Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Irina Kiseleva | -- | 1763 | 2022-06-30 09:16:05 | | | |
| 2 | Catherine Yang | Meta information modification | 1763 | 2022-06-30 09:50:33 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Kiseleva, I. WT Parent Virus for Effective LAIV. Encyclopedia. Available online: https://encyclopedia.pub/entry/24668 (accessed on 27 March 2026).
Kiseleva I. WT Parent Virus for Effective LAIV. Encyclopedia. Available at: https://encyclopedia.pub/entry/24668. Accessed March 27, 2026.
Kiseleva, Irina. "WT Parent Virus for Effective LAIV" Encyclopedia, https://encyclopedia.pub/entry/24668 (accessed March 27, 2026).
Kiseleva, I. (2022, June 30). WT Parent Virus for Effective LAIV. In Encyclopedia. https://encyclopedia.pub/entry/24668
Kiseleva, Irina. "WT Parent Virus for Effective LAIV." Encyclopedia. Web. 30 June, 2022.
Copy Citation
Current influenza vaccine candidates, for potential use in vaccine manufacturing, are reassortants of master donor virus (MDV) with wild-type (WT) virus that is antigenically similar to the recommended strain. MDVs have all the necessary characteristics for the type of vaccines of which they are intended. Two types of MDVs are used in the preparation of influenza vaccines—high-yielding donors for IIV and temperature-sensitive (ts) and cold-adapted (ca) donors of attenuation—for LAIV. There are a number of main features of WT influenza virus that may dramatically affect different aspects of the preparation of egg-derived live attenuated vaccine candidates and their effectiveness.
influenza vaccine
classical reassortment
reassortant
1. Naturally Occurring Temperature-Sensitive WT Influenza Viruses
An important characteristic of any virus is its non-ts phenotype—an ability for replication at the elevated temperatures of 38–40 °C, which exceed the upper limit of optimal values. In the past, it was thought that the typical WT virus is always non-ts and that this property determines viral virulence. In those times, primary screening of reassortant LAIV candidates was based on ts/ca attenuation markers [1][2]. Reassortants that did not contain suitable laboratory markers of attenuation (ts/ca phenotype) were screened out; only then, analyses of the genome composition, which at that time was quite complex, started. The ts-phenotype of the reassortant LAIV candidate is critical because ts viruses cannot multiply at the temperature of the lower respiratory tract.
The first mention of natural ts WT influenza viruses can be found in publications from the 1980s [3][4][5][6][7]. Later, it was found that changes in the ts/non-ts phenotype have a regular wave-like nature [8][9][10]. At the beginning of each influenza pandemic/epidemic cycle, the circulation of non-ts viruses was detected. Further evolution is leading to the change in non-ts with ts variants. The prevalence of ts strains in circulation indirectly indicates that novel non-ts viruses are expected to appear in circulation.
Thus, the permanent circulation of ts viruses can be considered as a precursor for the appearance of an antigenically distinct virus, seasonal or even pandemic. This assumption is supported by the fact that just before the 2009 pandemic, a kind of “calm before the storm” was noticed: only ts influenza A(H1N1), A(H3N2), and B viruses were detected in circulation [8].
The ts phenotype of the WT virus does not influence the efficiency of reassortment but interferes with the efficacy of primary screening of the egg-derived LAIV candidate, since one of the laboratory selective markers of attenuation, the ts marker, is lost. The problem of the existence of ts viruses lies elsewhere—since ts viruses are at the end of the ts wave of circulating viruses, they may be less immunogenic than their non-ts counterparts, whose circulation started this wave. It has been suggested that non-ts WT parent viruses may enhance the immunogenicity of LAIV and vice versa; LAIVs based on ts WT parent viruses were of low immunogenicity. Unfortunately, this study only tested a limited number of vaccines [10].
It seems reasonable that new potential candidates for vaccine strains should be evaluated not only in terms of the novelty of surface antigens but also taking into account temperature sensitivity in their replication.
2. Naturally Occurring Cold-Adapted WT Influenza Viruses
In nature, not only natural temperature-sensitive viruses circulate, which are numerous, but sometimes ca viruses also appear, which usually possess the non-ts phenotype [11]. Unlike natural ts viruses, there are so few that it is not possible to make any assumptions about the reasons and regularities in their appearance. In fact, the term “ca” (cold-adapted) is not quite appropriate in this case, since these viruses were not adapted to low temperatures by laboratory manipulations. It would be more accurate to talk about WT viruses that sufficiently replicate at low temperatures or WT viruses that are naturally resistant to low temperatures. The role of cold-resistance for replication of some natural isolates has not been studied yet; however, it can be assumed that non-ts/ca WT viruses, which can reproduce in a very wide temperature range, from 25 °C to 40 °C, can effectively infect both the upper and lower respiratory tract.
