
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Sudeepa Rajan | -- | 2025 | 2023-03-12 20:35:56 | | | |
| 2 | Camila Xu | -1 word(s) | 2024 | 2023-03-13 02:05:09 | | |
Video Upload Options
Actin is one of the key and highly conserved elements of the cytoskeleton. It is indispensable for driving many cellular processes, including cell migration, cytokinesis, vesicle transport, and contractile force generation. To perform diverse functions, actin filaments assemble into higher-order structures such as branched actin networks and actin bundles. This entry describes different types of actin bundles present in cells, their locations, and the bundling proteins involved in their formation.
1. Introduction
Actin, one of the key and highly conserved elements of the cytoskeleton, amounts to approximately 5–15% of total cell proteins [1][2]. It is indispensable for driving many cellular processes, including cell migration, cytokinesis, vesicle transport, and contractile force generation [3]. The globular actin monomers (G-actin) polymerize to form semi-flexible double-stranded helical filaments (F-actin), also known as microfilaments. These filaments assemble into higher-order structures, such as branched networks or bundles [5]. It is now recognized that complex actin networks and bundles are essential for many cellular functions. Lately, actin-bundling proteins have been attracting a lot of attention as their malfunction is linked to malignant cancers, muscular dystrophy, bone disease, and immunological disorders [4][5][6][7][8].
More than 150 actin-binding proteins (ABPs) are known to associate with the actin cytoskeleton, and many of them regulate actin functions [9]. These proteins are involved in: (1) regulation of actin assembly and disassembly, (2) actin-driven movements in cells, (3) connecting actin structures to