Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Bernhard Huchzermeyer | -- | 2671 | 2022-07-01 12:16:43 | | | |
| 2 | Camila Xu | Meta information modification | 2671 | 2022-07-04 03:13:22 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Koyro, H.; Huchzermeyer, B. Plant Performance and Interaction with Microorganisms. Encyclopedia. Available online: https://encyclopedia.pub/entry/24738 (accessed on 24 May 2026).
Koyro H, Huchzermeyer B. Plant Performance and Interaction with Microorganisms. Encyclopedia. Available at: https://encyclopedia.pub/entry/24738. Accessed May 24, 2026.
Koyro, Hans-Werner, Bernhard Huchzermeyer. "Plant Performance and Interaction with Microorganisms" Encyclopedia, https://encyclopedia.pub/entry/24738 (accessed May 24, 2026).
Koyro, H., & Huchzermeyer, B. (2022, July 01). Plant Performance and Interaction with Microorganisms. In Encyclopedia. https://encyclopedia.pub/entry/24738
Koyro, Hans-Werner and Bernhard Huchzermeyer. "Plant Performance and Interaction with Microorganisms." Encyclopedia. Web. 01 July, 2022.
Copy Citation
Plant performance can be improved under adverse environmental conditions by integrated soil–water–plant solutions or integrated soil fertility and plant nutrient management. Microorganisms living in the soil, the rhizobiome, are distinguished from endophytes living inside plants.
water withhold
salinity
stress amendments
biochar
plant-microorganism consortia
soil amendments
1. Plant Performance in the Field
Plants are sessile organisms. In the field, they have to cope with some fluctuating environmental factors. Therefore, plants have a certain window of resistance that allows for instant adaptation to the respective spectrum of environmental factors [1], with different resistances among plant species. In several experiments, it was observed that a narrow tolerance window correlates with a high growth rate, while broadening the window of resistance will result in reduced growth rates [2][3][4]. Therefore, it may be expected that at locations with only little fluctuation in environmental factors, and rare incidents of extreme weather conditions, the most abundant plant species may be characterized with a narrow window of resistance [5]. Several of our crops are characterized with a narrow window of resistance as compared with their wild ancestors, since breeding programs were aimed at high-yielding crops. In contrast to earlier breeding concepts, more recent projects are aimed at breeding for crop accessions that will produce a reliable yield, even under slightly suboptimal growth conditions [6][7][8][9][10].
Fluctuating illumination on a cloudy day, day–night changes, etc. are environmental changes that each plant has to tolerate. Plants will preferentially adapt by the regulation of enzyme activities rather than the production of new enzymes and modification of enzyme abundance. Therefore, these responses can occur almost instantly and easily escape detection in the field. Accordingly, most of the experiments that analyze these immediate responses were carried out in laboratories [11]. In general, these short-term events will not cause immediate damage, but may affect the resource use efficiency. On the other hand, long lasting periods of drought, heat, lack of nutrients, soil salinization, etc., will adversely affect plant performance at a significant degree. These conditions have been termed persistent stress. In accordance with Larcher [12], researchers have to distinguish between (i) mild stress, which allows stressed plants to thrive and subsequently re-adjust to a new physiological equilibrium, and (ii) severe, lethal stress, which is beyond the resistance level of the respective plant species (see Table 1).
In the literature, environmental stress is often referred to as abiotic stress to differentiate it from biotic stress, which is caused by pathogens and herbivores. With respect to its global economic importance, abiotic stress is estimated to cause 50 to 80% of yield losses in crop production [13][14]. The calculation is based on a comparison between the current annual crop yield and theoretical yield. This can be performed by scientists under completely controlled conditions in a greenhouse [15]. Therefore, the precise extent of yield reduction can be questioned. Nevertheless, the enormous economic impact of abiotic stress is beyond any discussion.
Plant response to abiotic stress is described as a multifactorial trait [16]. Moreover, plant species differ to a large degree in the preferential use of response patterns. This applies to any type of abiotic stress. Drought and salt stress are of primary importance for agriculture in arid areas. However, these two stresses can be found in other climatic zones, as well. Accordingly, many scientists have focused on analyzing the effects induced by the application of these two types of stress. The common trait between both stresses is that plant roots can sense low levels of available water. Therefore, some of the plant responses to these two types of abiotic stress are quite similar, such as osmolyte production (sugars, amino acids, and other organic molecules), in which the osmotic potential in plant roots adjusts to facilitate water uptake from the soil. In the case of soil salinity, some plants use an import of inorganic ions rather than osmolyte synthesis (on the expense of assimilate consumption) to control cellular osmotic potential [17][18][19].
Table 1. Plant response to environmental stress.
| Type of Stress | Application/Duration | Example | Reference |
|---|---|---|---|
| Eustress | whole plant priming | taking advantage of cross tolerance to different types of stress | Villagómez-Arande et al., 2022 [20] |
| Distress | fluctuating environmental conditions | day–night cycles; rain–sunshine cycles | Lichtenthaler, 1996 [1] |
| short time, hours to 4 days | laboratory experiments to identify stress-responsive traits (genes) | Miller et al., 2015 [21] | |
| series of adverse environmental conditions (weeks to months) | drought periods | Fahad et al., 2017 [22] | |
| poor growth conditions | soil salinity lack of nutrients deserts |
Zhao et al., 2020 [23] | |
| transfer to a different environment |
cultivation of plants not native to an area | Geppert et al., 2021 [24] |
All types of stress will affect the physiology of a plant as a whole. This indicates that stress perception is followed by signaling events (Figure 1). In the case of water shortage and salinity, ABA is one of the messengers. However, reactions other than closure of stomata and abscission of leaves can take place as a response to abiotic stress, as well. Among the stress responses, major physiological processes not under ABA control, such as nitrogen fixation, respiration, photosynthesis, and carbohydrate metabolism are included [25].
