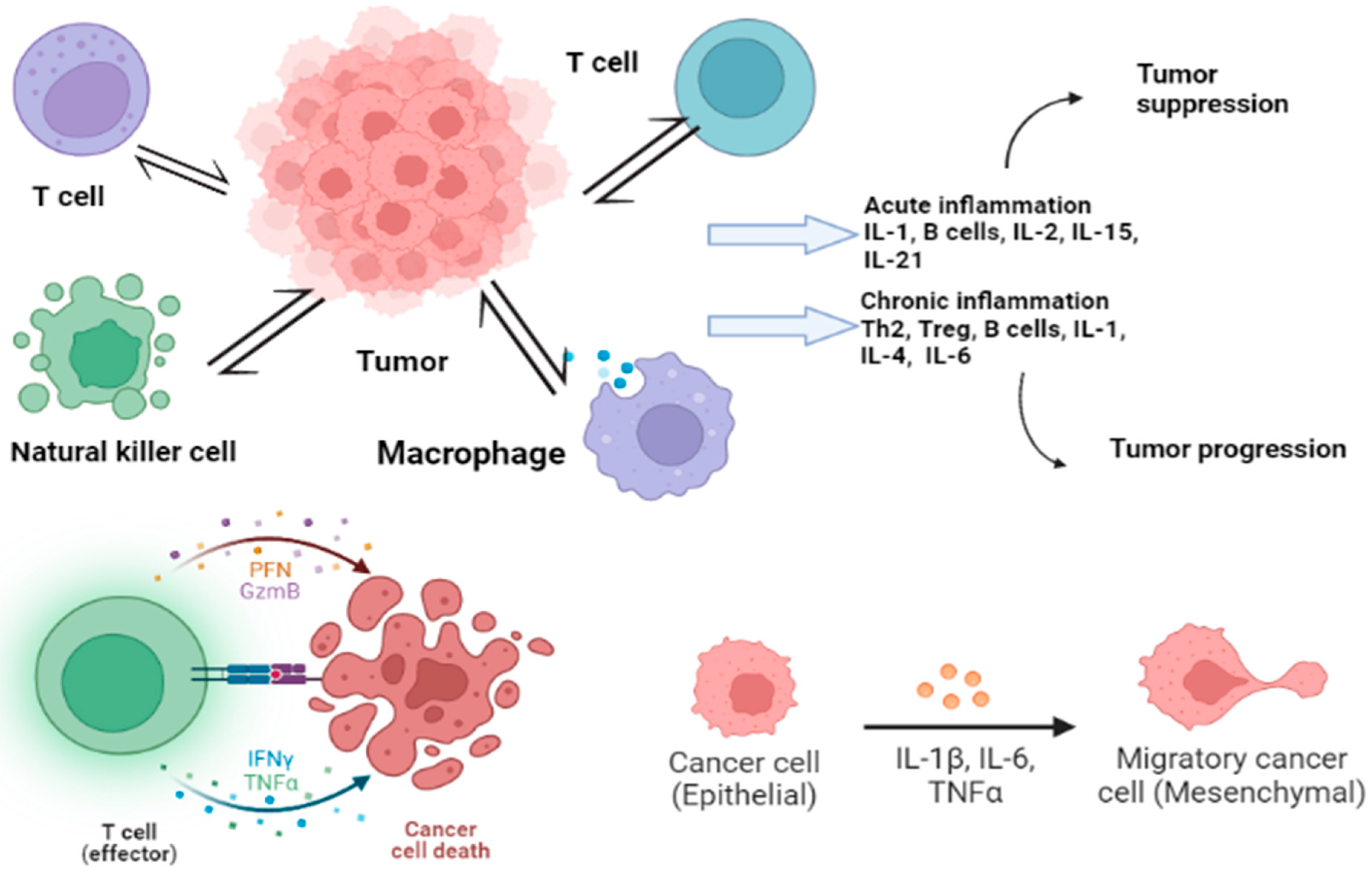
4. Cytokines’ Role in CRC
4.1. Forkhead Box P3 (FOXP3)
4.2. Tumor Necrosis Factor-α (TNF-α)
4.3. Interferon-Gamma (IFN-γ)
5. Interleukins in Colorectal Cancer
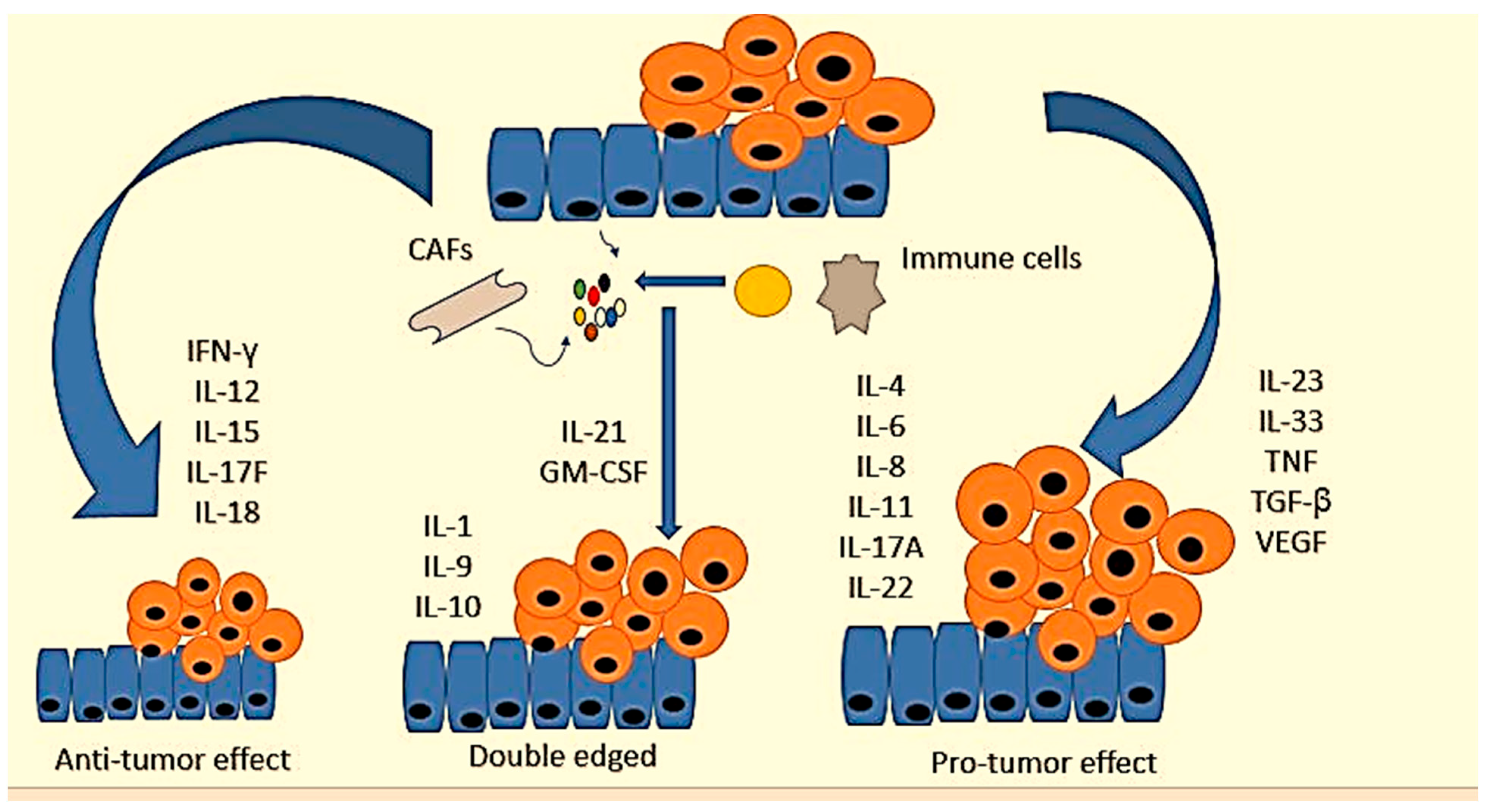
|
Cytokine |
Functional Effect in CRC |
Expression Patterns |
Reference |
|---|---|---|---|
|
IL-1α |
Promotes metastasis and the chemosensitivity |
|
|
|
IL-1β |
Promotes the proliferation of colon cancer cells, tumorigenesis, and alters the tumor microenvironment |
|
|
|
IL-18 |
Antitumorigenic properties and release of other signals |
|
|
|
IL-2, IL-7, IL-9, IL-15 |
Antitumor activity, promote EMT, proliferation, invasion, and metastasis |
IL-4, IL-7 upregulated, IL-9 downregulated, IL-2 in between |
|
|
IL-21 |
Activation of immune response biomarkers |
(Potential for biomarker) |
|
|
IL-6 |
Promotes mitosis, proliferation, metastasis, migration, and angiogenesis |
|
|
|
IL-11 |
Facilitates the proliferation of CRC |
|
|
|
IL-8 |
Promotes cell proliferation, angiogenesis, cancer metastasis, chemoresistance, antianoikis, maintains CCSC properties |
|
|
|
IL-10 |
Pathogenesis and progression |
|
|
|
IL-22 |
Dominant role in CRC tumorigenesis, antiapoptosis, and cell proliferation |
|
|
|
IL-17a |
Promotes cell cycle progression and angiogenesis |
|
[167] |
|
IL-17b |
Promotes tumor |
|
|
|
IL-4 |
Overexpressed in early CRC, tumor development |
|
[172] |
|
IL-23 |
Overexpressed in CRC tissue and predictive for CRC metastasis |
|
Upwards arrow just showed upregulation of the genes while downwards show suppression or downregulation.
References
- Safarzadeh, E.; Sandoghchian, S.; Baradaran, B. Herbal medicine as inducers of apoptosis in cancer treatment. Adv. Pharm. Bull. 2014, 4, 421–427.
- Shewach, D.S.; Kuchta, R.D. Introduction to cancer chemotherapeutics. Chem. Rev. 2009, 109, 2859–2861.
- Jemal, A.; Bray, F.; Center, M.M.; Ferlay, J.; Ward, E.; Forman, D. Global cancer statistics. CA Cancer J. Clin. 2011, 61, 69–90.
- Arnold, M.; Sierra, M.S.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global patterns and trends in colorectal cancer incidence and mortality. Gut 2017, 66, 683–691.
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424.
- Keum, N.; Giovannucci, E. Global burden of colorectal cancer: Emerging trends.; risk factors and prevention strategies. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 713–732.
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249.
- Williams, R.; White, P.; Nieto, J.; Vieira, D.; Francois, F.; Hamilton, F. Colorectal Cancer in African Americans: An Update. Clin. Transl. Gastroenterol. 2016, 7, e185.
- Jess, T.; Rungoe, C.; Peyrin-Biroulet, L. Risk of colorectal cancer in patients with ulcerative colitis: A meta-analysis of population-based cohort studies. Clin. Gastroenterol. Hepatol. 2012, 10, 639–645.
