Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Matthew A Nugent | + 2755 word(s) | 2755 | 2022-03-12 13:10:30 | | | |
| 2 | Vicky Zhou | Meta information modification | 2755 | 2022-03-22 09:58:33 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Nugent, M.A. SARS-CoV2 Spike Protein. Encyclopedia. Available online: https://encyclopedia.pub/entry/20827 (accessed on 21 May 2026).
Nugent MA. SARS-CoV2 Spike Protein. Encyclopedia. Available at: https://encyclopedia.pub/entry/20827. Accessed May 21, 2026.
Nugent, Matthew A. "SARS-CoV2 Spike Protein" Encyclopedia, https://encyclopedia.pub/entry/20827 (accessed May 21, 2026).
Nugent, M.A. (2022, March 21). SARS-CoV2 Spike Protein. In Encyclopedia. https://encyclopedia.pub/entry/20827
Nugent, Matthew A. "SARS-CoV2 Spike Protein." Encyclopedia. Web. 21 March, 2022.
Copy Citation
The SARS-CoV2 Spike protein protrudes from the surface of the virus where it plays a critical role in mediating attachment and entry into target human cells. Mutations in Spike have been implicated in the evolution of highly transmissible variants that have led to increased cases of COVID-19. Angiotensin converting enzyme 2 has been identified as the Spike receptor on target cells. Additional Spike binding sites on target cells, including heparan sulfate, have been shown to contribute to Spike-mediated SARS-CoV2 cell infection. It will be critical to define the molecular mechanisms used by Spike in order to design effective therapies to combat current and future variants of SARS-CoV2.
ACE2
angiotensin converting enzyme 2
co-receptors
coronavirus
heparan sulfate proteoglycans
heparin
HSPGs
Omicron
receptors
1. Spike Protein Structure and Function
Spike is a ~180 kDa protein that protrudes from the surface of the SARS-CoV2 particle where it mediates host-cell attachment and entry [1][2][3][4][5]. The SARS-CoV2 Spike is expressed as a trimer on the SARS-CoV2 surface, with approximately 25 Spike trimers per virus particle [6]. The human transmembrane glycoprotein angiotensin converting enzyme 2 (ACE2) has been identified as the cognate receptor for Spike [7][8][9][10]. Furthermore, ACE2 is expressed in various tissues, including prominently within type I and type II alveolar epithelial cells in the lungs, and in nasal and bronchial epithelium [11][12]. Although ACE2 expression within the respiratory system is consistent with the pathology of COVID-19, it is interesting to note that ACE2 is also highly expressed in oral epithelia, enterocytes within the small intestine and the vascular endothelium. While the broad expression pattern of ACE2 indicated that viral disease could spread through many routes, it has become clear that respiratory transmission is the dominant means. Hence, expression of ACE2 seems necessary for infection, yet it alone does not appear to be a sufficient predictor of which cells will be most susceptible to SARS-CoV2.
The binding of Spike to ACE2 positions Spike on the cell surface for cleavage by human proteases, particularly transmembrane protease serine 2 (TMPRSS2), that induces a conformational change from a compact structure to an extended “fusion” structure that facilitates the fusion of the viral and host cell membranes, thereby allowing viral RNA to gain entry to the cellular cytoplasm where it can be translated and replicated to produce new viral particles (Figure 1). The ability of TMPRSS2 to induce the conversion of Spike into the fusion conformation is enhanced if Spike is first cleaved by the protease furin during biosynthesis [13][14][15]. Thus, the expression and distribution of ACE2 determines the tissues that are infected by SARS-CoV2 and ultimately mediates COVID-19 pathogenesis. Interesting variations in ACE2 expression have been reported in individuals by age, gender, and disease, suggesting potential explanations for the highly variable clinical responses to SARS-CoV2 infection [16][17][18].
