
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Libero Nigro | -- | 1191 | 2022-04-18 10:32:40 | | | |
| 2 | Lindsay Dong | + 1691 word(s) | 2882 | 2022-04-18 11:28:37 | | | | |
| 3 | Lindsay Dong | + 76 word(s) | 2958 | 2022-04-19 03:05:26 | | | | |
| 4 | Lindsay Dong | -1 word(s) | 2957 | 2022-04-19 03:16:50 | | | | |
| 5 | Lindsay Dong | + 5 word(s) | 2962 | 2022-04-21 05:29:43 | | |
Video Upload Options
K-means is a well-known clustering algorithm often used for its simplicity and potential efficiency. K-means, though, suffers from computational problems when dealing with large datasets with many dimensions and great number of clusters.
1. Introduction
K-Means algorithm [1][2] is very often used in unsupervised clustering applications such as data mining, pattern recognition, image processing, medical informatics, genoma analysis and so forth. This is due to its simplicity and its linear complexity O(NKT) where N is the number of data points in the dataset, K is the number of assumed clusters and T is the number of iterations needed for convergence. Fundamental properties and limitations of K-Means have been deeply investigated by many theoretical/empirical works reported in the literature. Specific properties, though, like ensuring accurate clustering solutions through sophisticated initialization methods [3], can be difficult to achieve in practice when large datasets are involved. More in general, dealing with a huge number of data records, possibly with many dimensions, create computational problems to K-Means which motivate the use of a parallel organization/execution of the algorithm [4] either on a shared-nothing architecture, that is a distributed multi-computer context, or on a shared-memory multiprocessor. Examples of the former case include message-passing solutions, e.g., based on MPI [5], with a master/slave organization [6], or based on the functional framework MapReduce [7][8]. For the second case, notable are the experiments conducted by using OpenMP [9] or based on the GPU architecture [10]. Distributed solutions for K-Means can handle very large datasets which can be decomposed and mapped on the local memory of the various computing nodes. Shared memory solutions, on the other hand, assume the dataset to be allocated, either monolithically or in subblocks to be separately operated in parallel by multiple computing units.
2. K-Means
A dataset X={x1,x2,…,xN} is considered with N data points (records). Each data point has D coordinates (number of features or dimensions). Data points must be partitioned into K clusters in such a way to ensure similarity among the points of a same cluster and dissimilarity among points belonging to different clusters. Every cluster , , is represented by its centroid point , e.g., its mean or central point. The goal of K-Means is to assign points to clusters in such a way to minimize the sum of squared distances (SSD) objective function:
where is the number of points of X assigned to cluster and denotes the Euclidean distance between and . The problem is NP-hard and practically is approximated by heuristic algorithms. The standard method of Lloyd’s K-means [14] is articulated in the iterative steps shown in Figure 1.
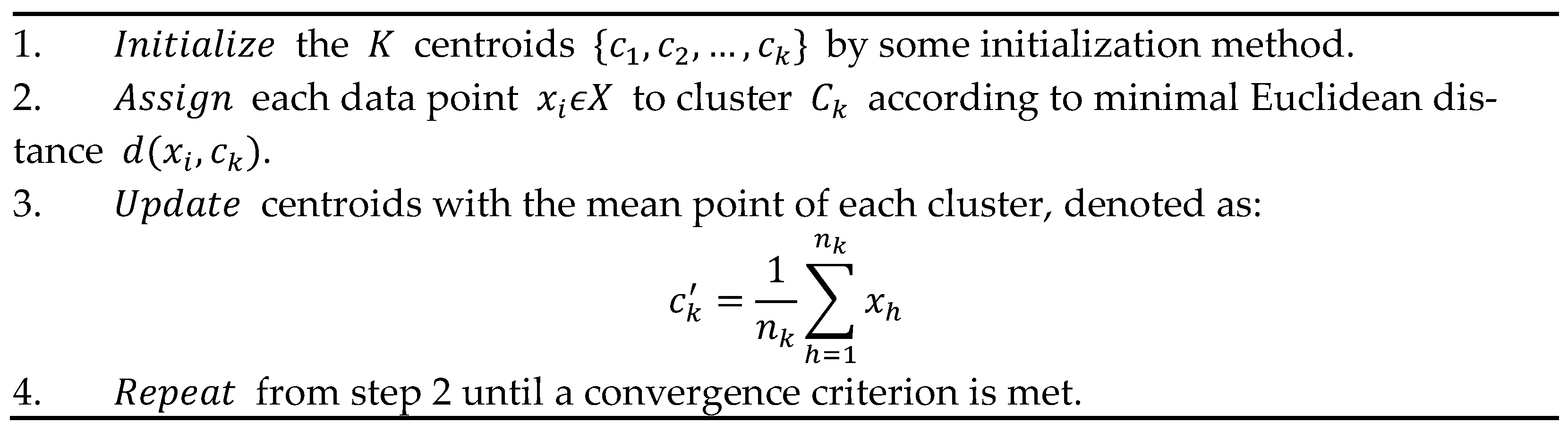
Figure 1. Sequential operation of K-Means.