- Marventano, S.; Forjaz, M.J.; Grosso, G.; Mistretta, A.; Giorgianni, G.; Platania, A.; Gangi, S.; Basile, F.; Biondi, A. Health related quality of life in colorectal cancer patients: State of the art. BMC Surg. 2013, 13, S15.
- Vacante, M.; Borzì, A.M.; Basile, F.; Biondi, A. Biomarkers in colorectal cancer: Current clinical utility and future perspectives. World J. Clin. Cases 2018, 6, 869–881.
- Mager, L.F.; Wasmer, M.H.; Rau, T.T.; Krebs, P. Cytokine-Induced Modulation of Colorectal Cancer. Front. Oncol. 2016, 6, 96.
- Lee, S.; Margolin, K. Cytokines in cancer immunotherapy. Cancers 2011, 3, 3856–3893.
- Koltsova, E.K.; Grivennikov, S.I. IL-22 Gets to the Stem of Colorectal Cancer. Immunity 2014, 40, 639–641.
- Wang, J.; Li, D.; Cang, H.; Guo, B. Crosstalk between cancer and immune cells: Role of tumor-associated macrophages in the tumor microenvironment. Cancer Med. 2019, 8, 4709–4721.
- Kawanishi, S.; Ohnishi, S.; Ma, N.; Hiraku, Y.; Murata, M. Crosstalk between DNA Damage and Inflammation in the Multiple Steps of Carcinogenesis. Int. J. Mol. Sci. 2017, 18, 1808.
- Li, J.; Huang, L.; Zhao, H.; Yan, Y.; Lu, J. The Role of Interleukins in Colorectal Cancer. Int. J. Biol. Sci. 2020, 16, 2323–2339.
- Veziant, J.; Villéger, R.; Barnich, N.; Bonnet, M. Gut Microbiota as Potential Biomarker and/or Therapeutic Target to Improve the Management of Cancer: Focus on Colibactin-Producing Escherichia coli in Colorectal Cancer. Cancers 2021, 13, 2215.
- Kim, J.; Lee, H.K. Potential Role of the Gut Microbiome In Colorectal Cancer Progression. Front. Immunol. 2022, 12, 807648.
- Olovo, C.V.; Huang, X.; Zheng, X.; Xu, M. Faecal microbial biomarkers in early diagnosis of colorectal cancer. J. Cell. Mol. Med. 2021, 25, 10783–10797.
- Rye, M.S.; Garrett, K.L.; Holt, R.A.; Platell, C.F.; McCoy, M.J. Fusobacterium nucleatum and Bacteroides fragilis detection in colorectal tumours: Optimal target site and correlation with total bacterial load. PLoS ONE 2022, 17, e0262416.
- Eklöf, V.; Löfgren-Burström, A.; Zingmark, C.; Edin, S.; Larsson, P.; Karling, P.; Alexeyev, O.; Rutegård, J.; Wikberg, M.L.; Palmqvist, R. Cancer-associated fecal microbial markers in colorectal cancer detection. Int. J. Cancer 2017, 141, 2528–2536.
- Nassar, F.J.; Msheik, Z.S.; Itani, M.M.; Helou, R.E.; Hadla, R.; Kreidieh, F.; Bejjany, R.; Mukherji, D.; Shamseddine, A.; Nasr, R.R.; et al. Circulating miRNA as Biomarkers for Colorectal Cancer Diagnosis and Liver Metastasis. Diagnostics 2021, 11, 341.
- Condrat, C.E.; Thompson, D.C.; Barbu, M.G.; Bugnar, O.L.; Boboc, A.; Cretoiu, D.; Suciu, N.; Cretoiu, S.M.; Voinea, S.C. miRNAs as Biomarkers in Disease: Latest Findings Regarding Their Role in Diagnosis and Prognosis. Cells 2020, 9, 276.
