
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Lamiaa Ahmed Shaala | -- | 2137 | 2023-01-10 09:21:36 | | | |
| 2 | Dean Liu | Meta information modification | 2137 | 2023-01-11 02:12:41 | | | | |
| 3 | Dean Liu | -2 word(s) | 2135 | 2023-01-16 06:43:35 | | |
Video Upload Options
Cyanobacteria ascribed to the genus Lyngbya (Family Oscillatoriaceae) represent a potential therapeutic gold mine of chemically and biologically diverse natural products that exhibit a wide array of biological properties. Phylogenetic analyses have established the Lyngbya ‘morpho-type’ as a highly polyphyletic group and have resulted in taxonomic revision and description of an additional six new cyanobacterial genera in the same family to date. Among the most prolific marine cyanobacterial producers of biologically active compounds are the species Moorena producens (previously L. majuscula, then Moorea producens), M. bouillonii (previously L. bouillonii), and L. confervoides. Over the years, compounding evidence from in vitro and in vivo studies in support of the significant pharmaceutical potential of ‘Lyngbya’-derived natural products has made the Lyngbya morphotype a significant target for biomedical research and novel drug leads development. Researchers concluded compounds with reported anti-infective activities through 2022 from the Lyngbya morphotype, including new genera arising from recent phylogenetic re-classification. So far, 72 anti-infective secondary metabolites have been isolated from various Dapis, Lyngbya, Moorea, and Okeania species.
1. Introduction
2. Compounds with Antibacterial Activities
| Compound | Source Organism | Collection Site | Targeted Bacteria | MIC/Inhibition Zone/IC50 | Reference |
|---|---|---|---|---|---|
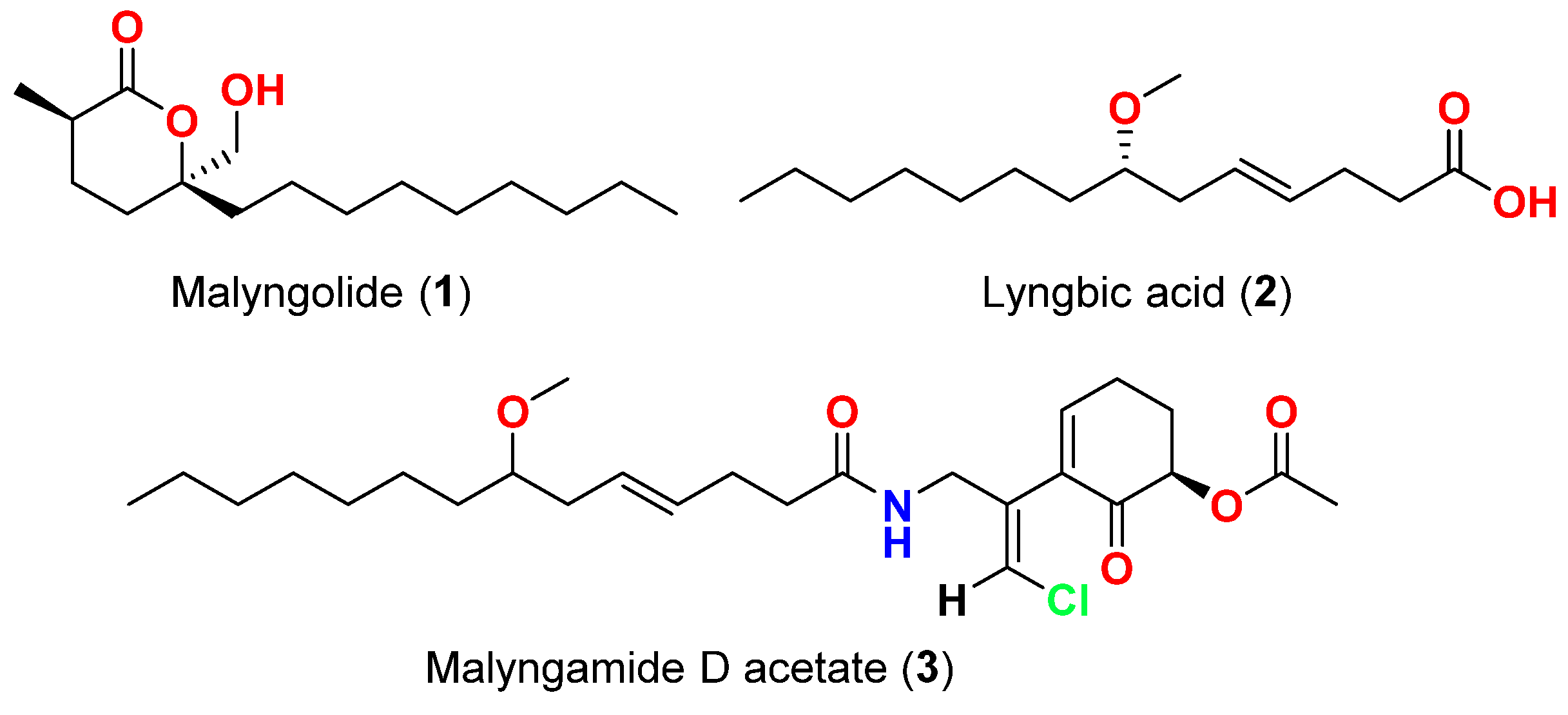
| Malyngolide (1) | L. majuscula | Hawaii, USA | M. smegmatis, S. pyogenes, S. aureus and B. subtilis | More active against M. smegmatis and S. pyogenes than S. aureus and B. subtilis | [12] |
| Lyngbic acid (2) | M. producens | Red Sea | M. tuberculosis H37Rv | 65% inhibition at 12.5 μg/mL | [13] |
| Lyngbic acid (2) | L. majuscula | Caribbean region | S. aureus and B. subtilis | Antibacterial activity | [14] |
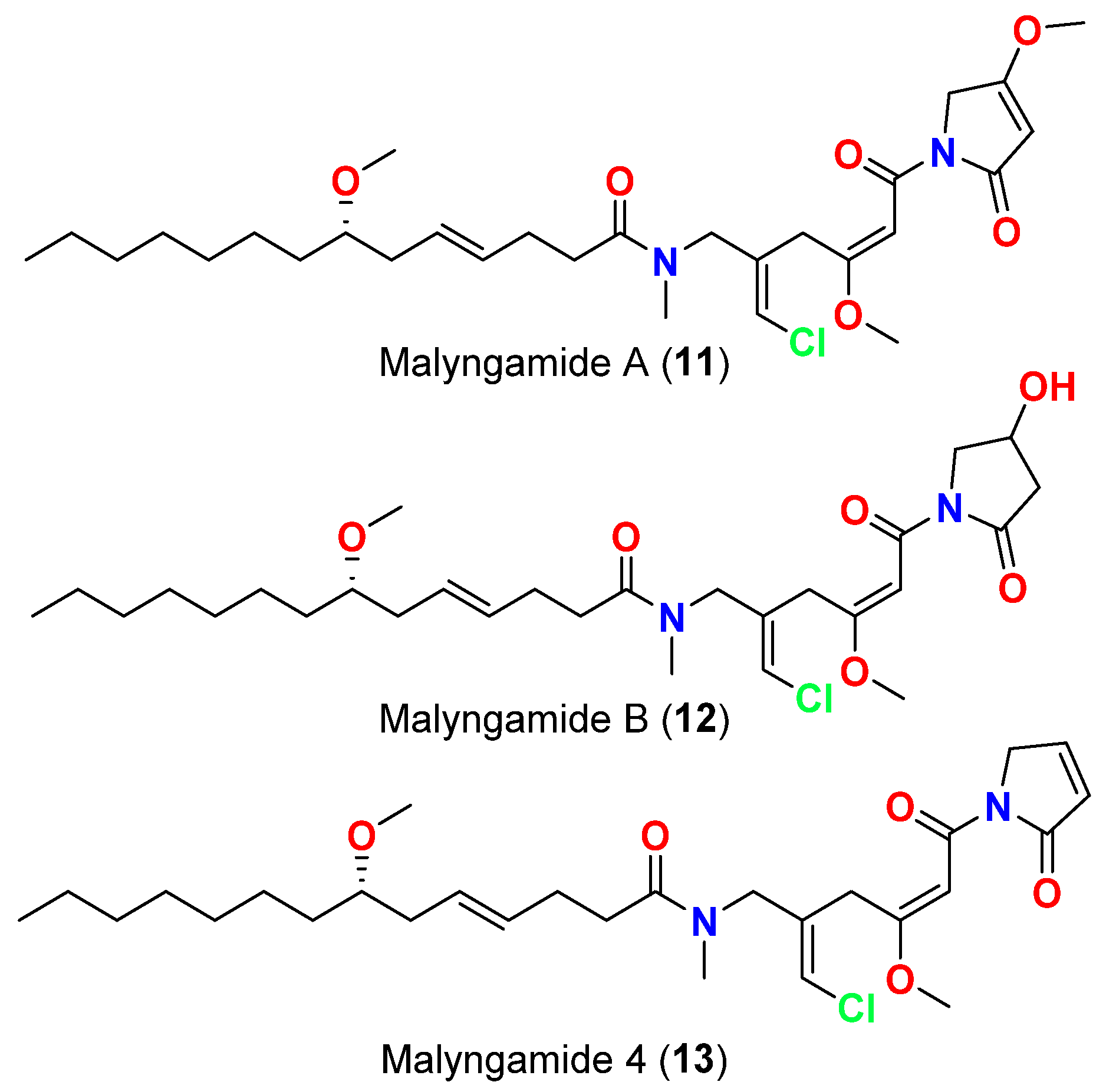
| Malyngamide D acetate (3) | L. majuscula | Caribbean region | S. aureus | Slight activity | [14] |
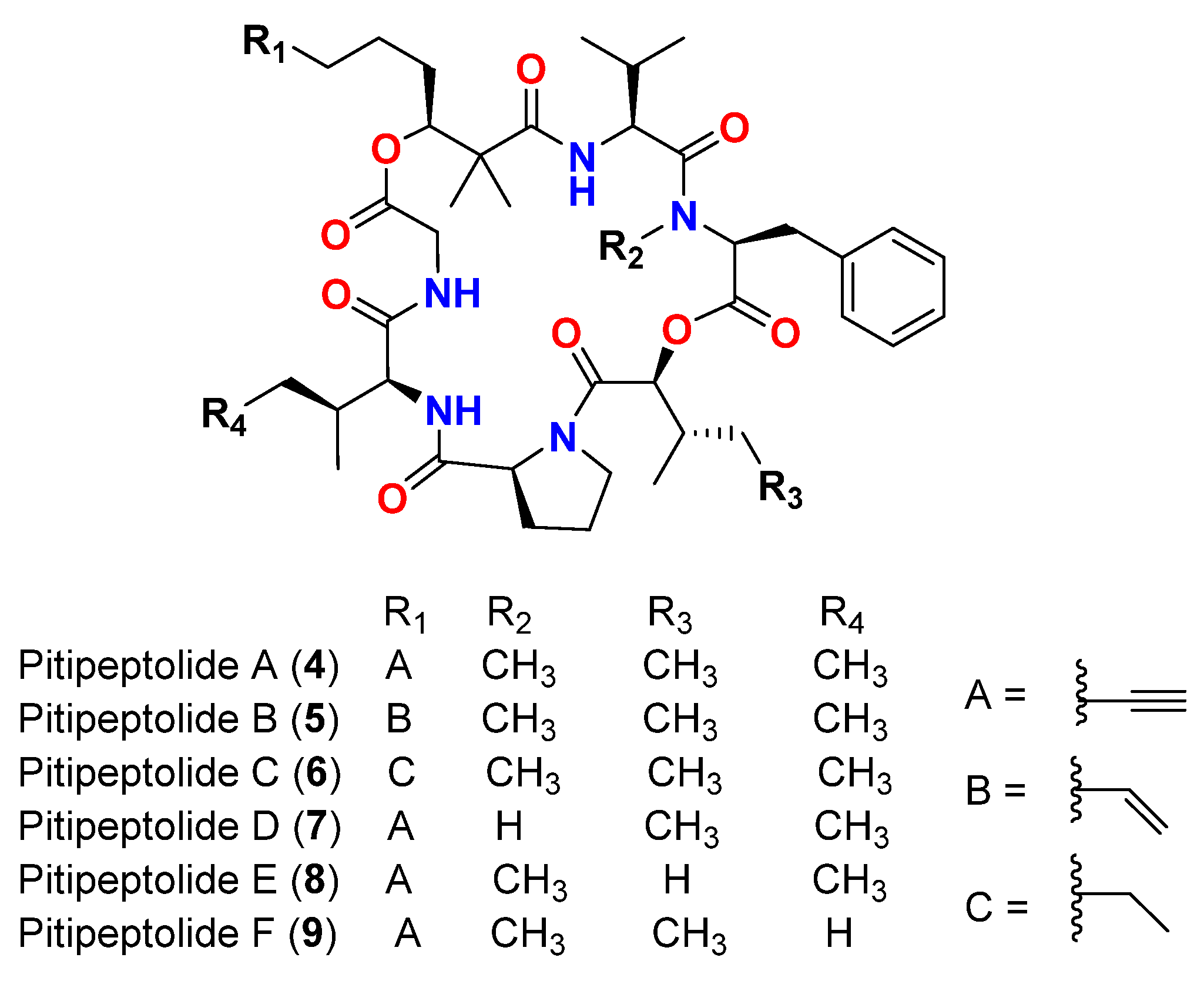
| Pitipeptolide A (4) | L. majuscula | Guam | M. tuberculosis ATCC 25177 | 25 mm at 100 µg 10 mm at 25 µg |
