K-means is a well-known clustering algorithm often used for its simplicity and potential efficiency. K-means, though, suffers from computational problems when dealing with large datasets with many dimensions and great number of clusters.
- parallel algorithms
- multi-core machines
- K-means clustering
- Java
1. Introduction
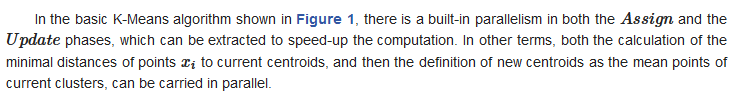
K-Means algorithm [1,2] is very often used in unsupervised clustering applications such as data mining, pattern recognition, image processing, medical informatics, genoma analysis and so forth. This is due to its simplicity and its linear complexity O(NKT) where N is the number of data points in the dataset, K is the number of assumed clusters and T is the number of iterations needed for convergence. Fundamental properties and limitations of K-Means have been deeply investigated by many theoretical/empirical works reported in the literature. Specific properties, though, like ensuring accurate clustering solutions through sophisticated initialization methods [3], can be difficult to achieve in practice when large datasets are involved. More in general, dealing with a huge number of data records, possibly with many dimensions, create computational problems to K-Means which motivate the use of a parallel organization/execution of the algorithm [4] either on a shared-nothing architecture, that is a distributed multi-computer context, or on a shared-memory multiprocessor. Examples of the former case include message-passing solutions, e.g., based on MPI [5], with a master/slave organization [6], or based on the functional framework MapReduce [7,8]. For the second case, notable are the experiments conducted by using OpenMP [9] or based on the GPU architecture [10]. Distributed solutions for K-Means can handle very large datasets which can be decomposed and mapped on the local memory of the various computing nodes. Shared memory solutions, on the other hand, assume the dataset to be allocated, either monolithically or in subblocks to be separately operated in parallel by multiple computing units.
2. K-Means
A dataset X={x1,x2,…,xN} is considered with N data points (records). Each data point has D coordinates (number of features or dimensions). Data points must be partitioned into K clusters in such a way to ensure similarity among the points of a same cluster and dissimilarity among points belonging to different clusters. Every cluster Ck, 1≤k≤K, is represented by its centroid point ck, e.g., its mean or central point. The goal of K-Means is to assign points to clusters in such a way to minimize the sum of squared distances (SSD) objective function:
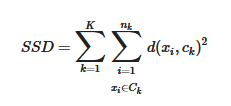
where nk is the number of points of X assigned to cluster Ck and d(xi,ck) denotes the Euclidean distance between xi and ck. The problem is NP-hard and practically is approximated by heuristic algorithms. The standard method of Lloyd’s K-means [14] is articulated in the iterative steps shown in Figure 1.
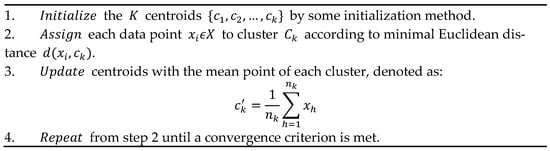

Figure 1. Sequential operation of K-Means.
Several initialization methods for the step 1 in Figure 1, either stochastic or deterministic (see also later in this section), have been considered and studied in the literature, see e.g., [3,15,16,17,18], where each one influences the evolution and the accuracy of K-Means. Often, also considering the complexity of some initialization algorithms, the random method is adopted which initializes the centroids as random selected points in the dataset, although it can imply K-Means evolves toward a local minima.
3. Parallel K-Means in Java
3.1. Supporting K-Means by Streams
A first solution to host parallel K-Means in Java is based on the use of streams, lambda expressions and a functional programming style [12], which were introduced since the Java 8 version. Streams are views (not copies) of collections (e.g., lists) of objects, which make it possible to express a fluent style of operations (method calls). Each operation works on a stream, transforms each object according to a lambda expression, and returns a new stream, ready to be handled by a new operation and so forth. In a fluent code segment, only the execution of the terminal operation actually triggers the execution of the intermediate operations.


Figure 4. A map/reduce schema for K-Means based on Java streams.