- Staiteieh, S.A.; Akil, L.; Al Khansa, R.; Nasr, R.; Sagheer, A.Z.; Houshaymi, B.; Merhi, R.A. Study of microRNA expression profiling as biomarkers for colorectal cancer patients in Lebanon. Mol. Clin. Oncol. 2022, 16, 39.
- Sur, D.; Advani, S.; Braithwaite, D. MicroRNA panels as diagnostic biomarkers for colorectal cancer: A systematic review and meta-analysis. Front. Med. 2022, 9, 915226.
- Precazzini, F.; Detassis, S.; Imperatori, A.S.; Denti, M.A.; Campomenosi, P. Measurements Methods for the Development of MicroRNA-Based Tests for Cancer Diagnosis. Int. J. Mol. Sci. 2021, 22, 1176.
- Kaur, J.; Preethi, M.; Srivastava, R.; Borse, V. Role of IL-6 and IL-8 biomarkers for optical and electrochemical based point-of-care detection of oral cancer. Biosens. Bioelectron. X 2022, 11, 100212.
- Liu, G.; Jiang, C.; Lin, X.; Yang, Y. Point-of-care detection of cytokines in cytokine storm management and beyond: Significance and challenges. View 2021, 2, 20210003.
- Chaudhry, H.; Zhou, J.; Zhong, Y.; Ali, M.M.; McGuire, F.; Nagarkatti, P.S.; Nagarkatti, M. Role of cytokines as a double-edged sword in sepsis. In Vivo 2013, 27, 669–684.
- Febbo, P.G.; Ladanyi, M.; Aldape, K.D.; De Marzo, A.M.; Hammond, M.E.; Hayes, D.F.; Lafrate, A.J.; Kelley, R.K.; Marcucci, G.; Ogino, S.; et al. NCCN Task Force report: Evaluating the clinical utility of tumor markers in oncology. J. Natl. Compr. Cancer Netw. 2011, 9, S1–S32.
- Kartikasari, A.E.R.; Huertas, C.S.; Mitchell, A.; Plebanski, M. Tumor-Induced Inflammatory Cytokines and the Emerging Diagnostic Devices for Cancer Detection and Prognosis. Front. Oncol. 2021, 11, 692142.
- Capone, F.; Guerriero, E.; Sorice, A.; Colonna, G.; Ciliberto, G.; Costantini, S. Serum Cytokinome Profile Evaluation: A Tool to Define New Diagnostic and Prognostic Markers of Cancer Using Multiplexed Bead-Based Immunoassays. Mediat. Inflamm. 2016, 2016, 3064643.
- Schett, G.; Elewaut, D.; McInnes, I.B.; Dayer, J.M.; Neurath, M.F. How cytokine networks fuel inflammation: Toward a cytokine-based disease taxonomy. Nat. Med. 2013, 19, 822–824.
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674.
- Jia, D.; Li, L.; Andrew, S.; Allan, D.; Li, X.; Lee, J.; Ji, Z.; Yao, Z.; Gadde, S.; Figeys, D.; et al. An autocrine inflammatory forward-feedback loop after chemotherapy withdrawal facilitates the repopulation of drug-resistant breast cancer cells. Cell Death Dis. 2017, 8, e2932.
- Nengroo, M.A.; Verma, A.; Datta, D. Cytokine chemokine network in tumor microenvironment: Impact on CSC properties and therapeutic applications. Cytokine 2022, 156, 155916.
- Furman, D.; Campisi, J.; Verdin, E.; Carrera-Bastos, P.; Targ, S.; Franceschi, C.; Ferrucci, L.; Gilroy, D.W.; Fasano, A.; Miller, G.W.; et al. Chronic inflammation in the etiology of disease across the life span. Nat. Med. 2019, 25, 1822–1832.
- Greten, F.R.; Grivennikov, S.I. Inflammation and Cancer: Triggers, Mechanisms, and Consequences. Immunity 2019, 51, 27–41.
- Sokolova, O.; Naumann, M. Crosstalk between DNA Damage and Inflammation in the Multiple Steps of Gastric Carcinogenesis. Curr. Top. Microbiol. Immunol. 2019, 421, 107–137.
- Yang, Z.H.; Dang, Y.Q.; Ji. G. Role of epigenetics in transformation of inflammation into colorectal cancer. World J. Gastroenterol. 2019, 25, 2863–2877.
- Martin, B.; Märkl, B. Immunologic Biomarkers and Biomarkers for Immunotherapies in Gastrointestinal Cancer. Visc. Med. 2019, 35, 3–10.
- Rhea, J.M.; Molinaro, R.J. Cancer biomarkers: Surviving the journey from bench to bedside. MLO Med. Lab. Obs. 2011, 43, 10–12.
- Nagata, H.; Ishihara, S.; Hata, K.; Murono, K.; Kaneko, M.; Yasuda, K.; Otani, K.; Watanabe, T. Survival and Prognostic Factors for Metachronous Peritoneal Metastasis in Patients with Colon Cancer. Ann. Surg. Oncol. 2017, 24, 1269–1280.
- Goossens, N.; Nakagawa, S.; Sun, X.; Hoshida, Y. Cancer biomarker discovery and validation. Transl. Cancer Res. 2015, 4, 256.
- Sarhadi, V.K.; Armengol, G. Molecular Biomarkers in Cancer. Biomolecules 2022, 12, 1021.
- Herceg, Z.; Hainaut, P. Genetic and epigenetic alterations as biomarkers for cancer detection, diagnosis and prognosis. Mol. Oncol. 2007, 1, 26–41.
- Harbaum, L.; Pollheimer, M.J.; Kornprat, P.; Lindtner, R.A.; Bokemeyer, C.; Langner, C. Peritumoral eosinophils predict recurrence in colorectal cancer. Mod. Pathol. 2015, 28, 403–413.
- HSU, C.-P.; Chung, Y.-C. Influence of Interleukin-6 on the Invasiveness of Human Colorectal Carcinoma. Anticancer. Res. 2006, 26, 4607–4614. Available online: https://ar.iiarjournals.org/content/anticanres/26/6B/4607.full.pdf (accessed on 17 March 2023).
- Waniczek, D.; Lorenc, Z.; Śnietura, M.; Wesecki, M.; Kopec, A.; Muc-Wierzgoń, M. Tumor-Associated Macrophages and Regulatory T Cells Infiltration and the Clinical Outcome in Colorectal Cancer. Arch. Immunol. Ther. Exp. 2017, 65, 445–454.
- Wikberg, M.L.; Ling, A.; Li, X.; Öberg, Å.; Edin, S.; Palmqvist, R. Neutrophil infiltration is a favorable prognostic factor in early stages of colon cancer. Hum. Pathol. 2017, 68, 193–202.
- Terzić, J.; Grivennikov, S.; Karin, E.; Karin. M. Inflammation and colon cancer. Gastroenterology 2010, 138, 2101–2114.e5.
- Calon, A.; Espinet, E.; Palomo-Ponce, S.; Tauriello, D.V.; Iglesias, M.; Céspedes, M.V. Dependency of colorectal cancer on a TGF-β-driven program in stromal cells for metastasis initiation. Cancer Cell 2012, 22, 571–584.
