
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Martin Kiening | -- | 3105 | 2022-07-26 15:57:24 | | | |
| 2 | Peter Tang | Meta information modification | 3105 | 2022-07-27 02:58:52 | | |
Video Upload Options
Mitochondria are essential organelles of mammalian cells, often emphasized for their function in energy production, iron metabolism and apoptosis as well as heme synthesis. The heme is an iron-loaded porphyrin behaving as a prosthetic group by its interactions with a wide variety of proteins. These complexes are termed hemoproteins and are usually vital to the whole cell comportment, such as the proteins hemoglobin, myoglobin or cytochromes, but also enzymes such as catalase and peroxidases. The building block of porphyrins is the 5-aminolevulinic acid, whose exogenous administration is able to stimulate the entire heme biosynthesis route. In neoplastic cells, this methodology repeatedly demonstrated an accumulation of the ultimate heme precursor, the fluorescent protoporphyrin IX photosensitizer, rather than in healthy tissues.
1. Introduction
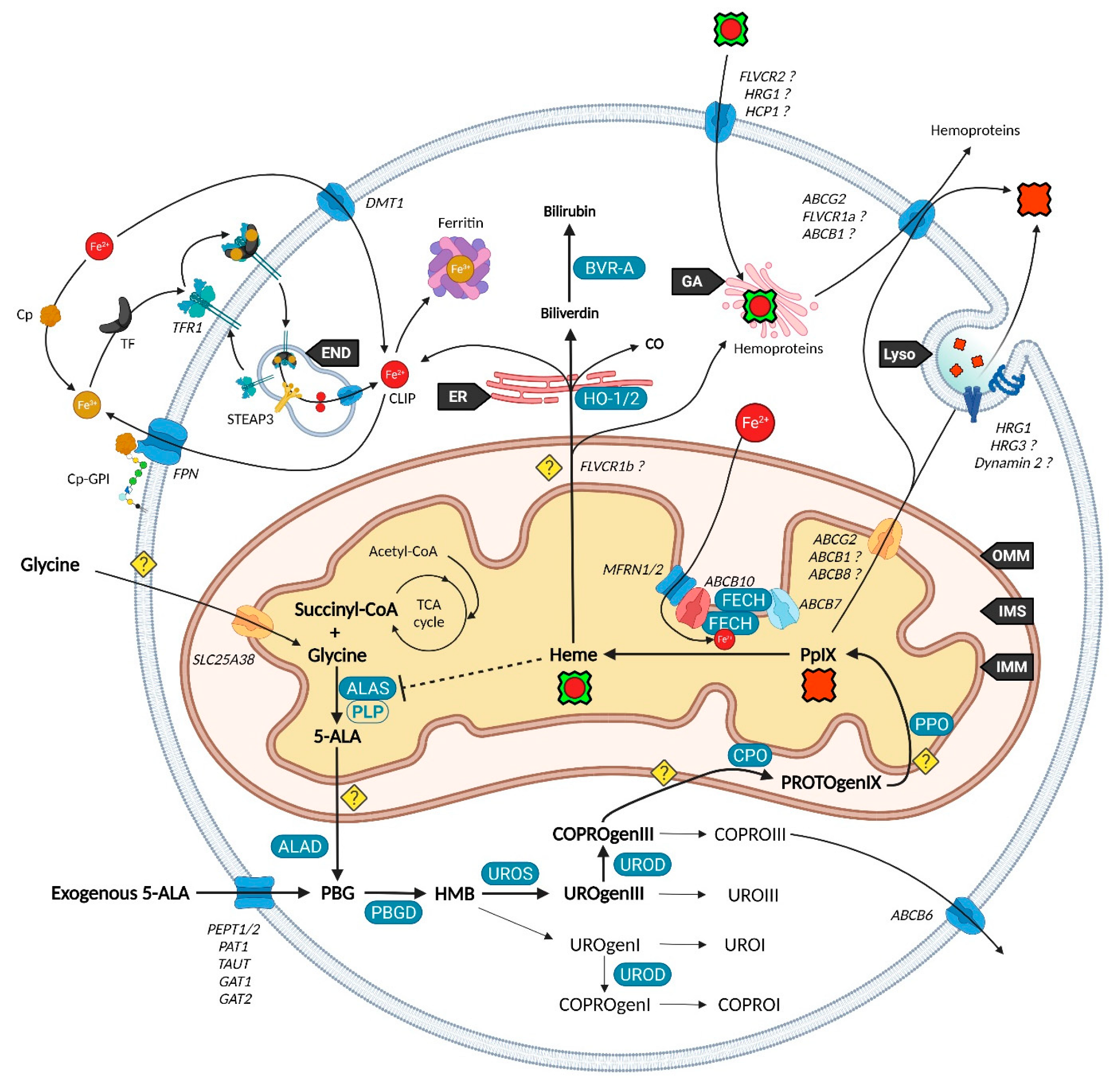
2. Heme Metabolism—A Complex Network Tightly Regulated by Enzymes, Transporters and Other Metabolites
2.1. Biosynthesis of Iron Protoporphyrin IX
2.2. Utilization and Degradation of Iron Protoporphyrin IX
2.3. Transporters of the Heme Metabolism: Where Are We Now?
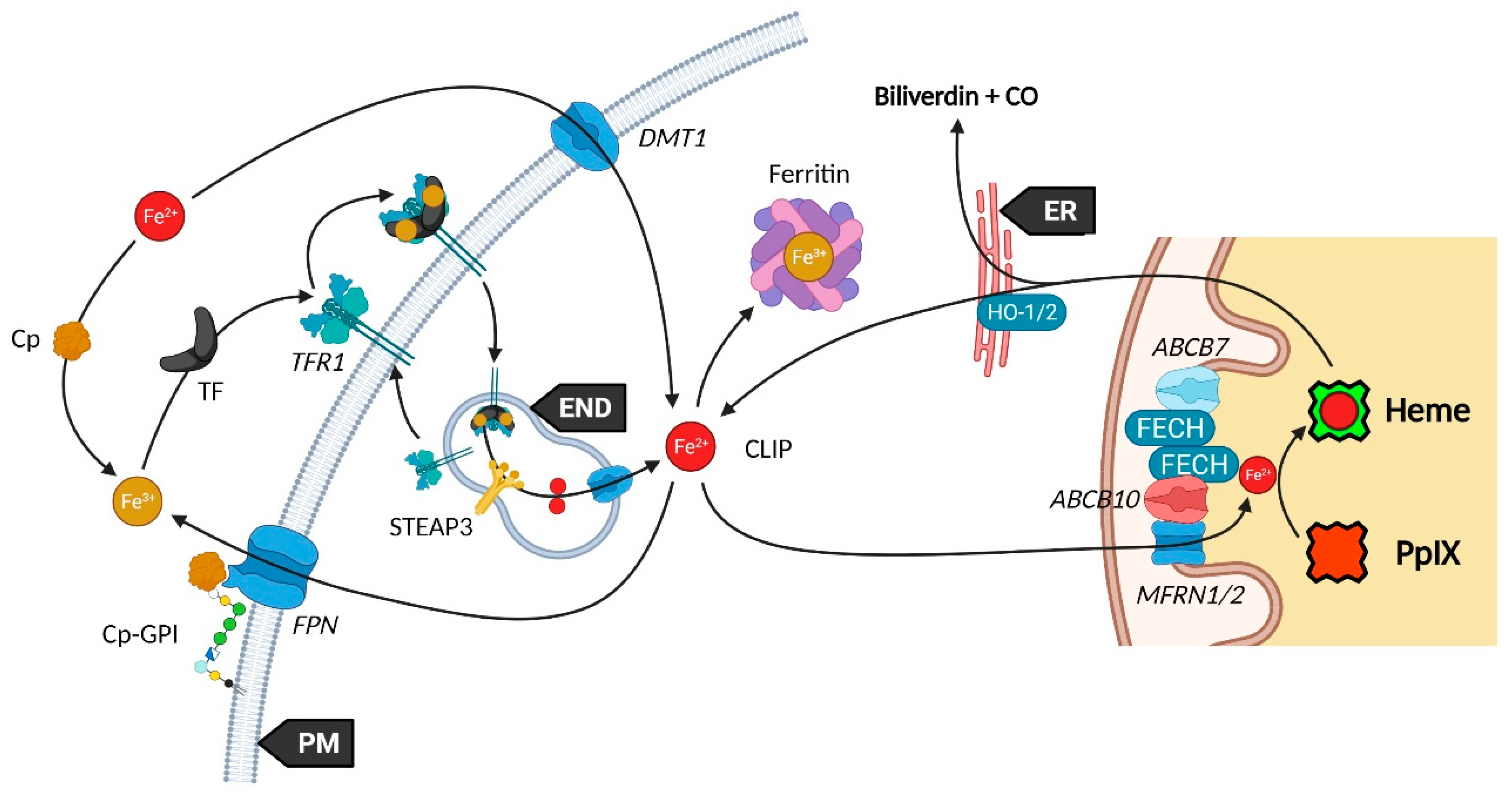
References
- Malik, Z.; Lugaci, H. Destruction of erythroleukaemic cells by photoactivation of endogenous porphyrins. Br. J. Cancer 1987, 56, 589–595.