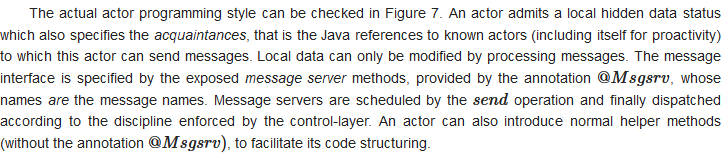
3.2. Actor-Based K-Means Using Theatre
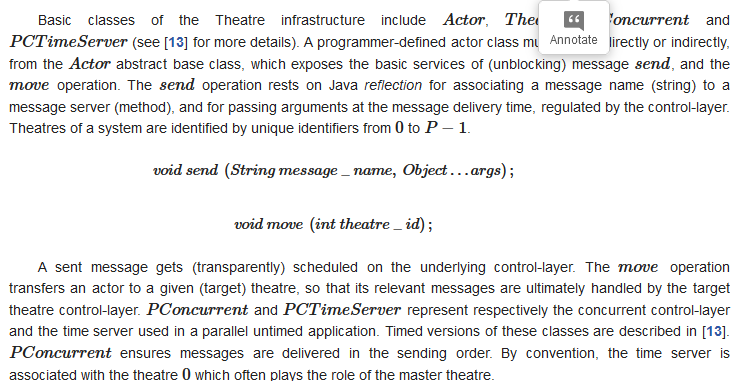
3.2.1. The Parallel Programming Model of Theatre
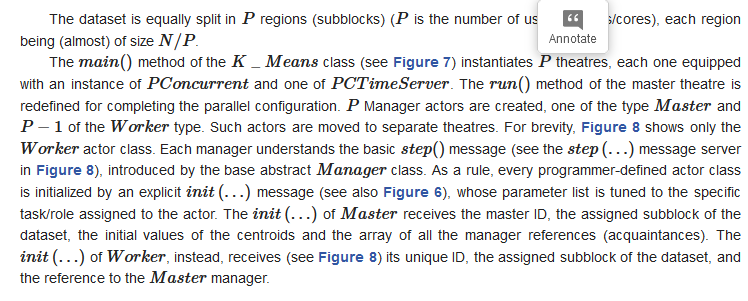
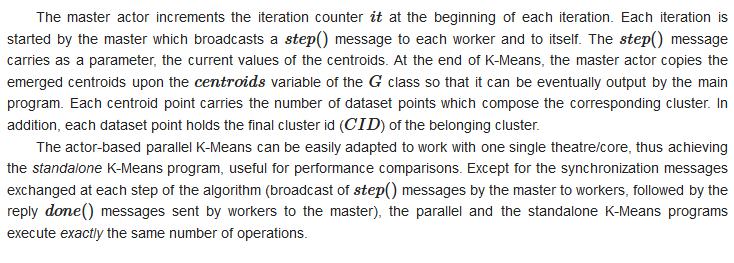
3.2.2. Mapping Parallel K-Means on Theatre
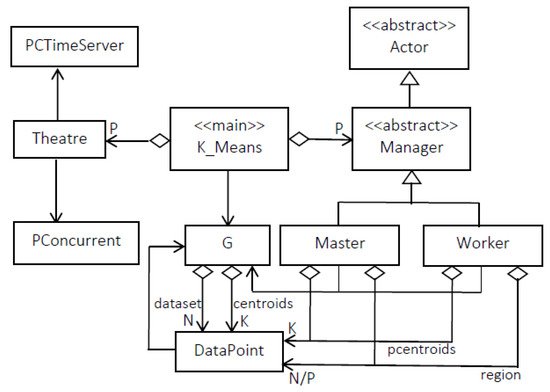


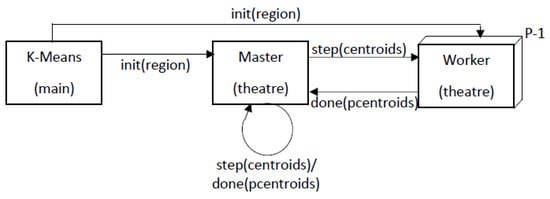





This entry is adapted from the peer-reviewed paper 10.3390/a15040117