[15] |
| Pitipeptolide A (4) | L. majuscula | Guam | M. tuberculosis ATCC 35818 | 15 mm at 100 µg 9 mm at 25 µg |
[15] |
| Pitipeptolide B (5) | L. majuscula | Guam | M. tuberculosis ATCC 25177 | 30 mm at 100 µg 15 mm at 25 µg |
[15] |
| Pitipeptolide B (5) | L. majuscula | Guam | M. tuberculosis ATCC 35818 | 15 mm at 100 µg 10 mm at 25 µg |
[15] |
| Pitipeptolide A (4) | L. majuscula | Guam | M. tuberculosis ATCC 25177 | 28 mm at 100 µg 23 mm at 50 µg 9 mm at 10 µg |
[16] |
| Pitipeptolide B (5) | L. majuscula | Guam | M. tuberculosis ATCC 25177 | 30 mm at 100 µg 24 mm at 50 µg 14 mm at 10 µg |
[16] |
| Pitipeptolide C (6) | L. majuscula | Guam | M. tuberculosis ATCC 25177 | 26 mm at 100 µg 21 mm at 50 µg 18 mm at 10 µg |
[16] |
| Pitipeptolide D (7) | L. majuscula | Guam | M. tuberculosis ATCC 25177 | 10 mm at 100 µg 0 mm at 50 µg 0 mm at 10 µg |
[16] |
| Pitipeptolide E (8) | L. majuscula | Guam | M. tuberculosis ATCC 25177 | 21 mm at 100 µg 15 mm at 50 µg 0 mm at 10 µg |
[16] |
| Pitipeptolide F (9) | L. majuscula | Guam | M. tuberculosis ATCC 25177 | 40 mm at 100 µg 30 mm at 50 µg 10 mm at 10 µg |
[16] |
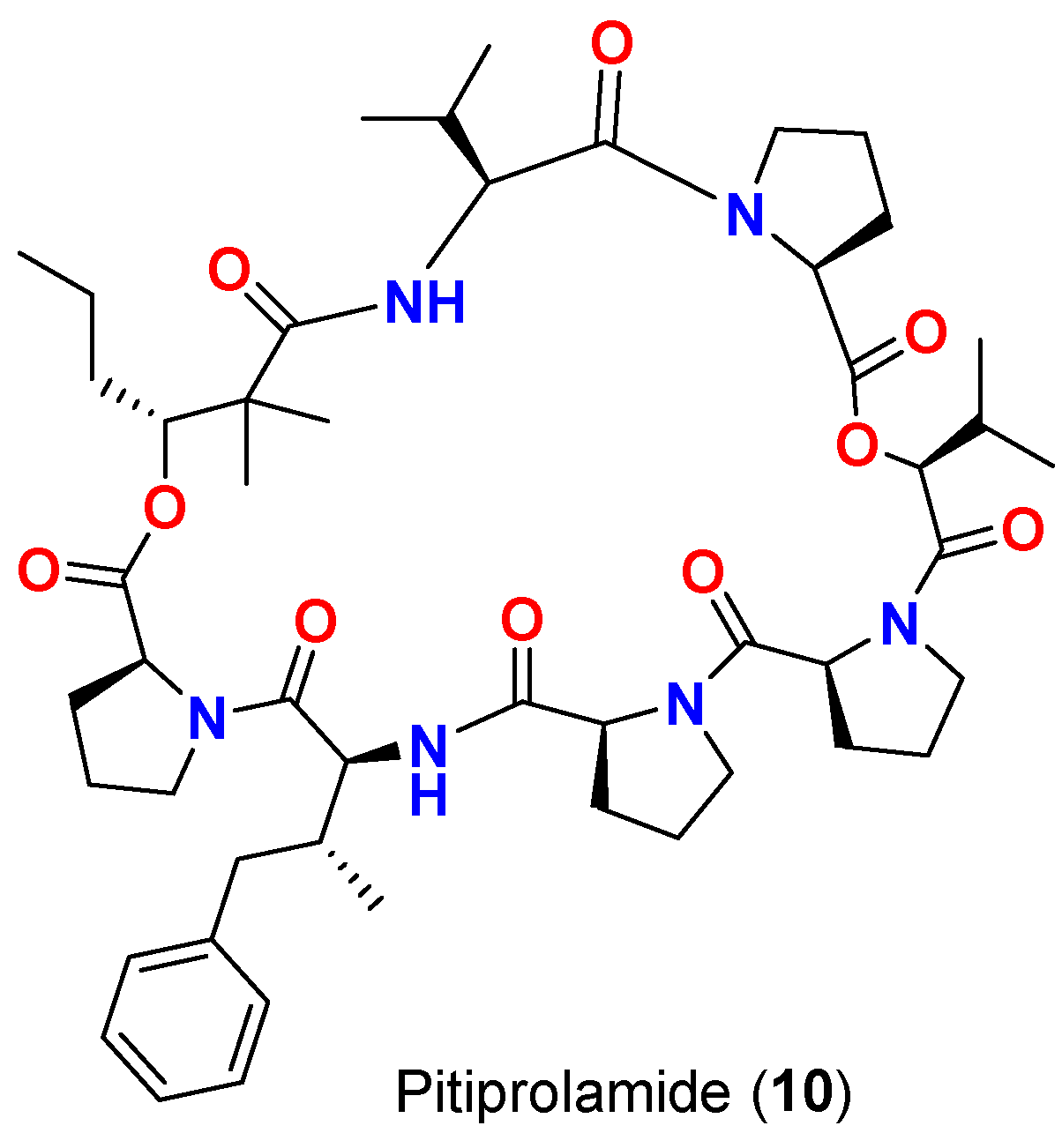
| Pitiprolamide (10) | L. majuscula | Guam | M. tuberculosis ATCC 25177 | 23 mm at 100 µg 13 mm at 50 µg 0 mm at 10 µg |