Unlike ts WT viruses whose ts phenotype does not influence the efficiency of reassortment, the ca phenotype of the WT parent dramatically disturbs the first steps in the reassortment process that are carried out at a low temperature of 25–26 °C. A loss of the ca-selective factor may lead to a significant increase in the total number of ca reassortants, but the overall number of reassortants with the desired 6:2 genome composition is decreasing.
3. Sensitivity of WT Viruses to Nonspecific Thermostable Serum γ-Inhibitors
WT influenza viruses exhibit marked differences in their sensitivity to nonspecific thermostable γ-inhibitors due to their distinguishable receptor specificity. H3N2 viruses, which preferentially bind the α-2,6 receptors, are very sensitive to serum thermostable γ-inhibitors, while H1N1 strains with α-2,3 or mixed α-2,3/α-2,6 specificity exhibit an inhibitor-resistant phenotype [12][13][14]. Before the 1970s–1980s, the majority of influenza B viruses possessed an inhibitor-resistant phenotype. In the 1980s, they diverged into two distinct genetic lineages, B/Victoria and B/Yamagata [15]. Since then, there have been two parallel evolutionary pathways of influenza type B in the human population [16]. After separation, the B/Victoria lineage viruses retained a high level of inhibitor resistance of past strains. Contrarily, viruses of the B/Yamagata lineage acquired high inhibitor sensitivity [17].
The standard scheme for the preparation of vaccine strains by the method of classical reassortment includes the use of anti-MDV serum [18][19][20]. This provides a selective advantage for reassortants in inheriting HA and NA from an antigenically relevant WT virus. However, the selection of 6:2 reassortants, based on inhibitor-susceptible WT viruses, can be complicated by nonspecific binding of their HA by γ-inhibitors, which are presented in the anti-serum against MDV [14].
The data presented in [14] were used for drawing Figure 1. Analysis of genome composition of 883 reassortants, obtained by classical reassortment in eggs of MDVs with 40 WT influenza viruses, which possessed a different degree of sensitivity to nonspecific γ-inhibitors, revealed the following consistent pattern: all reassortants inherited WT HA; nevertheless, the belonging of the remaining genes to WT or MDV parents varied [14]. The majority of reassortants based on inhibitor-resistant WT viruses (88.7%) inherited NA from the WT parent; also, the highest percentage of 6:2 reassortants (31.4%) was achieved (Figure 1, left panel).
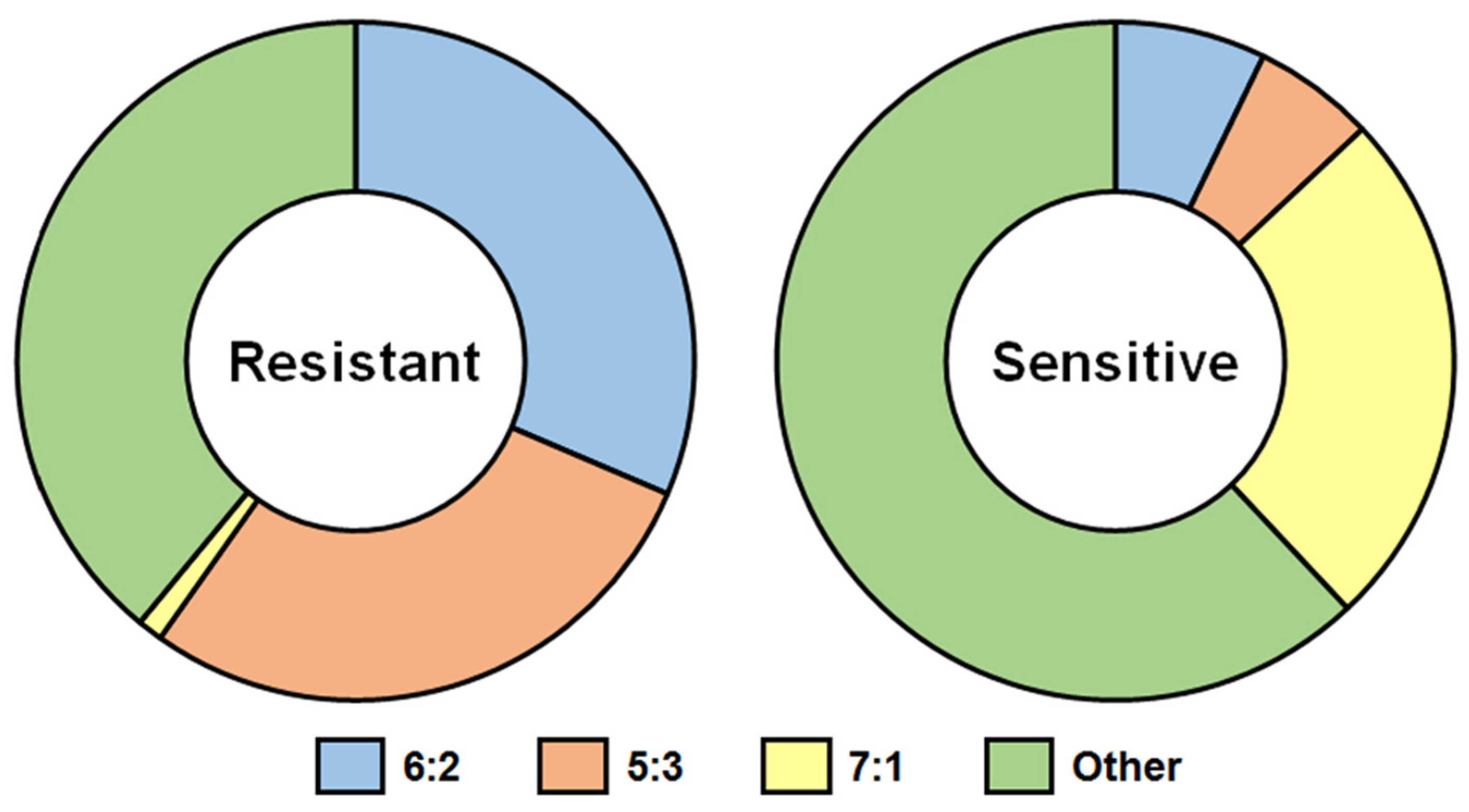