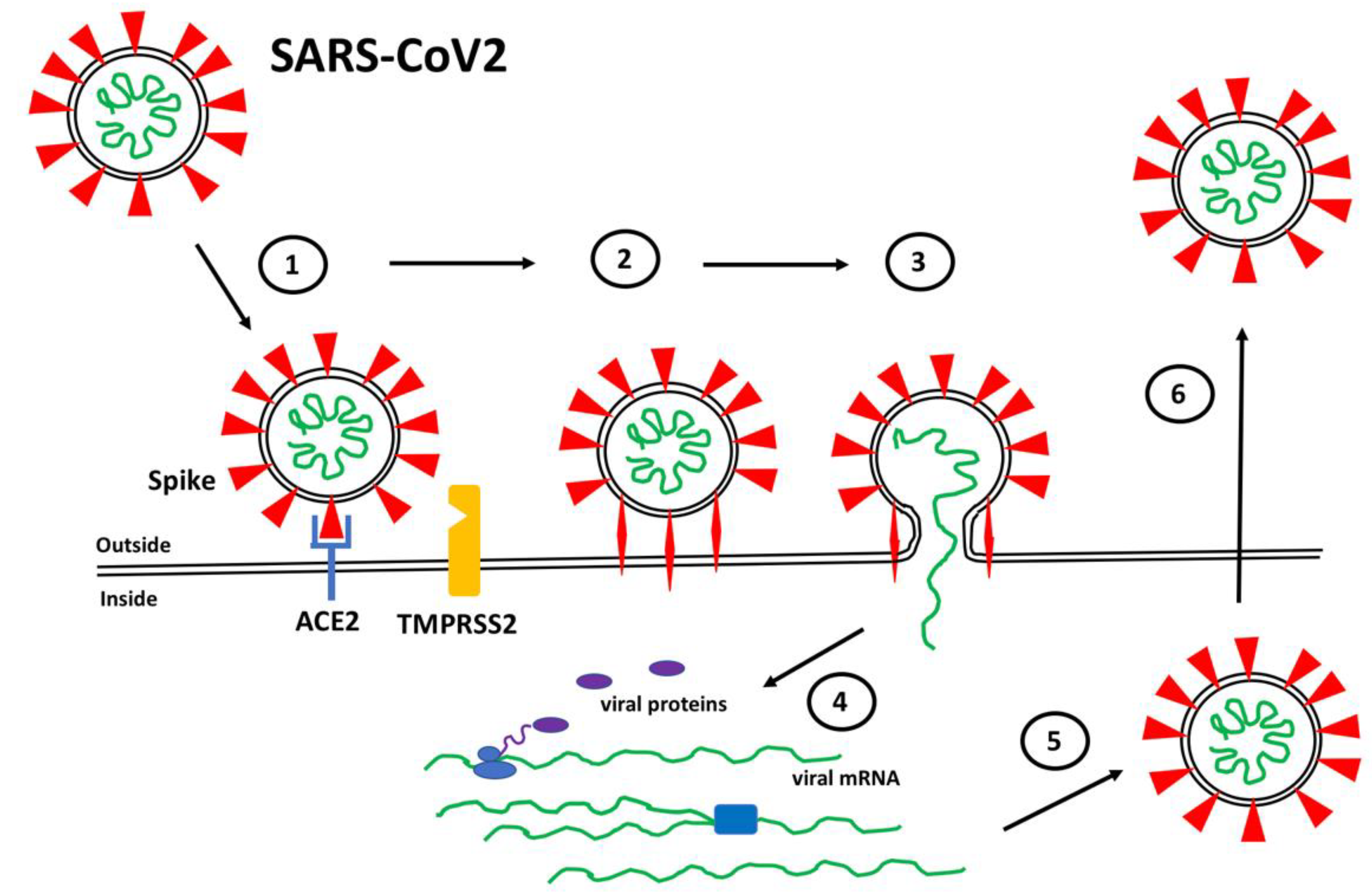
Figure 1. SARS-CoV2 Infection. (1) Spike proteins (red) on the SARS-CoV2 surface bind to ACE2 (blue) on the surface of target host cells. (2) Spike-ACE2 interactions position Spike for proteolytic processing on the cell surface by TMPRSS2 (orange) (or by cathepsin in endosomes after internalization). Cleavage of Spike releases a structural constraint that allows Spike to convert to its “fusion” structure that penetrates the host cell membrane. (3) “Fusion” Spike facilitates fusion between the viral and host membranes, allowing entry of viral RNA. (4) Viral RNA is translated and replicated within the host cell. (5) New virus particles are packaged in the host cell where furin cleaves Spike so as to prime it for later conversion to the “fusion” structure. (6) New SARS-CoV2 particles are released (or transferred to adjacent fused cells) to propagate further host cell infection.