[17] |
| Pitiprolamide (10) | L. majuscula | Guam | B. cereus ATCC 10987 | IC50 = 70 μM at 1 μM | [17] |
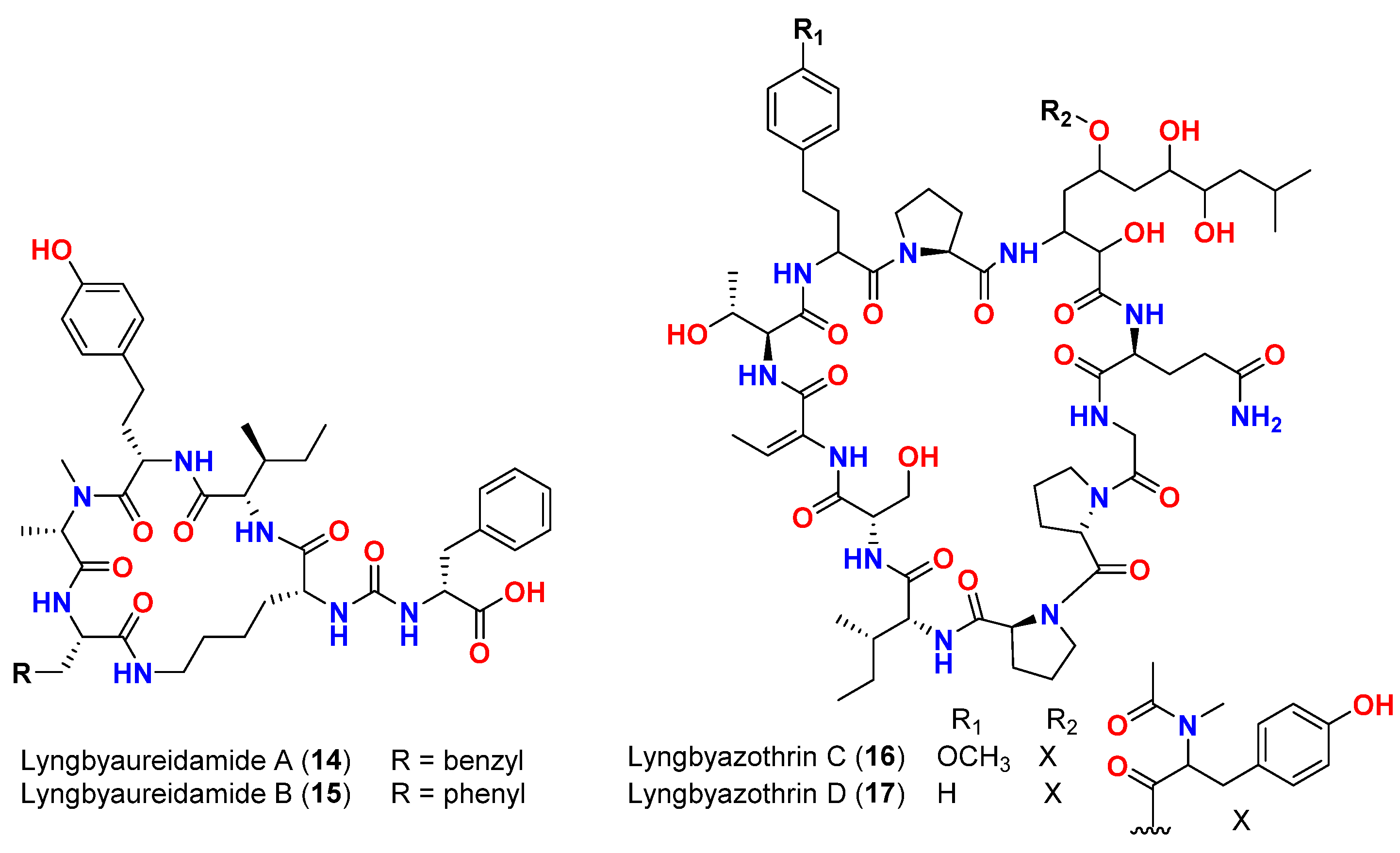
| Mixture of lyngbyazothrins A and B (14 and 15) | Lyngbya sp. | Germany (Culture) | M. flaVus SBUG 16 | 8 mm at 100 μg/disk | [18] |
| Mixture of lyngbyazothrins C (16) and D (17) | Lyngbya sp. | Germany (Culture) | B. subtilis SBUG 14 E. coli ATCC 11229 E. coli SBUG 13 P. aeruginosa ATCC 27853 S. marcescens SBUG 9 |
18 mm at 25 μg/disk 18 mm at 100 μg/disk 15 mm at 100 μg/disk 8 mm at 100 μg/disk 8 mm at 200 μg/disk |
[18] |
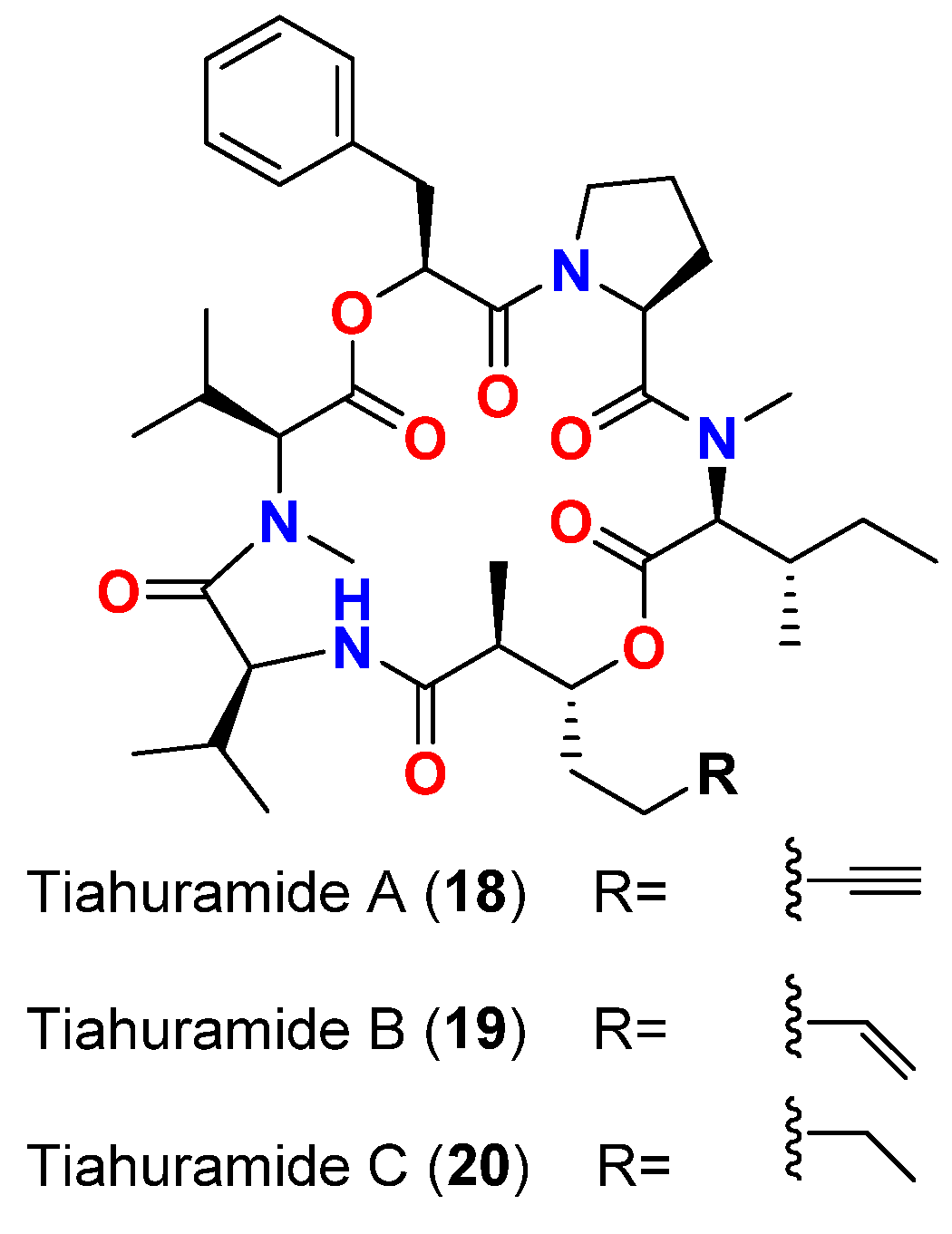
| Tiahuramide A (18) | L. majuscula | French Polynesia | A. salmonicida (CIP 103209T strain), V. anguillarum (CIP 63.36T), S. baltica (CIP 105850T), E. coli (CIP 54.8) and M. luteus (CIP A270) | MIC = 27, 33, >50, 35 and 47 μM, respectively | [19] |
| Tiahuramide B (19) | L. majuscula | French Polynesia | A. salmonicida (CIP 103209T strain), V. anguillarum (CIP 63.36T), S. baltica (CIP 105850T), E. coli (CIP 54.8) and M. luteus (CIP A270) | MIC = 9.4, 8.5, 22, 12 and 29 μM, respectively | [19] |
