Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Zeinab Ezzeddine | -- | 2027 | 2023-04-26 12:34:11 | | | |
| 2 | Camila Xu | Meta information modification | 2027 | 2023-04-27 03:20:01 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Chaaban, T.; Mohsen, Y.; Ezzeddine, Z.; Ghssein, G. Yersiniabactin and Yersinopine. Encyclopedia. Available online: https://encyclopedia.pub/entry/43521 (accessed on 08 May 2026).
Chaaban T, Mohsen Y, Ezzeddine Z, Ghssein G. Yersiniabactin and Yersinopine. Encyclopedia. Available at: https://encyclopedia.pub/entry/43521. Accessed May 08, 2026.
Chaaban, Taghrid, Yehya Mohsen, Zeinab Ezzeddine, Ghassan Ghssein. "Yersiniabactin and Yersinopine" Encyclopedia, https://encyclopedia.pub/entry/43521 (accessed May 08, 2026).
Chaaban, T., Mohsen, Y., Ezzeddine, Z., & Ghssein, G. (2023, April 26). Yersiniabactin and Yersinopine. In Encyclopedia. https://encyclopedia.pub/entry/43521
Chaaban, Taghrid, et al. "Yersiniabactin and Yersinopine." Encyclopedia. Web. 26 April, 2023.
Copy Citation
The pathogenic anaerobic bacteria Yersinia pestis (Y. pestis), which is well known as the plague causative agent, has the ability to escape or inhibit innate immune system responses, which can result in host death even before the activation of adaptive responses. Yersinia pestis produces two metallophores: yersiniabactin, for iron chelation (siderophore), and an opine type metallophore called yersinopine.
Yersinia pestis
pathogenesis
metallophores
metal ions
1. Introduction
The Gram-negative pathogen Yersinia pestis (Y. pestis), which is responsible for causing bubonic, pneumonic, and septicemic plague among humans, emerged from the enteropathogen Yersinia pseudotuberculosis [1]. This pathogen is considered a zoonosis because it is carried by rodents worldwide and transmitted within them through a flea vector (Xenopsylla cheopis) [2][3]. Effective inter-human transmission may occur through aerosols leading to the development of pneumonic plague [4][5]. However, recent evidence shows that inter-human transmission via aerosol is apparently insignificant [6] and may be due to body lice and fleas [1].
Y. pestis evolved multiple factors that enable it to colonize its hosts which include both mammals and insects [7]. An infected flea causes bubonic plague through its bite. Once deposited on the dermis, the bacterium spreads and colonizes the lymph nodes, leading to ‘‘bubo’’ which is the inflammation of the lymph nodes [8].
2. Yersiniabactin
The transition metal iron is crucial for almost all pathogenic bacteria including Y. pestis. In the host metal ions’ scarce environment, bacteria developed various strategies to sequester iron. One of these strategies is the synthesis and secretion of siderophores which are low molecular weight metabolites specific for obtaining ferric ions. The highly pathogenic Y. pestis produces the siderophore yersiniabactin (Ybt), which has shown an important role in the acquisition of iron and murine pathogenicity [9]. Moreover, it was demonstrated that Y. pestis strains with mutations in genes responsible for Ybt biosynthesis or uptake caused an almost complete inability to generate fatal bubonic and pneumonic plague [10][11]. These findings prove that Ybt is associated with virulence besides acquiring iron.
2.1. Yersiniabactin Biosynthesis
The siderophore Ybt is composed of thiazoline rings that are responsible for Fe3+ binding with a formation constant of 4 × 1036 [12]. The synthesis of Ybt is carried on by NRPS (nonribosomal peptide synthetase) mechanism combined with PKS (polyketide synthase). Ybt is constructed from salicylate, a linker malonyl group, three cysteines, and three methyl groups from S-adenosylmethionine [13]. These molecules yield a siderphore whose structure is composed of four rings (one salicylate, two thiazolines, and one thiazolidine) as shown in Figure 1.
