Video Upload Options
The genomes of polyomaviruses are characterized by their tripartite organization with an early region, a late region and a noncoding control region (NCCR). The early region encodes proteins involved in replication and transcription of the viral genome, while expression of the late region generates the capsid proteins. Transcription regulatory sequences for expression of the early and late genes, as well as the origin of replication are encompassed in the NCCR. Cell tropism of polyomaviruses not only depends on the appropriate receptors on the host cell, but cell-specific expression of the viral genes is also governed by the NCCR.
Note: The following contents are extract from your paper. The entry will be online only after author check and submit it.
1. Introduction: Human Polyomaviruses
Polyomaviruses (PyVs) are non-enveloped viruses that are typically 40–45 nm in diameter, and that possess a double-stranded circular genome of around 5000 base-pairs. Birds and mammals, including humans, are natural hosts for PyVs [1][2]. Recently, PyVs have also been isolated from fish [3][4]. So far, 15 different polyomaviruses have been isolated from human samples. The first human polyomaviruses, BKPyV and JCPyV, were identified in 1971 [5][6]. In 2007, two new human polyomaviruses (Karolinska Institute PyV (KIPyV) [7] and Washington University PyV (WUPyV) [8] were detected, and in the following years, Merkel cell PyV (MCPyV) [9], HPyV6 [10], HPyV7 [10], Trichodisplasia spinulosa PyV (TSPyV) [11], HPyV9 [12], HPyV10 [13], Saint Louis PyV (STLPyV) [14], HPyV12 [15], New Jersey PyV (NJPyV) [16], Lyon IARC PyV (LIPyV) [17], and Quebec PyV [18] have been described. Their original source of isolation and association with human diseases is summarized in Table 1.
Table 1. The novel human polyomaviruses, their original source of isolation and their association with human diseases.
|
Virus |
Original Source |
Associated Disease |
Reference |
|
KIPyV |
Nasopharyngeal aspirate |
None |
[7] |
|
WUPyV |
Bronchoavelar lavage |
None |
[8] |
|
MCPyV |
Merkel cell carcinoma |
None |
[9] |
|
HPyV6 |
Healthy skin |
Pruritic skin eruption in immunocompromised patients |
[10] |
|
HPyV7 |
Healthy skin |
Pruritic skin eruption in immunocompromised patients |
[10] |
|
TSPyV |
Trichodysplasia spinulosa spicules |
Trichodysplasia spinulosa |
[11] |
|
HPyV9 |
Serum from renal transplant recipient |
None |
[12] |
|
HPyV10 |
Condyloma specimens from a patient with WHIM * syndrome |
None |
[13] |
|
STLPyV |
Stool sample from a healthy 15-month-old child |
None |
[14] |
|
HPyV12 |
Liver sample from patient with malignant disease |
None |
[15] |
|
NJPyV |
Muscle biopsy from a pancreatic transplant patient |
None |
[16] |
|
LIPyV |
Skin swab |
None |
[17] |
|
QPyV |
Stool sample from 85-year old hospital patient |
None |
[18] |
* warts, hypogammaglobulinemia, infections, and myelokathexis.
Whether all of these are genuine human polyomaviruses (HPyVs) remains to be determined. BKPyV, JCPyV, KIPyV, WUPyV, MCPyV, HPyV6, HPyV, TSPyV, HPyV9, HPyV10, STLPyV, HPyV12, and NJPyV are classified as human polyomaviruses by the International Committee of Taxonomy of Viruses [19][20], LIPyV has only been very recently described, while LIPyV DNA was originally detected in human skin [17]. LIPyV seroreactivity in healthy individuals is ~5% in healthy individuals and much lower than the seroprevalence of the other HPyVs, which is between 50–100% [21]. Accordingly, LIPyV DNA was not detected or present in <2% of examined skin, eyebrow hair, gargle samples and tonsil samples [22][23]. Moreover, LIPyV DNA was frequently detected in the feces of cats [24], suggesting that it may be a feline PyV rather than a HPyV. QPyV DNA was detected in the feces of one patient [18], and the seroprevalence of this PyV has not been examined. Despite its original identification in human liver, gastro-intestinal tract and colon tissue and a VP1 seropositivity (respectively LT seropositivity) between ~20–90% (respectively 30–40%) in healthy adults or malignant and non-malignant gastro-intestinal tract patients [25], HPyV12 DNA could not be detected in numerous human samples from different sources [26][27][28][29][30]. Moreover, the group of Feltkamp reported that HPyV12 seroprevalence is only around 5% [31]. A nearly identical HPyV12 variant was isolated from shrew, suggesting that HPyV12 may be transmitted from shrew to humans, or that human HPyV12 positive samples were contaminated [32].