plasma membrane/cell organelles or other cytoskeleton proteins, and (4) organizing actin filaments (by their crosslinking) into higher-order structures, such as branched actin networks or actin bundles [1][10][11]. The proteins forming higher-order structures are among the most represented and diverse functional families of actin-binding proteins.
2. Actin Bundles in the Cell
In the cells, actin is present either in monomeric, filamentous, or higher-order three-dimensional structures, such as bundles and branched networks [10]. The length of filaments varies from dozens of nanometers (e.g., in branched networks) to several dozen micrometers (e.g., in stress fibers, filopodia, and stereocilia) [12][13][14][15][16]. Together, they form a continuum of systems enabling the reception and transduction of mechanical stimuli across the cell and providing mechanical support for the shape and polarity of cells. The necessity of forming higher-order actin structures is dictated by the immense variety of cellular functions supported by the actin cytoskeleton and a broad range of mechanical forces required to carry them out. Forces generated by these higher-order actin networks, ranging from piconewtons to nanonewtons, aid in cell migration and invasion, internal vesicle movements, endocytosis, exocytosis, phagocytosis, and cell division [17][18][19].
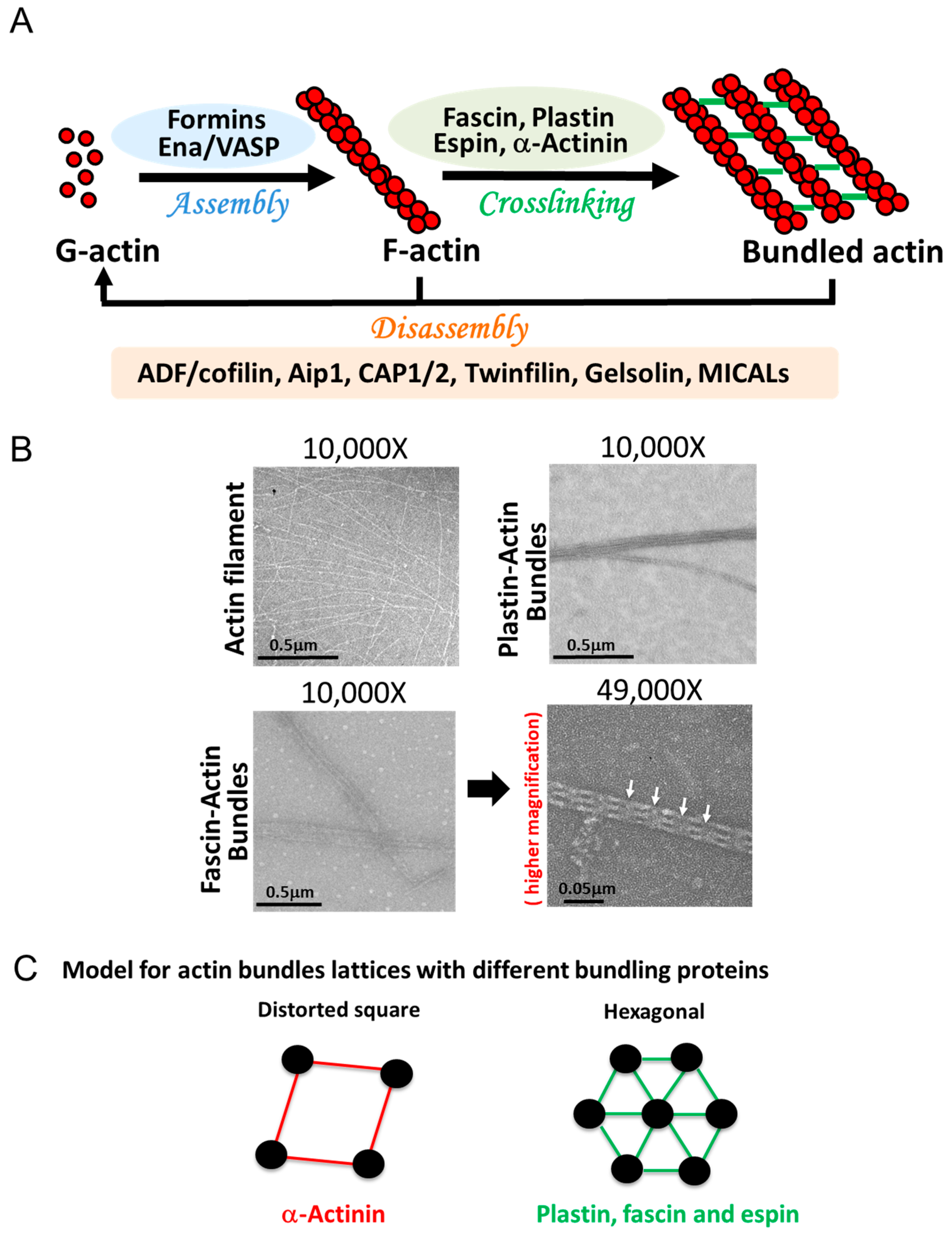
Actin bundles are linear arrays of actin filaments crosslinked by one or, more often, several different actin-bundling proteins (Figure 1 A,B). The length and width of such bundles, and the number of filaments present in them are dictated by their unique set of proteins and by kinetic conditions under which these bundles are formed [20][21], giving each bundled complex a specific structure with different mechanical properties [19][22], Figure 