
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Ping Zhang | -- | 1935 | 2022-08-29 11:48:34 | | | |
| 2 | Camila Xu | Meta information modification | 1935 | 2022-08-30 02:25:10 | | | | |
| 3 | Camila Xu | -59 word(s) | 1876 | 2022-08-30 02:25:30 | | |
Video Upload Options
The gut microbiome, or gut microbiota, also termed commensal, refers to the entire microbial community that populates the mammalian gastrointestinal (GI) tract, with the majority residing in the colon. Alterations of the gut microbiome are implicated in many gastrointestinal diseases, such as inflammatory bowel disease (IBD). Food components in our diet provide not only nutrients to our body but also substrates for the gut microbial flora. What we eat shapes the structure, composition, and function of the gut microbiome, which interacts with the gut epithelium and mucosal immune system and maintains intestinal homeostasis in a healthy state. There is growing interest in nutritional therapy to target the gut microbiome in IBD.
1. Introduction
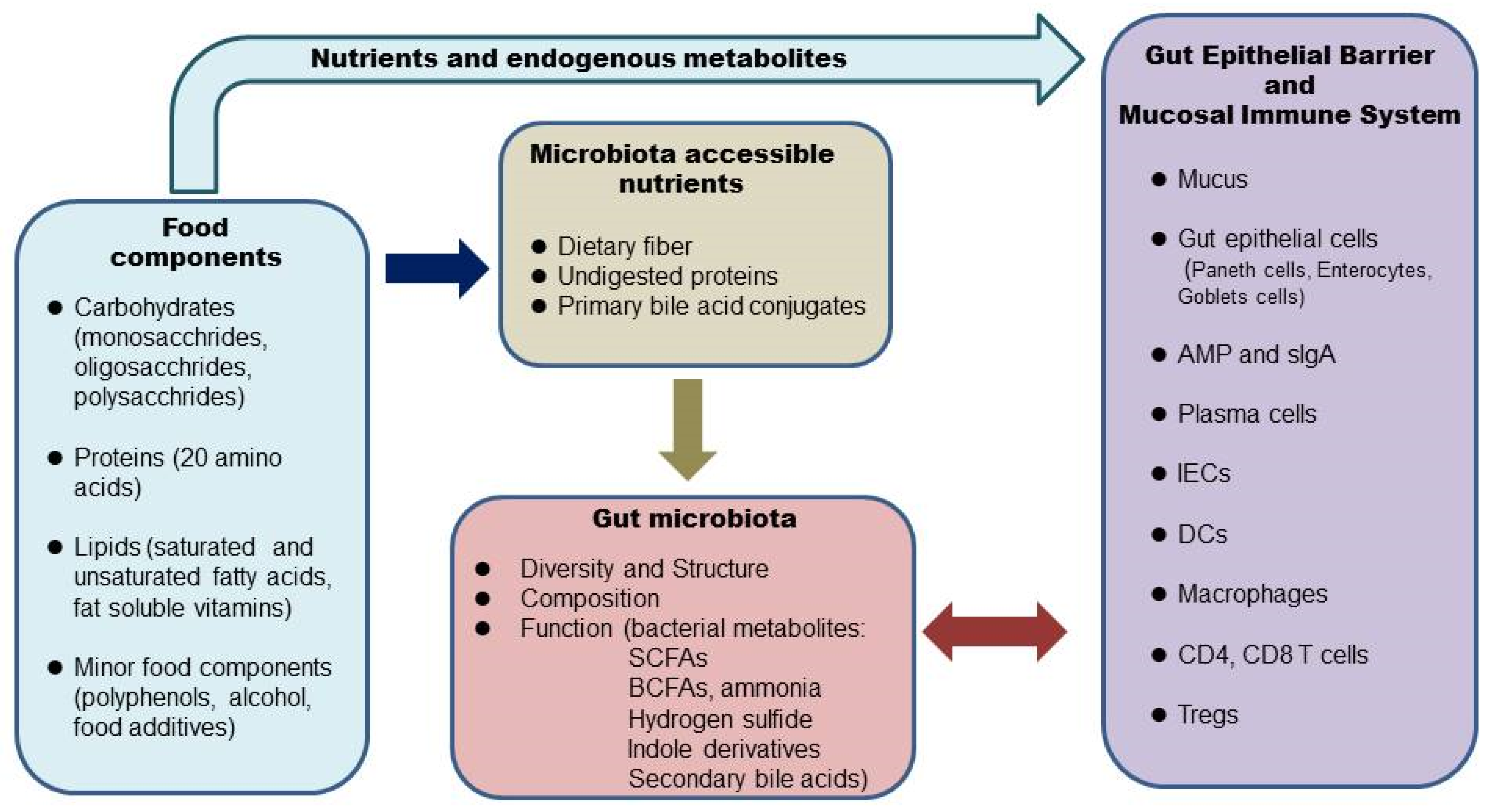
2. Diet, the Microbiome, and the Intestinal Barrier
3. Diet, the Microbiome, and the Intestinal Mucosal Immune System
References
- Sender, R.; Fuchs, S.; Milo, R. Revised estimates for the number of human and bacteria cells in the body. PLoS Biol. 2016, 14, e1002533.