2. The Polyomavirus Genome: The Noncoding Control Region
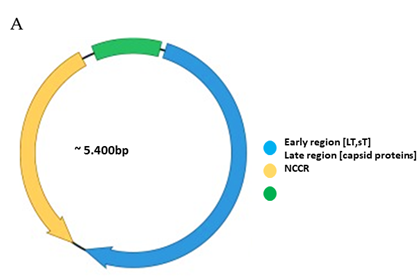
Functionally, the PyV genome is tripartite consisting of the early region, the late region, and the noncoding control region (NCCR) (Figure 1A). The early region codes for regulatory proteins involved in replication and transcription of the viral genome. The major early proteins are large T-antigen (LT) and small t-antigen (sT). The late region codes for the structural proteins VP1, VP2 and VP3 that form the capsid. VP1 is the major capsid protein, while VP2 and VP3 are the minor capsid proteins. However, not all PyVs express VP3 [33]. Interspersed between the early and late region are sequences that do not code for viral proteins, and is referred to as the NCCR.
B
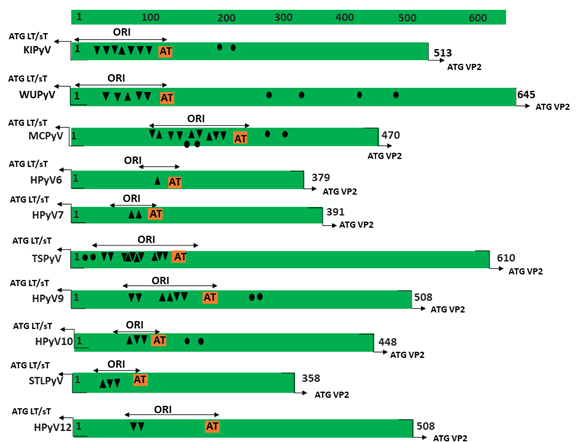
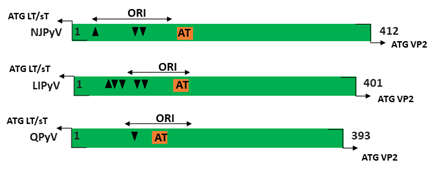
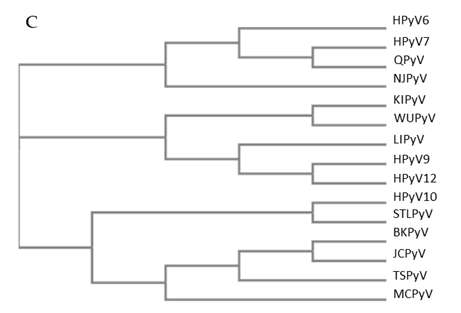
Figure 1. Genomic organization of the human polyomaviruses (HPyVs) genome and the structure of the noncoding control region (NCCR). (A) The circular dsDNA genome consists of the early and late regions that encode regulatory and structural proteins, respectively. Interspersed is the NCCR. (B) Schematic presentation of the NCCR of the novel HPyVs. The NCCR is the region between the start codon of Large T antigen (LT) and Small T antigen (sT) and the start codon of VP2. The AT-rich region (AT), repeated sequences (black dots), and LT binding motifs (upward pointing triangle = 5′-GRGGC-3′; downward pointing triangle = 5′-GCCYC-3′) are shown. (C) Phylogenetic tree bases on NCCR sequences of the different HPyVs. This is a neighbor-joining tree without distance corrections using Clustal Omega multiple sequence alignment [34].
Studies with simian virus 40 (SV40 or Macaca mulatta polyomavirus 1) and murine polyomaviruses have been pivotal in unveiling the functions of this region. The SV40 NCCR contains the origin of replication (ori), which consists of GRGGC motifs to which LT binds and is flanked by an AT-rich sequence and an easily denaturated imperfect palindrome [35][36]. Binding of LT to these motifs is also involved in regulation of viral transcription [37][38]. The NCCR also contains promoter and enhancer elements that control early and late transcription [39][40]. SV40 directly isolated from its natural host, rhesus monkey, has a NCCR that consists of an AT-rich region, triple GC-rich 21 base-pairs (bp) repeats, and a single 72 bp element. The 21 bp repeats contain the LT binding motif (GRGGC; [41][42]). This NCCR organization is known as the archetype. SV40 adapted to grow in cell culture has a duplication of this 72 bp element, with this type of NCCR referred to as prototype [43][44]. SV40 isolated from human tumors usually contain a single 72 bp repeat [43]. Rearrangements in the SV40 NCCR affect viral transcription and replication, as well as oncogenic properties of the virus [45][46] The Mouse polyomavirus (Mus musculus polyomavirus 1; MPyV) NCCR encompasses the ori consisting of an AT-tract and a GC-rich (LT binding motifs) inverted repeat, and the transcription regulatory domains A (or α) and B (or β), C and D [47][48][49][50]. Alterations in the MPyV NCCR have an effect on viral replication in cell culture and in the host, the host range, and in vitro transformation[51][52][53][54][55].