Several initialization methods for the step 1 in Figure 1, either stochastic or deterministic (see also later in this section), have been considered and studied in the literature, see e.g., [3][11][12][13][14], where each one influences the evolution and the accuracy of K-Means. Often, also considering the complexity of some initialization algorithms, the random method is adopted which initializes the centroids as random selected points in the dataset, although it can imply K-Means evolves toward a local minima.
3. Parallel K-Means in Java
In the basic K-Means algorithm shown in Figure 1, there is a built-in parallelism in both the and the phases, which can be extracted to speed-up the computation. In other terms, both the calculation of the minimal distances of points to current centroids, and then the definition of new centroids as the mean points of current clusters, can be carried in parallel.
3.1. Supporting K-Means by Streams
A first solution to host parallel K-Means in Java is based on the use of streams, lambda expressions and a functional programming style [15], which were introduced since the Java 8 version. Streams are views (not copies) of collections (e.g., lists ) of objects, which make it possible to express a fluent style of operations (method calls). Each operation works on a stream, transforms each object according to a lambda expression, and returns a new stream, ready to be handled by a new operation and so forth. In a fluent code segment, only the execution of the terminal operation actually triggers the execution of the intermediate operations.
Figure 2 depicts the main part of the K-Means solution based on streams which can execute either serially or in parallel. The two modes are controlled by a global Boolean parameter depicts the main part of the K-Means solution based on streams which can execute either serially or in parallel. The two modes are controlled by a global Boolean parameter PARALLEL which can be false or true. The algorithm first loads the dataset and the initial centroids onto two global arrays of data points (of a class DataPoint), respectively dataset and centroids of length N and K, from which corresponding (possibly parallel) streams are built.

Figure 2. A map/reduce schema for K-Means based on Java streams.
A Map/Reduce schema was actually coded where the Map stage corresponds to the Assign step of Figure 1, and the Reduce stage realizes the Update step of Figure 1.
The DataPoint class describes a point in , that is with D coordinates, and exposes methods for calculating the Euclidean distance between two points, adding points (by adding the corresponding coordinates and counting the number of additions), calculating the mean of a point summation and so forth. DataPoint also admits a field CID (Cluster Identifier) which stores the ID (index of the centroids array) of the cluster the point belongs to. The CID field is handled by the getCID()/setCID() methods. The map() method on p_stream receives a lambda expression which accepts a DataPoint p and returns p mapped to the cluster centroid closer to p. Since map() is an intermediate operation, a final fictitious forEach() operation is added which activates the execution of map().
When p_stream is completely processed, a newCentroids array is reset (each new centroid has 0 as coordinates) so as to generate on it the updated version of centroids. Purposely, the CID field of every new centroid is set to itself.
The c_stream is actually extracted from the newCentroids array. The map() method on c_stream receives a lambda expression which accepts a new centroid point c and adds to c all the points of the dataset whose CID coincides with the CID of c. Following the summation of the points belonging to the same new centroid c, the mean() method is invoked on c to calculate the mean point of the resultant cluster.
The iteration counter it gets incremented at the end of each K-Means iteration. The termination() method returns true when the convergence was obtained or the maximum number of iterations T was reached. In any case, termination() ends by copying the contents of newCentroids onto the centroids array. Convergence is sensed when all the distances among new centroids and current centroids fall under the threshold THR.
It is worthy of note that all the globals: the parameters N, K, D, T, THR, E, it, MP, PARALLEL …, the dataset, centroids/newCentroids data point arrays, the implemented centroid initialization methods (all the ones, either stochastic or deterministic, discussed in [3]), some auxiliary methods for calculating the SSD cost of a clustering solution or evaluating its quality by, e.g., the Silhouette index [3] and so forth, are introduced as static entities of a G class, which are statically imported in applicative classes for a direct access to its entities.