Spike protein from wild-type SARS-CoV2 contains 1273 amino acid residues arranged into two subunits, S1 (residues 14-685) and S2 (residues 686-1273) [1][3][5][19]. The receptor binding domain (RBD) is within the S1 subunit (residues 319-541) and contains the region that interacts directly with ACE2, called the receptor binding motif (RBM) (residues 437-507). The protease cleavage site responsible for the transition of Spike into the “fusion” conformation are within the S2 subunit. The SARS-CoV2 Spike protein also contains a unique insertion of a furin cleavage site, PRRA, at the S1/S2 junction that allows furin to prime Spike during biosynthesis for future cleavage by TMPRRS2 (or cathepsin in late endosomes). Thus, considerable attention has been focused on the particular characteristics of the Spike RBD and furin cleavage site to gain insight into how SARS-CoV2 has become so highly transmissible among the human population.
Spike protein is heavily glycosylated, with the exception of the RBD, providing extensive shielding from antibody recognition [20][21][22][23] (Figure 2). Interestingly, the RBD is structurally flexible and can dynamically switch between an open (up) and closed (down) conformation with the closed conformation being the preferred state. Spike can only bind to ACE2 when its RBD is in the open conformation. Within a given trimeric Spike, the three RBDs dynamically interact to stabilize the closed state such that it is rare to find more than one RBD in the open state at a time. Indeed, a Cryo-EM study of Spike protein indicates that, on average, an RBD will be in the open conformation in only ~20% of Spike trimers, while 80% will have all three RBDs in the closed state [3][24]. Thus, the relative amount of time that the RBD spends in the open state increases the probability that an encounter between Spike and ACE2 will lead to the formation of a bound complex while also increasing the chance that Spike will be recognized by host antibodies. Hence, Spike has likely evolved this conformational dynamic process to balance between these two competing events as a means of enhancing its ability to infect. Interestingly, the glycosylation of Spike also participates actively in the transition between the open versus closed state of the RBD [25][26]. In a unique mechanism, glycans at N165 and N234 stabilize RBD when it is in the open conformation while the glycan at N343 physically pushes the RBD up into the open conformation. Mutating any of these three residues reduces binding of Spike to ACE2 with the loss of the glycan at N343 causing a greater than 50% reduction in binding [26].
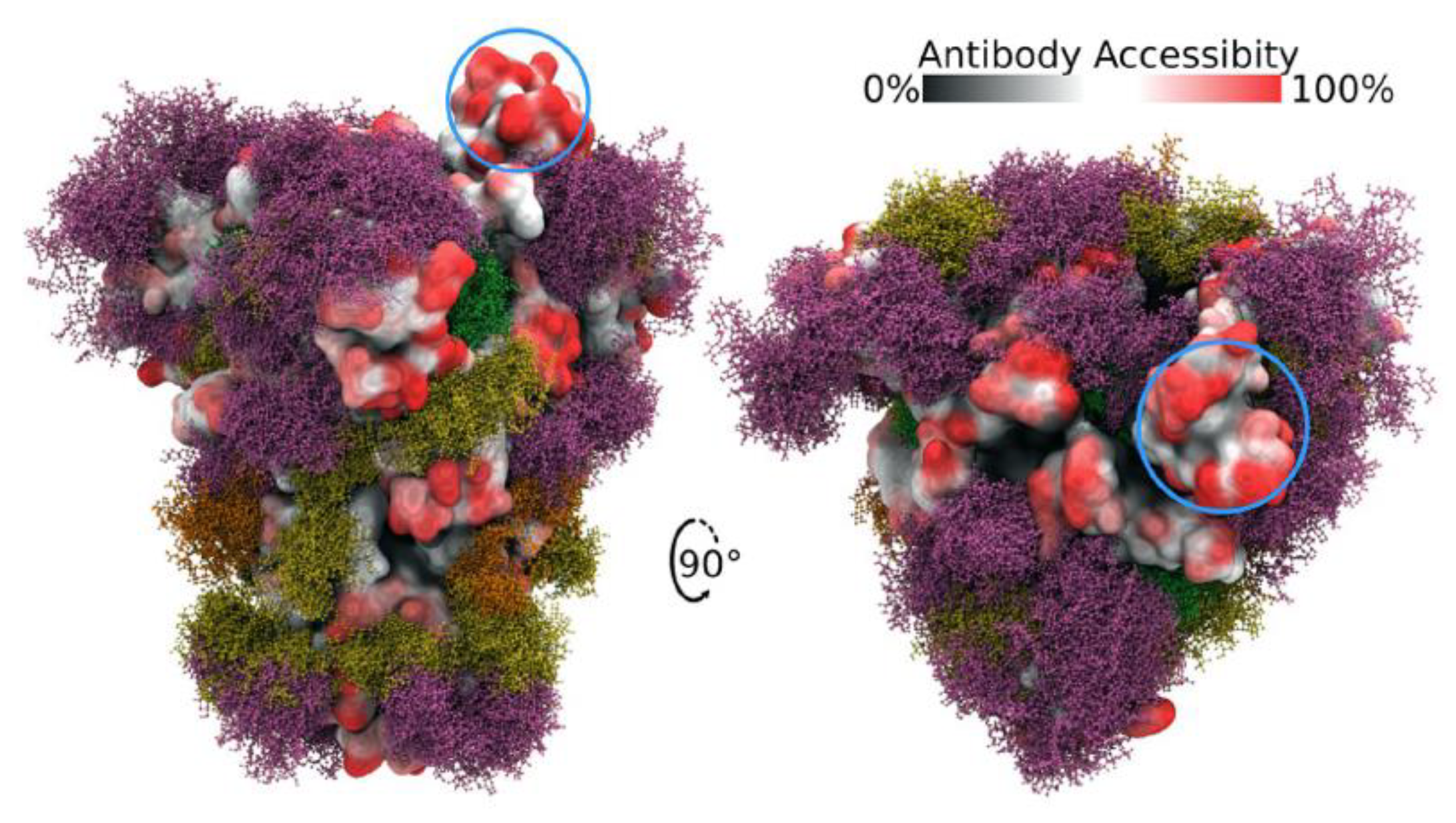