The NCCR of the HPyVs varies between 267 bp (JCPyV CY-strain; accession number AB038249) to 645 bp (WUPyV prototype; accession number NC_009539) (see Figure S1 for the NCCR sequences of the novel HPyV), and similar to the NCCR of SV40 and MPyV, the NCCR of HPyVs also contain the origin of replication, LT binding motifs, and AT-rich region (Figure 1B). This region of the genome displays little or no sequence identity between the different HPyV species (Figure S2). A neighbor-joining tree without distance corrections shows which NCCRs are most closely related (Figure 1C).
The diversification based on the presence of a certain NCCR rearranged structure contributed to determining HPyVs strains as “archetype” or prototype”. The importance of the NCCR rearrangements during HPyVs infection became obvious when different strains of JCPyV were examined. The archetype JCPyV NCCR strain (CY) is divided into six boxes named A (36 bp), B (23 bp), C (55 bp), D (66 bp), E (18 bp), and F (69 bp) and contains the origin of replication (ORI), the promoter and the enhancer elements [56]. The NCCR harbored transcription factor binding sites such as the nuclear transcription factor-1 (NF1), a JCPyV cell-specific regulator of promoter and enhancer activity [57][58], the activating protein 1 (AP1), involved in JCPyV early gene expression [59], and the specificity protein-1 (SP1) able to regulate JCPyV transcription [60]. The archetype NCCR is considered the transmissible form of the virus among the population, and could be released into the urine of healthy individuals due to periodic and subclinical reactivation in the kidney [61][62]. In contrast, in the context of immunosuppression or during immunomodulatory therapy or in AIDS patients, JCPyV can reactivate from latency to cause a fatal pathology of the central nervous system (CNS), known as progressive multifocal leukoencephalopathy (PML) [61]. JCPyV variants carrying rearranged NCCR were usually isolated from PML patients. The prototype Mad-1 strain is the most studied variant of JCPyV and is characterized by 98-bp tandem repeats in the NCCR late proximal region (arranged as ORI-A-C-E-A-C-E-F), and is able to increase viral gene expression in human glial cells, thereby indicating that it is involved in controlling cell gene expression [63][64][65]. The enhancer repeats found in the Mad-1 strain are lacking in the archetype JCPyV strains isolated from the urine of healthy individuals [64]. Additional NCCR rearrangements are implicated in the development of the JCPyV pathogenic strains. In fact, in a significant proportion of JCPyV archetype isolates, short deletions or duplications were observed, corroborating that this region is highly unstable [66][67]. Therefore, it is possible to assume that subsequent archetypal NCRR rearrangements could determine the onset of PML strains, such as Mad-1 [68].
Based on the occurrence in the NCRR of transcriptional enhancer repeat elements, BKPyV isolates can also be identified as archetype and prototype strains. The archetype BKPyV WW strain, characterized by five blocks named O (35 bp), which includes the origin of replication and a TATA-box, P (68 bp), Q (39 bp), R (63 bp), and S (63 bp), containing TATA-like elements and the regulatory regions for early and late genes expression, is considered the infectious strain, shed in the urine of immunocompetent individuals [69][70][71]. Approximately 30 transcription factor binding sites are in silico predicted: SP1 has been the most extensively studied [72][73][74], although the additional role played by other transcription factors such as NF1, ETS1, NFκB, the glucocorticoid and progesterone receptors, and CREB were evidenced in several studies [75][76][77].
Similarly to JCPyV, the plausible instability of the archetype BKPyV NCRR could contribute to the development of the prototype strains, which is able to cause polyomavirus-associated nephropathy in kidney transplant recipients and hemorrhagic cystitis in hematopoietic stem cell transplant recipients [78][79][80][81]. The Dunlop strain, the most salient prototype strain, was isolated from a kidney transplant recipient with ureteral stenosis [82]
Additional NCCR structures have been described for both viruses [85][86][87][88]. In particular, the presence of a common pattern of JCPyV NCCR rearrangement, such as the D-box deletion, can be considered a hallmark needed for the initial NCCR rearrangements critical co-factor for the development of PML in immunosuppressed individuals [88][89]. Besides the triplication of the P region, rearrangements of BKPyV NCCR involve the adjacent O and Q blocks. Differently, the S block is always retained, hence highlighting the importance of these nucleotide sequences. NCCR mutations were also observed during in vitro JCPyV and BKPyV cultivation, confirming that NCRR variants could arise after prolonged propagation of the viruses in cells [90][91]. The mechanisms by which both viruses determine relevant human diseases are not established, but it is accepted that the regulation of gene expression in HPyVs plays a role in determining the viral tropism, and in the promotion of pathogenesis progression [92].
Little is known about the genetic diversity of the NCCRs from the novel HPyVs and the biological relevance in terms of viral transcription, replication, and possible pathogenic properties. In this review, we provide an overview of the mutations in the NCCR, which is defined as the sequence between the start codon of the LT/sT gene and the start codon of the VP2 gene, of the novel HPyVs and their known effect on promoter activity. We discuss how NCCR rearrangements may affect the binding of putative transcription factors, and whether specific NCCR configurations are associated with disease.