The Java code in Figure 2 automatically adapts itself to a multi-threaded execution, when the p_stream and c_stream are turned to a parallel version. In this case, behind the scene, Java uses multiple threads of a thread-pool to accomplish in parallel, on selected data segments, the underlying operations required by the Map and Reduce phases.
A basic feature of the Java code is its independence from the order of processed data points and the absence of side-effects which can cause interference problems among threads when accessing to shared data. During the Map stage, the lambda expression operates on distinct data points and affects their CID without any race condition. Similarly, multiple threads act on distinct new centroid points during the Reduce phase, and safely accumulate data points of a same cluster.
A specific parameter in the G class is INIT_METHOD which takes values in an enumeration with the constants RANDOM, MAXIMIN, KMEANSPP, KAUFMAN, ROBIN, DKMEANSPP, to select a corresponding centroid initialization method [3]. Other initialization methods can be added as well. The ROBIN method, which can be configured to operate stochastically or deterministically, relies concretely on heap-sorting (through a PriorityQueue collection) the dataset in descending order of the minimal distances from the existing centroids and initially from the reference point. In addition, the nearest MP-neighborhood of a data point , considered in sorted order, is determined by moving around , and registering in a sorted data structure the distances of nearest points to . Movement is stopped as soon as a point is encountered whose distance from xi is greater than that of any already met nearest point, and the cardinality of previous visited nearest points is found to be ≥ MP.