2, Table 1. The size of the bundling proteins plays an important role in the topology and the compactness of the bundles (Figure 1 B,C). For example, fascin, plastins/fimbrins, and small espins are small sized actin-bundling proteins that crosslink actin filaments to form compact and tighly packed bundles, whereas α-actinin is a larger-sized crosslinker inducing widely spaced bundles with distorted square lattice (Figure 1 B,C).
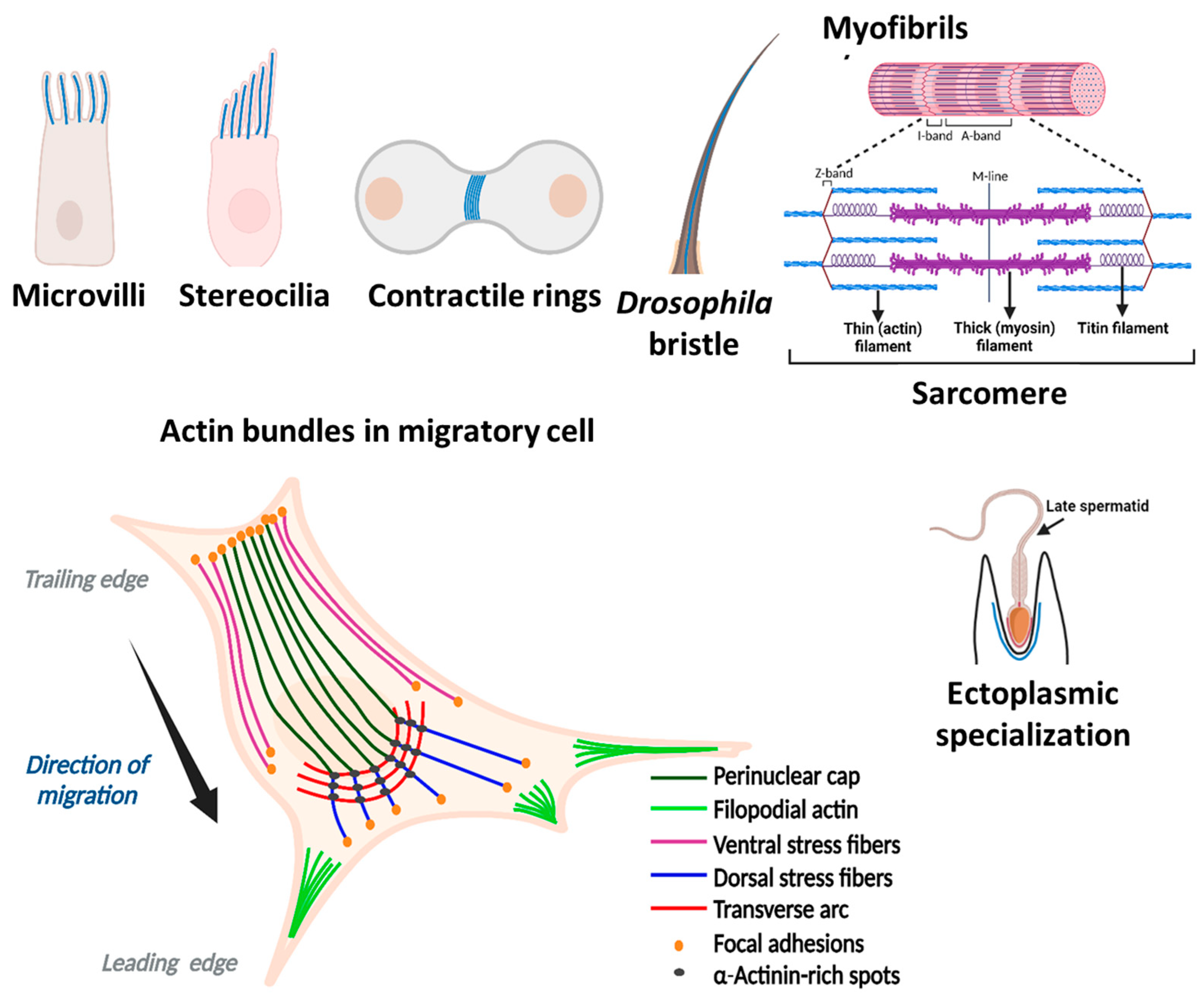
There are two main types of actin bundles, with either parallel or mixed polarity filament orientations. In parallel (or uniform polarity) actin bundles, the filaments are ordered with consistent polarity, allowing them to conduct work (e.g., membrane deformation) due to the directional F-actin elongation. Parallel actin bundles are present in chemosensory and mechanosensory cell protrusions (microvilli) of most cell types [22][23], stereocilia of inner ear hair cells [24], bristles in the thorax of Drosophila melanogaster [25], and in ectoplasmic specializations of Sertoli cells (Figure 2, [26]). Filopodia, microspikes, focal adhesions, and distal ends of dorsal stress fibers, found in most cell types, consist also of parallel actin bundles (Figure 2, [7][27]). These bundles create the force for localized membrane protrusions, while helping cells to resist compressive forces from the membrane [28]. They facilitate cell movement in response to extracellular stimuli or intracellular signaling [29]. The length (1 to 100 μm) and number (one to hundreds) of these bundles per cell, their diameter and the number of actin filaments in them (a few to ~1000) vary depending on the cell type and the structures they support (microvilli, stereocilia, bristles, filopodia, (Figure 2, Table 1), and invadosomes (podosomes/invadopodia) [19][25][27][30][31][32][33][34].