- Postler, T.S.; Ghosh, S. Understanding the holobiont: How microbial metabolites affect human health and shape the immune system. Cell Metab. 2017, 26, 110–130.
- Arumugam, M.; Raes, J.; Pelletier, E.; le Paslier, D.; Yamada, T.; Mende, D.R.; Fernandes, G.R.; Tap, J.; Bruls, T.; Batto, J.M.; et al. Enterotypes of the human gut microbiome. Nature 2011, 473, 174–180.
- Frank, D.N.; St. Amand, A.L.; Feidman, R.A.; Boedeker, E.C.; Harpaz, N.; Pace, N.R. Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc. Natl. Acad. Sci. USA 2007, 104, 13780–13785.
- Schroeder, B.; Bäckhed, F. Signals from the gut microbiota to distant organs in physiology and disease. Nat. Med. 2016, 22, 1079–1089.
- Singh, R.K.; Chang, H.W.; Yan, D.; Lee, K.M.; Ucmak, D.; Wong, K.; Abrouk, M.; Farahnik, B.; Nakamura, M.; Zhu, T.H.; et al. Influence of diet on the gut microbiome and implications for human health. J. Transl. Med. 2017, 15, 73.
- Smits, S.A.; Leach, J.; Sonnenburg, E.D.; Gonzalez, C.G.; Lichtman, J.S.; Reid, G.; Knight, R.; Manjurano, A.; Changalucha, J.; Elias, J.E.; et al. Seasonal cycling in the gut microbiome of the Hadaz hunter-gatherers of Tanzania. Science 2016, 357, 802–806.
- Jha, A.R.; Davenport, E.R.; Gautam, Y.; Bhandari, D.; Tandukar, S.; Ng, K.M.; Fragiadakis, G.K.; Holmes, S.; Gautam, G.P.; Leach, J.; et al. Gut microbiome transition across a lifestyle gradient in Himalaya. PLoS Biol. 2018, 16, e2005396.
- Deehan, E.C.; Yang, C.; Perez-Muñoz, M.E.; Nguyen, N.K.; Cheng, C.C.; Triador, L.; Zhang, Z.; Bakal, J.A.; Walter, J. Precision microbiome modulation with discrete dietary fiber structures directs short-chain fatty acids production. Cell Host Microbe 2020, 27, 389–404.
- Wastyk, H.; Fragiadakis, G.K.; Perelman, D.; Dahan, D.; Merrill, B.D.; Yu, F.B.; Topf, M.; Gonzalez, C.G.; Treuren, W.V.; Han, S. Gut-microbiota-targeted diets modulate human immune status. Cell 2021, 184, 4137–4153.
- Rinninella, E.; Cintoni, M.; Raoul, P.; Lopetuso, L.R.; Scaldafferri, F.; Pulcini, G.; Miggiano, G.A.D.; Gasbarrini, A.; Mele, G.P. Food components and dietary habits: Keys for a healthy gut microbiota composition. Nutrients 2019, 11, 2393.
- Lane, E.R.; Zisman, T.L.; Suskind, D.L. The microbiota in inflammatory bowel disease: Current and therapeutic insights. J. Inflamm. Res. 2017, 10, 63–73.
- Sugihara, K.; Kamada, N. Diet-microbiota interactions in inflammatory bowel disease. Nutrients 2021, 13, 1533.
- Staley, C.; Weingarden, A.R.; Khoruts, A.; Sadowsky, M.J. Interaction of gut microbiota with bile acid metabolism and its influence on disease states. Appl. Microbiol. Biotechnol. 2017, 101, 47–64.
- De Santis, S.; Cavalcanti, E.; Mastronardi, M.; Jirillo, E.; Chieppa, M. Nutritional keys for intestinal barrier modulation. Front. Immunol. 2015, 6, 612.
- Gerbe, F.; Legraverend, C.; Jay, P. The intestinal epithelium tuft cells: Specification and function. Cell Mol. Life Sci. 2012, 69, 2907–2917.
- Pelaseyed, T.; Bergström, J.H.; Gustafsson, J.K.; Ermund, A.; Birchenough, G.M.H.; Schütte, A.; van der Post, S.; Svensson, F.; Rodríguez-Piñeiro, A.M.; Nyström, E.E.L.; et al. The mucus and mucins of the goblet cells and enterocytes provide the first defense line of the gastrointestinal tract and interact with the immune system. Immonol. Rev. 2014, 260, 8–20.
- Desal, M.S.; Seekatz, A.M.; Koropatkin, N.K.; Kamada, N.; Hickey, C.A.; Wolter, M.; Pudlo, N.A.; Kitamoto, S.; Terrapon, N.; Muller, A.; et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell 2016, 167, 1339–1353.
- Mu, Q.; Kirby, J.; Reilly, C.M.; Luo, X.M. Leaky gut as a danger signal for autoimmune diseases. Front. Immonol. 2017, 8, 598.
