
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Francisco Javier Navas González | -- | 2884 | 2022-04-18 11:27:50 | | | |
| 2 | Peter Tang | Meta information modification | 2884 | 2022-04-18 11:40:41 | | | | |
| 3 | Peter Tang | Meta information modification | 2884 | 2022-04-18 11:41:11 | | |
Video Upload Options
Despite their pivotal position as relevant sources for high-quality proteins in particularly hard environmental contexts, the domestic goat has not benefited from the advances made in genomics compared to other livestock species. Genetic analysis based on the study of candidate genes is considered an appropriate approach to elucidate the physiological mechanisms involved in the regulation of the expression of functional traits. This is especially relevant when such functional traits are linked to economic interest. The knowledge of candidate genes, their location on the goat genetic map and the specific phenotypic outcomes that may arise due to the regulation of their expression act as a catalyzer for the efficiency and accuracy of goat-breeding policies, which in turn translates into a greater competitiveness and sustainable profit for goats worldwide.
1. Introduction

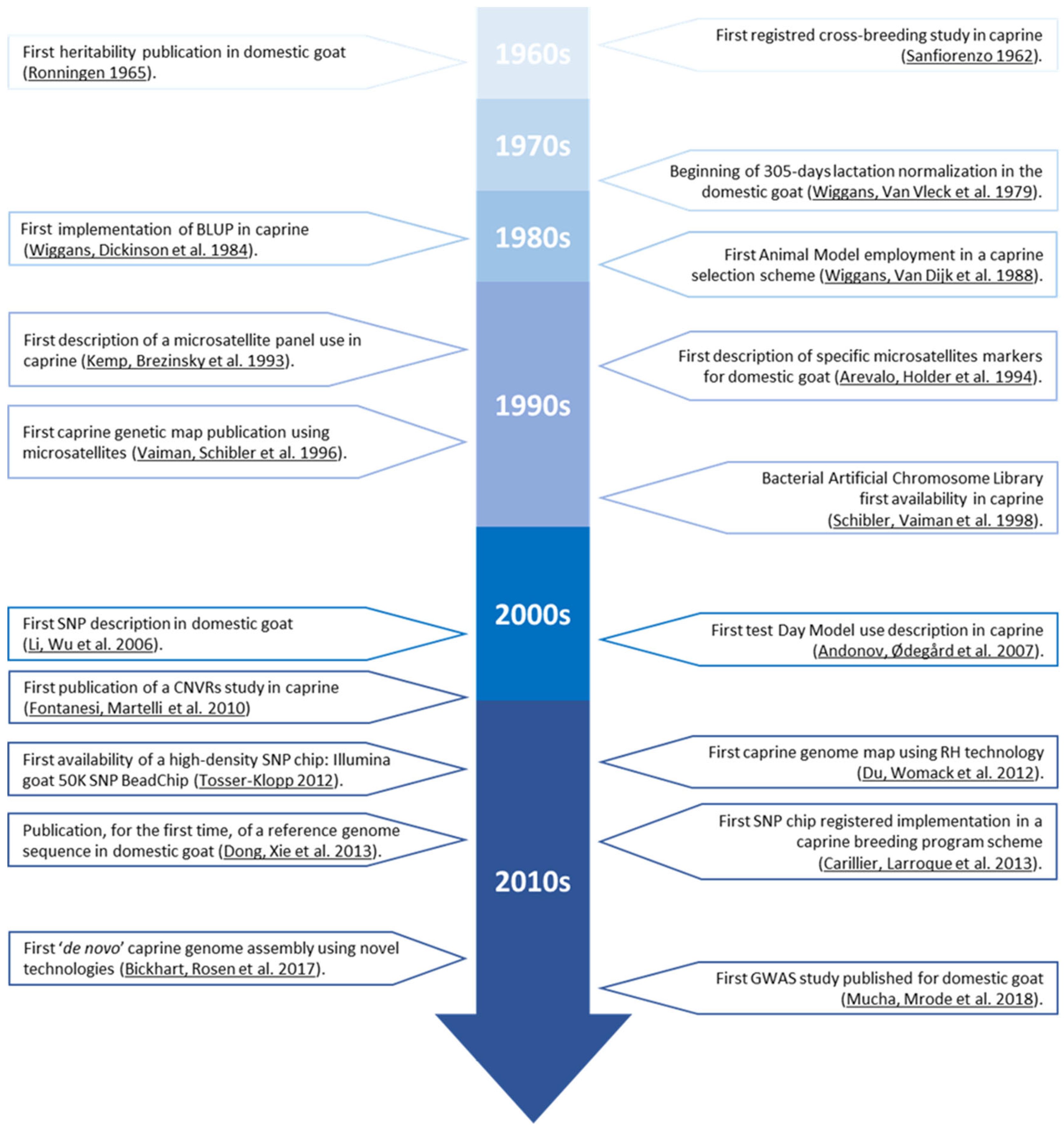
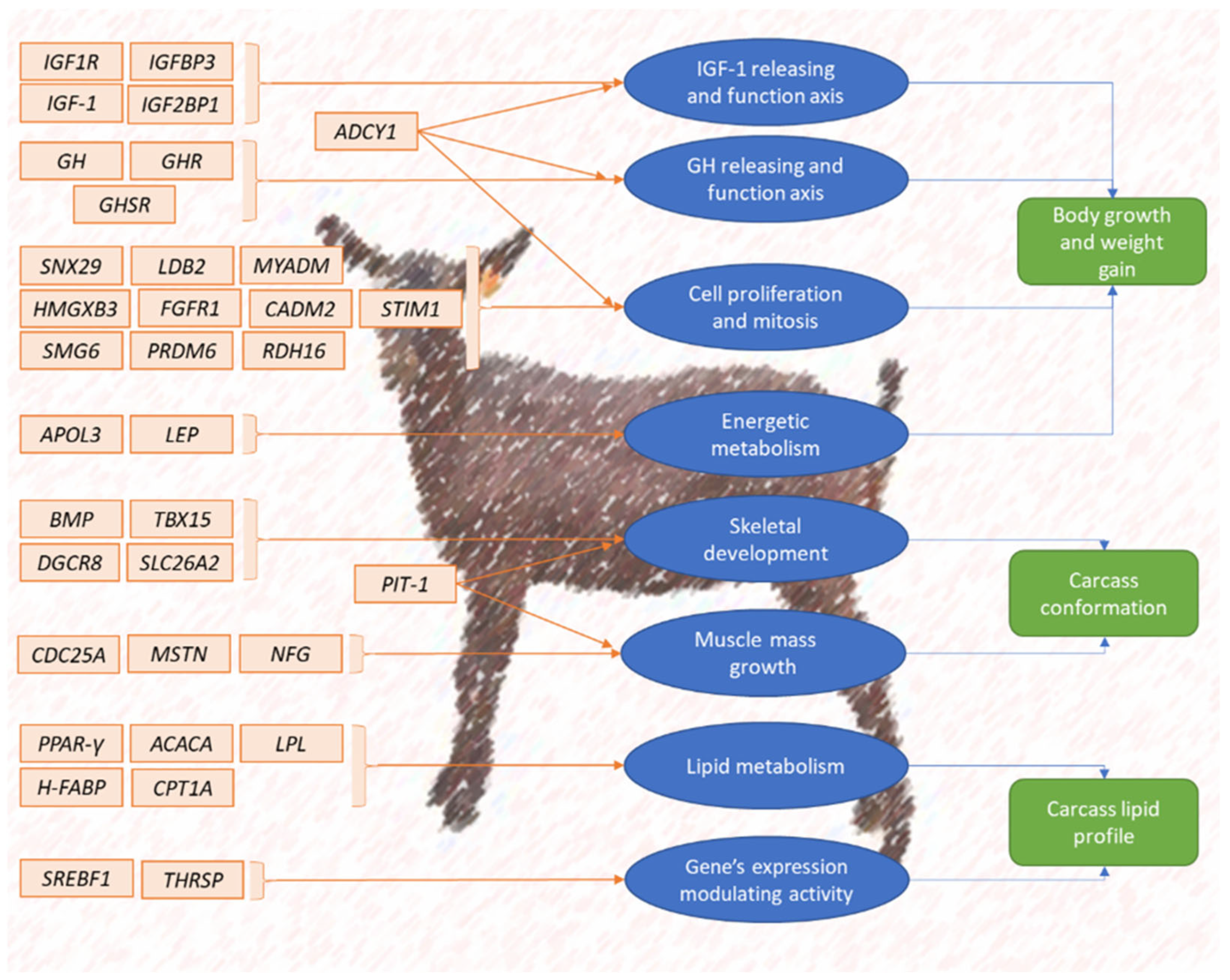
2. Candidate Genes Regulating the Expression of Economically Relevant Traits in Caprine