More than two actin-bundling proteins are often associated with these actin structures. Filopodia, a thin actin-rich plasma-membrane protrusions, help in chemosensing during cell migration, wound healing, and cell adhesion to the extracellular matrix [33]. They are enriched in fascin, but additional bundling proteins, such as α-actinin, fimbrin/plastin, filamin, and espin are also present in them under certain conditions [33][35]. Podosomes are actin-based dynamic structures near the plasma membrane of various cells (such as monocytic, endothelial, and smooth muscle cells). They contribute to cell migration, matrix invasiveness, bone remodeling, and mechanosensing [34][36][37] and contain fascin, L-plastin (a hematopoietic cell-specific plastin isoform), and α-actinin [34][38][39][40]. Invadopodia are functionally and structurally similar to podosomes of normal cells, but they are present in tumor cells [34]. Additionally, podosomes and invadopodia are unique in their ability to degrade ECM material by locally releasing proteolytic enzymes [34]. Ectoplasmic specializations of Sertoli cells, which are hybrid testis-specific cell–cell contacts (contributing to the blood–testis barrier), contain espin and T-plastin (a most abundant plastin isoform found in cells of most solid tissues) [19][41][42][43]. Microvilli—the finger-like projections on the surface of several types of cells - increase the total cell surface without substantially increasing its volume [19][23]. Most microvilli contain I-plastin (a plastin isoform found in the intestinal and kidney microvilli and stereocilia of the inner ear), small isoforms of espin, and villin. Similarly, stereocilia are the modified microvilli that transduce mechanical signals into stimulus-dependent electrical signals. They predominantly contain I-plastin and fascin, but also have isoforms of espin and other proteins expressed in smaller quantities [44][45].
| Location | Microvilli | Stereocilia | Filopodia | Stress fibers | Bristles (drosophila) |
|---|---|---|---|---|---|
| Cell type | Most cells (frequent in epithelial cells) | Auditory and vestibular sensory cells | Motile cells | Most cells (prominent in fibroblasts, smooth muscle, and endothelial cells) | Sensory organ precursor cells |
| Function | Increase apical surface area for absorption | Mechano-electrical signaling | Sensory and guiding | Contraction and adhesions | Mechanosensing |
| Length | 100 nm to 2 µm | 1.5–15 µm | ≤10 µm | ≥2 µm | Macrochaetes: 250–300 μm, Microchaetes: 70 μm (non-continuous 1–5 µm units) |
| Diameter | 50–100 nm | ~200 nm | 20–200 nm | Varies from cell to cell | Varies |
| Number of actin filaments | 30–40 | ~400–3000 | 10–30 | 10–30 | 7–18 bundles with hundreds of filaments |
| Actin filament organization | Parallel (unipolar) | Parallel (unipolar) | Parallel (unipolar) | Mixed (bipolar) | Parallel |
| Bundling proteins | Espin, plastin, villin | Fascin, espin, plastin | Fascin, α-actinin, plastin, espin | α-Actinin, fascin filamin |
singed (fascin), forked (espin) |
References
- Pollard, T.D. Actin and Actin-Binding Proteins. Cold Spring Harb. Perspect. Biol. 2016, 8, a018226.
- Lodish, H.; Berk, A.; Kaiser, C.A.; Kaiser, C.; Krieger, M.; Scott, M.P.; Bretscher, A.; Ploegh, H.; Matsudaira, P. Molecular Cell Biology; Macmillan: New York, NY, USA, 2008.
- Pollard, T.D.; Cooper, J.A. Actin, a central player in cell shape and movement. Science 2009, 326, 1208–1212.
- Liu, H.; Zhang, Y.; Li, L.; Cao, J.; Guo, Y.; Wu, Y.; Gao, W. Fascin actin-bundling protein 1 in human cancer: Promising biomarker or therapeutic target? Mol. Ther. Oncolytics 2021, 20, 240–264.
- Cheng, Y.W.; Zeng, F.M.; Li, D.J.; Wang, S.H.; He, J.Z.; Guo, Z.C.; Nie, P.J.; Wu, Z.Y.; Shi, W.Q.; Wen, B.; et al. P300/CBP-associated factor (PCAF)-mediated acetylation of Fascin at lysine 471 inhibits its actin-bundling activity and tumor metastasis in esophageal cancer. Cancer Commun. 2021, 41, 1398–1416.
- Morley, S.C. The actin-bundling protein L-plastin supports T-cell motility and activation. Immunol. Rev. 2013, 256, 48–62.
- Stevenson, R.P.; Veltman, D.; Machesky, L.M. Actin-bundling proteins in cancer progression at a glance. J. Cell Sci. 2012, 125, 1073–1079.