References
- DeCaprio, J.A.; Garcea, R.L. A cornucopia of human polyomaviruses. Nat. Rev. Microbiol. 2013, 11, 264–276.
- Moens, U.; Krumbholz, A.; Ehlers, B.; Zell, R.; Johne, R.; Calvignac-Spencer, S.; Lauber, C. Biology, evolution, and medical importance of polyomaviruses: An update. Infect. Genet. Evol. 2017, 54, 18–38.
- Peretti, A.; FitzGerald, P.C.; Bliskovsky, V.; Pastrana, D.V.; Buck, C.B. Genome Sequence of a Fish-Associated Polyomavirus, Black Sea Bass (Centropristis striata) Polyomavirus 1. Genome Announc. 2015, 3, e01476-14.
- Buck, C.B.; Van Doorslaer, K.; Peretti, A.; Geoghegan, E.M.; Tisza, M.J.; An, P.; Katz, J.P.; Pipas, J.M.; McBride, A.A.; Camus, A.C.; et al. The Ancient Evolutionary History of Polyomaviruses. PLoS Pathog. 2016, 12, e1005574.
- Gardner, S.D.; Field, A.M.; Coleman, D.V.; Hulme, B. New human papovavirus (B.K.) isolated from urine after renal transplantation. Lancet 1971, 1, 1253–1257.
- Padgett, B.L.; Walker, D.L.; ZuRhein, G.M.; Eckroade, R.J.; Dessel, B.H. Cultivation of papova-like virus from human brain with progressive multifocal leucoencephalopathy. Lancet 1971, 1, 1257–1260.
- Allander, T.; Andreasson, K.; Gupta, S.; Bjerkner, A.; Bogdanovic, G.; Persson, M.A.; Dalianis, T.; Ramqvist, T.; Andersson, B. Identification of a third human polyomavirus. J. Virol. 2007, 81, 4130–4136.
- Gaynor, A.M.; Nissen, M.D.; Whiley, D.M.; Mackay, I.M.; Lambert, S.B.; Wu, G.; Brennan, D.C.; Storch, G.A.; Sloots, T.P.; Wang, D. Identification of a novel polyomavirus from patients with acute respiratory tract infections. PLoS Pathog. 2007, 3, e64.
- Feng, H.; Shuda, M.; Chang, Y.; Moore, P.S. Clonal integration of a polyomavirus in human Merkel cell carcinoma. Science 2008, 319, 1096–1100.
- Schowalter, R.M.; Pastrana, D.V.; Pumphrey, K.A.; Moyer, A.L.; Buck, C.B. Merkel cell polyomavirus and two previously unknown polyomaviruses are chronically shed from human skin. Cell Host Microbe 2010, 7, 509–515.
- van der Meijden, E.; Janssens, R.W.; Lauber, C.; Bouwes Bavinck, J.N.; Gorbalenya, A.E.; Feltkamp, M.C. Discovery of a new human polyomavirus associated with trichodysplasia spinulosa in an immunocompromized patient. PLoS Pathog. 2010, 6, e1001024.
- Scuda, N.; Hofmann, J.; Calvignac-Spencer, S.; Ruprecht, K.; Liman, P.; Kühn, J.; Hengel, H.; Ehlers, B. A novel human polyomavirus closely related to the african green monkey-derived lymphotropic polyomavirus. J. Virol. 2011, 85, 4586–4590.
- Buck, C.B.; Phan, G.Q.; Raiji, M.T.; Murphy, P.M.; McDermott, D.H.; McBride, A.A. Complete genome sequence of a tenth human polyomavirus. J. Virol. 2012, 86, 10887.
- Lim, E.S.; Reyes, A.; Antonio, M.; Saha, D.; Ikumapayi, U.N.; Adeyemi, M.; Stine, O.C.; Skelton, R.; Brennan, D.C.; Mkakosya, R.S.; et al. Discovery of STL polyomavirus, a polyomavirus of ancestral recombinant origin that encodes a unique T antigen by alternative splicing. Virology 2013, 436, 295–303.
- Korup, S.; Rietscher, J.; Calvignac-Spencer, S.; Trusch, F.; Hofmann, J.; Moens, U.; Sauer, I.; Voigt, S.; Schmuck, R.; Ehlers, B. Identification of a novel human polyomavirus in organs of the gastrointestinal tract. PLoS ONE 2013, 8, e58021.
- Mishra, N.; Pereira, M.; Rhodes, R.H.; An, P.; Pipas, J.M.; Jain, K.; Kapoor, A.; Briese, T.; Faust, P.L.; Lipkin, W.I. Identification of a novel polyomavirus in a pancreatic transplant recipient with retinal blindness and vasculitic myopathy. J. Infect. Dis. 2014, 210, 1595–1599.