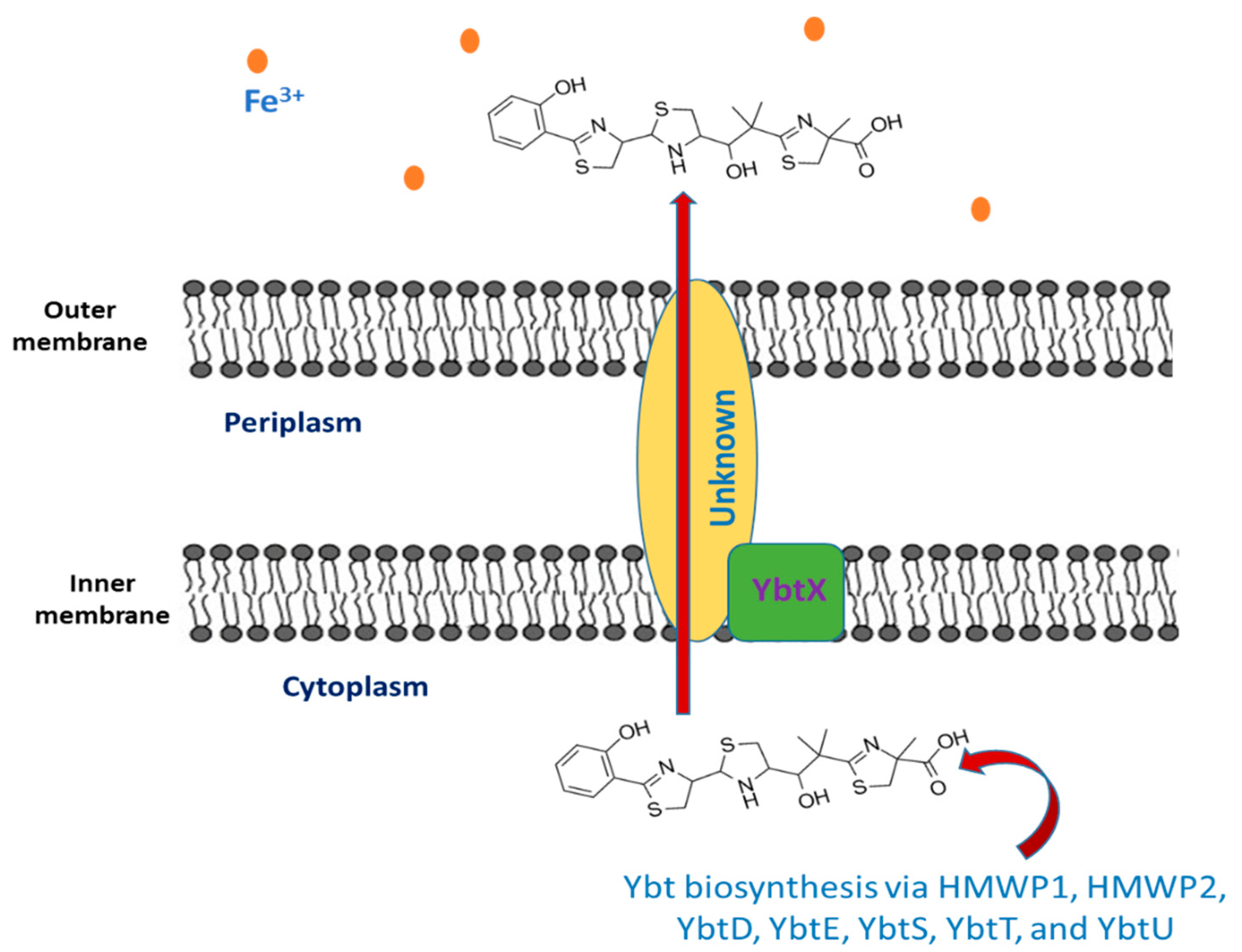
Figure 1. The export of yersiniabactin.
Seven proteins (high molecular weight proteins 1 HMWP1, high molecular weight proteins 2 HMWP2, YbtD, YbtE, YbtS, YbtT, and YbtU) are involved in the synthesis of Ybt. The activated components of Ybt are bound to HMWP1 and HMWP2 via the coenzyme A moiety 4′-phosphopantetheine that is added by YbtD phosphopantetheinyl transferase to the carrier protein domains sites of these enzymes. Salicylate is synthesized from chorismate by YbtS, and then, it is activated through adenylation and transferred to HMWP2 by YbtE. Both HMWP1 and HMWP2 give the framing scaffold for the synthesis of Ybt. The NRPS enzymatic domains present in HMWP2 are responsible for cyclizing and condensing two cysteines, thus producing two thiazoline rings bonded to the salicylate component. The first four domains of the HMWP1 are the PKS module that performs the bis-methylation and the reduction in a malonyl linker moiety. The third molecule of cysteine that forms the final thiazoline ring is cyclized and condensed by two NRPS domains of HMWP1. As for the middle thiazoline ring, YbtU reduces it to thiazolidine. Then, the terminal HMWP1 thioesterase domain releases the synthesized siderophore. It is thought that the assumed type II thioesterase YbtT removes irregular molecules from the enzyme complex [14][15][16][17].
2.2. The Export of Ybt
Once the synthesis of Ybt is completed, it is exported outside the bacterial cell, but the mechanism underlying this process is still unknown. Y. pestis comprise YbtX, which is an inner membrane protein assigned to be a part of the Ybt efflux system because it resembles both AlcS and EntS exporters in alcaligin and enterobactin siderophores in Bordetella and E. coli respectively [18][19][20]. However, experiments showed that a Y. pestis ybtX mutant can secrete Ybt, uptake iron-loaded Ybt, and can grow under iron-scarce conditions as well [19]. The YbtX function is not fully understood, although it is involved in the secretion of Ybt, it is not essential in the process.