- Wolff, L.; Strathmann, E.A.; Müller, I.; Mählich, D.; Veltman, C.; Niehoff, A.; Wirth, B. Plastin 3 in health and disease: A matter of balance. Cell. Mol. Life Sci. 2021, 78, 5275–5301.
- Gupta, C.M.; Ambaru, B.; Bajaj, R. Emerging Functions of Actins and Actin Binding Proteins in Trypanosomatids. Front. Cell Dev. Biol. 2020, 8, 587685.
- Lappalainen, P. Actin-binding proteins: The long road to understanding the dynamic landscape of cellular actin networks. Mol. Biol. Cell 2016, 27, 2519–2522.
- Pollard, T.D.; Blanchoin, L.; Mullins, R.D. Molecular mechanisms controlling actin filament dynamics in nonmuscle cells. Annu. Rev. Biophys. Biomol. Struct. 2000, 29, 545–576.
- Livne, A.; Geiger, B. The inner workings of stress fibers—From contractile machinery to focal adhesions and back. J. Cell Sci. 2016, 129, 1293–1304.
- Svitkina, T.M.; Bulanova, E.A.; Chaga, O.Y.; Vignjevic, D.M.; Kojima, S.; Vasiliev, J.M.; Borisy, G.G. Mechanism of filopodia initiation by reorganization of a dendritic network. J. Cell Biol. 2003, 160, 409–421.
- Li, A.; Xue, J.; Peterson, E.H. Architecture of the mouse utricle: Macular organization and hair bundle heights. J. Neurophysiol. 2008, 99, 718–733.
- Tilney, L.G.; Saunders, J.C. Actin filaments, stereocilia, and hair cells of the bird cochlea. I. Length, number, width, and distribution of stereocilia of each hair cell are related to the position of the hair cell on the cochlea. J. Cell Biol. 1983, 96, 807–821.
- Svitkina, T.M.; Verkhovsky, A.B.; McQuade, K.M.; Borisy, G.G. Analysis of the actin-myosin II system in fish epidermal keratocytes: Mechanism of cell body translocation. J. Cell Biol. 1997, 139, 397–415.
- Thomas, C. Bundling actin filaments from membranes: Some novel players. Front. Plant Sci. 2012, 3, 188.
- Revenu, C.; Athman, R.; Robine, S.; Louvard, D. The co-workers of actin filaments: From cell structures to signals. Nat. Rev. Mol. Cell Biol. 2004, 5, 635–646.
- Bartles, J.R. Parallel actin bundles and their multiple actin-bundling proteins. Curr. Opin. Cell Biol. 2000, 12, 72–78.
- Falzone, T.T.; Lenz, M.; Kovar, D.R.; Gardel, M.L. Assembly kinetics determine the architecture of alpha-actinin crosslinked F-actin networks. Nat. Commun. 2012, 3, 861.
- Sherer, L.A.; Courtemanche, N. Cooperative bundling by fascin generates actin structures with architectures that depend on filament length. Front. Cell Dev. Biol. 2022, 10, 974047.
- DeRosier, D.J.; Tilney, L.G. F-actin bundles are derivatives of microvilli: What does this tell us about how bundles might form? J. Cell Biol. 2000, 148, 1–6.
- Orbach, R.; Su, X. Surfing on Membrane Waves: Microvilli, Curved Membranes, and Immune Signaling. Front. Immunol. 2020, 11, 2187.
- Tilney, L.G.; Derosier, D.J.; Mulroy, M.J. The organization of actin filaments in the stereocilia of cochlear hair cells. J. Cell Biol. 1980, 86, 244–259.
- Tilney, L.G.; Tilney, M.S.; Guild, G.M. F actin bundles in Drosophila bristles. I. Two filament cross-links are involved in bundling. J. Cell Biol. 1995, 130, 629–638.
- Bartles, J.R.; Wierda, A.; Zheng, L. Identification and characterization of espin, an actin-binding protein localized to the F-actin-rich junctional plaques of Sertoli cell ectoplasmic specializations. J. Cell Sci. 1996, 109, 1229–1239.
- Svitkina, T. The Actin Cytoskeleton and Actin-Based Motility. Cold Spring Harb. Perspect. Biol. 2018, 10, a018267.
- Gupta, M.; Doss, B.; Lim, C.T.; Voituriez, R.; Ladoux, B. Single cell rigidity sensing: A complex relationship between focal adhesion dynamics and large-scale actin cytoskeleton remodeling. Cell Adhes. Migr. 2016, 10, 554–567.