Figure 2. Side and top views of Spike trimer with site-specific glycosylation shown (moss surface) from molecular dynamics simulations [23]. The glycans are shown in ball-and-stick representation in green, dark yellow, orange and pink. The protein surface is colored according to antibody accessibility from black (least) to red (most accessible). One RBD is shown in the open conformation (circled in blue). Images are from [23] and are used under Creative Commons Attribution 4.0 International License: https://creativecommons.org/licenses/by/4.0/, accessed on 1 March 2022.
Since the original identification of SARS-CoV2, a number of variant forms have evolved and spread. Each of the variants show mutations in the Spike protein that have been linked to altered virulence and/or transmissibility. The Alpha (B.1.1.7), Beta (B.1.351) and Gamma (P.1) variants all carry an N501Y mutation within the RBM that has been demonstrated to increase the binding affinity to ACE2 by 3- to 7-fold, depending on the presence of additional mutations [27][28][29]. The recent emergence of the highly contagious Delta (B.1.1.617.2) and Omicron (B.1.1.529) variants reveal a number of additional important mutations in Spike [30]. The Delta variant has two mutations in its RBM, L452R and T478K, and in some instances has additional mutations K417N and E484K. The Omicron variant Spike protein, in contrast, carries as many as 30 point mutations, three deletions and one insertion. Strikingly, it has 15 mutations within its RBD with 10 tightly clustered within the RBM, namely N440K, G446S, S477N, T478K, E484A, Q493K, G496S, Q498R, N501Y and Y505H (Figure 3). It is interesting to note that these 10 mutations result in a significant change in the charged nature of the RBM with the substitution of four non-charged residues (N, T, and 2Qs) with positively charged residues (R and 3Ks), the loss or one negatively charged residue (E) and the addition of one histidine which can carry a positive charge depending on its local environment. A systematic analysis of each single Omicron mutation on ACE2 binding revealed that nine of the RBD mutations would be expected to decrease binding affinity while the other six should increase affinity [31]. Indeed, a comparative analysis of WT, Delta and Omicron revealed that the Delta Spike binds with higher affinity compared to WT and Omicron. Interestingly, the Omicron Spike does not show increased affinity for ACE2 compared to WT Spike [31]. Hence, it appears that the binding affinity for ACE2 is not the sole determinant of the effectiveness of Spike protein that dictates the enhanced transmission of SARS-CoV2 variants.