2.1. Candidate Genes Regulating the Expression of Goat Meat Breeding Criteria and Traits
|
Breeding Criteria |
Traits |
Reference |
|---|---|---|
|
Body weight |
Birth weight, body weight at 7, 14, 21 and 28 days, monthly weighing until 18 months of age |
|
|
Growth |
Average daily gain (ADG) before weaning, from 3 to 6 months of age, from 6 to 12 months of age |
[52] |
|
Conformation/corporal structure |
Body length, height at the withers, chest girth, shoulder width and pin-bone width |
[53] |
|
Carcass quality |
Hot (immediately after slaughter) and cold (before chilling) carcass weight; weights of the head, skin, heart, and thoracic and abdominal viscera; carcass dressing percentage; leg length and carcass width; and carcass fat coverage and lean meat and bone yield percentages |
|
|
Meat quality |
Muscle pH measurements, colour, water retention capacity, nutritional composition (protein, fat, and collagen percentages), and fatty acid profile in samples of longissimus thoracis et lumborum, triceps brachii and semimembranosus |
[51] |
2.2. Candidate Genes Regulating the Expression of Goat Dairy Breeding Criteria and Traits
References
- Miller, B.A.; Lu, C.D. Current status of global dairy goat production: An overview. Asian Australas. J. Anim. Sci. 2019, 32, 1219.
- Gama, L.; Bressan, M. Biotechnology applications for the sustainable management of goat genetic resources. Small Rumin. Res. 2011, 98, 133–146.