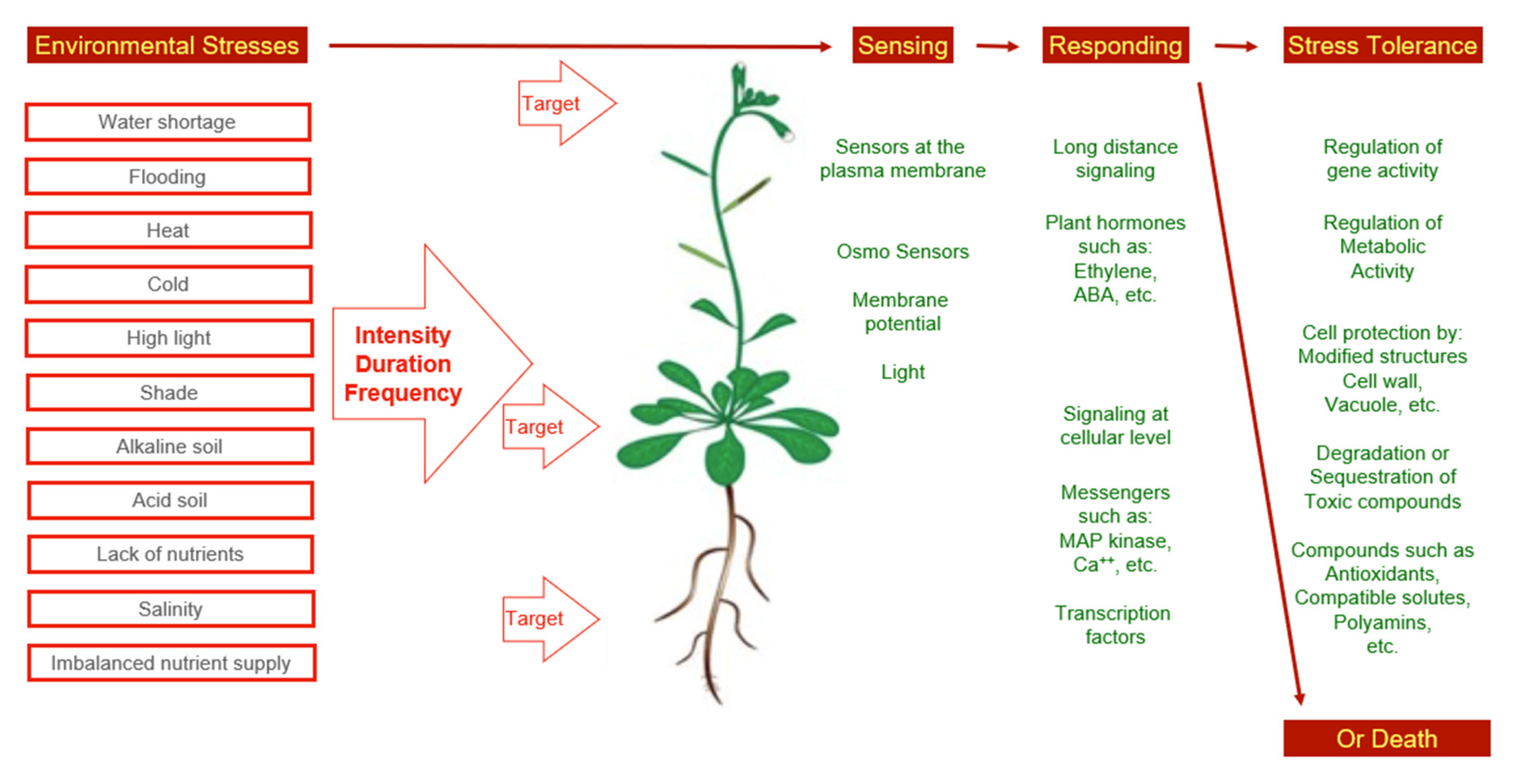 Figure 1. Environmental stress and observed concepts of stress adaptation.
Figure 1. Environmental stress and observed concepts of stress adaptation.From the description of ecosystems, it was evident that plant species differ to a large degree with respect to environmental preferences. In addition, this applies to the stress resistance level of plants. For instance, a puzzling observation was that subsequent to salinization a new plant population with a different frequency of species occupied abandoned areas. These highly salt-resistant species are referred to as halophytes [26][27][28]. Respective plant species turned into experimental plants in many research teams. The idea was to compare sensitive and resistant plant species to better understand concepts of stress perception, signaling, and responding. This approach requires the standardization of methods to allow for a comparison of the results. This need was addressed in a handbook that provides standardized protocols for the measurement of plant features, which reflect the ecological strategies of species [29]. After 10 years, the handbook has been updated to provide protocols on traits and methods that had proven to be useful in laboratories as well as in field studies. Furthermore, the new handbook contains additional protocols for functional traits of organs apart from leaves [30].
The traits described in the two handbooks were selected to predict long lasting effects at the ecosystem level. In addition, they might be used for prediction of annual crop yield, although remote sensing methods are preferentially used for these aspects. Moreover, the described standardized methods can be used to monitor the success of stress resistance breeding. However, they do not provide direct information on potential breeding targets at the metabolic and molecular level. Therefore, an impressive number of publications on genome, proteome, and metabolome analyses of plant species and accessions that differ in their stress resistance are available in the current literature [31][32][33]. Regulation of gene expression during the phase of stress adaptation has been analyzed, and the importance of individual genes for expression of a certain resistance feature has been analyzed by targeted mutation of experimental plants [34][35][36].
This was very helpful in the research of stress response, in which significant differences could be detected in accessions of the same plant species when harvested from areas with different local climates, soil conditions or the availability of water [37]. However, even when using plants as different as Arabidopsis and maize, differences in chromatin modification have been found [38][39].
Notably, the term stress does not exclude positive implications. Moderate stimuli have a positive impact (eustress), whereas excessive stimuli have a negative impact (distress) on plant response. Moreover, the positive effect of extra compounds as different as sugars, amines, elevated CO2 concentration, hormones, H2O2, UV radiation or nanomaterial has been tested in a concentration-dependent manner [40][41][42][43][44][45]. Several researchers reported that a repetitive application of moderate stress (with or without the addition of compounds) can modify metabolism, fluidity of biomembranes, the content of compatible solutes and ROS scavenging antioxidants, and enzymes [46]. Furthermore, the parameter value of the stress resistance window of treated plants correlated with the content of these beneficial compounds. Based on these observations, it is a matter of ongoing discussion, whether improved plant performance under stress in the presence of symbionts relies on similar regulatory mechanisms. In the meantime, economic interest has dramatically increased by the fact that (i) metabolites of pharmaceutical interest were among the compounds overproduced under eustress, and (ii) green algae and higher plants responded similarly to the applied eustress [47].
2. Interaction of Plants and Microorganisms
2.1. General Observations
In the field, plants, fungi, and bacteria form a well-structured community of organisms [48][49][50][51]. The microbial community of plants is called phyto-microbiome [52]. Microorganisms living in the soil, the rhizobiome, are distinguished from endophytes living inside plants. As growth conditions vary to a large degree between individual plant organs, specific populations of endophytic microorganisms can be found in each one [53]. Moreover, with respect to future agricultural application, beneficial microorganisms and pathogens have to be considered separately [54].
Research was hampered by the fact that most microorganisms of the phyto-microbiome cannot be cultured in vitro [55][56]. Therefore, information on species numbers and controlling effects, which are exerted on the microbial community by an individual plant genotype as well as the respective developmental stage of plants, was gained only when metagenomic methods became available [57][58][59]. Beneficial effects exerted on plant performance by rhizobia and mycorrhiza were observed early, and turned into a target of investigations [60][61] (Figure 2a). While it was possible to observe rhizobia-induced nodulation and the anatomy of mycorrhiza in the microscope, information on the interaction between plants and their microbial partners were mainly of indirect type. However, it was found that inoculation of low-fertility soil with plant growth promoting bacteria (PGPB) results in an increased production of biomass, an improved stress resistance, and particularly a reduced sensitivity to incidents of drought stress [45][62][63].