2.3. Iron Chelation by Ybt
As mentioned previously, Ybt contains a benzene ring, another thiazolidine ring, and two thiazoline rings. This structure provides five chiral centers (donor positions), enabling it to form a stable complex with trivalent ferric iron cations (Fe3+) with 1:1 complexes. The Ybt potential donor groups are: both the nitrogen found in the first thiazoline ring and the phenolic hydroxy group, either the nitrogen or the sulfur present in the thiazolidine moiety along with the aliphatic hydroxy group, as well as the carboxy group and the nitrogen of the terminal thiazoline ring [21].
2.4. The Intake of Iron-Loaded Ybt
After Ybt chelates iron, the bacterium uptakes the loaded siderophore. The first step includes binding to the TonB-dependent receptor Psn, which is followed by outer membrane translocation. In Gram-negative bacteria, the function of TonB is associated with ExbB and ExbD, yet it is not ascertained in Y. pestis. Then, the iron-loaded Ybt is transported from the periplasm into the cytoplasm via the YbtP-YbtQ ABC transporter. The latter are similar inner membrane proteins with both ATPase and permease domains and are usually components of Type I secretion systems [22][23]. In the cytoplasm, Fe is then released in the cytoplasm from the siderophore by a reduction in ferric ions to ferrous ions or by Ybt degradation. The process of Ybt-Fe3+ uptake is demonstrated in Figure 2. Genes that encode either type of iron release mechanisms are not found within the high pathogenicity island (HPI) that is responsible for encoding most ybt genes. Noteworthily, Y. pestis mutant strains in either psn or tonB demonstrated both growth and iron uptake defects. Moreover, despite the presence of a second TonB-like gene (hasB), it is unable to be an alternative to TonB in taking iron through the Ybt system [24][25]. Moreover, strains with ybtP or ybtQ mutations have a similar phenotype psn mutant and showed reduced iron uptake and growth but without a notable reduction in Ybt production [22].
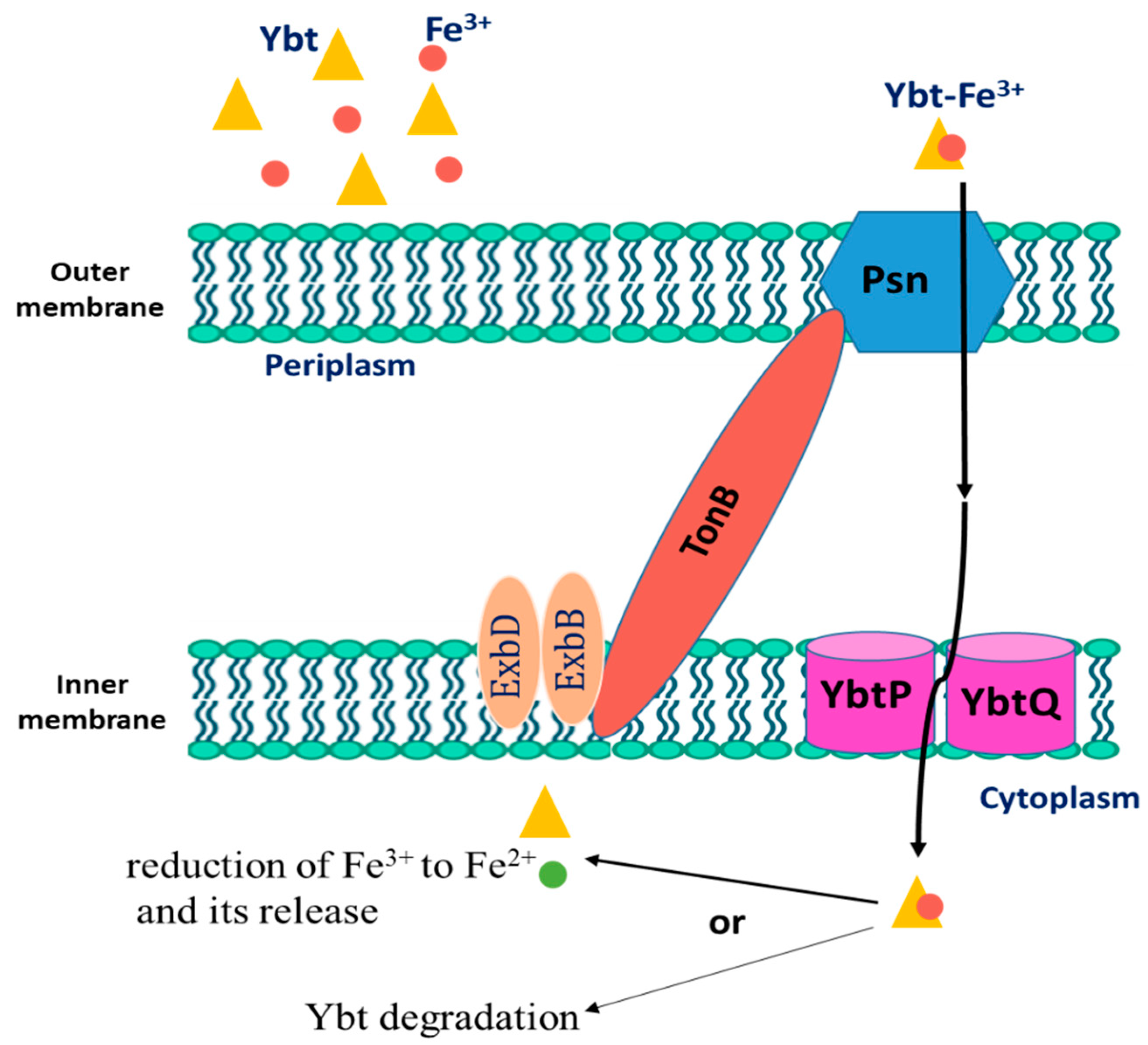
Figure 2. The uptake process of Fe3+-loaded Yersiniabactin (Ybt) by Y. pestis.