- Amills, M.; Capote, J.; Tosser-Klopp, G. Goat domestication and breeding: A jigsaw of historical, biological and molecular data with missing pieces. Anim. Genet. 2017, 48, 631–644.
- Fahmy, M.H.; Shrestha, J. Genetics for the improvement of goat meat production. In Proceedings of the 7th International Conference on Goats, Tours, France, 15–21 May 2000; pp. 187–190.
- Shrestha, J. Genetics and breeding of meat goats. In Goat Meat Production and Quality; Mahgoub, O., Kadim, I., Webb, E., Eds.; CAB International: London, UK, 2012; p. 52.
- Mazhangara, I.R.; Chivandi, E.; Mupangwa, J.F.; Muchenje, V. The potential of goat meat in the red meat industry. Sustainability 2019, 11, 3671.
- Daly, K.G.; Delser, P.M.; Mullin, V.E.; Scheu, A.; Mattiangeli, V.; Teasdale, M.D.; Hare, A.J.; Burger, J.; Verdugo, M.P.; Collins, M.J. Ancient goat genomes reveal mosaic domestication in the Fertile Crescent. Science 2018, 361, 85–88.
- Pidancier, N.; Jordan, S.; Luikart, G.; Taberlet, P. Evolutionary history of the genus Capra (Mammalia, Artiodactyla): Discordance between mitochondrial DNA and Y-chromosome phylogenies. Mol. Phylogenet. Evol. 2006, 40, 739–749.
- Mirkena, T.; Duguma, G.; Haile, A.; Tibbo, M.; Okeyo, A.; Wurzinger, M.; Sölkner, J. Genetics of adaptation in domestic farm animals: A review. Livest. Sci. 2010, 132, 1–12.