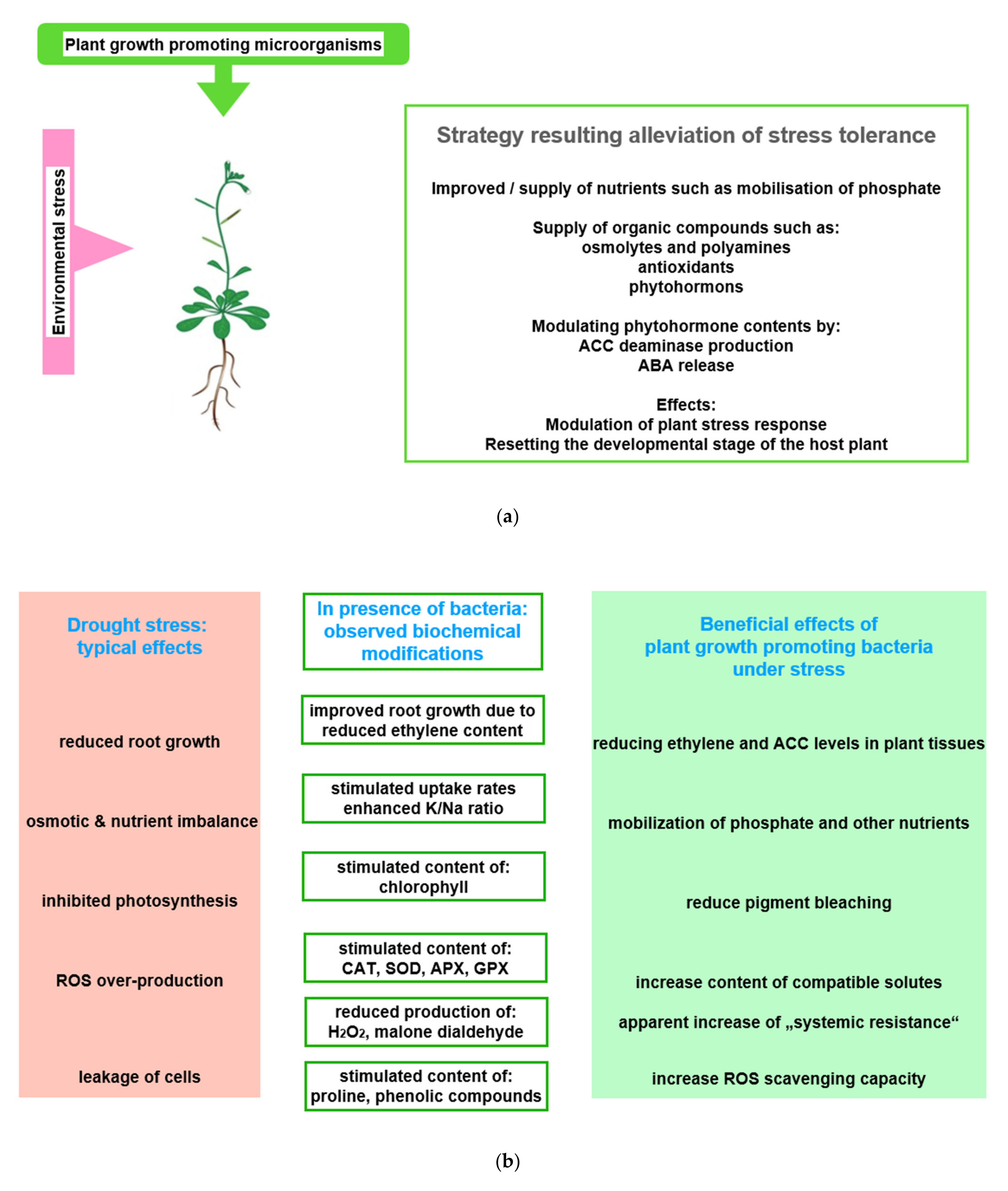
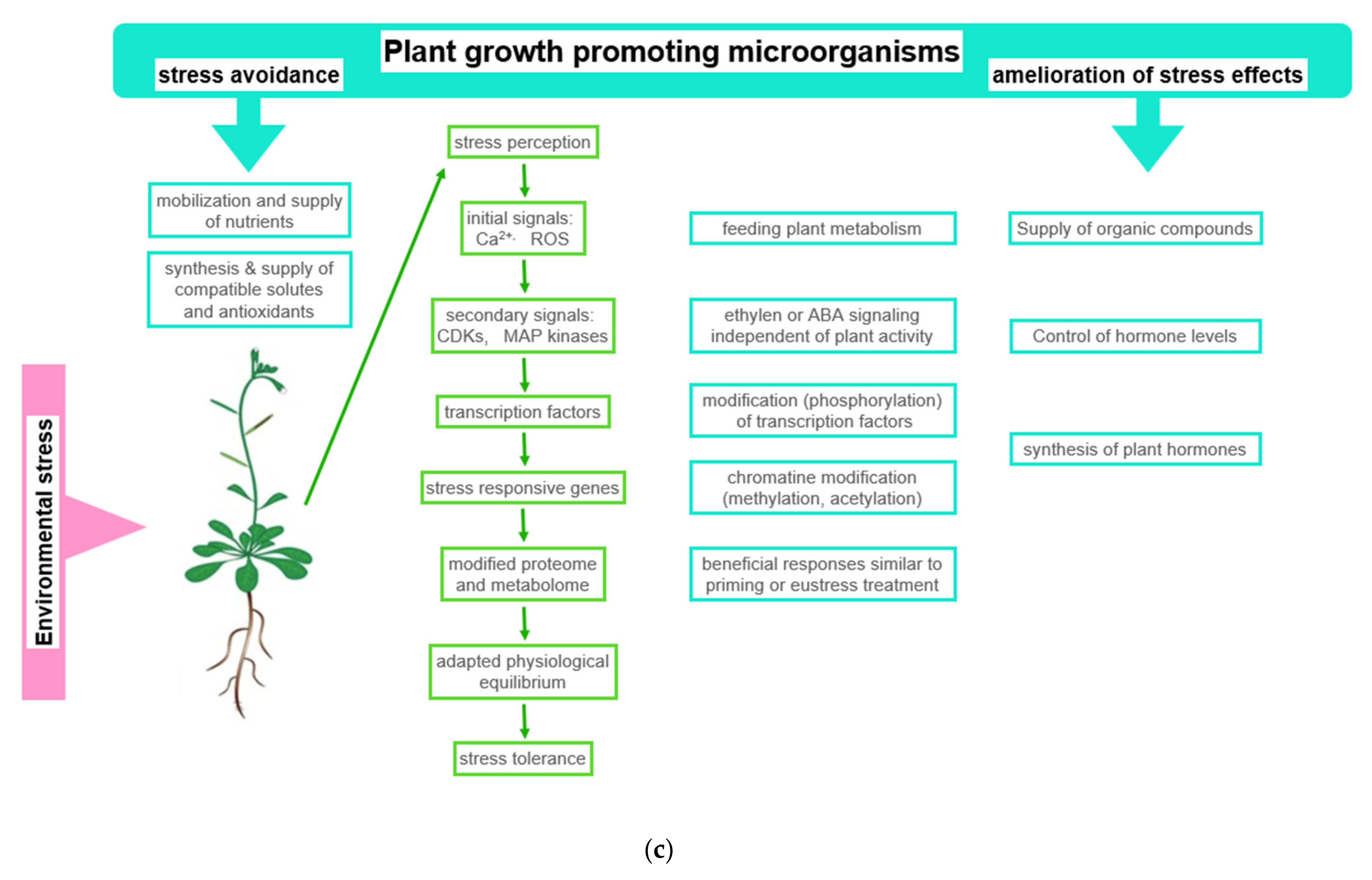 Figure 2. Beneficial effects of plant growth promoting microorganisms (PGPMs). (a) Observed stress ameliorating effects of PGPMs: The figure symbolizes the service offered by PGPMs, which helps in hosting plants to improve stress tolerance; (b) physiological basis of observed effects: Using drought stress as an example, typical stress effects are listed in the first column. Column two shows how PGPMs modify the biochemical stress response at biochemical level. In column three, the stress responses ameliorated by PGPMs are listed; (c) how PGPM activities interfere with the development of plant stress response: The boxes symbolize how PGPM activity may integrate into the reaction sequence, leading to plant stress tolerance. Abbreviations: APX: Ascorbate peroxidase; CAT: Catalase; GPX: Glutathione peroxidase; SOD: Superoxide dismutase.