- Gheit, T.; Dutta, S.; Oliver, J.; Robitaille, A.; Hampras, S.; Combes, J.D.; McKay-Chopin, S.; Le Calvez-Kelm, F.; Fenske, N.; Cherpelis, B.; et al. Isolation and characterization of a novel putative human polyomavirus. Virology 2017, 506, 45–54.
- Ondov, B.D.; Starrett, G.J.; Sappington, A.; Kostic, A.; Koren, S.; Buck, C.B.; Phillippy, A.M. Mash Screen: High-throughput sequence containment estimation for genome discovery. Genome Biol. 2019, 20, 232.
- Calvignac-Spencer, S.; Feltkamp, M.C.; Daugherty, M.D.; Moens, U.; Ramqvist, T.; Johne, R.; Ehlers, B. A taxonomy update for the family Polyomaviridae. Arch. Virol. 2016, 161, 1739–1750.
- Moens, U.; Calvignac-Spencer, S.; Lauber, C.; Ramqvist, T.; Feltkamp, M.C.W.; Daugherty, M.D.; Verschoor, E.J.; Ehlers, B. ICTV Report Consortium. ICTV Virus Taxonomy Profile: Polyomaviridae. J. Gen. Virol. 2017, 98, 1159–1160.
- Kamminga, S.; van der Meijden, E.; Feltkamp, M.C.W.; Zaaijer, H.L. Seroprevalence of fourteen human polyomaviruses determined in blood donors. PLoS ONE 2018, 13, e0206273.
- Kourieh, A.; Combes, J.D.; Tommasino, M.; Dalstein, V.; Clifford, G.M.; Lacau St Guily, J.; Clavel, C.; Franceschi, S.; Gheit, T. For The Split Study, G. Prevalence and risk factors of human polyomavirus infections in non-malignant tonsils and gargles: The SPLIT study. J. Gen. Virol. 2018, 99, 1686–1698.
- Wang, Y.; Keinonen, A.; Koskenmies, S.; Pitkänen, S.; Fyhrquist, N.; Sadeghi, M.; Mäkisalo, H.; Söderlund-Venermo, M.; Hedman, K. Occurrence of newly discovered human polyomaviruses in skin of liver transplant recipients and their relation with squamous cell carcinoma in situ and actinic keratosis—a single-center cohort study. Transpl. Int. 2019, 32, 516–522.
- Fahsbender, E.; Altan, E.; Estrada, M.; Seguin, M.A.; Young, P.; Leutenegger, C.M.; Delwart, E. Lyon-IARC Polyomavirus DNA in Feces of Diarrheic Cats. Microbiol. Resour. Announc. 2019, 8
- Gaboriaud, P.; Ferté, M.; Arnold, F.; Leblond, V.; Nicol, J.; Debare, H.; Le Meur, M.; Martini, F.; Tognon, M.; Touzé, A. Age-specific seroprevalence of human polyomavirus 12 and Saint Louis and New Jersey polyomaviruses. Emerg. Microbes Infect. 2018, 7, 22.
- Bialasiewicz, S.; Rockett, R.J.; Barraclough, K.A.; Leary, D.; Dudley, K.J.; Isbel, N.M.; Sloots, T.P. Detection of Recently Discovered Human Polyomaviruses in a Longitudinal Kidney Transplant Cohort. Am. J. Transpl. 2016, 16, 2734–2740.
- Li, K.; Zhang, C.; Zhao, R.; Xue, Y.; Yang, J.; Peng, J.; Jin, Q. The prevalence of STL polyomavirus in stool samples from Chinese children. J. Clin. 2015, 66, 19–23.
- Herberhold, S.; Hellmich, M.; Panning, M.; Bartok, E.; Silling, S.; Akgül, B.; Wieland, U. Human polyomavirus and human papillomavirus prevalence and viral load in non-malignant tonsillar tissue and tonsillar carcinoma. Med. Microbiol. Immunol. 2017, 206, 93–103.
- Bergallo, M.; Daprà, V.; Fava, P.; Ponti, R.; Calvi, C.; Montanari, P.; Novelli, M.; Quaglino, P.; Galliano, I.; Fierro, M.T. DNA from Human Polyomaviruses, MWPyV, HPyV6, HPyV7, HPyV9 and HPyV12 in Cutaneous T-cell Lymphomas. Anticancer Res. 2018, 38, 4111–4114.
- Daprà, V.; Galliano, I.; Rassu, M.; Calvi, C.; Montanari, P.; Merlino, C.; Bergallo, M. Lack of detection of HPyV12 DNA using real-time PCR in Italian infants with diarrhea. Minerva Pediatr. 2020, doi:10.23736/s0026-4946.20.05738-2.
- Kamminga, S.; van der Meijden, E.; Wunderink, H.F.; Touzé, A.; Zaaijer, H.L.; Feltkamp, M.C.W. Development and Evaluation of a Broad Bead-Based Multiplex Immunoassay To Measure IgG Seroreactivity against Human Polyomaviruses. J. Clin. Microbiol. 2018, 56, e01566-17.