| Tiahuramide C (20) | L. majuscula | French Polynesia | A. salmonicida (CIP 103209T strain), V. anguillarum (CIP 63.36T), S. baltica (CIP 105850T), E. coli (CIP 54.8) and M. luteus (CIP A270) | MIC = 6.7, 7.4, 16, 14 and 17 μM, respectively | [19] |
| Compound | Source Organism | Collection Site | Targeted Bacteria | MIC/Inhibition Zone/IC50 | Reference |
|---|---|---|---|---|---|
| Malyngolide (1) | L. majuscula | Hawaii, USA | M. smegmatis, S. pyogenes, S. aureus and B. subtilis | More active against M. smegmatis and S. pyogenes than S. aureus and B. subtilis | [12] |
| Lyngbic acid (2) | M. producens | Red Sea | M. tuberculosis H37Rv | 65% inhibition at 12.5 μg/mL | [13] |
| Lyngbic acid (2) | L. majuscula | Caribbean region | S. aureus and B. subtilis | Antibacterial activity | [14] |
| Malyngamide D acetate (3) | L. majuscula | Caribbean region | S. aureus | Slight activity | [14] |
| Pitipeptolide A (4) | L. majuscula | Guam | M. tuberculosis ATCC 25177 | 25 mm at 100 µg 10 mm at 25 µg |
[15] |
| Pitipeptolide A (4) | L. majuscula | Guam | M. tuberculosis ATCC 35818 | 15 mm at 100 µg 9 mm at 25 µg |
[15] |
| Pitipeptolide B (5) | L. majuscula | Guam | M. tuberculosis ATCC 25177 | 30 mm at 100 µg 15 mm at 25 µg |
[15] |
| Pitipeptolide B (5) | L. majuscula | Guam | M. tuberculosis ATCC 35818 | 15 mm at 100 µg 10 mm at 25 µg |