Figure 1. Genome composition (%) of reassortants derived by classical reassortment of ca MDVs with resistant or sensitive to nonspecific thermostable γ-inhibitors A(H1N1), A(H2N2), A(H3N2), A(H5N1), B/Victoria lineage, and B/Yamagata lineage WT influenza viruses [14]). Left panel—328 reassortants of ca MDVs with 20 WT viruses that are resistant to nonspecific thermostable γ-inhibitors were analyzed; 555 reassortants of ca MDVs with 20 WT viruses that are sensitive to nonspecific thermostable γ-inhibitors were analyzed. 6:2 genome composition—HA and NA are inherited from the WT parent, and 6 internal genes are inherited from MDV; 5:3 genome composition—HA, NA and one of the internal genes are inherited from the WT parent, the other five internal genes are inherited from MDV; 7:1 genome composition—HA is inherited from WT parent, all internal genes and NA are inherited from MDV.
In contrast, the efficiency of obtaining 6:2 reassortants was much lower (7.2%) if the WT parent virus possessed a high degree of sensitivity to nonspecific thermostable γ-inhibitors; clones with the 7:1 genotype (25%) prevailed among the obtained reassortants. Corruption of the constellation of genes encoding HA and NA was observed—only a quarter of all reassortants inherited both WT HA and WT NA and three-quarters had WT HA + MDV NA [14]) (Figure 1, right panel).
Thus, the inhibitor sensitivity of WT viruses becomes an obstacle to the effective preparation of vaccine reassortants for LAIV by classical reassortment, since immune serum against MDV is involved in the selection process/screening of vaccine candidates. Contrarily, the inhibitor resistance guarantees a faster and more stable result in the preparation of vaccine strains. The development of LAIV based on the classical reassortment method would benefit from the recommendation of viruses with a high level of resistance to inhibitors. On the other hand, inhibitor-sensitive viruses retain a preference for α-2,6-linked residues. For now, the question, “what should be the best vaccine strain—inhibitor-sensitive or inhibitor-resistant?”, remains open.
4. Infectivity of WT Viruses and LAIV Candidates
One of the key indicators of the quality of reassortant candidates for IIV is their high HA titer. There has been up to a 512-fold increase in HA titers of PR8-based vaccine reassortant observed as compared to the respective WT parent virus [21]. Reassortants that produce high HA titers do not always have a high yield of infectious viruses [22] but infectious viral titers of reassortant candidates are not so critical for IIV. For example, reassortants prepared on a high-yielding PR8 donor, NIBRG-23 (H5N1) and VN/PR/CDC-RG (H5N1), displayed rather low infectious viral titers, which did not exceed 6.2 log10 EID50/mL and 7.7 log10 EID50/mL, correspondingly [23][24].