- D’Hallewin, M.-A.; Bezdetnaya, L.; Guillemin, F. Fluorescence Detection of Bladder Cancer: A Review. Eur. Urol. 2002, 42, 417–425.
- Collaud, S.; Juzeniene, A.; Moan, J.; Lange, N. On the selectivity of 5-aminolevulinic acid-induced protoporphyrin IX formation. Curr. Med. Chem. Anticancer Agents 2004, 4, 301–316.
- Diez Valle, R.; Hadjipanayis, C.G.; Stummer, W. Established and emerging uses of 5-ALA in the brain: An overview. J. Neuro-Oncol. 2019, 141, 487–494.
- Kim, H.I.; Wilson, B.C. Photodynamic Diagnosis and Therapy for Peritoneal Carcinomatosis from Gastrointestinal Cancers: Status, Opportunities, and Challenges. J. Gastric Cancer 2020, 20, 355–375.
- Fotinos, N.; Campo, M.A.; Popowycz, F.; Gurny, R.; Lange, N. 5-Aminolevulinic Acid Derivatives in Photomedicine: Characteristics, Application and Perspectives. Photochem. Photobiol. 2006, 82, 994–1015.
- Shemin, D.; Rittenberg, D. The biological utilization of glycine for the synthesis of the protoporphyrin of hemoglobin. J. Biol. Chem. 1946, 166, 621–625.
- Bali, S.; Palmer, D.J.; Schroeder, S.; Ferguson, S.J.; Warren, M.J. Recent advances in the biosynthesis of modified tetrapyrroles: The discovery of an alternative pathway for the formation of heme and heme d 1. Cell. Mol. Life Sci. 2014, 71, 2837–2863.
- Heinemann, I.U.; Jahn, M.; Jahn, D. The biochemistry of heme biosynthesis. Arch. Biochem. Biophys. 2008, 474, 238–251.
- Hunter, G.A.; Ferreira, G.C. Molecular enzymology of 5-aminolevulinate synthase, the gatekeeper of heme biosynthesis. Biochim. Biophys. Acta 2011, 1814, 1467–1473.
- Medlock, A.E.; Shiferaw, M.T.; Marcero, J.R.; Vashisht, A.A.; Wohlschlegel, J.A.; Phillips, J.D.; Dailey, H.A. Identification of the Mitochondrial Heme Metabolism Complex. PLoS ONE 2015, 10, e0135896.
- Fortian, A.; Castaño, D.; Gonzalez, E.; Laín, A.; Falcon-Perez, J.M.; Millet, O. Structural, thermodynamic, and mechanistical studies in uroporphyrinogen III synthase: Molecular basis of congenital erythropoietic porphyria. Adv. Protein Chem. Struct. Biol. 2011, 83, 43–74.
- Kumar, S.; Bandyopadhyay, U. Free heme toxicity and its detoxification systems in human. Toxicol. Lett. 2005, 157, 175–188.