2.5. Genetic Regulation of Yersiniabactin (Ybt)
In Y. pestis, the ybt locus which is composed of four operons is responsible for Ybt biosynthesis, uptake, and possibly export. Out of the four operons, two are monocistronic, where one of them contains five genes encoding the biosynthetic enzymes, while the fourth operon genes encode the proteins involved in Ybt uptake and biosynthesis. Except for ybtD, fur, and the transport components (tonB, exbB, and exbC), all other genes involved in the function or regulation or function of the Ybt system are encoded within this locus [26]. The ybt locus is found within the high pathogenicity island (HPI) that was identified, for the first time, in the three Yersinia species causing pathogenesis in mammals. The Y. pestis HPI is found within the pigmentation (pgm) locus and comprises approximately one-third of it. The latter is an unstable wide area of the Y. pestis chromosome consisting of an HPI segment linked to a pigmentation segment [27]. The HPI is distributed widely among the Enterobacteriaceae family members. The IS100 related to HPI of Y. pestis is not found in the other Enterobacteriaceae. The majority of the organisms having the HPI produce the proteins HMWP1 and HMWP2 in iron-scarce conditions and synthesize the siderophore Ybt [26].
As in other Gram-negative bacteria, Y. pestis produces iron uptake regulation protein (Fur), which is responsible for repressing the transcription of the promoters associated with a Fur binding sequence (FBS) once iron is in excess [27]. All of the four ybt operons that lie within the HPI possess FBSs promoters and are repressed via Fur approximately 12-fold (psn), 8-fold (irp2-irp1-ybtUTE), 11-fold (ybtA), and 55-fold (ybtPQXS) upon the growth with 10 μM iron compared to the growth in deferrated, defined media (PMH/PMH2) without additional iron [24][25][26][27][28]. ybtD, on the contrary, that is encoded outside the HPI and pgm locus is not Fur regulated.
2.6. The Role of Yersiniabactin in Overcoming Zinc Restriction
It was recently demonstrated that Ybt binds. thus enhancing the growth Y. pestis lacking ZnuABC transporter in zinc scarce medium. These findings suggest that Ybt may have a role in overcoming zinc nutritional immunity in addition to its role in iron sequestering. To test this assumption, Price et al. used an iron-mediated nutritional immunity defective mouse and demonstrated the contribution of Ybt to virulence independent of iron. In addition, they found that Y. pestis utilizes Ybt to compete for zinc with calprotectin. Furthermore, they discovered that in flea midgut, this pathogen relies on Ybt to survive zinc limitation [29]. Older studies showed that the irp2 gene of Y. pestis, responsible for HMWP2 encoding, is needed for its growth in zinc-deficient medium for strains lacking ZnuABC [30]. Additionally, they showed that YbtX is required as well for the uptake of Ybt-dependent Zn2+ while both Psn and TonB, along with YbtPQ, are not required for zinc acquisition [31]. These results were further proved by Bobrov et al. who demonstrated that strains lacking both ybtX and znu genes were avirulent for bubonic and pneumonic plague in mice models at the time the virulence ability was conserved in strains mutant only for ybtX [32]. All these findings prove that Ybt has a role in the mechanism of zinc acquisition in Y. pestis, enabling it to overcome zinc limitation upon infecting both insects and mammals.
3. Yersinopine
Plants produce the metabolite nicotianamine, which is used in metal ions homeostasis such as zinc, iron, and nickel [33]. It is also synthesized by filamentous fungi [34] along with some mosses [35]. On the other hand, it was found that bacteria produce opine-type metallophores resembling nicotianamine, [36] in particular Staphylococcus aureus, Pseudomonas aeruginosa, and Yersinia pestis. These bacteria produce, respectively, staphylopine, pseudopaline, and yersinopine containing an imidazole ring and three carboxylic groups [37][38]. The structure of yersinopine is shown in Figure 3.