Figure 2. Beneficial effects of plant growth promoting microorganisms (PGPMs). (a) Observed stress ameliorating effects of PGPMs: The figure symbolizes the service offered by PGPMs, which helps in hosting plants to improve stress tolerance; (b) physiological basis of observed effects: Using drought stress as an example, typical stress effects are listed in the first column. Column two shows how PGPMs modify the biochemical stress response at biochemical level. In column three, the stress responses ameliorated by PGPMs are listed; (c) how PGPM activities interfere with the development of plant stress response: The boxes symbolize how PGPM activity may integrate into the reaction sequence, leading to plant stress tolerance. Abbreviations: APX: Ascorbate peroxidase; CAT: Catalase; GPX: Glutathione peroxidase; SOD: Superoxide dismutase.In this context, for instance, the degree of beneficial effects was observed to be correlated with the improvement of nitrogen use [64][65]. Recently, in projects that rank the success of inoculation with PGPB, the concentration of intermediates of the TCA cycle, the Calvin–Benson cycle, and photorespiration, which easily could be measured in a laboratory, proved to be useful parameters [145]. The concentration of these compounds and antioxidants significantly increased in response to osmotic stress [66]. The modulation of these metabolite concentrations, among others, and the alteration of beneficial enzyme activities were interpreted as possible reasons for improved plant growth (Figure 2b). Among the more visible symptoms are enhanced seed germination, an expanded and elongated root system, and an increased chlorophyll content. All of the listed parameters are indicators of plant species-specific mechanism, which relieve the adverse effects of abiotic stress [67][68].
However, data concerning the TCA cycle can sometimes be misinterpreted and misleading. For instance, antioxidants will be degraded under prolonged and intensive stress. However, when monitored over time, the concentration will show a maximum and this maximum will be shifted in the presence of PGPB [69]. While overproduction of these compounds is preferentially achieved by stimulation of plant metabolism, it is assumed that increased concentrations of metabolites of the indol pathway are based on the import of precursors delivered by PGPB. This applies for shikimic, quinic, and salicylic acids [70]. The latter compound functions as a messenger in the hosting plant.
2.2. Acquisition of Symbionts
When a plant faces unfavorable conditions, it re-shapes physiological and biochemical parameters, allowing for a modified plant–microbe and microbe–microbe interactions [71][72]. Plants adapt to the patterns and abundance of suitable species of the rhizo-microbiome by the release of various metabolites, such as signaling molecules as well as organic C and N sources for microorganisms to feed on [50][52][71][73][74][75][76][77]. The composition of compounds released by the plants can vary with both, the growth conditions and the developmental stage of an individual plant species [78]. The amount of released carbohydrates may resemble 10% of the assimilate production [79]. This explains why the plant rhizosphere contains a significantly higher number of microorganisms as compared with the same soil type in the absence of any plants [80]. The spectrum of microbial species attracted to the plant roots will vary accordingly [81][82][83]. Moreover, it has been observed that the service of the attracted rhizo-microbiome is fine-tuned to the needs of the hosting plant [84][85][86][87][88]. Correspondingly, a modified abundance of microbial species occurs in the root-free soil areas and in samples collected from the root surface, respectively.
In response to compounds released by the plant, microorganisms release signaling compounds and metabolites that are beneficial to plants [78]. This signaling interplay is coordinated between both partners during adaptation to changing environmental conditions [89][90] (Figure 2c). This symbiosis can lead to the segregation of other organisms. Some invasive plant species are capable of modifying the abundance of beneficial microorganisms in the soil in favor of organisms that preferentially support the invading species [88].
2.3. Two Examples of Improved Nutrient Supply Provided by Plant Growth Promoting Microorganisms
As previously described, triose phosphates are exported from the chloroplast exclusively in exchange for phosphate. Therefore, a lack of phosphate inhibits the supply of the plant with assimilates, and indirectly prevents the use of absorbed light energy in photosynthesis. Nitrate reduction is performed using reducing equivalents and ATP. The primary product of nitrate assimilation is glutamine. Lack of water and salinity inhibit growth and the need for assimilates. An excess of energy arises, manifested as an over-reduction of the electron transport chain. As a result, a good nitrate supply can relieve the system by discharging ATP and reducing equivalents for nitrate reduction. In fact, it has been found that PGPMs can improve the plant supply of phosphate and nitrate.
2.3.1. Mobilization of Plant-Inaccessible Phosphate
Nutrient supply and use efficiency of crops have received increased attention due to the disappearance of plant available mineral resources and raw material stocks. Therefore, inoculation experiments using plant growth promoting bacteria in combination with conditions for an efficient nutrient use and minimal nutrient losses, were a promising approach to ensure a sustainable nutrient supply. The expectation was an improved nutrient uptake of the plants and growth promoting effects [91][92][93]. The PGPM inoculation leads to changes in the root architecture. This effect was explained by the release of phytohormones, such as IAA to the bacteria hosting plants [94][95][96].
Fertilization with the macronutrient phosphorus is one of the most important strategies to improve crop yield. However, only monobasic (H2PO4−) and dibasic (HPO42−) phosphates are resorbed by plant roots (Perez-Montano et al., 2014 [xx]) and a significant portion of soil phosphorus is unavailable for plants [97][98]. Phosphate solubilization capability of soil microorganisms is an important feature for plant growth enhancement under moderate fertility conditions [97][99][100][101]. Based on their ability to secrete organic acids, acid phosphatase, and alkaline phosphatase, phosphate-solubilizing bacteria are able to lower the pH in the rhizosphere, and in this way, increase the soluble and plant available phosphate fraction [102][103][104]. Notably, as abovementioned, these phosphate-releasing bacteria promote plant growth by offering further compounds in addition to mobile phosphates.