- Yanatori, I.; Richardson, D.R.; Toyokuni, S.; Kishi, F. How iron is handled in the course of heme catabolism: Integration of heme oxygenase with intracellular iron transport mechanisms mediated by poly (rC)-binding protein-2. Arch. Biochem. Biophys. 2019, 672, 108071.
- Stocker, R.; Yamamoto, Y.; McDonagh, A.F.; Glazer, A.N.; Ames, B.N. Bilirubin Is an Antioxidant of Possible Physiological Importance. Science 1987, 235, 1043–1046.
- Thomas, M.; Hardikar, W.; Greaves, R.F.; Tingay, D.G.; Loh, T.P.; Ignjatovic, V.; Newall, F.; Rajapaksa, A.E. Mechanism of bilirubin elimination in urine: Insights and prospects for neonatal jaundice. Clin. Chem. Lab. Med. 2021, 59, 1025–1033.
- Sumida, K.; Kawana, M.; Kouno, E.; Itoh, T.; Takano, S.; Narawa, T.; Tukey, R.H.; Fujiwara, R. Importance of UDP-glucuronosyltransferase 1A1 expression in skin and its induction by UVB in neonatal hyperbilirubinemia. Mol. Pharmacol. 2013, 84, 679–686.
- Kutty, R.K.; Maines, M.D. Purification and characterization of biliverdin reductase from rat liver. J. Biol. Chem. 1981, 256, 3956–3962.
- Kutty, R.K.; Maines, M.D. Hepatic heme metabolism: Possible role of biliverdin in the regulation of heme oxygenase activity. Biochem. Biophys. Res. Commun. 1984, 122, 40–46.
- Lunetti, P.; Damiano, F.; De Benedetto, G.; Siculella, L.; Pennetta, A.; Muto, L.; Paradies, E.; Marobbio, C.M.; Dolce, V.; Capobianco, L. Characterization of Human and Yeast Mitochondrial Glycine Carriers with Implications for Heme Biosynthesis and Anemia. J. Biol. Chem. 2016, 291, 19746–19759.
- Guernsey, D.L.; Jiang, H.; Campagna, D.R.; Evans, S.C.; Ferguson, M.; Kellogg, M.D.; Lachance, M.; Matsuoka, M.; Nightingale, M.; Rideout, A.; et al. Mutations in mitochondrial carrier family gene SLC25A38 cause nonsyndromic autosomal recessive congenital sideroblastic anemia. Nat. Genet. 2009, 41, 651–653.
- Bayeva, M.; Khechaduri, A.; Wu, R.; Burke, M.A.; Wasserstrom, J.A.; Singh, N.; Liesa, M.; Shirihai, O.S.; Langer, N.B.; Paw, B.H.; et al. ATP-binding cassette B10 regulates early steps of heme synthesis. Circ. Res. 2013, 113, 279–287.
- Seguin, A.; Takahashi-Makise, N.; Yien, Y.Y.; Huston, N.C.; Whitman, J.C.; Musso, G.; Wallace, J.A.; Bradley, T.; Bergonia, H.A.; Kafina, M.D.; et al. Reductions in the mitochondrial ABC transporter Abcb10 affect the transcriptional profile of heme biosynthesis genes. J. Biol. Chem. 2017, 292, 16284–16299.
- Frolund, S.; Marquez, O.C.; Larsen, M.; Brodin, B.; Nielsen, C.U. Delta-aminolevulinic acid is a substrate for the amino acid transporter SLC36A1 (hPAT1). Br. J. Pharmacol. 2010, 159, 1339–1353.
- Tran, T.T.; Mu, A.; Adachi, Y.; Adachi, Y.; Taketani, S. Neurotransmitter transporter family including SLC6A6 and SLC6A13 contributes to the 5-aminolevulinic acid (ALA)-induced accumulation of protoporphyrin IX and photodamage, through uptake of ALA by cancerous cells. Photochem. Photobiol. 2014, 90, 1136–1143.