The G class is extended by a further parameter P which specifies the degree of parallelism, that is the number of the underlying cores (threads) exploitable in the execution experiments.
3.2. Actor-Based K-Means Using Theatre
3.2.1. The Parallel Programming Model of Theatre
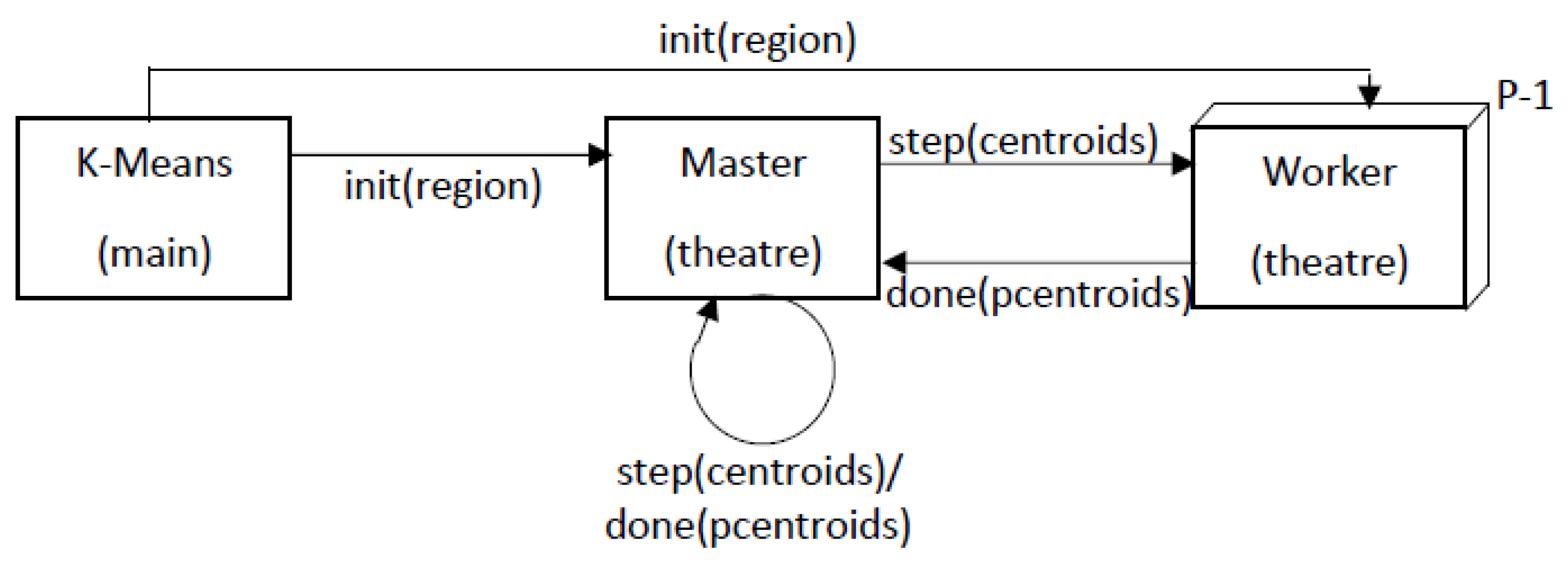
3.2.2. Mapping Parallel K-Means on Theatre
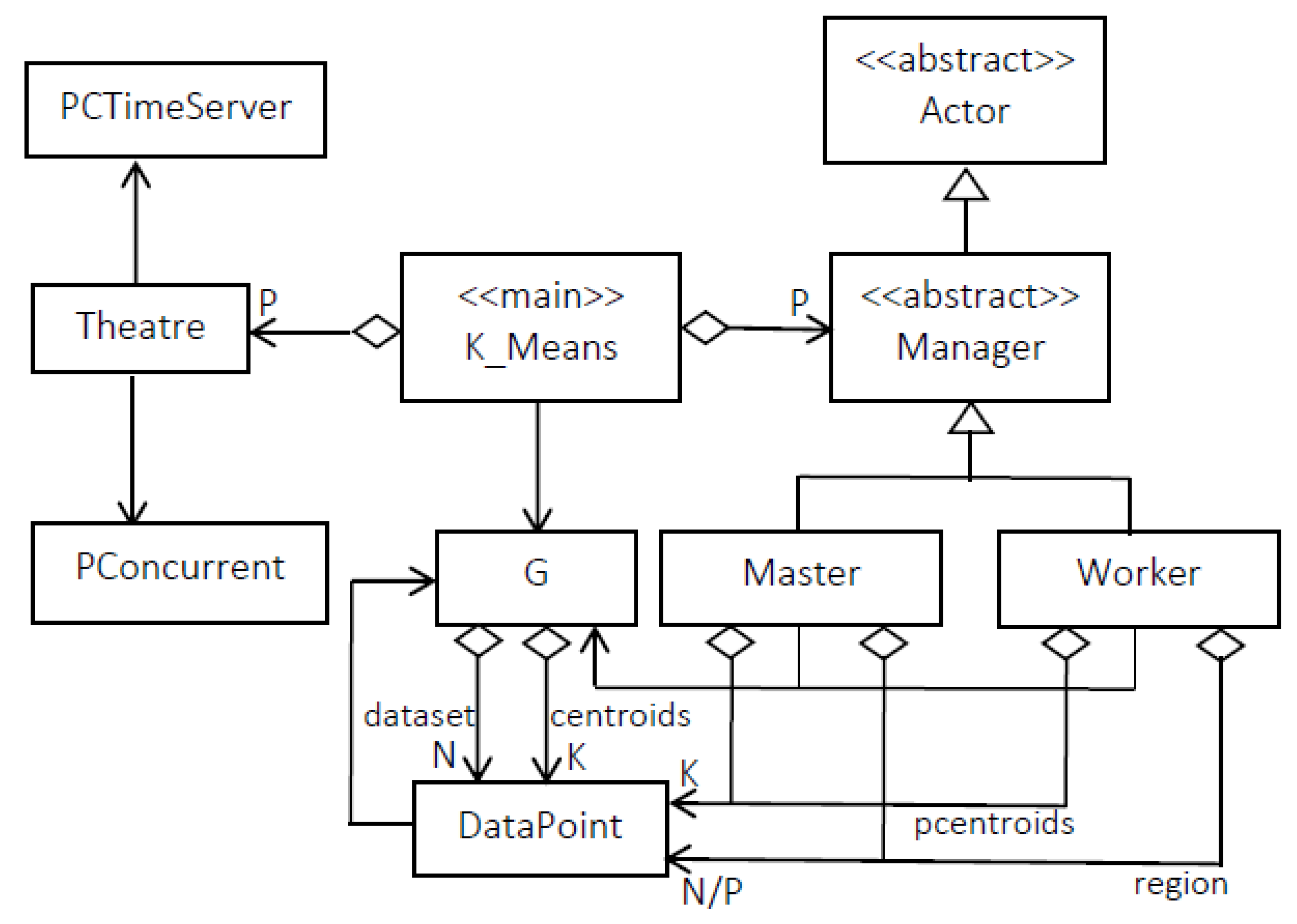
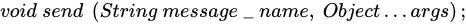
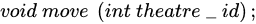


| Parameter | Value | Meaning |
|---|---|---|
| P | 16 | Number of used underlying threads |
| N | 2,458,285 | Size of the Census1990 data set |
| D | 68 | Dimensions of the dataset records |
| K | e.g., 80 | Number of assumed centroids |
| T | 1000 | Maximum number of iterations |
| THR | 1 × 10−10 | Threshold for assessing convergence |
| INIT_METHOD | ROBIN | The centroid initialization method |
| E | 0.05 | Tolerance in the Local Outlier Factor (LOF) detection (ROBIN) |
| MP | e.g., 15 | Size of the MP-neighborhood of data points (ROBIN) |
References
- MacQueen, J. Some methods for classification and analysis of multivariate observations. In Proceedings of the 5th Berkeley Symposium on Mathematical Statistics and Probability, Berkeley, CA, USA, 21 June–18 July 1965 and 27 December 1965–7 January 1966; University of California Press: Berkeley, CA, USA, 1967; pp. 281–297.