On the contrary, infectivity is critical for LAIV. Whereas the WT parent virus typically has relatively low infectious titers (6.2–7.7 log10 EID50/mL [24]), the titers of LAIV candidates on the backbone of ca MDV are usually 8.7–10.2 log10 EID50/mL [18][23][24].
As for the viruses to be recommended, typically, a reference strain and a few reference strain-like viruses that are similar in antigenic properties to the reference virus are recommended. Sometimes reference strain-like viruses appear to be less or more effective in the development of reassortant vaccine candidates than reference strains. For instance, based on our experience, the reassortant LAIV candidate of A/Leningrad/134/17/57 MDV with A/Brisbane/34/2018 (H3N2) WT parent (A/Kansas/14/2017-like virus recommended for use in 2019-2020 Northern Hemisphere influenza season) displayed ~ 1 log10 EID50/mL higher infectious activity than the reassortant candidate based on the A/Kansas/14/2017 (H3N2) reference strain, respectively. In contrast, the reassortant LAIV candidate of A/Leningrad/134/17/57 MDV with the A/Michigan/173/2020 (H3N2) WT parent (A/Darwin/9/2021-like virus recommended for use in 2022-2023 Northern Hemisphere influenza season) displayed ~ 1.0–1.5 lg10 EID50/mL lower infectious activity than the reassortant candidate based on the A/Darwin/9/2021 (H3N2) reference strain, respectively (I. Kiseleva, E. Bazhenova, E. Stepanova, N. Larionova and L. Rudenko. Personal communications).
Interestingly, the reassortment of ca MDV with PR8-based vaccine strains for IIV of relatively low infection titers led to a dramatic increase in infectivity of the resulting reassortants [23][24]. Unfortunately, there are cases when the presence of genes from an attenuated ca MDV does not significantly increase the infectious viral titers of reassortants. This has been observed in recent years for A(H3N2) influenza viruses and may be related to the receptor specificity of these viruses. If A(H1N1) and A(H1N1)pdm09 influenza viruses possess α-2,3 or α-2,3/α-2,6 specificity, due to which they multiply well in eggs without prior adaptation, then A(H3N2) influenza viruses retaining a preference for α-2,6 specificity [12] have always been a problem for reproduction in eggs, becoming even more serious recently. Sometimes, national influenza centers that conduct year-round surveillance for influenza were not able to isolate A(H3N2) viruses in eggs to be recommended for seasonal vaccines in a timely manner.
References
- Mills, J.; Chanock, V.; Chanock, R.M. Temperature-sensitive mutants of influenza virus. I. Behavior in tissue culture and in experimental animals. J. Infect. Dis. 1971, 123, 145–157.
- Maassab, H.F.; DeBorde, D.C. Development and characterization of cold-adapted viruses for use as live virus vaccines. Vaccine 1985, 3, 355–369.
- Oxford, J.S.; Corcoran, T.; Schild, G.C. Naturally occurring temperature-sensitive influenza A viruses of the H1N1 and H3N2 subtypes. J. Gen. Virol. 1980, 48, 383–389.
- Chu, C.M.; Tian, S.F.; Ren, G.F.; Zhang, Y.M.; Zhang, L.X.; Liu, G.Q. Occurrence of temperature-sensitive influenza A viruses in nature. J. Virol. 1982, 41, 353–359.
- Polezhaev, F.I.; Aleksandrova, G.I. Isolation of temperature-sensitive strains of the influenza virus in the epidemic caused by the A/Victoria virus in 1975–1976. Vopr. Virusol. 1979, 4, 430.
- Zhang, Y.M.; Tian, S.F.; Zhu, J.M. Identification of naturally occurring temperature-sensitive strains of influenza A virus and location of their genetic lesions. Sci. Sin. B 1982, 25, 411–419.
- Richman, D.D.; Murphy, B.R. The association of the temperature-sensitive phenotype with viral attenuation in animals and humans: Implications for the development and use of live virus vaccines. Rev. Infect. Dis. 1979, 1, 413–433.
- Kiseleva, I.; Larionova, N. (Eds.) Influenza virus ecology and evolution. In Influenza: A Century of Research; Bentham Science Publisher: Sharjah, United Arab Emirates, 2021; pp. 63–97.
- Larionova, N.V.; Kiseleva, I.V.; Rudenko, L.G. Evolution of influenza viruses based on sensitivity to temperature of replication. J. Microbiol. Epidemiol. Immunobiol. 2019, 6, 47–55.
- Rudenko, L.G.; Kiseleva, I.V.; Larionova, N.V.; Grigorieva, E.P.; Naikhin, A.N. Analysis of some factors influencing immunogenicity of live cold–adapted reassortant influenza vaccines. In Proceedings of the Options for the Control of Influenza V, Okinawa, Japan, 6–9 October 2003; pp. 542–546.