- Iwaki, K.; Fujiwara, T.; Ito, T.; Suzuki, C.; Sasaki, K.; Ono, K.; Saito, K.; Fukuhara, N.; Onishi, Y.; Yokoyama, H.; et al. Flow Cytometry-Based Photodynamic Diagnosis with 5-Aminolevulinic Acid for the Detection of Minimal Residual Disease in Multiple Myeloma. Tohoku J. Exp. Med. 2019, 249, 19–28.
- Rodriguez, L.; Batlle, A.; Di Venosa, G.; MacRobert, A.J.; Battah, S.; Daniel, H.; Casas, A. Study of the mechanisms of uptake of 5-aminolevulinic acid derivatives by PEPT1 and PEPT2 transporters as a tool to improve photodynamic therapy of tumours. Int. J. Biochem. Cell Biol. 2006, 38, 1530–1539.
- Ning, S.; Kang, Q.; Fan, D.; Liu, J.; Xue, C.; Zhang, X.; Ding, C.; Zhang, J.; Peng, Q.; Ji, Z. Protein 4.1R is Involved in the Transport of 5-Aminolevulinic Acid by Interaction with GATs in MEF Cells. Photochem. Photobiol. 2018, 94, 173–178.
- Bermudez Moretti, M.; Correa Garcia, S.; Perotti, C.; Batlle, A.; Casas, A. Delta-Aminolevulinic acid transport in murine mammary adenocarcinoma cells is mediated by beta transporters. Br. J. Cancer 2002, 87, 471–474.
- Rud, E.; Gederaas, O.; Høgset, A.; Berg, K. 5-Aminolevulinic Acid, but not 5-Aminolevulinic Acid Esters, is Transported into Adenocarcinoma Cells by System BETA Transporters. Photochem. Photobiol. 2000, 71, 640–647.
- Krishnamurthy, P.C.; Du, G.; Fukuda, Y.; Sun, D.; Sampath, J.; Mercer, K.E.; Wang, J.; Sosa-Pineda, B.; Murti, K.G.; Schuetz, J.D. Identification of a mammalian mitochondrial porphyrin transporter. Nature 2006, 443, 586–589.
- Zhao, S.G.; Chen, X.F.; Wang, L.G.; Yang, G.; Han, D.Y.; Teng, L.; Yang, M.C.; Wang, D.Y.; Shi, C.; Liu, Y.H.; et al. Increased expression of ABCB6 enhances protoporphyrin IX accumulation and photodynamic effect in human glioma. Ann. Surg. Oncol. 2013, 20, 4379–4388.
- Bergam, P.; Reisecker, J.M.; Rakvacs, Z.; Kucsma, N.; Raposo, G.; Szakacs, G.; van Niel, G. ABCB6 Resides in Melanosomes and Regulates Early Steps of Melanogenesis Required for PMEL Amyloid Matrix Formation. J. Mol. Biol. 2018, 430, 3802–3818.
- Kiss, K.; Brozik, A.; Kucsma, N.; Toth, A.; Gera, M.; Berry, L.; Vallentin, A.; Vial, H.; Vidal, M.; Szakacs, G. Shifting the paradigm: The putative mitochondrial protein ABCB6 resides in the lysosomes of cells and in the plasma membrane of erythrocytes. PLoS ONE 2012, 7, e37378.
- Paterson, J.K.; Shukla, S.; Black, C.M.; Tachiwada, T.; Garfield, S.; Wincovitch, S.; Ernst, D.N.; Agadir, A.; Li, X.; Ambudkar, S.V.; et al. Human ABCB6 Localizes to Both the Outer Mitochondrial Membrane and the Plasma Membrane. Biochemistry 2007, 46, 9443–9452.
- Matsumoto, K.; Hagiya, Y.; Endo, Y.; Nakajima, M.; Ishizuka, M.; Tanaka, T.; Ogura, S. Effects of plasma membrane ABCB6 on 5-aminolevulinic acid (ALA)-induced porphyrin accumulation in vitro: Tumor cell response to hypoxia. Photodiagn. Photodyn. Ther. 2015, 12, 45–51.