- Gootwine, E. Genetics and breeding of sheep and goats. In Animal Agriculture; Elsevier: College Station, TX, USA, 2020; pp. 183–198.
- Ferrando, A.; Manunza, A.; Jordana, J.; Capote, J.; Pons, A.; Pais, J.; Delgado, T.; Atoche, P.; Cabrera, B.; Martínez, A. A mitochondrial analysis reveals distinct founder effect signatures in Canarian and Balearic goats. Anim. Genet. 2015, 46, 452–456.
- Yalcin, B. Sheep and Goats in Turkey; FAO: Rome, Italy, 1986.
- Lauvergne, J.; Renieri, C.; Machado, T.; Acharya, R. Goat genetic resources. In Proceedings of the 5th International Goat Conference, New Delhi, India, 2–8 March 1992.
- Jenot, F. Goat unions, FNEC and technical institutes, an intersecting destiny: Trajectories and dynamics since 1950 in France. Ethnozootechnie 2011, 91, 123–133.
- Casey, N.; Van Niekerk, W. The Boer goat. I. Origin, adaptability, performance testing, reproduction and milk production. Small Rumin. Res. 1988, 1, 291–302.
- Sanfiorenzo, J.H. Evaluation of six breeding lines of milk goats. J. Agric. Univ. P. R. 1962, 46, 205–212.
- Ronningen, K. Causes of variation in the flavour intensity of goat milk. Acta Agric. Scand. 1965, 15, 301–342.
- Wiggans, G.; Van Vleck, L.D.; Dickinson, F. Projection factors for goat lactation records. J. Dairy Sci. 1979, 62, 797–801.
- Gipson, T.A. Recent advances in breeding and genetics for dairy goats. Asian Australas. J. Anim. Sci. 2019, 32, 1275.
- Wiggans, G.; Dickinson, F.; King, G.; Weller, J. Genetic evaluation of dairy goat bucks for daughter milk and fat. J. Dairy Sci. 1984, 67, 201–207.
- Wiggans, G.; Van Dijk, J.; Misztal, I. Genetic evaluation of dairy goats for milk and fat yield with an animal model. J. Dairy Sci. 1988, 71, 1330–1337.
- Kemp, S.J.; Brezinsky, L.; Teale, A. A panel of bovine, ovine and caprine polymorphic microsatellites. Anim. Genet. 1993, 24, 363–365.
- Vaiman, D.; Schibler, L.; Bourgeois, F.; Oustry, A.; Amigues, Y.; Cribiu, E.P. A genetic linkage map of the male goat genome. Genetics 1996, 144, 279–305.
- Arevalo, E.; Holder, D.; Derr, J.; Bhebhe, E.; Linn, R. Caprine microsatellite dinucleotide repeat polymorphisms at the Sr-Crsp-1, Sr-Crsp-2, Sr-Crsp-3, Sr-Crsp-4 and Sr-Crsp-5 loci. Anim. Genet. 1994, 25, 202.
- Schibler, L.; Vaiman, D.; Oustry, A.; Guinec, N.; Dangy-Caye, A.L.; Billault, A.; Cribiu, E.P. Construction and extensive characterization of a goat bacterial artificial chromosome library with threefold genome coverage. Mamm. Genome 1998, 9, 119–124.
- Mucha, S.; Mrode, R.; Coffey, M.; Kizilaslan, M.; Desire, S.; Conington, J. Genome-wide association study of conformation and milk yield in mixed-breed dairy goats. J. Dairy Sci. 2018, 101, 2213–2225.
- Bickhart, D.M.; Rosen, B.D.; Koren, S.; Sayre, B.L.; Hastie, A.R.; Chan, S.; Lee, J.; Lam, E.T.; Liachko, I.; Sullivan, S.T. Single-molecule sequencing and chromatin conformation capture enable de novo reference assembly of the domestic goat genome. Nat. Genet. 2017, 49, 643–650.
- Carillier, C.; Larroque, H.; Palhière, I.; Clément, V.; Rupp, R.; Robert-Granié, C. A first step toward genomic selection in the multi-breed French dairy goat population. J. Dairy Sci. 2013, 96, 7294–7305.