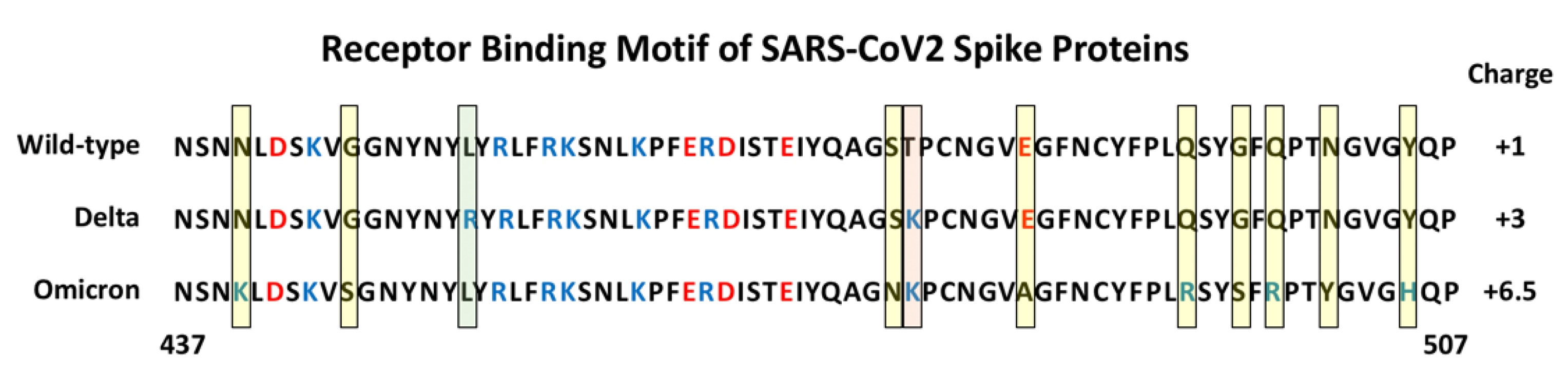
Figure 3. Amino Acid sequence of the Receptor Binding Motif of Spike Proteins. The sequences from position 437 to 507 are shown for the wild-type, and Delta and Omicron variants. The yellow shaded boxes indicate unique mutations in Omicron, the green box indicates a unique mutation in Delta, the pink box indicates a mutation shared by Delta and Omicron. Basic amino acids are colored blue and acidic amino acids red. The estimated charge for each RBM is indicated on the right (+0.5 was assigned to H505 in Omicron Spike).
3. Heparan Sulfate Proteoglycans (HSPGs) and Spike Interactions
HSPGs are a class of macromolecules expressed on cell surfaces and within the extracellular matrix of nearly all mammalian cells and tissues [32][33][34]. HSPGs are characterized by a core protein with covalently attached heparan sulfate (HS) chains. HS is a class of negatively charged linear polysaccharides characterized by repeating disaccharide units of alternating N-substituted glucosamine and hexuronic acid residues subject to selective modification including sulfation of the N-position as well as the C-6 and C-3 O-positions of the glucosamine and the C-2 O-position of the uronic acid. Thus, the 32 (or more) potential unique disaccharide units and their grouping into structural motifs make this class of compounds one of the most information-dense in biology, providing a wide-array of protein-binding sites [35][36][37]. As such, HSPGs have been shown to serve as co-receptors to assist growth factor binding to its tyrosine kinase receptors [38][39][40], and have also been demonstrated to provide attachment and entry sites for a number of viruses [41][42]. The high density of HSPG expression on cell surfaces combined with the large flexible HS chains are features that make this class of molecule ideal as initial attachment sites. Consequently, proteins and viruses can use HSPG to scan large regions of the cell surface to increase the probability of encountering lower density specific receptors. Moreover, as has been well characterized for heparin-binding growth factors, the presence of distinct HS and receptor binding sites on proteins allow HS to contribute to the formation of high-affinity ternary complexes that enhance ligand-receptor binding and signaling [39][40].