- Fukuda, Y.; Cheong, P.L.; Lynch, J.; Brighton, C.; Frase, S.; Kargas, V.; Rampersaud, E.; Wang, Y.; Sankaran, V.G.; Yu, B.; et al. The severity of hereditary porphyria is modulated by the porphyrin exporter and Lan antigen ABCB6. Nat. Commun. 2016, 7, 12353.
- Yien, Y.Y.; Robledo, R.F.; Schultz, I.J.; Takahashi-Makise, N.; Gwynn, B.; Bauer, D.E.; Dass, A.; Yi, G.; Li, L.; Hildick-Smith, G.J.; et al. TMEM14C is required for erythroid mitochondrial heme metabolism. J. Clin. Investig. 2014, 124, 4294–4304.
- Kobuchi, H.; Moriya, K.; Ogino, T.; Fujita, H.; Inoue, K.; Shuin, T.; Yasuda, T.; Utsumi, K.; Utsumi, T. Mitochondrial localization of ABC transporter ABCG2 and its function in 5-aminolevulinic acid-mediated protoporphyrin IX accumulation. PLoS ONE 2012, 7, e50082.
- Partha, K.; John, D.S. The Role of ABCG2 and ABCB6 in Porphyrin Metabolism and Cell Survival. Curr. Pharm. Biotechnol. 2011, 12, 647–655.
- Kitajima, Y.; Ishii, T.; Kohda, T.; Ishizuka, M.; Yamazaki, K.; Nishimura, Y.; Tanaka, T.; Dan, S.; Nakajima, M. Mechanistic study of PpIX accumulation using the JFCR39 cell panel revealed a role for dynamin 2-mediated exocytosis. Sci. Rep. 2019, 9, 8666.
- Yoshioka, E.; Chelakkot, V.S.; Licursi, M.; Rutihinda, S.G.; Som, J.; Derwish, L.; King, J.J.; Pongnopparat, T.; Mearow, K.; Larijani, M. Enhancement of Cancer-Specific Protoporphyrin IX Fluorescence by Targeting Oncogenic Ras/MEK Pathway. Theranostics 2018, 8, 2134–2146.
- Shono, K.; Mizobuchi, Y.; Yamaguchi, I.; Nakajima, K.; Fujiwara, Y.; Fujihara, T.; Kitazato, K.; Matsuzaki, K.; Uto, Y.; Sampetrean, O.; et al. Elevated cellular PpIX potentiates sonodynamic therapy in a mouse glioma stem cell-bearing glioma model by downregulating the Akt/NF-kappaB/MDR1 pathway. Sci. Rep. 2021, 11, 15105.
- Maio, N.; Kim, K.S.; Holmes-Hampton, G.; Singh, A.; Rouault, T.A. Dimeric ferrochelatase bridges ABCB7 and ABCB10 homodimers in an architecturally defined molecular complex required for heme biosynthesis. Haematologica 2019, 104, 1756–1767.
- Chen, W.; Paradkar, P.N.; Li, L.; Pierce, E.L.; Langer, N.B.; Takahashi-Makise, N.; Hyde, B.B.; Shirihai, O.S.; Ward, D.M.; Kaplan, J.; et al. Abcb10 physically interacts with mitoferrin-1 (Slc25a37) to enhance its stability and function in the erythroid mitochondria. Proc. Natl. Acad. Sci. USA 2009, 106, 16263–16268.
- Blajan, I.; Miersch, H.; Schmidt, D.; Kristiansen, G.; Perner, S.; Ritter, M.; Ellinger, J.; Klumper, N. Comprehensive Analysis of the ATP-binding Cassette Subfamily B Across Renal Cancers Identifies ABCB8 Overexpression in Phenotypically Aggressive Clear Cell Renal Cell Carcinoma. Eur. Urol. Focus 2021, 7, 1121–1129.
- Ichikawa, Y.; Bayeva, M.; Ghanefar, M.; Potini, V.; Sun, L.; Mutharasan, R.K.; Wu, R.; Khechaduri, A.; Jairaj, N.T.; Ardehali, H. Disruption of ATP-binding cassette B8 in mice leads to cardiomyopathy through a decrease in mitochondrial iron export. Proc. Natl. Acad. Sci. USA 2012, 109, 4152–4157.