- Dong, Y.; Xie, M.; Jiang, Y.; Xiao, N.; Du, X.; Zhang, W.; Tosser-Klopp, G.; Wang, J.; Yang, S.; Liang, J. Sequencing and automated whole-genome optical mapping of the genome of a domestic goat (Capra hircus). Nat. Biotechnol. 2013, 31, 135–141.
- Tosser-Klopp, G. Goat genome assembly, Availability of an international 50K SNP chip and RH panel: An update of the International Goat Genome Consortium projects. In Proceedings of the International Plant and Animal Genome XX Conference, Town and Country Hotel, San Diego, CA, USA, 14–18 January 2012.
- Fontanesi, L.; Martelli, P.L.; Beretti, F.; Riggio, V.; Dall’Olio, S.; Colombo, M.; Casadio, R.; Russo, V.; Portolano, B. An initial comparative map of copy number variations in the goat (Capra hircus) genome. BMC Genom. 2010, 11, 639.
- Li, X.-L.; Wu, Z.-L.; Liu, Z.-Z.; Gong, Y.-F.; Zhou, R.-Y.; Zheng, G.-R. SNP identification and analysis in part of intron 2 of goat MSTN gene and variation within and among species. J. Hered. 2006, 97, 285–289.
- Andonov, S.; Ødegård, J.; Svendsen, M.; Ådnøy, T.; Vegara, M.; Klemetsdal, G. Comparison of random regression and repeatability models to predict breeding values from test-day records of Norwegian goats. J. Dairy Sci. 2013, 96, 1834–1843.
- Gebreselassie, G.; Berihulay, H.; Jiang, L.; Ma, Y. Review on genomic regions and candidate genes associated with economically important production and reproduction traits in sheep (Ovies aries). Animals 2020, 10, 33.
- Zhang, H.; Wang, Z.; Wang, S.; Li, H. Progress of genome wide association study in domestic animals. J. Anim. Sci. Biotechnol. 2012, 3, 26.
- Lashmar, S.; Visser, C.; van Marle-Köster, E. Validation of the 50k Illumina goat SNP chip in the South African Angora goat. S. Afr. J. Anim. Sci. 2015, 45, 56–59.
- Li, X.; Wu, Z.L.; Gong, Y.; Liu, Y.; Liu, Z.; Wang, X.; Xin, T.; Ji, Q. Single-nucleotide polymorphism identification in the caprine myostatin gene. J. Anim. Breed. 2006, 123, 141–144.
- Andonov, S.; Ødegård, J.; Boman, I.; Svendsen, M.; Holme, I.; Ådnøy, T.; Vukovic, V.; Klemetsdal, G. Validation of test-day models for genetic evaluation of dairy goats in Norway. J. Dairy Sci. 2007, 90, 4863–4871.
- Pizarro Inostroza, M.G.; Navas González, F.J.; Landi, V.; León Jurado, J.M.; Delgado Bermejo, J.V.; Fernández Álvarez, J.; Martínez Martínez, M.d.A. Software-Automatized Individual Lactation Model Fitting, Peak and Persistence and Bayesian Criteria Comparison for Milk Yield Genetic Studies in Murciano-Granadina Goats. Mathematics 2020, 8, 1505.
- Tosser-Klopp, G.; Bardou, P.; Bouchez, O.; Cabau, C.; Crooijmans, R.; Dong, Y.; Donnadieu-Tonon, C.; Eggen, A.; Heuven, H.C.; Jamli, S. Design and characterization of a 52K SNP chip for goats. PLoS ONE 2014, 9, e86227.
- Onzima, R.; Upadhyay, M.; Mukiibi, R.; Kanis, E.; Groenen, M.; Crooijmans, R. Genome-wide population structure and admixture analysis reveals weak differentiation among Ugandan goat breeds. Anim. Genet. 2018, 49, 59–70.
- Schibler, L.; Vaiman, D.; Oustry, A.; Giraud-Delville, C.; Cribiu, E.P. Comparative gene mapping: A fine-scale survey of chromosome rearrangements between ruminants and humans. Genome Res. 1998, 8, 901–915.