[15] |
| Pitipeptolide A (4) | L. majuscula | Guam | M. tuberculosis ATCC 25177 | 28 mm at 100 µg 23 mm at 50 µg 9 mm at 10 µg |
[16] |
| Pitipeptolide B (5) | L. majuscula | Guam | M. tuberculosis ATCC 25177 | 30 mm at 100 µg 24 mm at 50 µg 14 mm at 10 µg |
[16] |
| Pitipeptolide C (6) | L. majuscula | Guam | M. tuberculosis ATCC 25177 | 26 mm at 100 µg 21 mm at 50 µg 18 mm at 10 µg |
[16] |
| Pitipeptolide D (7) | L. majuscula | Guam | M. tuberculosis ATCC 25177 | 10 mm at 100 µg 0 mm at 50 µg 0 mm at 10 µg |
[16] |
| Pitipeptolide E (8) | L. majuscula | Guam | M. tuberculosis ATCC 25177 | 21 mm at 100 µg 15 mm at 50 µg 0 mm at 10 µg |
[16] |
| Pitipeptolide F (9) | L. majuscula | Guam | M. tuberculosis ATCC 25177 | 40 mm at 100 µg 30 mm at 50 µg 10 mm at 10 µg |
[16] |
| Pitiprolamide (10) | L. majuscula | Guam | M. tuberculosis ATCC 25177 | 23 mm at 100 µg 13 mm at 50 µg 0 mm at 10 µg |
[17] |
| Pitiprolamide (10) | L. majuscula | Guam | B. cereus ATCC 10987 | IC50 = 70 μM at 1 μM | [17] |
| Mixture of lyngbyazothrins A and B (14 and 15) | Lyngbya sp. | Germany (Culture) | M. flaVus SBUG 16 | 8 mm at 100 μg/disk | [18] |
| Mixture of lyngbyazothrins C (16) and D (17) | Lyngbya sp. | Germany (Culture) | B. subtilis SBUG 14 E. coli ATCC 11229 E. coli SBUG 13 P. aeruginosa ATCC 27853 S. marcescens SBUG 9 |