Additionally, the SARS-CoV2 Spike protein has been shown to bind HS and heparin (a highly sulfated form of HS expressed by mast cells), which is thought to be reflective of Spike-HS interactions on the cell surface that promote virus entry [43][44][45][46][47]. Indeed, removal of cell surface HS with heparinase has been shown to reduce SARS-CoV2 infection in cell culture models [43][45]. Spike has also been shown to bind to ACE2 and heparin independently in vitro such that a ternary complex can be generated involving all three molecules [44]. Computational docking and molecular dynamics simulations have identified a putative heparin/HS binding domain on Spike, a long positively charged patch that overlaps the S1/S2 junction and extends throughout the RBD [44][48]. Electron microscopic analysis indicates that heparin enhances the open conformation of the RBD suggesting a model whereby Spike binding to HS facilitates Spike-ACE2 interactions through a variety of mechanisms [44]. These findings, combined with the clinical observations that COVID-19 patients treated with heparin show improved outcomes [49][50], have led to a number of studies aimed at targeting Spike-HS interactions as a means of blocking SARS-CoV2 infection [51]. Most of these approaches are based on a model whereby soluble and/or extracellular heparin will bind to Spike protein and competitively inhibit binding to HS on the cell surface [47]. As such, there are more than 100 active clinical trials in the US exploring the use of heparin to treat COVID-19, including trials that are evaluating the delivery of heparin via nasal sprays or nebulizers to protect susceptible tissues against SARS-CoV2 infection (see: https://www.clinicaltrials.gov/ct2/results?cond=COVID-19&term=heparin&cntry=&state=&city=&dist=, accessed on 1 March 2022).
The presence of HS generally presents protein-binding sites in the range of 106–107 per cell such that an initial collision of SARS-CoV2 with a cell might now be predicted to lead to hundreds to thousands of virus encounters with HS binding sites. Moreover, the binding of proteins to HS often show very fast kinetics such that proteins can bind, release, and rebind extensively, preventing their release from the cell surface [52][53]. This rapid binding mechanism provides a means for a protein, or a bound virus particle, to scan a large area of the cell surface to increase the chance of encountering and binding to lower density receptors. Thus, a SARS-CoV2 collision with the cell surface might be expected to result in Spike-HS interactions that ultimately facilitate the formation of stable HS-Spike-ACE2 complexes (Figure 4). Considering this HS co-receptor mechanism, the expression level and binding affinity of ACE2 on the cell surface might not be the key factors that limit SARS-CoV2 infection. Instead, the ability of Spike to interact with HS, or its susceptibility to proteolytic cleavage by TMPRSS2 might be the rate-determining factors.