- Guo, S.; Deng, C.X. Effect of Stromal Cells in Tumor Microenvironment on Metastasis Initiation. Int. J. Biol. Sci. 2018, 14, 2083–2093.
- Nguyen, L.H.; Goel, A.; Chung, D.C. Pathways of Colorectal Carcinogenesis. Gastroenterology 2020, 158, 291–302.
- Pino, M.S.; Chung, D.C. The chromosomal instability pathway in colon cancer. Gastroenterology 2010, 138, 2059–2072.
- Powell, D.W.; Adegboyega, P.A.; Di Mari, J.F.; Mifflin, R.C. Epithelial cells and their neighbors I. Role of intestinal myofibroblasts in development repair and cancer. Am. J. Physiol. Gastrointest. Liver Physiol. 2005, 289, G2–G7.
- Koliaraki, V.; Pallangyo, C.K.; Greten, F.R.; Kollias, G. Mesenchymal Cells in Colon Cancer. Gastroenterology 2017, 152, 964–979.
- Kryczek, I.; Lin, Y.; Nagarsheth, N.; Peng, D.; Zhao, L.; Zhao, E.; Vatan, L.; Szeliga, W.; Dou, Y.; Owens, S.; et al. IL-22(+) CD4(+) T cells promote colorectal cancer stemness via STAT3 transcription factor activation and induction of the methyltransferase DOT1L. Immunity 2014, 40, 772–784.
- Todaro, M.; Gaggianesi, M.; Catalano, V.; Benfante, A.; Iovino, F.; Biffoni, M.; Apuzzo, T.; Sperduti, I.; Volpe, S.; Cocorullo, G.; et al. CD44v6 is a marker of constitutive and reprogrammed cancer stem cells driving colon cancer metastasis. Cell Stem Cell 2014, 14, 342–356.
- Lotti, F.; Jarrar, A.M.; Pai, R.K.; Hitomi, M.; Lathia, J.; Mace, A.; Rich, J.N. Chemotherapy activates cancer-associated fibroblasts to maintain colorectal cancer-initiating cells by IL-17A. J. Exp. Med. 2013, 210, 2851–2872.
- Chen, K.; Liu, Q.; Tsang, L.L.; Ye, Q.; Chan, H.C.; Sun, Y.; Jiang, X. Human MSCs promotes colorectal cancer epithelial–mesenchymal transition and progression via CCL5/β-catenin/Slug pathway. Cell Death Dis. 2017, 8, e2819.
- Vermeulen, L.; De Sousa, E.M.F.; van der Heijden, M.; Cameron, K.; de Jong, J.H.; Borovski, T.; Tuynman, J.B.; Todaro, M.; Merz, C.; Rodermond, H.; et al. Wnt activity defines colon cancer stem cells and is regulated by the microenvironment. Nat. Cell Biol. 2010, 12, 468–476.
- Schulte, W.; Jürgen, B.; Richard, B. Cytokines in sepsis: Potent immunoregulators and potential therapeutic targets—An updated view. Mediat. Inflamm. 2013, 2013, 165974.
- Sprague, A.H.; Khalil, R.A. Inflammatory cytokines in vascular dysfunction and vascular disease. Biochem. Pharmacol. 2009, 78, 539–552.
- Rea, I.M.; Gibson, D.S.; McGilligan, V.; McNerlan, S.E. Alexander HD, Ross OA. Age and Age-Related Diseases: Role of Inflammation Triggers and Cytokines. Front. Immunol. 2018, 9, 586.
- Nedoszytko, B.; Sokołowska-Wojdyło, M.; Ruckemann-Dziurdzińska, K.; Roszkiewicz, J.; Nowicki, R.J. Chemokines and cytokines network in the pathogenesis of the inflammatory skin diseases: Atopic dermatitis, psoriasis and skin mastocytosis. Postępy Dermatol. I Alergol. 2014, 31, 84–91.
- Kabel, A.M. Relationship between cancer and cytokines. J. Cancer Res. Treat. 2014, 2, 41–43.
- Comella-Del-Barrio, P.; Abellana, R.; Villar-Hernández, R.; Jean Coute, M.D.; Sallés Mingels, B.; Canales Aliaga, L.; Narcisse, M.; Gautier, J.; Ascaso, C.; Latorre, I.; et al. A model based on the combination of IFN-γ, IP-10, ferritin and 25-hydroxyvitamin D for discriminating latent from active tuberculosis in children. Front. Microbiol. 2019, 10, 1855.
- Pugliese, N.R.; Fabiani, I.; Conte, L.; Nesti, L.; Masi, S.; Natali, A.; Colombo, P.C.; Pedrinelli, R.; Dini, F.L. Persistent congestion, renal dysfunction and inflammatory cytokines in acute heart failure: A prognosis study. J. Cardiovasc. Med. 2020, 21, 494–502.
- Shimabukuro-Vornhagen, A.; Gödel, P.; Subklewe, M.; Stemmler, H.J.; Schlößer, H.A.; Schlaak, M.; Kochanek, M.; Böll, B.; von Bergwelt-Baildon, M.S. Cytokine release syndrome. J. Immunother. Cancer 2018, 6, 56.
- Moore, J.B.; Carl, H.J. Cytokine release syndrome in severe COVID-19. Science 2020, 368, 473–474.
- Simpson, S.; Kaislasuo, J.; Guller, S.; Pal, L. Thermal stability of cytokines: A review. Cytokine 2020, 125, 154829.
- Liu, G.; Qi, M.; Hutchinson, M.R.; Yang, G.; Goldys, E.M. Recent advances in cytokine detection by immunosensing. Biosens. Bioelectron. 2016, 79, 810–821.
- Monastero, R.N.; Srinivas, P. Cytokines as biomarkers and their respective clinical cutoff levels. Int. J. Inflam. 2017, 2017, 4309485.
- Kouwenhoven, M.; Ozenci, V.; Teleshova, N.; Hussein, Y.; Huang, Y.M.; Eusebio, A.; Link, H. Enzyme-linked immunospot assays provide a sensitive tool for detection of cytokine secretion by monocytes. Clin. Diagn. Lab. Immunol. 2001, 8, 1248–1257.
- Stenken, J.A.; Poschenrieder, A.J. Bioanalytical chemistry of cytokines–a review. Anal. Chim. Acta 2015, 853, 95–115.
- Zhang, K.; Baratta, M.V.; Liu, G.; Frank, M.G.; Leslie, N.R.; Watkins, L.R.; Maier, S.F.; Hutchinson, M.R.; Goldys, E.M. A novel platform for in vivo detection of cytokine release within discrete brain regions. Brain Behav. Immun. 2018, 71, 18–22.
- Arman, A.; Deng, F.; Goldys, E.M.; Liu, G.; Hutchinson, M.R. In vivo intrathecal IL-1β quantification in rats: Monitoring the molecular signals of neuropathic pain. Brain Behav. Immun. 2020, 88, 442–450.
- Tertis, M.; Leva, P.I.; Bogdan, D.; Suciu, M.; Graur, F.; Cristea, C. Impedimetric aptasensor for the label-free and selective detection of Interleukin-6 for colorectal cancer screening. Biosens. Bioelectron. 2019, 137, 123–132.