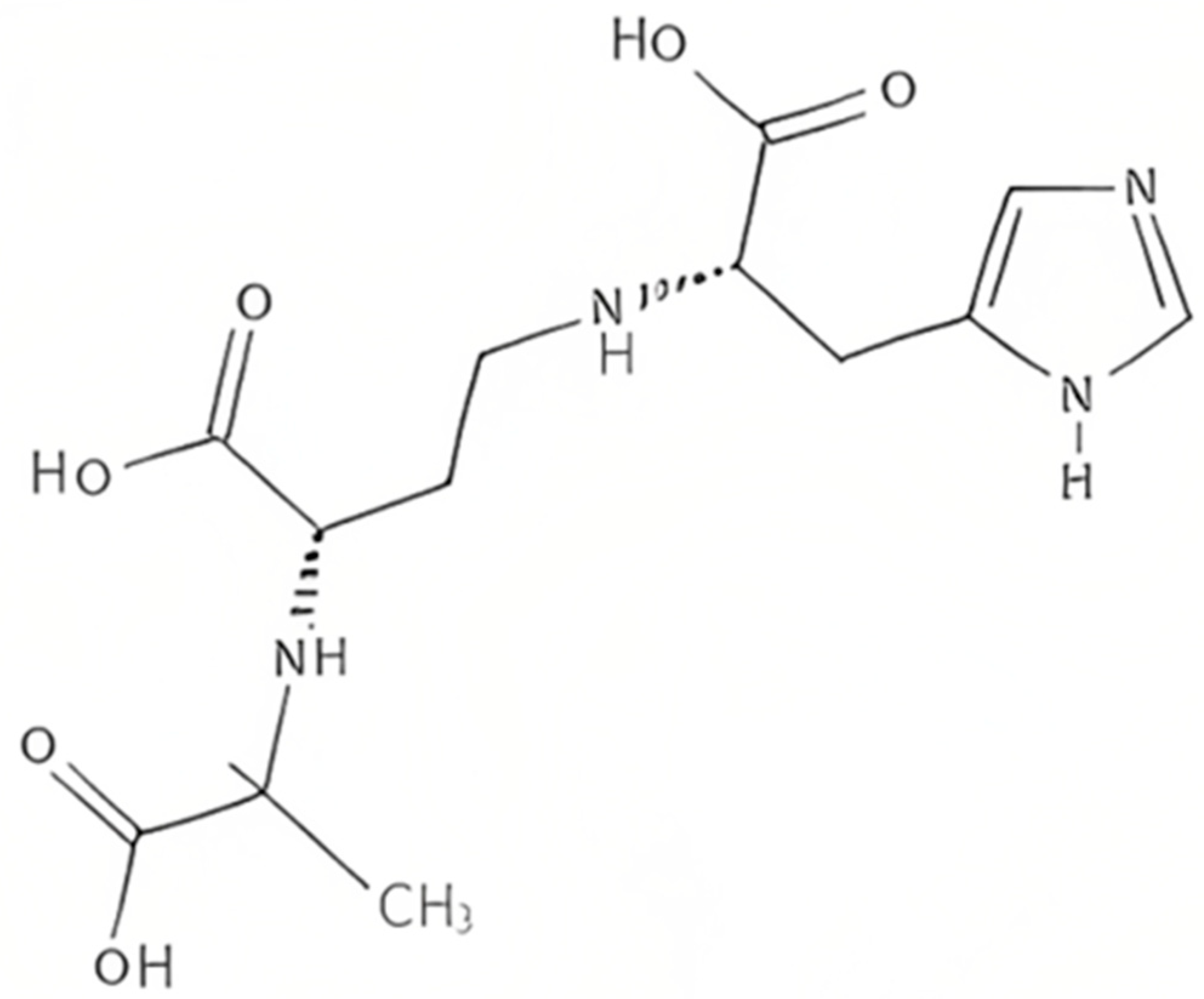
Figure 3. The structure of yersinopine.
Two enzymes are involved in the biosynthesis of opine metallophores, the first is CntL, nicotianamine synthase (NAS), and the second is CntM, an opine dehydrogenase (ODH). The aminoalkyl transferase NAS which is S-adenosyl-L-methionine (SAM)-dependent forms a secondary amine between the aminobutyrate SAM component and amino acid [39]. In P. aeruginosa, the NAS uses L-histidine together with SAM to produce L-His-nicotianamine (L-HisNA), the substrate for P. aeruginosa ODH [40]. However, histidine racemase (CntK), which is a third enzyme produced by S. aureus, generates D-histidine used by the S. aureus NAS to produce D-His-nicotianamine (D-HisNA), the substrate for Staphylococcus aureus ODH [33]. Each one of these ODH binds to its HisNA substrate to perform a reductive condensation with an α-keto acid, which is followed by the reduction in NAD(P)H to generate the final opine metallophore. There exists a different substrate specificity between yersinopine dehydrogenase (YpODH) produced by Y. pestis compared to pseudopaline dehydrogenases. Pyruvate is selected by YpODH as its primary substrate, which is the case with S. aureus ODH while P. aeruginosa ODH is specific for α-ketoglutarate [36]. Such diversity in the opine-type metallophore is a result of possessing or lacking the cntK gene responsible for histidine racemase encoding [36].