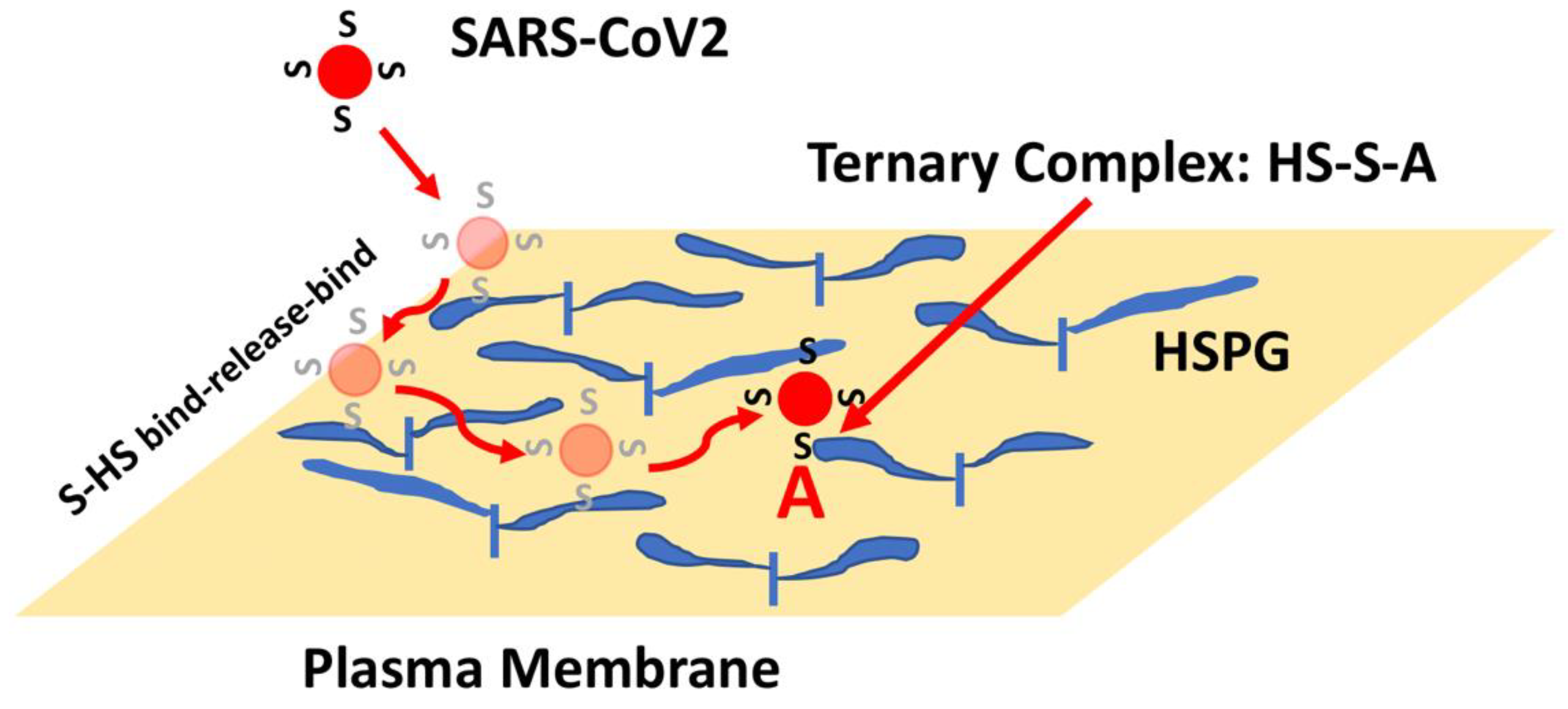
Figure 4. HSPG-mediated Binding of Spike Leads to Increased Binding to ACE2. The initial collision of SARS-CoV2 with the cell surface is likely to result in encounters with high-density and large volume HS chains. Binding of Spike (S) to HS can result in binding, release and re-binding to adjacent HS chains allowing for viral particle movement from one HS chain to another until Spike engages ACE2 (A). Spike-ACE2 interactions are stabilized by the combined binding of HS within a ternary complex.
In addition to HS, there are several reports indicating that Spike binds to other potential co-receptors such as neuropilin 1 and extracellular matrix metalloproteinase inducer (EMMPRIN, also known as basigin and CD147) [54][55][56]. Neuropilin 1 is a transmembrane glycoprotein that function as a co-receptor for semaphorins and vascular endothelial growth factor (VEGF) family members to regulate neurogenesis and angiogenesis [57]. Furthermore, neuropilin 1 has been demonstrated to collaborate with HS to form high affinity multimeric complexes with VEGF and its receptors [58]. Recently, neuropilin 1 was found to potentiate SARS-CoV2 infectivity using HEK-293T cells transiently expressing ACE2, TMPRSS2 and neuropilin 1. Furthermore, an analysis of human COVID-19 autopsies revealed that SARS-CoV2 infected neuropilin 1 positive cells in the nasal cavity [59].
EMMPRIN is an integral plasma membrane glycoprotein that is enriched on the surface of tumor cells where it stimulates the production of several matrix metalloproteinases by adjacent stromal cells [60][61]. EMMPRIN has also been implicated in oral cancers and inflammatory disorders, suggesting a connection to sites of SARS-CoV2 infection. Blocking EMMPRIN with antibodies or through knockdown in Vero E6 cells has been shown to inhibit SARS-CoV2 amplification while the expression of EMMPRIN in non-susceptible BHK-21 cells allows virus entry [56]. In addition, transgenic mice expressing human EMMPRIN showed high viral loads in the lungs, while wild-type mice did not. However, there remains some debate over whether direct Spike interactions with EMMPRIN are responsible for the observed enhanced SARS-CoV2 cell entry [62][63]. Nevertheless, the characterization of secondary Spike binding sites, in addition to ACE2, is revealing a broad range of mechanisms that Spike might employ to mediate SARS-CoV2 infection of cells.