- Aldo, P.; Marusov, G.; Svancara, D.; David, J.; Mor, G. Simple Plex™: A novel multi-analyte, automated microfluidic immunoassay platform for the detection of human and mouse cytokines and chemokines. Am. J. Reprod. Immunol. 2016, 75, 6.
- Zhong, Y.; Tang, X.; Li, J.; Lan, Q.; Min, L.; Ren, C.; Hu, X.; Torrente, R.M.; Gao, W.; Yang, Z. A nanozyme tag enabled chemiluminescence imaging immunoassay for multiplexed cytokine monitoring. Chem. Commun. 2018, 54, 13813–13816.
- Emoto, S.; Ishigami, H.; Yamashita, H.; Yamaguchi, H.; Kaisaki, S.; Kitayama, J. Clinical significance of CA125 and CA72-4 in gastric cancer with peritoneal dissemination. Gastric Cancer 2012, 15, 154–161.
- Liang, Y.; Wang, W.; Fang, C.; Raj, S.S.; Hu, W.M.; Li, Q.W.; Zhou, Z.W. Clinical significance and diagnostic value of serum CEA, CA19-9 and CA72-4 in patients with gastric cancer. Oncotarget 2016, 7, 49565–49573.
- Ulusoy, H.; Kangalgil, M.; Küçük, A.O.; Özdemir, A.; Karahan, S.C.; Yaman, S.Ö.; Yavuz, H.B.; Ok, Ü. Effects of different lipid emulsions on serum adipokines, inflammatory markers and mortality in critically ill patients with sepsis: A prospective observational cohort study. Clin. Nutr. 2021, 40, 4569–4578.
- He, R.; Zhu, Q.; Wang, Y.; Chen, G.; Chen, S.; Wang, Y. Influence of respiratory function training under the mode of mutual-assisted patients on postoperative pulmonary infection and immune function on lung cancer. Am. J. Transl. Res. 2021, 13, 9260–9268.
- Diamandis, E.P.; Hoffman, B.R.; Sturgeon, C.M. National academy of clinical biochemistry laboratory medicine practice guidelines for the use of tumor markers. Clin. Chem. 2008, 54, 1935–1939.
- Hing, J.X.; Mok, C.W.; Tan, P.T.; Sudhakar, S.S.; Seah, C.M.; Lee, W.P.; Tan, S.M. Clinical utility of tumour marker velocity of cancer antigen 15-3 (CA 15-3) and carcinoembryonic antigen (CEA) in breast cancer surveillance. Breast 2020, 52, 95–101.
- Diaz, C.B.; Marono, S.H.; Olivas, M.L.G.; Sosa, M.A.; Soto, R.G.; Arrieta, O. Association of neurologic manifestations and CEA levels with the diagnosis of brain metastases in lung cancer patients. Clin. Transl. Oncol. 2019, 21, 1538–1542.
- Song, X.; Liang, B.; Wang, C.; Shi, S. Clinical value of color Doppler ultrasound combined with serum CA153, CEA and TSGF detection in the diagnosis of breast cancer. Exp. Ther. Med. 2020, 20, 1822–1828.
- Sakamoto, T.; Saito, H.; Amisaki, M.; Tokuyasu, N.; Honjo, S.; Fujiwara, Y. Combined preoperative platelet-to-lymphocyte ratio and serum carbohydrate antigen 19-9 level as a prognostic factor in patients with resected pancreatic cancer. Hepatobiliary Pancreat. Dis. Int. 2019, 18, 278–284.
- Peng, H.X.; Yang, L.; He, B.S.; Pan, Y.Q.; Ying, H.Q.; Sun, H.L.; Lin, K.; Hu, X.X.; Xu, T.; Wang, S.K. Combination of preoperative NLR, PLR and CEA could increase the diagnostic efficacy for I-III stage CRC. J. Clin. Lab. Anal. 2017, 31, e22075.
- Yan, X.; Han, L.; Zhao, R.; Fatima, S.; Zhao, L.; Gao, F. Prognosis value of IL-6, IL-8, and IL-1β in serum of patients with lung cancer: A fresh look at interleukins as a biomarker. Heliyon 2022, 8, e09953.
- Xi, C.; Zhang, G.Q.; Sun, Z.K.; Song, H.J.; Shen, C.T.; Chen, X.Y.; Sun, J.W.; Qiu, Z.L.; Luo, Q.Y. Interleukins in Thyroid Cancer: From Basic Researches to Applications in Clinical Practice. Front. Immunol. 2020, 11, 1124.
- Provatopoulou, X.; Georgiadou, D.; Sergentanis, T.N.; Kalogera, E.; Spyridakis, J.; Gounaris, A.; Zografos, G.N. Interleukins as markers of inflammation in malignant and benign thyroid disease. Inflamm. Res. 2014, 63, 667–674.
- Rébé, C.; Ghiringhelli, F. Interleukin-1β and Cancer. Cancers 2020, 12, 1791.
- Valeri, M.; Raffatellu, M. Cytokines IL-17 and IL-22 in the host response to infection. Pathog. Dis. 2016, 74, ftw111.
- Anestakis, D.; Petanidis, S.; Kalyvas, S.; Nday, C.M.; Tsave, O.; Kioseoglou, E.; Salifoglou, A. Mechanisms and Applications of Ιnterleukins in Cancer Immunotherapy. Int. J. Mol. Sci. 2015, 16, 1691–1710.
- Zarogoulidis, P.; Lampaki, S.; Yarmus, L.; Kioumis, I.; Pitsiou, G.; Katsikogiannis, N.; Hohenforst-Schmidt, W.; Li, Q.; Huang, H.; Sakkas, A. Interleukin-7 and interleukin-15 for cancer. J. Cancer 2014, 5, 765–773.
- Todoric, J.; Antonucci, L.; Karin, M. Targeting Inflammation in Cancer Prevention and Therapy. Cancer Prev. Res. 2016, 9, 895.
- Coussens, L.M.; Werb, Z. Inflammation and cancer. Nature 2002, 420, 860–867.
- Mantovani, A.; Allavena, P.; Sica, A.; Balkwill, F. Cancer-related inflammation. Nature 2008, 454, 436–444.
- Ilyin, S.E.; Belkowski, S.M.; Plata-Salamán, C.R. Biomarker discovery and validation: Technologies and integrative approaches. Trends Biotechnol. 2004, 22, 411–416.
- Brunkow, M.E.; Jeffery, E.W.; Hjerrild, K.A.; Paeper, B.; Clark, L.B.; Yasayko, S.A.; Wilkinson, J.E.; Galas, D.; Ziegler, S.F.; Ramsdell, F. Disruption of a new forkhead/winged-helix protein.; scurfin.; results in the fatal lymphoproliferative disorder of the scurfy mouse. Nat. Genet. 2001, 27, 68–73.
- Katoh, M.; Igarashi, M.; Fukuda, H.; Nakagama, H.; Katoh, M. Cancer genetics and genomics of human FOX family genes. Cancer Lett. 2013, 328, 198–206.
- Tian, T.; Wang, M.; Zheng, Y.; Yang, T.; Zhu, W.; Li, H.; Lin, S.; Liu, K.; Xu, P.; Deng, Y.; et al. Association of two FOXP3 polymorphisms with breast cancer susceptibility in Chinese Han women. Cancer Manag. Res. 2018, 10, 867–872.