2.3.2. Support of Nitrogen Uptake and Fixation
Few plant growth promoting bacteria species are able to improve nitrogen use efficiency. They are not only involved in N cycling in the rhizosphere, but can also enter plant tissues and modify plant anatomy [105]. Transfer to plant tissues includes incorporation into seeds, thus promoting the development of next generation seedlings [88]. As these bacteria lack the nifH gene, enhanced nitrogen assimilation will occur through mechanisms other than nitrogen fixation [106]. Their bacterial activity correlates with plant species-specific polyamine production [107]. Interestingly, it was observed that the patterns of accumulated polyamines, amino acids, and urea in an experimental plant was modified by the application of stresses, independent of the growth promoting bacteria species present in the test [108][109].
References
- Lichtenthaler, H.K. Vegetation stress: An introduction to the stress concept in plants. J. Plant Physiol. 1996, 148, 4–14.
- Fischer, K.S.; Fukai, S.; Kumar, A.; Leung, H.; Jongdee, B. Field phenotyping strategies and breeding for adaptation of rice to drought. Front. Physiol. 2012, 3, 282.
- Mickelbart, M.V.; Hasegawa, P.M.; Bailey-Serres, J. Genetic mechanisms of abiotic stress tolerance that translate to crop yield stability. Nat. Rev. Genet. 2015, 16, 237–251.
- Senapati, N.; Semenov, M.A. Large genetic yield potential and genetic yield gap estimated for wheat in europe. Glob. Food Secur. 2020, 24, 100340.
- Lobell, D.B.; Roberts, M.J.; Schlenker, W.; Braun, N.; Little, B.B.; Rejesus, R.M.; Hammer, G.L. Greater sensitivity to drought accompanies maize yield increase in the u.s. midwest. Science 2014, 344, 516–519.
- Tollenaar, M.; Lee, E.A. Yield potential, yield stability and stress tolerance in maize. Field Crops Res. 2002, 75, 161–169.
- Witcombe, J.R.; Hollington, P.A.; Howarth, C.J.; Reader, S.; Steele, K.A. Breeding for abiotic stresses for sustainable agriculture. Philos. Trans. R. Soc. B 2008, 363, 703–716.
- Zhang, H.; Li, Y.; Zhu, J.-K. Developing naturally stress-resistant crops for a sustainable agriculture. Nat. Plants 2018, 4, 989–996.
- Godoy, F.; Olivos-Hernández, K.; Stange, C.; Handford, M. Abiotic stress in crop species: Improving tolerance by applying plant metabolites. Plants 2021, 10, 186.
- Mondal, S.; Sallam, A.; Sehgal, D.; Sukumaran, S.; Farhad, M.d.; Krishnan, J.N.; Kumar, U.; Biswal, A. Advances in breeding for abiotic stress tolerance in wheat. In Genomic Designing for Abiotic Stress Resistant Cereal Crops; Kole, C., Ed.; Springer: Cham, Switzerland, 2021; Volume 2, pp. 71–103.
- Lebaudy, A.; Vavasseur, A.; Hosy, E.; Dreyer, I.; Leonhardt, N.; Thibaud, J.-B.; Very, A.-A.; Simonneau, T.; Sentenac, H. Plant adaptation to fluctuating environment and biomass production are strongly dependent on guard cell potassiu channels. Proc. Natl. Acad. Sci. USA 2008, 105, 5271–5276.
- Larcher, W. Physiological Plant Ecology: Ecophysiology and Stress Physiology of Functional Groups; Springer: Berlin/Heidelberg, Germany, 2003; ISBN 978-3-540-43516-7.
- Huang, J.; Rozelle, S. Environmental stress and grain yields in china. Am. J. Agric. Econ. 1995, 77, 853–864.
- Yadav, S.; Modi, P.; Dave, A.; Vijapura, A.; Patel, D.; Patel, M. Effect of abiotic stress on crops. In Sustainable Crop Production; Hasanuzzaman, M., Teixeira Filho, M.C.M., Masayuki Fujita, M., Rodrigues Nogueira, T.A., Eds.; IntechOpen: London, UK, 2020.
- Ashraf, M.; Wu, L. Breeding for salinity tolerance in plants. Crit. Rev. Plant Sci. 1994, 13, 17–42.
- Munns, R. Genes and salt tolerance: Bringing them together. New Phytol. 2005, 167, 645–663.
- Guerrier, G. Fluxes of Na+. K+ and Cl− and osmotic adjustment in Lycopersicon pimpinellifolium and L. esculentum during short- and long-term exposures to NaCl. Physiol. Plant. 1996, 97, 583–591.
- Yousfi, N.; Slama, I.; Ghnaya, T.; Savoure, A.; Abdelly, C. Effects of water deficit stress on growth, water relations and osmolyte accumulation in Medicago truncatula and M. laciniata populations. Comptes Rendus Biol. 2010, 333, 205–213.
- Slama, S.; Abdelly, C.; Bouchereau, A.; Flowers, T.; Savouré, A. Diversity, distribution and roles of osmoprotective compounds accumulated in halophytes under abiotic stress. Ann. Bot. 2015, 115, 433–447.
- Villagómez-Aranda, A.L.; Feregrino-Pérez, A.A.; García-Ortega, L.F.; González-Chavira, M.M.; Torres-Pacheco, I.; Guevara-González, R.G. Activating stress memory: Eustressors as potential tools for plant breeding. Plant Cell Rep. 2022.
- Miller, M.; Qingxin Song, Q.; Shi, X.; Juenger, T.E.; Chen, Z.J. Natural variation in timing of stress-responsive gene expression predicts heterosis in intraspecific hybrids of Arabidopsis. Nat. Commun. 2015, 6, 7453.
- Fahad, S.; Bajwa, A.A.; Nazir, U.; Anjum, S.A.; Farooq, A.; Zohaib, A.; Sadia, S.; Nasim, W.; Adkins, S.; Saud, S.; et al. Crop production under drought and heat stress: Plant responses and management options. Front. Plant Sci. 2017, 8, 1147.
- Zhao, C.; Zhang, H.; Song, C.; Zhu, J.-K.; Shabala, S. Mechanisms of plant responses and adaptation to soil alinity. Innovation 2020, 1, 100017.
- Geppert, C.; Boscutti, F.; La Bella, G.; De Marchi, V.; Corcos, D.; Filippi, A.; Marini, L. Contrasting response of native and non-native plants to disturbance and herbivory in mountain environments. Front. Plant Sci. 2021, 8, 1147.
- Smirnoff, N.; Cumbes, Q.J. Hydroxyl radical scavenging activity of compatible solutes. Phytochemistry 1989, 28, 1057–1060.
- Stocker, O. Das Halophytenproblem. Ergeb. Biol. 1928, 3, 265–353.