- Mogilner, A.; Rubinstein, B. The physics of filopodial protrusion. Biophys. J. 2005, 89, 782–795.
- Shin, J.B.; Krey, J.F.; Hassan, A.; Metlagel, Z.; Tauscher, A.N.; Pagana, J.M.; Sherman, N.E.; Jeffery, E.D.; Spinelli, K.J.; Zhao, H.; et al. Molecular architecture of the chick vestibular hair bundle. Nat. Neurosci. 2013, 16, 365–374.
- Zheng, L.; Sekerkova, G.; Vranich, K.; Tilney, L.G.; Mugnaini, E.; Bartles, J.R. The deaf jerker mouse has a mutation in the gene encoding the espin actin-bundling proteins of hair cell stereocilia and lacks espins. Cell 2000, 102, 377–385.
- Lewis, A.K.; Bridgman, P.C. Nerve growth cone lamellipodia contain two populations of actin filaments that differ in organization and polarity. J. Cell Biol. 1992, 119, 1219–1243.
- Gallop, J.L. Filopodia and their links with membrane traffic and cell adhesion. Semin. Cell Dev. Biol. 2020, 102, 81–89.
- Linder, S.; Cervero, P.; Eddy, R.; Condeelis, J. Mechanisms and roles of podosomes and invadopodia. Nat. Rev. Mol. Cell Biol. 2023, 24, 86–106.
- Mellor, H. The role of formins in filopodia formation. Biochim. Biophys. Acta 2010, 1803, 191–200.
- Hu, F.; Zhu, D.; Dong, H.; Zhang, P.; Xing, F.; Li, W.; Yan, R.; Zhou, J.; Xu, K.; Pan, L.; et al. Super-resolution microscopy reveals nanoscale architecture and regulation of podosome clusters in primary macrophages. iScience 2022, 25, 105514.
- Alonso, F.; Spuul, P.; Daubon, T.; Kramer, I.; Genot, E. Variations on the theme of podosomes: A matter of context. Biochim. Biophys. Acta Mol. Cell Res. 2019, 1866, 545–553.
- Murphy, D.A.; Courtneidge, S.A. The ‘ins’ and ‘outs’ of podosomes and invadopodia: Characteristics, formation and function. Nat. Rev. Mol. Cell Biol. 2011, 12, 413–426.
- Van Audenhove, I.; Debeuf, N.; Boucherie, C.; Gettemans, J. Fascin actin bundling controls podosome turnover and disassembly while cortactin is involved in podosome assembly by its SH3 domain in THP-1 macrophages and dendritic cells. Biochim. Biophys. Acta 2015, 1853, 940–952.
- De Clercq, S.; Boucherie, C.; Vandekerckhove, J.; Gettemans, J.; Guillabert, A. L-plastin nanobodies perturb matrix degradation, podosome formation, stability and lifetime in THP-1 macrophages. PLoS ONE 2013, 8, e78108.
- Li, N.; Mruk, D.D.; Wong, C.K.; Lee, W.M.; Han, D.; Cheng, C.Y. Actin-bundling protein plastin 3 is a regulator of ectoplasmic specialization dynamics during spermatogenesis in the rat testis. FASEB J. 2015, 29, 3788–3805.
- Berruti, G.; Paiardi, C. The dynamic of the apical ectoplasmic specialization between spermatids and Sertoli cells: The case of the small GTPase Rap1. Biomed. Res. Int. 2014, 2014, 635979.
- Kopera, I.A.; Bilinska, B.; Cheng, C.Y.; Mruk, D.D. Sertoli-germ cell junctions in the testis: A review of recent data. Philos. Trans. R Soc. B Biol. Sci. 2010, 365, 1593–1605.
- Krey, J.F.; Krystofiak, E.S.; Dumont, R.A.; Vijayakumar, S.; Choi, D.; Rivero, F.; Kachar, B.; Jones, S.M.; Barr-Gillespie, P.G. Plastin 1 widens stereocilia by transforming actin filament packing from hexagonal to liquid. J. Cell Biol. 2016, 215, 467–482.
- Krey, J.F.; Barr-Gillespie, P.G. Molecular Composition of Vestibular Hair Bundles. Cold Spring Harb. Perspect. Med. 2019, 9, a033209.