- Luo, Q.; Zhang, S.; Wei, H.; Pang, X.; Zhang, H. Roles of Foxp3 in the occurrence and development of cervical cancer. Int. J. Clin. Exp. Pathol. 2015, 8, 8717–8730.
- Sun, X.; Feng, Z.; Wang, Y.; Qu, Y.; Gai, Y. Expression of Foxp3 and its prognostic significance in colorectal cancer. Int. J. Immunopathol. Pharmacol. 2017, 30, 201–206.
- Liu, Y.; Lan, Q.; Lu, L.; Chen, M.; Xia, Z.; Ma, J.; Wang, J.; Fan, H.; Shen, Y.; Ryffel, B. Phenotypic and functional characteristic of a newly identified CD8+ Foxp3− CD103+ regulatory T cells. J. Mol. Cell Biol. 2014, 6, 81–92.
- Kim, M.; Grimmig, T.; Grimm, M.; Lazariotou, M.; Meier, E.; Rosenwald, A.; Tsaur, I.; Blaheta, R.; Heemann, U.; Germer, C.T. Expression of Foxp3 in colorectal cancer but not in Treg cells correlates with disease progression in patients with colorectal cancer. PLoS ONE 2013, 8, e53630.
- Kuwahara, T.; Hazama, S.; Suzuki, N.; Yoshida, S.; Tomochika, S.; Nakagami, Y.; Matsui, H.; Shindo, Y.; Kanekiyo, S.; Tokumitsu, Y. Intratumoural-infiltrating CD4 + and FOXP3 + T cells as strong positive predictive markers for the prognosis of resectable colorectal cancer. Br. J. Cancer 2019, 121, 659–665.
- Grimmig, T.; Kim, M.; Germer, C.-T.; Gasser, M.; Maria Waaga-Gasser, A. The role of FOXP3 in disease progression in colorectal cancer patients. Oncoimmunology 2013, 2, e24521.
- Bessis, N.; Chiocchia, G.; Kollias, G.; Minty, A.; Fournier, C.; Fradelizi, D.; Boissier, M.C. Modulation of proinflammatory cytokine production in tumour necrosis factor-alpha (TNF-alpha)-transgenic mice by treatment with cells engineered to secrete IL-4.; IL-10 or IL-13. Clin. Exp. Immunol. 1998, 111, 391–396.
- Bates, R.C.; Mercurio, A.M. Tumor necrosis factor-alpha stimulates the epithelial-to-mesenchymal transition of human colonic organoids. Mol. Biol. Cell. 2003, 14, 1790–1800.
- Ou, B.; Zhao, J.; Guan, S.; Feng, H.; Wangpu, X.; Zhu, C.; Zong, Y.; Ma, J.; Sun, J.; Shen, X.; et al. CCR4 promotes metastasis via ERK/NF-κB/MMP13 pathway and acts downstream of TNF-α in colorectal cancer. Oncotarget 2016, 7, 47637.
- Jovanovic, D.V.; Di Battista, J.A.; Martel-Pelletier, J.; Jolicoeur, F.C.; He, Y.; Zhang, M.; Mineau, F.; Pelletier, J.P. IL-17 stimulates the production and expression of proinflammatory cytokines.; IL-beta and TNF-alpha. by human macrophages. J. Immunol. 1998, 160, 3513–3521.
- Hunter, K. Host genetics influence tumour metastasis. Nat. Rev. Cancer 2006, 6, 141–146.
- Al Obeed, O.A.; Alkhayal, K.A.; Al Sheikh, A.; Zubaidi, A.M.; Vaali-Mohammed, M.A.; Boushey, R.; Mckerrow, J.H.; Abdulla, M.H. Increased expression of tumor necrosis factor-α is associated with advanced colorectal cancer stages. World J. Gastroenterol. 2014, 20, 18390–18396.
- Grimm, M.; Kim, M.; Rosenwald, A.; von Raden, B.H.; Tsaur, I.; Meier, E.; Heemann, U.; Germer, C.T.; Gasser, M.; Waaga-Gasser, A.M. Tumour-mediated TRAIL-Receptor expression indicates effective apoptotic depletion of infiltrating CD8+ immune cells in clinical colorectal cancer. Eur. J. Cancer 2010, 46, 2314–2323.
- Stanilov, N.; Miteva, L.; Dobreva, Z.; Stanilova, S. Colorectal cancer severity and survival in correlation with tumour necrosis factor-alpha. Biotechnol. Biotechnol. Equip. 2014, 28, 911–917.
- Warsinggih; Limanu, F.; Labeda, I.; Lusikooy, R.E.; Mappincara; Faruk, M. The relationship of tumor necrosis factor alpha levels in plasma toward the stage and differentiation degree in colorectal cancer. Med. Clínica Práctica 2021, 4, 100224.
- Wang, L.; Wang, Y.; Song, Z.; Chu, J.; Qu, X. Deficiency of interferon-gamma or its receptor promotes colorectal cancer development. J. Interferon Cytokine Res. 2015, 35, 273–280.
- Kosmidis, C.; Sapalidis, K.; Koletsa, T.; Kosmidou, M.; Efthimiadis, C.; Anthimidis, G.; Varsamis, N.; Michalopoulos, N.; Koulouris, C.; Atmatzidis, S.; et al. Interferon-γ and Colorectal Cancer: An up-to date. J. Cancer 2018, 9, 232–238.
- Liu, S.; Yu, X.; Wang, Q.; Liu, Z.; Xiao, Q.; Hou, P.; Hu, Y.; Hou, W.; Yang, Z.; Guo, D.; et al. Specific expression of interferon-γ induced by synergistic activation mediator-derived systems activates innate immunity and inhibits tumorigenesis. J. Microbiol. Biotechnol. 2017, 27, 1855–1866.
- Schroder, K.; Hertzog, P.J.; Ravasi, T.; Hume, D.A. Interferon-gamma: An overview of signals.; mechanisms and functions. J. Leukoc. Biol. 2004, 75, 163–189.
- Jorgovanovic, D.; Song, M.; Wang, L.; Zhang, Y. Roles of IFN-γ in tumor progression and regression: A review. Biomark. Res. 2020, 8, 49.
- Bellucci, R.; Martin, A.; Bommarito, D.; Wang, K.; Hansen, S.H.; Freeman, G.J.; Ritz, J. Interferon-γ-induced activation of JAK1 and JAK2 suppresses tumor cell susceptibility to NK cells through upregulation of PD-L1 expression. Oncoimmunology 2015, 4, e1008824.
- Mimura, K.; The, J.L.; Okayama, H.; Shiraishi, K.; Kua, L.F.; Koh, V.; Smoot, D.T.; Ashktorab, H.; Oike, T.; Suzuki, Y.; et al. PD-L1 expression is mainly regulated by interferon gamma associated with JAK-STAT pathway in gastric cancer. Cancer Sci. 2018, 109, 43–53.
- Zhao, T.; Li, Y.; Zhang, J.; Zhang, B. PD-L1 expression increased by IFN-γ via JAK2-STAT1 signaling and predicts a poor survival in colorectal cancer. Oncol. Lett. 2020, 20, 1127–1134.
- Calik, I.; Calik, M.; Turken, G.; Ozercan, I.H.; Dagli, A.F.; Artas, G.; Sarikaya, B. Intratumoral Cytotoxic T-Lymphocyte Density and PD-L1 Expression Are Prognostic Biomarkers for Patients with Colorectal Cancer. Medicina 2019, 55, 723.