18 mm at 25 μg/disk 18 mm at 100 μg/disk 15 mm at 100 μg/disk 8 mm at 100 μg/disk 8 mm at 200 μg/disk |
[18] |
| Tiahuramide A (18) | L. majuscula | French Polynesia | A. salmonicida (CIP 103209T strain), V. anguillarum (CIP 63.36T), S. baltica (CIP 105850T), E. coli (CIP 54.8) and M. luteus (CIP A270) | MIC = 27, 33, >50, 35 and 47 μM, respectively | [19] |
| Tiahuramide B (19) | L. majuscula | French Polynesia | A. salmonicida (CIP 103209T strain), V. anguillarum (CIP 63.36T), S. baltica (CIP 105850T), E. coli (CIP 54.8) and M. luteus (CIP A270) | MIC = 9.4, 8.5, 22, 12 and 29 μM, respectively | [19] |
| Tiahuramide C (20) | L. majuscula | French Polynesia | A. salmonicida (CIP 103209T strain), V. anguillarum (CIP 63.36T), S. baltica (CIP 105850T), E. coli (CIP 54.8) and M. luteus (CIP A270) | MIC = 6.7, 7.4, 16, 14 and 17 μM, respectively | [19] |
References
- Schopf, J.W.; Packer, B.M. Early Archean (3.3-billion to 3.5-billion-yearold) microfossils from Warrawoona Group, Australia. Science 1987, 237, 70–73.
- Whitton, B.A.; Potts, M. The Ecology of Cyanobacteria: Their Diversity in Time and Space; Springer: Dordrecht, The Netherlands, 2000.