- Sulpice, E.; Plouet, J.; Berge, M.; Allanic, D.; Tobelem, G.; Merkulova-Rainon, T. Neuropilin-1 and neuropilin-2 act as coreceptors, potentiating proangiogenic activity. Blood 2008, 111, 2036–2045.
- Stasikowska-Kanicka, O.; Wągrowska-Danilewicz, M.; Danilewicz, M. Immunohistochemical study on neuropilin 1 (NRP1) immunoexpression in oral squamous cell carcinoma. Folia Histochem. Cytobiol. 2018, 56, 98–105.
- Stephenson, J.M.; Banerjee, S.; Saxena, N.K.; Cherian, R.; Banerjee, S.K. Neuropilin-1 is differentially expressed in myoepithelial cells and vascular smooth muscle cells in preneoplastic and neoplastic human breast: A possible marker for the progression of breast cancer. Int. J. Cancer 2002, 101, 409–414.
- Hong, T.M.; Chen, Y.L.; Wu, Y.Y.; Yuan, A.; Chao, Y.C.; Chung, Y.C.; Wu, M.H.; Yang, S.C.; Pan, S.H.; Shih, J.Y.; et al. Targeting neuropilin 1 as an antitumor strategy in lung cancer. Clin. Cancer Res. 2007, 13, 4759–4768.
- Issitt, T.; Bosseboeuf, E.; De Winter, N.; Dufton, N.; Gestri, G.; Senatore, V.; Chikh, A.; Randi, A.M.; Raimondi, C. Neuropilin-1 Controls Endothelial Homeostasis by Regulating Mitochondrial Function and Iron-Dependent Oxidative Stress. iScience 2019, 11, 205–223.
- Litwack, G. Micronutrients (Metals and Iodine). Hum. Biochem. 2018, 591–643.
- Dietz, J.V.; Fox, J.L.; Khalimonchuk, O. Down the Iron Path: Mitochondrial Iron Homeostasis and Beyond. Cells 2021, 10, 2198.
- Wolff, N.A.; Ghio, A.J.; Garrick, L.M.; Garrick, M.D.; Zhao, L.; Fenton, R.A.; Thevenod, F. Evidence for mitochondrial localization of divalent metal transporter 1 (DMT1). FASEB J. 2014, 28, 2134–2145.
- Paul, B.T.; Manz, D.H.; Torti, F.M.; Torti, S.V. Mitochondria and Iron: Current questions. Expert Rev./Hematol. 2017, 10, 65–79.
- Osaki, S.; Johnson, D.A.; Frieden, E. The Possible Significance of the Ferrous Oxidase Activity of Ceruloplasmin in Normal Human Serum. J. Biol. Chem. 1966, 241, 2746–2751.
- Jeong, S.Y.; David, S. Glycosylphosphatidylinositol-anchored ceruloplasmin is required for iron efflux from cells in the central nervous system. J. Biol. Chem. 2003, 278, 27144–27148.
- Ponka, P.; Sheftel, A.D.; English, A.M.; Scott Bohle, D.; Garcia-Santos, D. Do Mammalian Cells Really Need to Export and Import Heme? Trends Biochem. Sci. 2017, 42, 395–406.
- Chiabrando, D.; Marro, S.; Mercurio, S.; Giorgi, C.; Petrillo, S.; Vinchi, F.; Fiorito, V.; Fagoonee, S.; Camporeale, A.; Turco, E.; et al. The mitochondrial heme exporter FLVCR1b mediates erythroid differentiation. J. Clin. Investig. 2012, 122, 4569–4579.
- Tupta, B.; Stuehr, E.; Sumi, M.P.; Sweeny, E.A.; Smith, B.; Stuehr, D.J.; Ghosh, A. GAPDH is involved in the heme-maturation of myoglobin and hemoglobin. FASEB J. 2022, 36, e22099.
- Quigley, J.G.; Yang, Z.; Worthington, M.T.; Phillips, J.D.; Sabo, K.M.; Sabath, D.E.; Berg, C.L.; Sassa, S.; Wood, B.L.; Abkowitz, J.L. Identification of a human heme exporter that is essential for erythropoiesis. Cell 2004, 118, 757–766.