- Carson, W.E.; Dierksheide, J.E.; Jabbour, S.; Anghelina, M.; Bouchard, P.; Ku, G.; Yu, H.; Baumann, H.; Shah, M.H.; Cooper, M.A.; et al. Coadministration of interleukin-18 and interleukin-12 induces a fatal inflammatory response in mice: Critical role of natural killer cell interferon-γ production and STAT-mediated signal transduction. Blood 2000, 96, 1465–1473.
- Chaisavaneeyakorn, S.; Othoro, C.; Shi, Y.P.; Otieno, J.; Chaiyaroj, S.C.; Lal, A.A.; Udhayakumar, V. Relationship between plasma Interleukin-12 (IL-12) and IL-18 levels and severe malarial anemia in an area of holoendemicity in western Kenya. Clin. Diagn. Lab. Immunol. 2003, 10, 362–366.
- Feng, X.; Zhang, Z.; Sun, P.; Song, G.; Wang, L.; Sun, Z.; Yuan, N.; Wang, Q.; Lun, L. Interleukin-18 Is a Prognostic Marker and Plays a Tumor Suppressive Role in Colon Cancer. Disease Markers 2020, 2020, 6439614.
- Braumüller, H.; Mauerer, B.; Andris, J.; Berlin, C.; Wieder, T.; Kesselring, R. The Cytokine Network in Colorectal Cancer: Implications for New Treatment Strategies. Cells 2023, 12, 138.
- Briukhovetska, D.; Dörr, J.; Endres, S.; Libby, P.; Dinarello, C.A.; Kobold, S. Interleukins in cancer: From biology to therapy. Nat. Rev. Cancer 2021, 21, 481–499.
- Chen, X.W.; Zhou, S.F. Inflammation, cytokines, the IL-17/IL-6/STAT3/NF-κB axis, and tumorigenesis. Drug. Des. DevelTher. 2015, 9, 2941–2946.
- Cheng, K.J.; Mejia, M.E.H.; Khong, T.L.; Mohd, Z.S.; Thavagnanam, S.; Ibrahim, Z.A. IL-1α and colorectal cancer pathogenesis: Enthralling candidate for anti-cancer therapy. Crit. Rev. Oncol. Hematol. 2021, 163, 103398.
- Li, Y.; Wang, L.; Pappan, L. IL-1β promotes stemness and invasiveness of colon cancer cells through Zeb1 activation. Mol. Cancer 2012, 11, 87.
- Lubberink, M.; Golla, S.S.; Jonasson, M.; Rubin, K.; Glimelius, B.; Sörensen, J.; Nygren, P. (15)O-Water PET Study of the Effect of Imatinib.; a Selective Platelet-Derived Growth Factor Receptor Inhibitor.; Versus Anakinra.; an IL-1R Antagonist.; on Water-Perfusable Tissue Fraction in Colorectal Cancer Metastases. J. Nucl. Med. 2015, 56, 1144–1149.
- Pastille, E.; Wasmer, M.H.; Adamczyk, A.; Vu, V.P.; Mager, L.F.; Phuong, N.N.T.; Palmieri, V.; Simillion, C.; Hansen, W.; Kasper, S.; et al. The IL-33/ST2 pathway shapes the regulatory T cell phenotype to promote intestinal cancer. Mucosal Immunol. 2019, 12, 990–1003.
- Bergman, M.; Levin, G.S.; Bessler, H.; Djaldetti, M.; Salman, H. Resveratrol affects the cross talk between immune and colon cancer cells. Biomed. Pharmacother. 2013, 67, 43–47.
- Idris, A.; Ghazali, N.B.; Koh, D. Interleukin 1β—A Potential Salivary Biomarker for Cancer Progression? Biomark. Cancer 2015, 7, BIC.S25375.
- Li, B.; Wang, F.; Ma, C.; Hao, T.; Geng, L.; Jiang, H. Predictive value of IL-18 and IL-10 in the prognosis of patients with colorectal cancer. Oncol. Lett. 2019, 18, 713–719.
- Nikiteas, N.; Yannopoulos, A.; Chatzitheofylaktou, A.; Tsigris, C. Heterozygosity for Interleukin-18 -607 A/C Polymorphism is Associated with Risk for Colorectal Cancer. Anticancer Res. 2007, 27, 3849–3853. Available online: https://ar.iiarjournals.org/content/anticanres/27/6B/3849.full.pdf (accessed on 17 March 2023).
- Misa, B.I.; Diakowska, D.; Korpacka, K.M. Local and Systemic IL-7 Concentration in Gastrointestinal-Tract Cancers. Medicina 2019, 55, 262.
- Chen, J.; Gong, C.; Mao, H.; Li, Z.; Fang, Z.; Chen, Q.; Lin, M.; Jiang, X.; Hu, Y.; Wang, W.; et al. E2F1/SP3/STAT6 axis is required for IL-4-induced epithelial-mesenchymal transition of colorectal cancer cells. Int. J. Oncol. 2018, 53, 567–578.
- Liu, X.; Zhu, L.; Lu, X.; Bian, H.; Wu, X.; Yang, W.; Qin, Q. IL-33/ST2 pathway contributes to metastasis of human colorectal cancer. Biochem. Biophys. Res. Commun. 2014, 453, 486–492.
- Rocca, Y.S.; Roberti, M.P.; Juliá, E.P.; Pampena, M.B.; Bruno, L.; Rivero, S.; Huertas, E.; Sánchez Loria, F.; Pairola, A.; Caignard, A.; et al. Phenotypic and Functional Dysregulated Blood NK Cells in Colorectal Cancer Patients Can Be Activated by Cetuximab Plus IL-2 or IL-15. Front. Immunol. 2016, 7, 413.
- Wang, C.; Lu, Y.; Chen, L.; Gao, T.; Yang, Q.; Zhu, C.; Chen, Y. Th9 cells are subjected to PD-1/PD-L1-mediated inhibition and are capable of promoting CD8 T cell expansion through IL-9R in colorectal cancer. Int. Immunopharmacol. 2020, 78, 106019.
- Fukushima, Y.; Iinuma, H.; Tsukamoto, M.; Matsuda, K.; Hashiguchi, Y. Clinical significance of microRNA-21 as a biomarker in each Dukes’ stage of colorectal cancer. Oncol. Rep. 2015, 33, 573–582.
- Steele, N.; Anthony, A.; Saunders, M.; Esmarck, B.; Ehrnrooth, E.; Kristjansen, P.E.; Nihlén, A.; Hansen, L.T.; Cassidy, J. A phase 1 trial of recombinant human IL-21 in combination with cetuximab in patients with metastatic colorectal cancer. Br. J. Cancer 2012, 106, 793–798.
- Stolfi, C.; Rizzo, A.; Franzè, E.; Rotondi, A.; Fantini, M.C.; Sarra, M.; Caruso, R.; Monteleone, I.; Sileri, P.; Franceschilli, L. Involvement of interleukin-21 in the regulation of colitis-associated colon cancer. J. Exp. Med. 2011, 208, 2279–2290.
- Belluco, C.; Nitti, D.; Frantz, M.; Toppan, P.; Basso, D.; Plebani, M.; Lise, M.; Jessup, J.M. Interleukin-6 Blood Level Is Associated with Circulating Carcinoembryonic Antigen and Prognosis in Patients with Colorectal Cancer. Ann. Surg. Oncol. 2000, 7, 133–138.