- Jain, A.K. Data clustering: 50 years beyond k-means. Pattern Recognit. Lett. 2010, 31, 651–666.
- Vouros, A.; Langdell, S.; Croucher, M.; Vasilaki, E. An empirical comparison between stochastic and deterministic centroid initialisation for K-means variations. Mach. Learn. 2021, 110, 1975–2003.
- Qiu, X.; Fox, G.C.; Yuan, H.; Bae, S.H.; Chrysanthakopoulos, G.; Nielsen, H.F. Parallel clustering and dimensional scaling on multicore systems. In Proceedings of the High Performance Computing & Simulation (HPCS 2008), Nicosia, Cyprus, 3–6 June 2008; p. 67.
- Zhang, J.; Wu, G.; Hu, X.; Li, S.; Hao, S. A parallel k-means clustering algorithm with MPI. In Proceedings of the IEEE Fourth International Symposium on Parallel Architectures, Algorithms and Programming, NW Washington, DC, USA, 9–11 December 2011; pp. 60–64.
- Kantabutra, S.; Couch, A.L. Parallel K-means clustering algorithm on NOWs. NECTEC Tech. J. 2000, 1, 243–247.
- Zhao, W.; Ma, H.; He, Q. Parallel K-Means clustering based on MapReduce. In Proceedings of the IEEE International Conference on Cloud Computing, NW Washington, DC, USA, 21–25 September 2009; Springer: Berlin/Heidelberg, Germany, 2009; pp. 674–679.
- Bodoia, M. MapReduce Algorithms for k-Means Clustering. Available online: https://stanford.edu/~rezab/classes/cme323/S16/projects_reports/bodoia.pdf (accessed on 1 January 2022).
- Naik, D.S.B.; Kumar, S.D.; Ramakrishna, S.V. Parallel processing of enhanced K-Means using OpenMP. In Proceedings of the IEEE International Conference on Computational Intelligence and Computing Research, Madurai, India, 26–28 December 2013; pp. 1–4.
- Cuomo, S.; De Angelis, V.; Farina, G.; Marcellino, L.; Toraldo, G. A GPU-accelerated parallel K-means algorithm. Comput. Electr. Eng. 2019, 75, 262–274.
- Franti, P.; Sieranoja, S. K-means properties on six clustering benchmark datasets. Appl. Intell. 2018, 48, 4743–4759.
- Al Hasan, M.; Chaoji, V.; Salem, S.; Zaki, M.J. Robust partitional clustering by outlier and density insensitive seeding. Pattern Recognit. Lett. 2009, 30, 994–1002.
- Celebi, M.E.; Kingravi, H.A.; Vela, P.A. A comparative study of efficient initialization methods for the k-means clustering algorithm. Expert Syst. Appl. 2013, 40, 200–210.
- Franti, P.; Sieranoja, S. How much can k-means be improved by using better initialization and repeats? Pattern Recognit. 2019, 93, 95–112.
- Subramaniam, V. Functional Programming in Java—Harnessing the Power of Java 8 Lambda Expressions; The Pragmatic Programmers, LLC: Raleigh, NC, USA, 2014.
- Nigro, L.; Sciammarella, P.F. Qualitative and quantitative model checking of distributed probabilistic timed actors. Simul. Model. Pract. Theory 2018, 87, 343–368.
- Cicirelli, F.; Nigro, L. A development methodology for cyber-physical systems based on deterministic Theatre with hybrid actors. TASK Q. Spec. Issue Cyber-Phys. Syst. 2021, 25, 233–261.
- Agha, G. Actors: A Model of Concurrent Computation in Distributed Systems. Ph.D. Thesis, MIT Artificial Intelligence Laboratory, Cambridge, MA, USA, 1986.
- Karmani, R.K.; Agha, G. Actors; Springer: Boston, MA, USA, 2011; pp. 1–11.
- Nigro, L. Parallel Theatre: A Java actor-framework for high-performance computing. Simul. Model. Pract. Theory 2021, 106, 102189.