- Destefanis, F.; Fiorito, V.; Altruda, F.; Tolosano, E. Investigating the Connection Between Endogenous Heme Accumulation and COX2 Activity in Cancer Cells. Front. Oncol. 2019, 9, 162.
- Duffy, S.P.; Shing, J.; Saraon, P.; Berger, L.C.; Eiden, M.V.; Wilde, A.; Tailor, C.S. The Fowler syndrome-associated protein FLVCR2 is an importer of heme. Mol. Cell. Biol. 2010, 30, 5318–5324.
- Yuan, X.; Protchenko, O.; Philpott, C.C.; Hamza, I. Topologically conserved residues direct heme transport in HRG-1-related proteins. J. Biol. Chem. 2012, 287, 4914–4924.
- Li, Y.; Ivica, N.A.; Dong, T.; Papageorgiou, D.P.; He, Y.; Brown, D.R.; Kleyman, M.; Hu, G.; Chen, W.W.; Sullivan, L.B.; et al. MFSD7C switches mitochondrial ATP synthesis to thermogenesis in response to heme. Nat. Commun. 2020, 11, 4837.
- Rajagopal, A.; Rao, A.U.; Amigo, J.; Tian, M.; Upadhyay, S.K.; Hall, C.; Uhm, S.; Mathew, M.K.; Fleming, M.D.; Paw, B.H.; et al. Haem homeostasis is regulated by the conserved and concerted functions of HRG-1 proteins. Nature 2008, 453, 1127–1131.
- Pek, R.H.; Yuan, X.; Rietzschel, N.; Zhang, J.; Jackson, L.; Nishibori, E.; Ribeiro, A.; Simmons, W.; Jagadeesh, J.; Sugimoto, H.; et al. Hemozoin produced by mammals confers heme tolerance. Elife 2019, 8, e49503.
- Fogarty, F.M.; O’Keeffe, J.; Zhadanov, A.; Papkovsky, D.; Ayllon, V.; O’Connor, R. HRG-1 enhances cancer cell invasive potential and couples glucose metabolism to cytosolic/extracellular pH gradient regulation by the vacuolar-H(+) ATPase. Oncogene 2014, 33, 4653–4663.
- Shayeghi, M.; Latunde-Dada, G.O.; Oakhill, J.S.; Laftah, A.H.; Takeuchi, K.; Halliday, N.; Khan, Y.; Warley, A.; McCann, F.E.; Hider, R.C.; et al. Identification of an intestinal heme transporter. Cell 2005, 122, 789–801.
- Li, H.; Wang, D.; Wu, H.; Shen, H.; Lv, D.; Zhang, Y.; Lu, H.; Yang, J.; Tang, Y.; Li, M. SLC46A1 contributes to hepatic iron metabolism by importing heme in hepatocytes. Metabolism 2020, 110, 154306.
- Piel, R.B., 3rd; Shiferaw, M.T.; Vashisht, A.A.; Marcero, J.R.; Praissman, J.L.; Phillips, J.D.; Wohlschlegel, J.A.; Medlock, A.E. A Novel Role for Progesterone Receptor Membrane Component 1 (PGRMC1): A Partner and Regulator of Ferrochelatase. Biochemistry 2016, 55, 5204–5217.
- Kaluka, D.; Batabyal, D.; Chiang, B.Y.; Poulos, T.L.; Yeh, S.R. Spectroscopic and mutagenesis studies of human PGRMC1. Biochemistry 2015, 54, 1638–1647.
- McGuire, M.R.; Mukhopadhyay, D.; Myers, S.L.; Mosher, E.P.; Brookheart, R.T.; Kammers, K.; Sehgal, A.; Selen, E.S.; Wolfgang, M.J.; Bumpus, N.N.; et al. Progesterone receptor membrane component 1 (PGRMC1) binds and stabilizes cytochromes P450 through a heme-independent mechanism. J. Biol. Chem. 2021, 297, 101316.