It was confirmed that there is a strong link between bacterial infection and opine-type metallophores production in both S. aureus and P. aeruginosa. Starting with S. aureus, it was found that CntA, the receptor of staphylopine, is essential for optimal urease functioning as well as bacterium virulence. The deletion of this receptor causes a decrease in bacteremia and urinary tract infections in murine [41]. Concerning P. aeruginosa, the pseudopaline biosynthetic genes were found to be overexpressed in human burn wound infections [42]. Such overexpression allows surviving metal ions limitations during nutritional immunity. In addition, CntI, the exporter of pseudopaline, plays a crucial role in the growth and survival of P. aeruginosa in cystic fibrosis airway. Similarly, this exporter deletion leads to respiratory infection attenuation [43]. Moreover, the staphylopine exporter is important for S. aureus fitness in abscesses [44]. As for Y. pestis, there is no data currently available indicating a link between the production of yersinopine and virulence and the discovery of this metallophore is so far restricted to studies performed in vitro [45].
References
- Yang, R.; Atkinson, S.; Chen, Z.; Cui, Y.; Du, Z.; Han, Y.; Sebbane, F.; Slavin, P.; Song, Y.; Yan, Y.; et al. Yersinia pestis and Plague: Some Knowns and Unknowns. Zoonoses 2023, 3.
- Eisen, R.J.; Bearden, S.W.; Wilder, A.P.; Montenieri, J.A.; Antolin, M.F.; Gage, K.L. Early-phase transmission of Yersinia pestis by unblocked fleas as a mechanism explaining rapidly spreading plague epizootics. Proc. Natl. Acad. Sci. USA 2006, 103, 15380–15385.
- Hinnebusch, B.J.; Jarrett, C.O.; Bland, D.M. “Fleaing” the plague: Adaptations of Yersinia pestis to its insect vector that lead to transmission. Annu. Rev. Microbiol. 2017, 71, 215–232.
- Lathem, W.W.; Crosby, S.D.; Miller, V.L.; Goldman, W.E. Progression of primary pneumonic plague: A mouse model of infection, pathology, and bacterial transcriptional activity. Proc. Natl. Acad. Sci. USA 2005, 102, 17786–17791.
- Pechous, R.D.; Sivaraman, V.; Stasulli, N.M.; Goldman, W.E. Pneumonic plague: The darker side of Yersinia pestis. Trends Microbiol. 2016, 24, 190–197.
- Dean, K.R.; Krauer, F.; Walløe, L.; Lingjærde, O.C.; Bramanti, B.; Stenseth, N.C.; Schmid, B.V. Human ectoparasites and the spread of plague in Europe during the Second Pandemic. Proc. Natl. Acad. Sci. USA 2018, 115, 1304–1309.
- Demeure, C.E.; Dussurget, O.; Fiol, G.M.; Le Guern, A.-S.; Savin, C.; Pizarro-Cerdá, J. Yersinia pestis and plague: An updated view on evolution, virulence determinants, immune subversion, vaccination, and diagnostics. Genes Immun. 2019, 20, 357–370.
- Montminy, S.W.; Khan, N.; McGrath, S.; Walkowicz, M.J.; Sharp, F.; Conlon, J.E.; Fukase, K.; Kusumoto, S.; Sweet, C.; Miyake, K.; et al. Virulence factors of Yersinia pestis are overcome by a strong lipopolysaccharide response. Nat. Immunol. 2006, 7, 1066–1073.
- Rakin, A.; Saken, E.; Harmsen, D.; Heesemann, J. The pesticin receptor of Yersinia enterocolitica: A novel virulence factor with dual function. Mol. Microbiol. 1994, 13, 253–263.
- Fetherston, J.D.; Kirillina, O.; Bobrov, A.G.; Paulley, J.T.; Perry, R.D. The yersiniabactin transport system is critical for the pathogenesis of bubonic and pneumonic plague. Infect. Immun. 2010, 70, 2045–2052.
- Lee-Lewis, H.; Anderson, D.M. Absence of inflammation and pneumonia during infection with nonpigmented Yersinia pestis reveals a new role for the pgm locus in pathogenesis. Infect. Immun. 2009, 78, 220–230.
- Perry, R.D.; Fetherston, J.D. Yersiniabactin iron uptake: Mechanisms and role in Yersinia pestis pathogenesis. Microbes Infect. 2011, 13, 808–817.