- Nagle, D.G.; Paul, V.J. Production of secondary metabolites by filamentous tropical marine cyanobacteria: Ecological functions of the compounds. J. Phycol. 1999, 35, 1412–1421.
- Berry, J.P.; Gantar, M.; Perez, M.H.; Berry, G.; Noriega, F.G. Cyanobacterial toxins as allelochemicals with potential applications as algaecides, herbicides and insecticides. Mar. Drugs 2008, 6, 117–146.
- Burja, A.M.; Banaigs, B.; Abou-Mansour, E.; Burgess, J.G.; Wright, P.C. Marine cyanobacteria—A prolific source of natural products. Tetrahedron 2001, 57, 9347–9377.
- Engene, N.; Rottacker, E.C.; Kas, J.; Byrum, T.; Gerwick, W.H.; Choi, H.; Ellisman, M.H. Moorea producens Gen. Nov., Sp. Nov. and Moorea bouillonii Comb. Nov., Tropical marine cyanobacteria rich in bioactive secondary metabolites. Int. J. Syst. Evol. Microbiol. 2012, 62 Pt 5, 1171–1178.
- Engene, N.; Tronholm, A.; Paul, V.J. Uncovering cryptic diversity of Lyngbya: The new tropical marine cyanobacterial genus Dapis (Oscillatoriales). J. Phycol. 2018, 54, 435–446.
- Jiří, K.; Eliška, Z.; Jan, Š.; Jiří, K.; Jason, W.; Brett, A.N.; Jaroslava, K. Polyphasic evaluation of Limnoraphis robusta, a water—Bloom forming cyanobacterium from Lake Atitlán, Guatemala, with a description of Limnoraphis gen. nov. J. Czech Phycol. Soc. 2013, 13, 39–52.
- Tronholm, A.; Engene, N. Moorena Gen. Nov., a valid name for “Moorea Engene & Al.” Nom. Inval. (Oscillatoriaceae, Cyanobacteria). Not. Algarum 2019, 122, 1–2.
- Mcgregor, G.B.; Sendall, B.C. Phylogeny and toxicology of Lyngbya wollei (Cyanobacteria, Oscillatoriales) from North-Eastern Australia, with description of Microseira gen. nov. J. Phycol. 2015, 51, 109–119.
- Engene, N.; Paul, V.J.; Byrum, T.; Gerwick, W.H.; Thor, A.; Ellisman, M.H. Five chemically rich species of tropical marine cyanobacteria of the genus Okeania gen. nov. (Oscillatoriales, Cyanoprokaryota). J. Phycol. 2013, 49, 1095–1106.
- Cardllina, J.H.C.; Moore, R.E.; Arnold, E.V.; Clardy, J. Structure and absolute configuration of malyngolide, an antibiotic from the marine blue-green alga Lyngbya majuscula Gomont. J. Org. Chem. 1979, 44, 4039–4042.
- Shaala, L.A.; Youssef, D.T.A.; McPhail, K.L.; Elbandy, M. Malyngamide 4, a new lipopeptide from the Red Sea marine cyanobacterium Moorea producens (Formerly Lyngbya majuscula). Phytochem. Lett. 2013, 6, 183–188.
- Gekwick, W.H.; Reyes, S.; Alvarado, B. Two malyngamides from the Caribbean cyanobacterium Lyngbya majuscula. Phytochemistry 1987, 26, 1701–1704.
- Luesch, H.; Pangilinan, R.; Yoshida, W.Y.; Moore, R.E.; Paul, V.J. Pitipeptolides A and B, new cyclodepsipeptides from the marine cyanobacterium Lyngbya majuscula. J. Nat. Prod. 2001, 64, 304–307.
- Montaser, R.; Paul, V.J.; Luesch, H. Pitipeptolides C-F, antimycobacterial cyclodepsipeptides from the marine cyanobacterium Lyngbya majuscula from Guam. Phytochemistry 2011, 72, 2068–2074.
- Montaser, R.; Abboud, K.A.; Paul, V.J.; Luesch, H. Pitiprolamide, a proline-rich dolastatin 16 analogue from the marine cyanobacterium Lyngbya majuscula from Guam. J. Nat. Prod. 2011, 74, 109–112.
- Zainuddin, E.N.; Jansen, R.; Nimtz, M.; Wray, V.; Preisitsch, M.; Lalk, M.; Mundt, S. Lyngbyazothrins A-D, antimicrobial cyclic undecapeptides from the cultured cyanobacterium Lyngbya sp. J. Nat. Prod. 2009, 72, 2080.
- Levert, A.; Alvarin, R.; Bornancin, L.; Mansour, E.A.; Burja, A.M.; Genevie, A.; Bonnard, I.; Alonso, E.; Botana, L.; Banaigs, B. Structures and activities of tiahuramides A−C, cyclic depsipeptides from a Tahitian collection of the marine cyanobacterium Lyngbya majuscula. J. Nat. Prod. 2018, 81, 1301–1310.