- Blanchoin, L.; Boujemaa-Paterski, R.; Sykes, C.; Plastino, J. Actin dynamics, architecture, and mechanics in cell motility. Physiol. Rev. 2014, 94, 235–263.
- Tojkander, S.; Gateva, G.; Lappalainen, P. Actin stress fibers--assembly, dynamics and biological roles. J. Cell Sci. 2012, 125, 1855–1864.
- Laporte, D.; Ojkic, N.; Vavylonis, D.; Wu, J.Q. Alpha-Actinin and fimbrin cooperate with myosin II to organize actomyosin bundles during contractile-ring assembly. Mol. Biol. Cell 2012, 23, 3094–3110.
- Skau, C.T.; Courson, D.S.; Bestul, A.J.; Winkelman, J.D.; Rock, R.S.; Sirotkin, V.; Kovar, D.R. Actin filament bundling by fimbrin is important for endocytosis, cytokinesis, and polarization in fission yeast. J. Biol. Chem. 2011, 286, 26964–26977.
- Mei, L.; Reynolds, M.J.; Garbett, D.; Gong, R.; Meyer, T.; Alushin, G.M. Structural mechanism for bidirectional actin cross-linking by T-plastin. Proc. Natl. Acad. Sci. USA 2022, 119, e2205370119.
- Hotulainen, P.; Lappalainen, P. Stress fibers are generated by two distinct actin assembly mechanisms in motile cells. J. Cell Biol. 2006, 173, 383–394.
- Letort, G.; Ennomani, H.; Gressin, L.; Théry, M.; Blanchoin, L. Dynamic reorganization of the actin cytoskeleton. F1000Res. 2015, 4, 940.
- Cramer, L.P.; Siebert, M.; Mitchison, T.J. Identification of novel graded polarity actin filament bundles in locomoting heart fibroblasts: Implications for the generation of motile force. J. Cell Biol. 1997, 136, 1287–1305.
- Burnette, D.T.; Manley, S.; Sengupta, P.; Sougrat, R.; Davidson, M.W.; Kachar, B.; Lippincott-Schwartz, J. A role for actin arcs in the leading-edge advance of migrating cells. Nat. Cell Biol. 2011, 13, 371–381.
- Khatau, S.B.; Hale, C.M.; Stewart-Hutchinson, P.J.; Patel, M.S.; Stewart, C.L.; Searson, P.C.; Hodzic, D.; Wirtz, D. A perinuclear actin cap regulates nuclear shape. Proc. Natl. Acad. Sci. USA 2009, 106, 19017–19022.
- Pfisterer, K.; Jayo, A.; Parsons, M. Control of nuclear organization by F-actin binding proteins. Nucleus 2017, 8, 126–133.
- Dasbiswas, K.; Hu, S.; Schnorrer, F.; Safran, S.A.; Bershadsky, A.D. Ordering of myosin II filaments driven by mechanical forces: Experiments and theory. Philos. Trans. R Soc. Lond. B Biol. Sci. 2018, 373, 20170114.
- Fenix, A.M.; Neininger, A.C.; Taneja, N.; Hyde, K.; Visetsouk, M.R.; Garde, R.J.; Liu, B.; Nixon, B.R.; Manalo, A.E.; Becker, J.R.; et al. Muscle-specific stress fibers give rise to sarcomeres in cardiomyocytes. Elife 2018, 7, e42144.
- Mangione, M.C.; Gould, K.L. Molecular form and function of the cytokinetic ring. J. Cell Sci. 2019, 132, jcs226928.
- Field, C.M.; Alberts, B.M. Anillin, a contractile ring protein that cycles from the nucleus to the cell cortex. J. Cell Biol. 1995, 131, 165–178.
- Matsuda, K.; Sugawa, M.; Yamagishi, M.; Kodera, N.; Yajima, J. Visualizing dynamic actin cross-linking processes driven by the actin-binding protein anillin. FEBS Lett. 2020, 594, 1237–1247.
- Chugh, P.; Paluch, E.K. The actin cortex at a glance. J. Cell Sci. 2018, 131, jcs186254.
- Svitkina, T.M. Ultrastructure of the actin cytoskeleton. Curr. Opin. Cell Biol. 2018, 54, 1–8.