- Kalina, U.; Koyama, N.; Hosoda, T.; Nuernberger, H.; Sato, K.; Hoelzer, D.; Herweck, F.; Manigold, T.; Singer, M.V.; Rossol, S.; et al. Enhanced production of IL-18 in butyrate-treated intestinal epithelium by stimulation of the proximal promoter region. Eur. J. Immunol. 2002, 32, 2635–2643.
- Zhao, F.; Zhou, G.; Liu, X.; Song, S.; Xu, X.; Hooiveld, G.; Müller, M.; Liu, L.; Kristiansen, K.; Li, C. Dietary protein sources differentially affect the growth of akkermansia muciniphila and maintenance of the gut mucus barrier in mice. Mol. Nutr. Food Res. 2019, 63, 1900589.
- Gruber, L.; Kisling, S.; Lichti, P.; Martin, F.P.; May, S.; Klingenspor, M.; Lichtenegger, M.; Rychlik, M.; Haller, D. High fat diet accelerates pathogenesis of murine crohn’s disease-like ileitis independently of obesity. PLoS ONE 2013, 8, e71161.
- Lam, Y.Y.; Ha, C.W.Y.; Campbell, C.R.; Mitchell, A.J.; Dinudom, A.; Oscarsson, J.; Cook, D.I.; Hunt, N.H.; Caterson, I.D.; Holmes, A.J.; et al. Increased gut permeability and microbioa change associate with mesenteric fat inflammation and metabolic dysfunction in diet-induced obese mice. PLoS ONE 2012, 7, e34233.
- Khan, S.; Waliullah, S.; Godfrey, V.; Khan, M.A.W.; Ramachandran, R.A.; Cantarel, B.; Behrendt, C.; Peng, L.; Hooper, L.V.; Zaki, H. Dietary simple sugars alter microbial ecology in the gut and promote colitis in mice. Sci. Transl. Med. 2020, 12, eaay6218.
- Do, M.H.; Lee, E.; Oh, M.J.; Kim, Y.; Park, H.Y. High-glucose or -fructose diet cause changes of the gut microbiota and metabolic disorders in mice without body weight changes. Nutrients 2018, 10, 761.
- Chassaing, B.; Koren, O.; Goodrich, J.K.; Poole, A.C.; Srinivasan, S.; Ley, R.E.; Gewirtz, A.T. Dietary emulsifiers impact the mouse gut microbiota promoting colitis and metabolic syndrome. Nature 2015, 519, 92–96.
- Panduro, M.; Benoist, C.; Mathis, D. Tissue-Tregs. Annu. Rev. Immunol. 2016, 34, 609–633.
- Tanoue, T.; Atarashi, K.; Honda, K. Development and maintenance of intestinal regulatory T cells. Nat. Rev. Immunol. 2016, 16, 295–309.
- Smith, P.M.; Howitt, M.R.; Panikov, N.; Michaud, M.; Gallini, C.A.; Bohlooly-Y, M.; Glickman, J.N.; Garrett, W.S. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 2013, 341, 569–573.
- Furusawa, Y.; Obata, Y.; Fukuda, S.; Endo, T.A.; Nakato, G.; Takahashi, D.; Nakanishi, Y.; Uetake, C.; Kato, K.; Kato, T.; et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 2013, 504, 446–450.
- Campbell, C.; McKenney, P.T.; Konstantinovsky, D.; Isaeva, O.I.; Schizas, M.; Verter, J.; Mai, C.; Jin, W.B.; Guo, C.J.; Violante, S. Bacterial metabolism of bile acids promotes generation of peripheral regulatory T cells. Nature 2022, 581, 475–479.
- Song, X.; Sun, X.; Oh, S.F.; Wu, M.; Zhang, Y.; Zheng, W.; Geva-Zatorysky, N.; Jupp, R.; Mathis, D.; Benoist, C.; et al. Microbial bile acid metabolites modulate gut RORγ+ regulatory T cell homeostasis. Nature 2020, 577, 410–415.
- Lamas, B.; Richard, M.L.; Leducaq, V.; Pham, H.-P.; Michel, M.-L.; da Costa, G.; Bridonneau, C.; Jegou, S.; Hoffmann, T.W.; Natividad, J.M.; et al. CARD9 impacts colitis by altering gut microbiota metabolism of tryptophan into aryl hydrocarbon receptor ligands. Nat. Med. 2016, 22, 598–605.
- Geva-Zatorsky, N.; Sefik, E.; Kua, L.; Pasman, L.; Tan, T.G.; Ortiz-Lopez, A.; Yanortsang, T.B.; Yang, L.; Jupp, R.; Mathis, D. Mining the human gut microbiota for immunomodulatory organisms. Cell 2017, 168, 928–943.
- Fontenot, J.D.; Gavin, M.A.; Rudensky, A.Y. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat. Immunol. 2003, 4, 330–336.
- Sakai, R.; Komai, K.; Iizuka-Koga, M.; Yoshimura, A.; Ito, M. Regulatory T cells: Pathophysiological roles and clinical applications. Keio J. Med. 2020, 69, 1–15.