- Gedvilaite, A.; Tryland, M.; Ulrich, R.G.; Schneider, J.; Kurmauskaite, V.; Moens, U.; Preugschas, H.; Calvignac-Spencer, S.; Ehlers, B. Novel polyomaviruses in shrews (Soricidae) with close similarity to human polyomavirus 12. J. Gen. Virol. 2017, 98, 3060–3067.
- Schowalter, R.M.; Buck, C.B. The Merkel cell polyomavirus minor capsid protein. PLoS Pathog. 2013, 9, e1003558.
- Madeira, F.; Park, Y.M.; Lee, J.; Buso, N.; Gur, T.; Madhusoodanan, N.; Basutkar, P.; Tivey, A.R.N.; Potter, S.C.; Finn, R.D.; Lopez, R. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res 2019, 47, W636-W641.
- Fanning, E.; Zhao, K. SV40 DNA replication: From the A gene to a nanomachine. Virology 2009, 384, 352–359.
- Kelly, T.J. SV40 DNA replication. J. Biol. Chem. 1988, 263, 17889–17892.
- Farmerie, W.G.; Folk, W.R. Regulation of polyomavirus transcription by large tumor antigen. Proc. Natl. Acad. Sci. USA 1984, 81, 6919–6923.
- Tjian, R. Regulation of viral transcription and DNA replication by the SV40 large T antigen. Curr. Top Microbiol. Immunol. 1981, 93, 5–24.
- Zenke, M.; Grundström, T.; Matthes, H.; Wintzerith, M.; Schatz, C.; Wildeman, A.; Chambon, P. Multiple sequence motifs are involved in SV40 enhancer function. EMBO J. 1986, 5, 387–397.
- Rio, D.C.; Tjian, R. Multiple control elements involved in the initiation of SV40 late transcription. J. Mol. Appl. Gen. 1984, 2, 423–435.
- Cowie, A.; Kamen, R. Multiple binding sites for polyomavirus large T antigen within regulatory sequences of polyomavirus DNA. J. Virol. 1984, 52, 750–760.
- Jones, K.A.; Tjian, R. Essential contact residues within SV40 large T antigen binding sites I and II identified by alkylation-interference. Cell 1984, 36, 155–162.
- Lednicky, J.A.; Butel, J.S. Simian virus 40 regulatory region structural diversity and the association of viral archetypal regulatory regions with human brain tumors. Semin. Cancer Biol. 2001, 11, 39–47.
- O’Neill, F.J.; Greenlee, J.E.; Carney, H. The archetype enhancer of simian virus 40 DNA is duplicated during virus growth in human cells and rhesus monkey kidney cells but not in green monkey kidney cells. Virology 2003, 310, 173–182.
- Lednicky, J.A.; Wong, C.; Butel, J.S. Artificial modification of the viral regulatory region improves tissue culture growth of SV40 strain 776. Virus Res. 1995, 35, 143–153.
- Sroller, V.; Vilchez, R.A.; Stewart, A.R.; Wong, C.; Butel, J.S. Influence of the viral regulatory region on tumor induction by simian virus 40 in hamsters. J. Virol. 2008, 82, 871–879.
- Muller, W.J.; Mueller, C.R.; Mes, A.M.; Hassell, J.A. Polyomavirus origin for DNA replication comprises multiple genetic elements. J. Virol. 1983, 47, 586–599.
- Pomerantz, B.J.; Hassell, J.A. Polyomavirus and simian virus 40 large T antigens bind to common DNA sequences. J. Virol. 1984, 49, 925–937.
- Herbomel, P.; Bourachot, B.; Yaniv, M. Two distinct enhancers with different cell specificities coexist in the regulatory region of polyoma. Cell 1984, 39, 653–662.
- Iacoangeli, A.; Melucci-Vigo, G.; Risuleo, G.; Santi, E. Role of mouse polyomavirus late region in the control of viral DNA replication: A review. Biochimie 1995, 77, 780–786.
- Jat, P.; Novak, U.; Cowie, A.; Tyndall, C.; Kamen, R. DNA sequences required for specific and efficient initiation of transcription at the polyoma virus early promoter. Mol. Cell Biol. 1982, 2, 737–751.
- Mueller, C.R.; Mes-Masson, A.M.; Bouvier, M.; Hassell, J.A. Location of sequences in polyomavirus DNA that are required for early gene expression in vivo and in vitro. Mol. Cell Biol. 1984, 4, 2594–2609.
- Rochford, R.; Campbell, B.A.; Villarreal, L.P. Genetic analysis of the enhancer requirements for polyomavirus DNA replication in mice. J. Virol. 1990, 64, 476–485.