- Chambers, C.E.; McIntyre, D.D.; Mouck, M.; Sokol, P.A. Physical and structural characterization of yersiniophore, a siderophore produced by clinical isolates of Yersinia enterocolitica. Biometals 1996, 9, 157–167.
- Bearden, S.W.; Fetherston, J.D.; Perry, R.D. Genetic organization of the yersiniabactin biosynthetic region and construction of avirulent mutants in Yersinia pestis. Infect. Immun. 1997, 65, 1659–1668.
- Gehring, A.M.; DeMoll, E.; Fetherston, J.D.; Mori, I.; Mayhew, G.F.; Blattner, F.R.; Walsh, C.T.; Perry, R.D. Iron acquisition in plague: Modular logic in enzymatic biogenesis of yersiniabactin by Yersinia pestis. Chem. Biol. 1998, 5, 573–586.
- Bobrov, A.G.; Geoffroy, V.A.; Perry, R.D. Yersiniabactin production requires the thioesterase domain of HMWP2 and YbtD, a putative phosphopantetheinylate transferase. Infect. Immun. 2002, 70, 4204–4214.
- Geoffroy, V.A.; Fetherston, J.D.; Perry, R.D. Yersinia pestis YbtU and YbtT are involved in synthesis of the siderophore yersiniabactin but have different effects on regulation. Infect. Immun. 2000, 68, 4452–4461.
- Miller, M.C.; Fetherston, J.D.; Pickett, C.L.; Bobrov, A.G.; Weaver, R.H.; DeMoll, E.; Perry, R.D. Reduced synthesis of the Ybt siderophore or production of aberrant Ybt-like molecules activates transcription of yersiniabactin genes in Yersinia pestis. Microbiology 2010, 156, 2226–2238.
- Furrer, J.L.; Sanders, D.N.; Hook-Barnard, I.G.; McIntosh, M.A. Export of the siderophore enterobactin in Escherichia coli: Involvement of a 43 kDa membrane exporter. Mol. Microbiol. 2002, 44, 1225–1234.
- Brickman, T.J.; Armstrong, S.K. Bordetella AlcS transporter functions in alcaligin siderophore export and is central to inducer sensing in positive regulation of alcaligin system gene expression. J. Bacteriol. 2005, 187, 3650–3661.
- Drechsel, H.; Stephan, H.; Lotz, R.; Haag, H.; Zähner, H.; Hantke, K.; Jung, G. Structure elucidation of yersiniabactin, a siderophore from highly virulent Yersinia strains. Liebigs Ann. 1995, 10, 1727–1733.
- Fetherston, J.D.; Bertolino, V.J.; Perry, R.D. YbtP and YbtQ: Two ABC transporters required for iron uptake in Yersinia pestis. Mol. Microbiol. 1999, 32, 289–299.
- Buchanan, S.K. Type I secretion and multidrug efflux: Transport through the TolC channel-tunnel. Trends Biochem. Sci. 2001, 28, 3–6.
- Fetherston, J.D.; Lillard, J.W.; Perry, R.D. Analysis of the pesticin receptor from Yersinia pestis: Role in iron-deficient growth and possible regulation by its siderophore. J. Bacteriol. 1995, 177, 1824–1833.
- Perry, R.D.; Shah, J.; Bearden, S.W.; Thompson, J.M.; Fetherston, J.D. Yersinia pestis TonB: Role in iron, heme and hemoprotein utilization. Infect. Immun. 2003, 71, 4159–4162.
- Lesic, B.; Carniel, E. Yersinia Molecular and Cellular Biology. In Horizon Bioscience; Carniel, E., Hinnebusch, B.J., Eds.; UCL Press: Norfolk, UK, 2004; pp. 285–306.
- Leal-Balbino, T.C.; Leal, N.C.; Nascimento, M.G.M.D.; Oliveira, M.B.M.D.; Balbino, V.D.Q.; Almeida, A.M.P.D. The pgm locus and pigmentation phenotype in Yersinia pestis. Genet. Mol. Biol. 2006, 29, 126–131.
- Gao, H.; Zhou, D.; Li, Y.; Guo, Z.; Han, Y.; Song, Y.; Zhai, J.; Du, Z.; Wang, X.; Lu, J.; et al. The ironresponsive Fur regulon in Yersinia pestis. J. Bacteriol. 2008, 190, 3063–3075.
- Price, S.L.; Vadyvaloo, V.; DeMarco, J.K.; Brady, A.; Gray, P.A.; Kehl-Fie, T.E.; Garneau-Tsodikova, S.; Perry, R.D.; Lawrenz, M.B. Yersiniabactin contributes to overcoming zinc restriction during Yersinia pestis infection of mammalian and insect hosts. Proc. Natl. Acad. Sci. USA 2021, 118, 2104073118.