- Byrt, C.S.; Munns, R. Living with salinity. New Phytol. 2008, 179, 903–905.
- Huchzermeyer, B.; Flowers, T. Putting halophytes to work—Genetics, biochemistry and physiology. Funct. Plant Biol. 2013, 40, 5–8.
- Cornelissen, J.H.C.; Lavorel, S.; Garnier, E.; Diaz, S.; Buchmann, N.; Gurvich, D.E.; Reich, P.B.; ter Steege, H.; Morgan, H.D.; van der Heijden, M.G.A.; et al. A handbook of protocols for standardized and easy measurements of plant functional traits worldwide. Aust. J. Bot. 2003, 51, 335–380.
- Pérez-Harguindeguy, N.; Díaz, S.; Garnier, E.; Lavorel, S.; Poorter, H.; Jaureguiberry, P.; Bret-Harte, M.S.; Cornwell, W.K.; Craine, J.M.; Gurvich, D.E.; et al. New handbook for standardised measurement of plant functional traits worldwide. Aust. J. Bot. 2013, 61, 167–234.
- Ahmed, M.Z.; Shimazaki, T.; Gulzar, S.; Kikuchi, A.B.; Khan, M.A.; Koyro, H.-W.; Huchzermeyer, B.; Watanabe, K.N. The influence of genes regulating transmembrane transport of Na+ on the salt resistance of Aeluropus lagopoides. Funct. Plant Biol. 2013, 40, 860–871.
- Kosová, K.; Prášil, I.T.; Vítámvás, P. Protein contribution to plant salinity response and tolerance acquisition. Int. J. Mol. Sci. 2013, 14, 6757–6789.
- Koyro, H.-W.; Zörb, C.; Debez, A.; Huchzermeyer, B. The effect of hyper-osmotic salinity on protein pattern and enzyme activities of halophytes. Funct. Plant Biol. 2013, 40, 787–804.
- Bartels, D.; Dinakar, C. Balancing salinity stress responses in halophytes and non-halophytes: A comparison between Thellungiella and Arabidopsis thaliana. Funct. Plant Biol. 2013, 40, 819–831.
- Kranner, I.; Seal, C. Salt stress, signalling and redox control in seeds. Funct. Plant Biol. 2013, 40, 848–859.
- Ozgur, R.; Turkan, I.; Uzilday, B.; Sekmen, A.H. Endoplasmic reticulum stress triggers ROS signalling, changes the redox state, and regulates the antioxidant defence of Arabidopsis thaliana. J. Exp. Bot. 2014, 65, 1377–1390.
- Ben Amora, N.; Jiménez, A.; Megdiche, W.; Lundqvist, M.; Sevilla, F.; Abdelly, C. Response of antioxidant systems to NaCl stress in the halophyte Cakile maritima. Physiol. Plant. 2006, 126, 446–457.
- Lämke, J.; Bäuerle, I. Epigenetik and chromatin-based mechanisms in environmental stress adaptation and stress memory in plants. Genome Biol. 2017, 18, 124.
- Turgut-Kara, N.; Arikan, B.; Celik, H. Epigenetic memory and priming in plants. Genetica 2020, 148, 47–54.
- Farooq, M.; Wahid, A.; Kobayashi, N.; Fujita, D.; Basra, S.M.A. Plant drought stress: Effects, mechanisms and management. Agronomy 2009, 29, 185–212.
- Latef, A.A.H.A.; Kordrostami, M.; Zakir, A.; Zaki, H.; Saleh, O.M. Eustress with H2O2 facilitates plant growth by improving tolerance to salt stress in two wheat cultivars. Plants 2019, 8, 303.
- Amritha, M.S.; Sridharan, K.; Puthur, J.T.; Dhankher, O.P. Priming with nanoscale materials for boosting abiotic stress tolerance in crop plants. J. Agric. Food Chem. 2021, 69, 10017–10035.
- Barickman, T.C.; Adhikari, B.; Sehgal, A.; Walne, C.H.; Reddy, K.R.; Gao, W. Drought and elevated carbon dioxide impact the morphophysiological profile of basil (Ocimum basilicum L.). Crops 2021, 1, 118–128.
- Lataef, A.A.A.; Hasanuzzaman, M.; Tahjib-ul-Arif, M. Mitigation of salinity stress by exogenous application of cytokinin in faba bean (Vicia faba L.). Not. Bot. Horti Agrobot. 2021, 49, 12192.
- Zhang, X.; Li, C.; Tie, D.; Quan, J.; Yue, M.; Liu, X. Epigenetic memory and growth responses of the clonal plant Glechoma longituba to parental recurrent UV-B stress. Funct. Plant Biol. 2021, 48, 827–838.
- Zheng, Y.; Xia, Z.; Wu, J.; Ma, H. Effects of repeated drought stress on the physiological characteristics and lipid metabolism of Bombax ceiba L. during subsequent drought and heat stresses. BMC Plant Biol. 2021, 21, 467.
- Roach, T.; Stöggl, W.; Baur, T.; Kranner, I. Distress and eustress of reactive electrophiles and relevance to light stress acclimation via stimulation of thiol/disulphide-based redox defences. Free Radic. Biol. Med. 2018, 122, 65–73.
- Lundberg, D.S.; Lebeis, S.L.; Paredes, S.H.; Yourstone, S.; Gehring, J.; Malfatti, S.; Tremblay, J.; Engelbrektson, A.; Kunin, V.; del Rio, T.G.; et al. Defining the core Arabidopsis thaliana root microbiome. Nature 2012, 488, 86–90.
- Turner, T.R.; James, E.K.; Poole, P.S. The plant microbiome. Genome Biol. 2013, 14, 209.
- Chaparro, J.M.; Badri, D.V.; Vivanco, J.M. Rhizosphere microbiome assemblage is affected by plant development. ISME J. 2014, 8, 790–803.
- Bulgarelli, D.; Garrido-Oter, R.; Münch, P.C.; Weiman, A.; Dröge, J.; Pan, Y.; McHardy, A.C.; Schulze-Lefert, P. Structure and function of the bacterial root microbiota in wild and domesticated barley. Cell Host Microbe 2015, 17, 392–403.
- Smith, D.L.; Gravel, V.; Yergeau, E. Editorial: Signaling in the phytomicrobiome. Front. Plant Sci. 2017, 8, 611.
- Gouda, S.; Das, G.; Sen, S.K.; Shin, H.-S.; Patra, J.K. Endophytes: A treasure house of bioactive compounds of medicinal importance. Front. Microbiol. 2016, 7, 1538.
- Nadarajah, K.; Abdul Rahman, N.S.N. Plant–Microbe Interaction: Above ground to below ground, from the good to the bad. Int. J. Mol. Sci. 2021, 22, 10388.
- Epstein, S. General model of microbial uncultivability. In Uncultivated Microorganisms; Epstein, S., Ed.; Springer: Berlin/Heidelberg, Germany, 2009; pp. 131–160.