- Sekikawa, K.; Levine, A.J. Isolation and characterization of polyoma host range mutants that replicate in nullipotential embryonal carcinoma cells. Proc. Natl. Acad. Sci. USA 1981, 78, 1100–1104.
- Veldman, G.M.; Lupton, S.; Kamen, R. Polyomavirus enhancer contains multiple redundant sequence elements that activate both DNA replication and gene expression. Mol. Cell Biol. 1985, 5, 649–658.
- White, M.K.; Khalili, K. Pathogenesis of progressive multifocal leukoencephalopathy—revisited. J. Infect. Dis. 2011, 203, 578–586.
- Assetta, B.; Atwood, W.J. The biology of JC polyomavirus. Biol. Chem. 2017, 398, 839–855.
- Chen, N.N.; Khalili, K. Transcriptional regulation of human JC polyomavirus promoters by cellular proteins YB-1 and Pur alpha in glial cells. J. Virol. 1995, 69, 5843–5848.
- Sadowska, B.; Barrucco, R.; Khalili, K.; Safak, M. Regulation of human polyomavirus JC virus gene transcription by AP-1 in glial cells. J. Virol. 2003, 77, 665–672.
- Romagnoli, L.; Sariyer ,I.K.; Tung, J.; Feliciano, M.; Sawaya, B.E.; Del Valle, L.; Ferrante, P.; Khalili, K.; Safak, M.; White, M.K. Early growth response-1 protein is induced by JC virus infection and binds and regulates the JC virus promoter. Virology 2008, 375, 331–341.
- Pietropaolo, V.; Prezioso, C.; Bagnato, F.; Antonelli, G. John Cunningham virus: An overview on biology and disease of the etiological agent of the progressive multifocal leukoencephalopathy. New Microbiol. 2018, 41, 179–186.
- Van Loy, T.; Thys, K.; Tritsmans, L.; Stuyver, L.J. Quasispecies analysis of JC virus DNA present in urine of healthy subjects. PLoS ONE 2013, 8, e70950.
- Kenney, S.; Natarajan, V.; Strike, D.; Khoury, G.; Salzman, N.P. JC virus enhancer-promoter active in human brain cells. Science 1984, 226, 1337–1339.
- Yogo, Y.; Kitamura, T.; Sugimoto, C.; Ueki, T.; Aso, Y.; Hara, K.; Taguchi, F. Isolation of a possible archetypal JC virus DNA sequence from nonimmunocompromised individuals. J. Virol. 1990, 64, 3139–3143.
- L’Honneur, A.S.; Leh, H.; Laurent-Tchenio, F.; Hazan, U.; Rozenberg, F.; Bury-Moné, S. Exploring the role of NCCR variation on JC polyomavirus expression from dual reporter minicircles. PLoS ONE 2018, 13, e0199171.
- Agostini, H.T.; Ryschkewitsch, C.F.; Stoner, G.L. Rearrangements of archetypal regulatory regions in JC virus genomes from urine. Res. Virol. 1998, 149, 163–170.
- Bofill-Mas, S.; Clemente-Casares, P.; Major, E.O.; Curfman, B.; Girones, R. Analysis of the excreted JC virus strains and their potential oral transmission. J. Neurovirol. 2003, 9, 498–507.
- Frisque, R.J.; Bream, G.L.; Cannella, M.T. Human polyomavirus JC virus genome. J. Virol. 1984, 51, 458–469.
- Markowitz, R.B.; Dynan, W.S. Binding of cellular proteins to the regulatory region of BK virus DNA. J. Virol. 1988, 62, 3388–3398.
- Moens, U.; Van Ghelue, M. Polymorphism in the genome of non-passaged human polyomavirus BK: Implications for cell tropism and the pathological role of the virus. Virology 2005, 331, 209–231.
- Rubinstein, R.; Pare, N.; Harley, E.H. Structure and function of the transcriptional control region of nonpassaged BK virus. J. Virol. 1987, 61, 1747–1750.
- Bethge, T.; Ajuh, E.; Hirsch, H.H. Imperfect Symmetry of Sp1 and Core Promoter Sequences Regulates Early and Late Virus Gene Expression of the Bidirectional BK Polyomavirus Noncoding Control Region. J. Virol. 2016, 90, 10083–10101.
- Bethge, T.; Hachemi, H.A.; Manzetti, J.; Gosert, R.; Schaffner, W.; Hirsch, H.H. Sp1 sites in the noncoding control region of BK polyomavirus are key regulators of bidirectional viral early and late gene expression. J. Virol. 2015, 89, 3396–3411.
- Helle, F.; Brochot, E.; Handala, L.; Martin, E.; Castelain, S.; Francois, C.; Duverlie, G. Biology of the BKPyV: An Update. Viruses 2017, 9, 327.
- Moens, U.; Sundsfjord, A.; Flaegstad, T.; Traavik, T. BK virus early RNA transcripts in stably transformed cells: Enhanced levels induced by dibutyryl cyclic AMP, forskolin and 12-O-tetradecanoylphorbol-13-acetate treatment. J. Gen. Virol. 1990, 71, 1461–1471.