- Mastropasqua, M.C.; D′Orazio, M.; Cerasi, M.; Pacello, F.; Gismondi, A.; Canini, A.; Canuti, L.; Consalvo, A.; Ciavardelli, D.; Chirullo, B.; et al. Growth of Pseudomonas aeruginosa in zinc poor environments is promoted by a nicotianamine-related metallophore. Mol. Microbiol. 2017, 106, 543–561.
- Bobrov, A.G.; Kirillina, O.; Fetherston, J.D.; Miller, M.C.; Burlison, J.A.; Perry, R.D. The Yersinia pestis siderophore, yersiniabactin, and the ZnuABC system both contribute to zinc acquisition and the development of lethal septicaemic plague in mice. Mol. Microbiol. 2014, 93, 759–775.
- Bobrov, A.G.; Kirillina, O.; Fosso, M.Y.; Fetherston, J.D.; Miller, M.C.; VanCleave, T.T.; Burlison, J.A.; Arnold, W.K.; Lawrenz, M.B.; Garneau-Tsodikova, S.; et al. Zinc transporters YbtX and ZnuABC are required for the virulence of Yersinia pestis in bubonic and pneumonic plague in mice. Metallomics 2017, 9, 757–772.
- Reichman, S.M.; Parker, D.R. Revisiting the metal-binding chemistry of nicotianamine and 2′-deoxymugineic acid. Implications for iron nutrition in strategy II plants. Plant Physiol. 2002, 129, 1435–1438.
- Trampczynska, A.; Böttcher, C.; Clemens, S. The transition metal chelator nicotianamine is synthesized by filamentous fungi. FEBS Lett. 2006, 580, 3173–3178.
- Burkhead, J.L.; Gogolin Reynolds, K.A.; Abdel-Ghany, S.E.; Cohu, C.M.; Pilon, M. Copper homeostasis. New Phytol. 2009, 182, 799–816.
- Ghssein, G.; Ezzeddine, Z. The Key Element Role of Metallophores in the Pathogenicity and Virulence of Staphylococcus aureus: A Review. Biology 2022, 11, 1525.
- Ghssein, G.; Ezzeddine, Z. A Review of Pseudomonas aeruginosa Metallophores: Pyoverdine, Pyochelin and Pseudopaline. Biology 2022, 11, 1711.
- Ghssein, G.; Brutesco, C.; Ouerdane, L.; Fojcik, C.; Izaute, A.; Wang, S.; Hajjar, C.; Lobinski, R.; Lemaire, D.; Richaud, P.; et al. Biosynthesis of a broad-spectrum nicotianamine-like metallophore in Staphylococcus aureus. Science 2016, 352, 1105–1109.
- McFarlane, J.S.; Lamb, A.L. Biosynthesis of an opine metallophore by Pseudomonas aeruginosa. Biochemistry 2017, 56, 5967–5971.
- Andrews, S.C.; Robinson, A.K.; Rodríguez Quiñones, F. Bacterial iron homeostasis. FEMS Microbiol. Rev. 2003, 27, 215–237.
- Laffont, C.; Brutesco, C.; Hajjar, C.; Cullia, G.; Fanelli, R.; Ouerdane, L.; Cavelier, F.; Arnoux, P. Simple rules govern the diversity of bacterial nicotianamine-like metallophores. Biochem. J. 2019, 476, 2221–2233.
- Remy, L.; Carrière, M.; Derré-Bobillot, A.; Martini, C.; Sanguinetti, M.; Borezée-Durant, E. The Staphylococcus aureus Opp1 ABC transporter imports nickel and cobalt in zinc depleted conditions and contributes to virulence: Nickel and cobalt uptake in Staphylococcus aureus. Mol. Microbiol. 2013, 87, 730–743.
- Bielecki, P.; Puchałka, J.; Wos-Oxley, M.L.; Loessner, H.; Glik, J.; Kawecki, M.; Nowak, M.; Tümmler, B.; Weiss, S.; dos Santos, V.A.P.M. In-Vivo Expression Profiling of Pseudomonas aeruginosa Infections Reveals Niche-Specific and Strain-Independent Transcriptional Programs. PLoS ONE 2011, 6, e24235.
- Gi, M.; Lee, K.M.; Kim, S.C.; Yoon, J.H.; Yoon, S.S.; Choi, J.Y. A novel siderophore system is essential for the growth of Pseudomonas aeruginosa in airway mucus. Sci. Rep. 2015, 5, 14644.
- Ding, Y.; Fu, Y.; Lee, J.C.; Hooper, D.C. Staphylococcus aureus NorD, a Putative Efflux Pump Coregulated with the Opp1 Oligopeptide Permease, Contributes Selectively to Fitness In Vivo. J. Bacteriol. 2012, 194, 6586–6593.
More
Information
Subjects:
Microbiology
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.2K
Revisions:
2 times
(View History)
Update Date:
27 Apr 2023
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