- Du, X.; Womack, J.; Owens, K.; Elliott, J.; Sayre, B.; Bottcher, P.; Milan, D.; Podesta, M.G.; Zhao, S.; Malek, M. A whole-genome radiation hybrid panel for goat. Small Rumin. Res. 2012, 105, 114–116.
- Webb, E.; Casey, N.; Simela, L. Growth, development and growth manipulation in goats. In Goat Meat Production and Quality; Mahgoub, O., Kadim, I.T., Cottington Webb, E., Ovid Technologies, Inc., Eds.; CAB International: Cambridge, MA, USA, 2012; pp. 196–199.
- Shrestha, J.; Fahmy, M. Breeding goats for meat production: 3. Selection and breeding strategies. Small Rumin. Res. 2007, 67, 113–125.
- Shrestha, J.; Fahmy, M. Breeding goats for meat production: 2. Crossbreeding and formation of composite population. Small Rumin. Res. 2007, 67, 93–112.
- Fernández Álvarez, J.; León Jurado, J.; Navas González, F.J.; Iglesias Pastrana, C.; Delgado Bermejo, J.V. CAPRIGRAN Linear Appraisal Evidences Dairy Selection Signs in Murciano-Granadina Goats and Bucks: Presentation of the New Linear Appraisal Scale. Arch. Zootecn. 2021, 70, 240–245.
- Webb, E.; Casey, N.; Simela, L. Goat meat quality. Small Rumin. Res. 2005, 60, 153–166.
- Webb, E. Goat meat production, composition, and quality. Anim. Front. 2014, 4, 33–37.
- Raes, K.; Balcaen, A.; Dirinck, P.; De Winne, A.; Claeys, E.; Demeyer, D.; De Smet, S. Meat quality, fatty acid composition and flavour analysis in Belgian retail beef. Meat Sci. 2003, 65, 1237–1246.
- Herrera, P.Z.; Bermejo, J.V.D.; Henríquez, A.A.; Vallejo, M.E.C.; Costa, R.G. Effects of extensive system versus semi-intensive and intensive systems on growth and carcass quality of dairy kids. Rev. Bras. Zootec. 2011, 40, 2613–2620.
- Jembere, T.; Dessie, T.; Rischkowsky, B.; Kebede, K.; Okeyo, A.; Haile, A. Meta-analysis of average estimates of genetic parameters for growth, reproduction and milk production traits in goats. Small Rumin. Res. 2017, 153, 71–80.
- Huson, H.; Sonstegard, T.; Silverstein, J.; Woodward Greene, J.; Masiga, C.; Muchadeyi, F.; Rees, J.; Sayre, B.; Elbeltagy, A.; Rothschild, M.F. Genetic and phenotypic characterization of African goat populations to prioritize conservation and production efforts for small-holder farmers in Sub-Saharan africa. In Proceedings of the 10th World Congress on Genetics Applied to Livestock Production, Vancouver, BC, Canada, 17–22 August 2014; American Society of Animal Science: Champaign, IL, USA, 2014.
- Zhang, N.; Teng, Z.; Qi, Q.; Hu, G.; Lian, H.; Gao, T. Carcass traits, meat quality characteristics, and lipid metabolism-related gene expression pattern of Yaoshan white goats raised in traditional extensive production system: Effects of slaughter age and meat cuts. Small Rumin. Res. 2020, 182, 29–36.
- FAO. FAOSTAT Statistical Database. 2019. Available online: http://www.fao.org/faostat/en/ (accessed on 17 July 2021).
- García, V.; Rovira, S.; Boutoial, K.; López, M. Improvements in goat milk quality: A review. Small Rumin. Res. 2014, 121, 51–57.
- Pazzola, M.; Stocco, G.; Dettori, M.L.; Bittante, G.; Vacca, G.M. Effect of goat milk composition on cheesemaking traits and daily cheese production. J. Dairy Sci. 2019, 102, 3947–3955.