- Berdy, B.; Spoering, A.; Ling, L.; Epstein, S. In situ cultivation of previously uncultivable microorganisms using the ichip. Nat. Protoc. 2017, 12, 2232–2242.
- Hirsch, P.R.; Mauchline, T.H. Who’s who in plant root microbiome? Nat. Biotechnol. 2012, 30, 961–962.
- Delaplace, P.; Delory, B.M.; Baudson, C.; Mendaluk-Saunier de Cazenave, M.; Spaepen, S.; Varin, S.; Brostaux, Y.; du Jardin, P. Influence of rhizobacterial volatiles on the root system architecture and the production and allocation of biomass in the model grass Brachypodium distachyon (L.) P. Beauv. BMC Plant Biol. 2015, 15, 195.
- Backer, R.; Rokem, J.S.; Ilangumaran, G.; Lamont, J.; Praslickova, D.; Ricci, E.; Subramanian, S.; Smith, D.L. Plant growth-promoting rhizobacteria: Context, mechanisms of action, and roadmap to commercialization of biostimulants for sustainable agriculture. Front. Plant Sci. 2018, 9, 1473.
- Simard, S.W.; Durall, D.M. Mycorrhizal networks: A review of their extent, function, and importance. Can. J. Bot. 2004, 82, 1140–1165.
- Gray, E.J.; Smith, D.L. Intracellular and extracellular PGPR: Commonalities and distinctions in the plant–bacterium signaling processes. Soil Biol. Biochem. 2005, 37, 395–412.
- Nelson, E.B. The seed microbiome: Origins, interactions, and impacts. Plant Soil 2017, 422, 7–34.
- Canellas, N.O.A.; Olivares, F.L.; Canellas, L.P. Metabolite fingerprints of maize and sugarcane seedlings: Searching for markers after inoculation with plant growth-promoting bacteria in humic acids. Chem. Biol. Technol. Agric. 2019, 6, 14.
- Canellas, L.P.; Silva, S.F.; Olk, D.; Olivares, F.L. Foliar application of Herbaspirillum seropedicae and humic acid increase maize yields. J. Food Agric. Environ. 2015, 13, 146–153.
- Olivares, F.L.; Busato, J.G.; Paula, A.M.; Lima, L.S.; Aguiar, N.O.; Canellas, L.P. Plant growth promoting bacteria and humic substances: Crop promotion and mechanisms of action. Chem. Biol. Technol. Agric. 2017, 4, 30.
- Koyro, H.-W.; Geissler, N.; Seenivasan, R.; Huchzermeyer, B. Plant stress physiology: Physiological and biochemical strategies allowing plants/crops to thrive under ionic stress. In Handbook of Plant and Crop Stress, 3rd ed.; Pessarakli, M., Ed.; CRC Press: Boca Raton, FL, USA, 2011; Volume 42, pp. 1051–1093.
- Nelson, L.M. Plant growth promoting rhizobacteria (PGPR): Prospects for new inoculants. Crop Manag. 2004, 3, 301–305.
- Sabki, M.H.; Ong, P.Y.; Ibrahim, N.; Lee, C.T.; Klemeš, J.J.; Li, C.; Gao, Y. A Review on abiotic stress tolerance and plant growth metabolite framework by plant growth-promoting bacteria for sustainable agriculture. Chem. Eng. Trans. 2021, 83, 367–372.
- Foyer, C.H.; Trebst, A.; Noctor, G. Protective and signaling functions of ascorbate, glutathione and tocopherol in chloroplasts. In Advances in Photosynthesis and Respiration: Photoprotection, Photoinhibition, Gene Regulation, and Environment; Demmig-Adams, B., Adams, W.W., Eds.; Springer Science Publishers: Dordrecht, The Netherlands, 2005; pp. 241–268.
- Dempsey, D.A.; Klessig, D.F. How does the multifaceted plant hormone sali-cylic acid combat disease in plants and are similar mechanisms utilized in humans? BMC Biol. 2017, 15, 23.
- Chaparro, J.M.; Dayakar, V.; Badri, D.V.; Matthew, G.; Bakker, M.G.; Akifumi Sugiyama, A.; Daniel, K.; Manter, D.K.; Jorge, M.; Vivanco, J.M. Root exudation of phytochemicals in arabidopsis follows specific patterns that are developmentally programmed and correlate with soil microbial functions. PLoS ONE 2013, 8, 1371.
- Uroz, S.; Courty, P.E.; Oger, P. Plant symbionts are engineers of the plant-associated microbiome. Trends Plant Sci. 2019, 24, 905–916.
- Liu, H.; Brettel, L.E.; Qiu, Z.; Sing, B.K. Microbiome-Mediated Stress Resistance in Plants. Trends Plant Sci. 2020, 25, 733–743.
- Cueva-Yesquén, L.G.; Goulart, M.C.; Attili de Angelis, D.; Nopper Alves, M.; Fantinatti-Garboggini, F. Multiple plant growth-promotion traits in endophytic bacteria retrieved in the vegetative stage from passionflower. Front. Plant Sci. 2021, 11, 621740.
- Chaparro, J.M.; Sheflin, A.M.; Manter, D.K.; Vivanco, J.M. Manipulating the soil microbiome to increase soil health and plant fertility. Biol. Fertil. Soils 2012, 48, 489–499.
- Nelson, M.S.; Sadowsky, M.J. Secretion systems and signal exchange between nitrogen-fixing rhizobia and legumes. Front. Plant Sci. 2015, 6, 491.
- Massalha, H.; Korenblum, E.; Malitsky, S.; Shapiro, O.H.; Aharon, A. Live imaging of root–bacteria interactions in a microfluidics setup. Proc. Natl. Acad. Sci. USA 2017, 114, 4549–4554.
- Nelson, E.B. Microbial dynamics and interactions in the spermosphere. Annu. Rev. Phytopathol. 2004, 42, 271–309.
- Pausch, J.; Kuzyakov, Y. Carbon input by roots into the soil: Quantification of rhizodeposition from root to ecosystem scale. Glob. Chang. Biol. 2018, 24, 1–12.
- Rovira, A.D. Interactions between plant roots and soil microorganisms. Annu. Rev. Microbiol. 1965, 19, 241–266.
- Lemanceau, P.; Corberand, T.; Gardan, L.; Latour, X.; Laguerre, G.; Boeufgras, J.-M.; Alabouvette, C. Effect of two plant species, flax (Linum usitatissnum L.) and tomato (Lycopersicon esculentum Mill.) on the diversity of soilborne populations of fluorescent Pseudomonas. Appl. Environ. Microbiol. 1995, 61, 1004–1012.
- Grayston, S.J.; Wang, S.; Campbell, C.D.; Edwards, A.C. Selective influence of plant species on microbial diversity in the rhizosphere. Soil Biol. Biochem. 1998, 30, 369–378.
- Miethling, R.; Wieland, G.; Backhaus, H. Variation of microbial rhizosphere communities in response to crop species, soil origin, and inoculation with Sinorhizobium meliloti L33. Microb. Ecol. 2000, 41, 43–56.