- Kiseleva, I.V.; Voeten, J.T.; Teley, L.C.; Larionova, N.V.; Drieszen-van der Cruijsen, S.K.; Basten, S.M.; Heldens, J.G.; van den Bosch, H.; Rudenko, L.G. PB2 and PA genes control the expression of the temperature-sensitive phenotype of cold-adapted B/USSR/60/69 influenza master donor virus. J. Gen. Virol. 2010, 91, 931–937.
- Rogers, G.N.; D’Souza, B.L. Receptor binding properties of human and animal H1 influenza virus isolates. Virology 1989, 173, 317–322.
- Rogers, G.N.; Pritchett, T.J.; Lane, J.L.; Paulson, J.C. Differential sensitivity of human, avian, and equine influenza A viruses to a glycoprotein inhibitor of infection: Selection of receptor specific variants. Virology 1983, 131, 394–408.
- Kiseleva, I.; Larionova, N.; Fedorova, E.; Bazhenova, E.; Dubrovina, I.; Isakova-Sivak, I.; Rudenko, L. Contribution of neuraminidase of influenza viruses to the sensitivity to sera inhibitors and reassortment efficiency. Open Microbiol. J. 2014, 8, 59–70.
- Yang, J.R.; Huang, Y.P.; Chang, F.Y.; Hsu, L.C.; Lin, Y.C.; Huang, H.Y.; Wu, F.T.; Wu, H.S.; Liu, M.T. Phylogenetic and evolutionary history of influenza B viruses, which caused a large epidemic in 2011–2012, Taiwan. PLoS ONE 2012, 7, e47179.
- Rota, P.A.; Wallis, T.R.; Harmon, M.W.; Rota, J.S.; Kendal, A.P.; Nerome, K. Cocirculation of two distinct evolutionary lineages of influenza type B virus since 1983. Virology 1990, 175, 59–68.
- Larionova, N.; Kiseleva, I.; Isakova, I.; Litvinova, O.; Rudenko, L. Naturally occuring temperature-sensitive strains of influenza B virus. Vopr. Virusol. 2006, 51, 38–41.
- Shcherbik, S.; Pearce, N.; Kiseleva, I.; Larionova, N.; Rudenko, L.; Xu, X.; Wentworth, D.E.; Bousse, T. Implementation of new approaches for generating conventional reassortants for live attenuated influenza vaccine based on Russian master donor viruses. J. Virol. Methods 2016, 227, 33–39.
- Wareing, M.D.; Marsh, G.A.; Tannock, G.A. Preparation and characterisation of attenuated cold-adapted influenza A reassortants derived from the A/Leningrad/134/17/57 donor strain. Vaccine 2002, 20, 2082–2090.
- Shcherbik, S.V.; Pearce, N.C.; Levine, M.L.; Klimov, A.I.; Villanueva, J.M.; Bousse, T.L. Rapid strategy for screening by pyrosequencing of influenza virus reassortants-candidates for live attenuated vaccines. PLoS ONE 2014, 9, e92580.
- Fulvini, A.A.; Ramanunninair, M.; Le, J.; Pokorny, B.A.; Arroyo, J.M.; Silverman, J.; Devis, R.; Bucher, D. Gene constellation of influenza A virus reassortants with high growth phenotype prepared as seed candidates for vaccine production. PLoS ONE 2011, 6, e20823.
- Trifkovic, S.; Gilbertson, B.; Fairmaid, E.; Cobbin, J.; Rockman, S.; Brown, L.E. Gene segment interactions can drive the emergence of dominant yet suboptimal gene constellations during influenza virus reassortment. Front. Microbiol. 2021, 12, 683152.
- Kiseleva, I.V.; Larionova, N.V.; Fedorova, E.A.; Isakova-Sivak, I.N.; Rudenko, L.G. New methodological approaches in the development of Russian live attenuated vaccine for pandemic influenza. Transl. Biomed. 2015, 6, 1–9.
- Larionova, N.; Kiseleva, I.; Isakova-Sivak, I.; Rekstin, A.; Dubrovina, I.; Bazhenova, E.; Ross, T.M.; Swayne, D.; Gubareva, L.; Tsvetnitsky, V.; et al. Live attenuated influenza vaccines against highly pathogenic H5N1 avian influenza: Development and preclinical characterization. J. Vaccines Vaccin. 2013, 4, 208.
More
Information
Subjects:
Infectious Diseases
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
874
Entry Collection:
Biopharmaceuticals Technology
Revisions:
2 times
(View History)
Update Date:
30 Jun 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