- Moens, U.; Subramaniam, N.; Johansen, B.; Johansen, T.; Traavik, T. A steroid hormone response unit in the late leader of the noncoding control region of the human polyomavirus BK confers enhanced host cell permissivity. J. Virol. 1994, 68, 2398–2408.
- Gorrill, T.S.; Khalili, K. Cooperative interaction of p65 and C/EBPbeta modulates transcription of BKV early promoter. Virology 2005, 335, 1–9.
- Anselmo, A.; Prezioso, C.; Saccà, F.A.; Di Lella, F.M.; Palmieri, G.; Tisone, G.; Pietropaolo, V.; Ciotti, M. Kidney graft failure induced by BKPyV replication despite a strong reduction of the immunosuppressive therapy. J. Med. Virol. 2019, 91, 1698–1701.
- Arthur, R.R.; Shah, K.V.; Baust, S.J.; Santos, G.W.; Saral, R. Association of BK viruria with hemorrhagic cystitis in recipients of bone marrow transplants. N. Engl. J. Med. 1986, 315, 230–234.
- Rosen, S.; Harmon, W.; Krensky, A.M.; Edelson, P.J.; Padgett, B.L.; Grinnell, B.W.; Rubino, M.J.; Walker, D.L. Tubulo-interstitial nephritis associated with polyomavirus (BK type) infection. N. Engl. J. Med. 1983, 308, 1192–1196.
- McIlroy, D.; Halary, F.; Bressollette-Bodin, C. Intra-patient viral evolution in polyomavirus-related diseases. Philos. Trans. R. Soc. B Biol. Sci. 2019, 374, 20180301.
- Seif, I.; Khoury, G.; Dhar, R. The genome of human papovavirus BKV. Cell 1979, 18, 963–977.
- Yang, J.F.; You, J. Regulation of Polyomavirus Transcription by Viral and Cellular Factors. Viruses 2020, 12, 1072.
- Olsen, G.H.; Hirsch, H.H.; Rinaldo, C.H. Functional analysis of polyomavirus BK non-coding control region quasispecies from kidney transplant recipients. J. Med. Virol. 2009, 81, 1959–1967.
- Degener, A.M.; Pietropaolo, V.; Di Taranto, C.; Jin, L.; Ameglio, F.; Cordiali-Fei, P.; Trento, E.; Sinibaldi, L.; Orsi, N. Identification of a new control region in the genome of the DDP strain of BK virus isolated from PBMC. J. Med. Virol. 1999, 58, 413–419.
- Pietropaolo, V.; Videtta, M.; Fioriti, D.; Mischitelli, M.; Arancio, A.; Orsi, N.; Degener, A.M. Rearrangement patterns of JC virus noncoding control region from different biological samples. J. Neurovirol. 2003, 9, 603–611.
- Mischitelli, M.; Fioriti, D.; Videtta, M.; Degener, A.M.; Antinori, A.; Cinque, P.; Giordano, A.; Pietropaolo, V. Investigation on the role of cell transcriptional factor Sp1 and HIV-1 TAT protein in PML onset or development. J. Cell Physiol. 2005, 204, 913–918.
- Ciardi, M.R.; Zingaropoli, M.A.; Iannetta, M.; Prezioso, C.; Perri, V.; Pasculli, P.; Lichtner, M.; D’Ettorre, G.; Altieri, M.; Conte, A.; et al. JCPyV NCCR analysis in PML patients with different risk factors: Exploring common rearrangements as essential changes for neuropathogenesis. Virol. J. 2020, 17, 23.
- Prezioso, C.; Zingaropoli, M.A.; Iannetta, M.; Rodio, D.M.; Altieri, M.; Conte, A.; Vullo, V.; Ciardi, M.R.; Palamara, A.T.; Pietropaolo, V. Which is the best PML risk stratification strategy in natalizumab-treated patients affected by multiple sclerosis? Mult. Scler. Relat. Disord. 2020, 41, 102008.
- Prezioso, C.; Scribano, D.; Bellizzi, A.; Anzivino, E.; Rodio, D.M.; Trancassini, M.; Palamara, A.T.; Pietropaolo, V. Efficient propagation of archetype JC polyomavirus in COS-7 cells: Evaluation of rearrangements within the NCCR structural organization after transfection. Arch. Virol. 2017, 162, 3745–3752.
- Prezioso, C.; Scribano, D.; Rodio, D.M.; Ambrosi, C.; Trancassini, M.; Palamara, A.T.; Pietropaolo, V. COS-7-based model: Methodological approach to study John Cunningham virus replication cycle. Virol. J. 2018, 15, 29.
- Jiang, M.; Abend, J.R.; Johnson, S.F.; Imperiale, M.J. The role of polyomaviruses in human disease. Virology 2009, 384, 266–273.