- Berg, G.; Smalla, K. Plant species and soil type cooperatively shape the structure and function of microbial communities in the rhizosphere. FEMS Microbiol. Ecol. 2009, 68, 1–13.
- Mendes, R.; Kruijt, M.; de Bruijn, I.; Dekkers, E.; van der Voort, M.; Schneider, J.H.M.; Piceno, Y.M.; DeSantis, T.Z.; Andersen, G.L.; Bakker, P.A.H.M.; et al. Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 2011, 332, 1097–1100.
- Berendsen, R.L.; Pieterse, C.M.J.; Bakker, P.A.H.M. The rhizosphere microbiome and plant health. Trends Plant Sci. 2012, 17, 478–486.
- Gaiero, J.R.; Mc Cali, C.M.; Thompson, K.A.; Day, N.J. Inside the root microbiome: Bacterial root endophytes and plant growth promotion. Am. J. Bot. 2013, 100, 1738–1750.
- Rout, M.E.; Southworth, A.S. The root microbiome influences scales from molecules to ecosystems: The unseen majority. Am. J. Bot. 2013, 100, 1689–1691.
- Leach, J.E.; Triplett, L.R.; Argueso, C.T.; Trivedi, P. Communication in the phytobiome. Cell 2017, 169, 587–596.
- Chauhan, H.; Bagyaraj, D.; Selvakumar, G.; Sundaram, S. Novel plant growth promoting rhizobacteria—Prospects and potential. Appl. Soil Ecol. 2015, 95, 38–53.
- Etesami, H.; Alikhani, H.A. Rhizosphere and endorhiza of oilseed rape (Brassica napus L.) plant harbor bacteria with multifaceted beneficial effects. Biol. Control 2016, 94, 11–24.
- Khalifa, A.Y.Z.; Alsyeeh, A.-M.; Almalki, M.A.; Saleh, F.A. Characterization of the plant growth promoting bacterium, Enterobacter cloacae MSR1, isolated from roots of non-nodulating Medicago sativa. Saudi J. Biol. Sci. 2016, 23, 79–86.
- Devi, K.A.; Pandey, G.; Rawat, A.K.S.; Sharma, G.D.; Pandey, P. The endophytic symbiont—Pseudomonas aeruginosa stimulates the antioxidant activity and growth of Achyranthes aspera L. Front. Microbiol. 2017, 8, 1897.
- Aloni, R.; Aloni, E.; Langhans, M.; Ullrich, C.I. Role of cytokinin and auxin in shaping root architecture: Regulating vascular differentiation, lateral root initiation, root apical dominance and root gravitropism. Ann. Bot. 2006, 97, 883–893.
- Vacheron, J.; Desbrosses, G.; Bouffaud, M.L.; Touraine, B.; Moenne-Loccoz, Y.; Muller, D.; Legendre, L.; Wisniewski-Dyè, F.; Prigent-Combaret, C. Plant growth-promoting rhizobacteria and root system functioning. Front. Plant Sci. 2013, 4, 19.
- Mahmood, A.; Kataoka, R. Metabolite profiling reveals a complex response of plants to application of plant growth-promoting endophytic bacteria. Microbiol. Res. 2020, 234, 126421.
- Zaidi, A.; Khan, M.S.; Ahemad, M.; Oves, M. Plant growth promotion by phosphate solubilizing bacteria. Acta Microbiol. Immunol. Hung. 2009, 56, 263–284.
- Campos, P.; Borie, F.; Cornejo, P.; Ráez, J.A.L.; López-Garciá, A.; Seguel, A. Phosphorus acquisition efficiency related to root traits: Is mycorrhizal symbiosis a key factor to wheat and barley cropping? Front. Plant Sci. 2018, 9, 752.
- Rodriguez, H.; Fraga, R. Phosphate solubilizing bacteria and their role in plant growth promotion. Biotechnol. Adv. 1999, 17, 319–339.
- Khan, M.S.; Zaidi, A.; Wani, P.A. Role of phosphate-solubilizing microorganisms in sustainable agriculture—A review. Agron. Sustain. Dev. 2007, 27, 29–43.
- Li, Y.; Liu, X.; Hao, T.; Chen, S. Colonization and maize growth promotion induced by phosphate solubilizing bacterial isolates. Int. J. Mol. Sci. 2017, 18, 1253.
- Goswami, D.; Dhandhukia, P.; Patel, P.; Thakker, J.N. Screening of PGPR from saline desert of Kutch: Growth promotion in Arachis hypogea by Bacillus licheniformis A2. Microbiol. Res. 2014, 169, 66–75.
- Oteino, N.; Lally, R.D.; Kiwanuka, S.; Lloyd, A.; Ryan, D.; Germaine, K.J.; Dowling, D.N. Plant growth promotion induced by phosphate solubilizing endophytic Pseudomonas isolates. Front. Microbiol. 2015, 6, 745.
- Chen, Q.; Liu, S. Identification and characterization of the phosphate-solubilizing bacterium Pantoea sp. S32 in Reclamation Soil in Shanxi, China. Front. Microbiol. 2019, 10, 21.
- Williams, S.T.; Vail, S.; Arcand, M.M. Nitrogen use efficiency in parent vs. hybrid canola under varying nitrogen availabilities. Plants 2021, 10, 2364.
- Paungfoo-Lonhienne, C.; Lonhienne, T.G.A.; Yeoh, Y.K.; Donose, B.C.; Webb, R.I.; Parsons, J.; Liao, W.; Sagulenko, E.; Lakshmanan, P.; Hugenholtz, P.; et al. Crosstalk between sugarcane and a plant-growth promoting Burkholderia species. Sci. Rep. 2016, 6, 37389.
- Xie, S.-S.; Wu, H.-J.; Zang, H.-Y.; Wu, L.-M.; Zhu, Q.-Q.; Gao, X.-W. Plant growth promotion by spermidine-producing Bacillus subtilis OKB105. Mol. Plant-Microbe Interact. 2014, 27, 655–663.
- Huang, C.Y.; Roessner, U.; Eickmeier, I.; Genc, Y.; Callahan, D.L.; Shirley, N.; Langridge, P.; Bacic, A. Metabolite profiling reveals distinct changes in carbon and nitrogen metabolism in phosphate-deficient barley plants (Hordeum vulgare L.). Plant Cell Physiol. 2008, 49, 691–703.
- Takagi, H.; Ishiga, Y.; Watanabe, S.; Konishi, T.; Egusa, M.; Akiyoshi, N.; Matsuura, T.; Mori, I.C.; Hirayama, T.; Kaminaka, H.; et al. Allantoin, a stress-related purine metabolite, can activate jasmonate signaling in a MYC2-regulated and abscisic acid-dependent manner. J. Exp. Bot. 2016, 67, 2519–2532.
More
Information
Subjects:
Plant Sciences
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.5K
Entry Collection:
Environmental Sciences
Revisions:
2 times
(View History)
Update Date:
04 Jul 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