- Knüpfer, H.; Preiss, R. Serum interleukin-6 levels in colorectal cancer patients—A summary of published results. Int. J. Color. Dis. 2010, 25, 135–140.
- St John, M.A.; Li, Y.; Zhou, X.; Denny, P.; Ho, C.M.; Montemagno, C.; Shi, W.; Qi, F.; Wu, B.; Sinha, U.; et al. Interleukin 6 and Interleukin 8 as Potential Biomarkers for Oral Cavity and Oropharyngeal Squamous Cell Carcinoma. Arch. Otolaryngol. Head Neck Surg. 2004, 130, 929–935.
- Vainer, N.; Dehlendorff, C.; Johansen, J.S. Systematic literature review of IL-6 as a biomarker or treatment target in patients with gastric, bile duct, pancreatic and colorectal cancer. Oncotarget 2018, 9, 29820–29841.
- Huynh, J.; Chand, A.; Ernst, M. IL-11 as a therapeutic target to treat colorectal cancer. Cancer Res. 2018, 78, 5733.
- Putoczki, T.L.; Thiem, S.; Loving, A.; Busuttil, R.A.; Wilson, N.J.; Ziegler, P.K.; Nguyen, P.M.; Preaudet, A.; Farid, R.; Edwards, K.M.; et al. Interleukin-11 is the dominant IL-6 family cytokine during gastrointestinal tumorigenesis and can be targeted therapeutically. Cancer Cell 2013, 24, 257–271.
- Yoshizaki, A.; Nakayama, T.; Yamazumi, K.; Yakata, Y.; Taba, M.; Sekine, I. Expression of interleukin (IL)-11 and IL-11 receptor in human colorectal adenocarcinoma: IL-11 up-regulation of the invasive and proliferative activity of human colorectal carcinoma cells. Int. J. Oncol. 2006, 29, 869–876.
- Sun, Q.; Sun, F.; Wang, B.; Liu, S.; Niu, W.; Liu, E.; Peng, C.; Wang, J.; Gao, H.; Liang, B.; et al. Interleukin-8 promotes cell migration through integrin αvβ6 upregulation in colorectal cancer. Cancer Lett. 2014, 354, 245–253.
- Shimizu, M.; Tanaka, N. IL-8-induced O-GlcNAc modification via GLUT3 and GFAT regulates cancer stem cell-like properties in colon and lung cancer cells. Oncogene 2019, 38, 1520–1533.
- Xia, W.; Chen, W.; Zhang, Z.; Wu, D.; Wu, P.; Chen, Z.; Li, C.; Huang, J. Prognostic value, clinicopathologic features and diagnostic accuracy of interleukin-8 in colorectal cancer: A meta-analysis. PLoS ONE 2015, 10, e0123484.
- Miteva, L.D.; Stanilov, N.S.; Deliysky, T.S.; Stanilova, S.A. Significance of−1082A/G polymorphism of IL10 gene for progression of colorectal cancer and IL-10 expression. Tumor Biol. 2014, 35, 12655–12664.
- O’Hara, R.J.; Greenman, J.; MacDonald, A.W.; Gaskell, K.M.; Topping, K.P.; Duthie, G.S.; Kerin, M.J.; Lee, P.W.; Monson, J.R.J. Advanced colorectal cancer is associated with impaired interleukin 12 and enhanced interleukin 10 production. Clin. Cancer Res. 1998, 4, 1943–1948. Available online: https://clincancerres.aacrjournals.org/content/clincanres/4/8/1943.full.pdf (accessed on 17 March 2023).
- Shouval, D.S.; Biswas, A.; Goettel, J.A.; McCann, K.; Conaway, E.; Redhu, N.S.; Mascanfroni, I.D.; Al Adham, Z.; Lavoie, S.; Ibourk, M.; et al. Interleukin-10 receptor signaling in innate immune cells regulates mucosal immune tolerance and anti-inflammatory macrophage function. Immunity 2014, 40, 706–719.
- Doulabi, H.; Rastin, M.; Shabahangh, H.; Maddah, G.; Abdollahi, A.; Nosratabadi, R.; Esmaeili, S.A.; Mahmoudi, M. Analysis of Th22.; Th17 and CD4(+)cells co-producing IL-17/IL-22 at different stages of human colon cancer. Biomed. Pharmacother. 2018, 103, 1101–1106.
- Yu, L.Z.; Wang, H.Y.; Yang, S.P.; Yuan, Z.P.; Xu, F.Y.; Sun, C.; Shi, R.H. Expression of interleukin-22/STAT3 signaling pathway in ulcerative colitis and related carcinogenesis. World J. Gastroenterol. WJG 2013, 19, 2638.
- Wang, C.; Gong, G.; Sheh, A.; Muthupalani, S.; Bryant, E.M.; Puglisi, D.A.; Holcombe, H.; Conaway, E.A.; Parry, N.A.P.; Bakthavatchalu, V.; et al. Interleukin-22 drives nitric oxide-dependent DNA damage and dysplasia in a murine model of colitis-associated cancer. Mucosal Immunol. 2017, 10, 1504–1517.
- Moseley, T.A.; Haudenschild, D.R.; Rose, L.; Reddi, A.H. Interleukin-17 family and IL-17 receptors. Cytokine Growth Factor Rev. 2013, 14, 155–174.
- Razi, S.; Baradaran, N.B.; Keshavarz-Fathi, M.; Rezaei, N. IL-17 and colorectal cancer: From carcinogenesis to treatment. Cytokine 2019, 116, 7–12.
- Alinejad, V.; Dolati, S.; Motallebnezhad, M.; Yousefi, M. The role of IL17B-IL17RB signaling pathway in breast cancer. Biomed. Pharmacother. 2017, 88, 795–803.
- Shamoun, L.; Skarstedt, M.; Andersson, R.E.; Wågsäter, D.; Dimberg, J. Association study on IL-4, IL-4Rα and IL-13 genetic polymorphisms in Swedish patients with colorectal cancer. Clin. Chim. Acta 2018, 487, 101–106.
- Neurath, M.F. IL-23 in inflammatory bowel diseases and colon cancer. Cytokine Growth Factor Rev. 2019, 45, 1–8.
- Zhang, L.; Li, J.; Li, L.; Zhang, J.; Wang, X.; Yang, C.; Li, Y.; Lan, F.; Lin, P. IL-23 selectively promotes the metastasis of colorectal carcinoma cells with impaired Socs3 expression via the STAT5 pathway. Carcinogenesis 2014, 35, 1330–1340.
- Suzuki, H.; Ogawa, H.; Miura, K.; Haneda, S.; Watanabe, K.; Ohnuma, S.; Sasaki, H.; Sase, T.; Kimura, S.; Kajiwara, T.; et al. IL-23 directly enhances the proliferative and invasive activities of colorectal carcinoma. Oncol. Lett. 2012, 4, 199–204.
- Hu, W.H.; Chang, C.D.; Liu, T.T.; Chen, H.H.; Hsiao, C.C.; Kang, H.Y.; Chuang, J.H. Association of sarcopenia and expression of interleukin-23 in colorectal cancer survival. Clin. Nutr. 2021, 40, 5322–5326.

