The evidence that HS, neuropilin-1 and EMMPRIN are critical mediators of SARS-CoV2 infectivity via the ability to bind Spike and, in the case of HS, by stabilizing Spike-ACE2 interactions, necessitates that the models presented in Figure 1 and Figure 4 be expanded to capture the full nature of the viral-cell interaction process.
4. Conclusions
Mutations in the Spike protein of SARS-CoV2 have been attributed to the evolution of highly transmissible variants that have led to waves of increased COVID-19 cases, hospitalizations and deaths. Each additional infection provides opportunity for additional mutations in Spike and other viral proteins that may lead to new variants that could elude vaccine-induced or natural immunity. Herein has identified elements of Spike protein functions and additional relevant interactions that contribute to the ability of the virus to gain entry into target cells. The answer to the question of how much better (or worse in terms of causing human disease) Spike can become remains open, yet it is important to recognize that there is a limit. The concepts presented here may help guide studies to identify the aspects of viral infection that are subject to evolutionary improvement. Future viral pandemics are inevitable. Thus, it is critical that the scientific, public health and medical communities work together to foresee what evolution may have in store so that effective vaccines and therapies will be ready.
References
- Walls, A.C.; Park, Y.-J.; Tortorici, M.A.; Wall, A.; McGuire, A.T.; Veesler, D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell 2020, 181, 281–292.e6.
- Huang, Y.; Yang, C.; Xu, X.-F.; Xu, W.; Liu, S.-W. Structural and functional properties of SARS-CoV-2 spike protein: Potential antivirus drug development for COVID-19. Acta Pharmacol. Sin. 2020, 41, 1141–1149.
- Ke, Z.; Oton, J.; Qu, K.; Cortese, M.; Zila, V.; McKeane, L.; Nakane, T.; Zivanov, J.; Neufeldt, C.J.; Cerikan, B.; et al. Structures and distributions of SARS-CoV-2 spike proteins on intact virions. Nature 2020, 588, 498–502.
- Kumavath, R.; Barh, D.; Andrade, B.S.; Imchen, M.; Aburjaile, F.F.; Ch, A.; Rodrigues, D.L.N.; Tiwari, S.; Alzahrani, K.J.; Góes-Neto, A.; et al. The Spike of SARS-CoV-2: Uniqueness and Applications. Front. Immunol. 2021, 12, 663912.
- Zhang, J.; Xiao, T.; Cai, Y.; Chen, B. Structure of SARS-CoV-2 spike protein. Curr. Opin. Virol. 2021, 50, 173–182.
- Laue, M.; Kauter, A.; Hoffmann, T.; Möller, L.; Michel, J.; Nitsche, A. Morphometry of SARS-CoV and SARS-CoV-2 particles in ultrathin plastic sections of infected Vero cell cultures. Sci. Rep. 2021, 11, 3515.
- Yan, R.; Zhang, Y.; Li, Y.; Xia, L.; Guo, Y.; Zhou, Q. Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 2020, 367, 1444–1448.
- Letko, M.; Marzi, A.; Munster, V. Functional assessment of cell entry and receptor usage for SARS-CoV-2 and other lineage B betacoronaviruses. Nat. Microbiol. 2020, 5, 562–569.
- V’Kovski, P.; Kratzel, A.; Steiner, S.; Stalder, H.; Thiel, V. Coronavirus biology and replication: Implications for SARS-CoV-2. Nat. Rev. Microbiol. 2021, 19, 155–170.
- Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.-R.; Zhu, Y.; Li, B.; Huang, C.-L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273.
- Gembardt, F.; Sterner-Kock, A.; Imboden, H.; Spalteholz, M.; Reibitz, F.; Schultheiss, H.-P.; Siems, W.-E.; Walther, T. Organ-specific distribution of ACE2 mRNA and correlating peptidase activity in rodents. Peptides 2005, 26, 1270–1277.
- Hamming, I.; Timens, W.; Bulthuis, M.L.C.; Lely, A.T.; Navis, G.J.; van Goor, H. Tissue distribution of ACE2 protein, the functional receptor for SARS coronavirus. A first step in understanding SARS pathogenesis. J. Pathol. 2004, 203, 631–637.
- Bestle, D.; Heindl, M.R.; Limburg, H.; Van, T.V.L.; Pilgram, O.; Moulton, H.; Stein, D.A.; Hardes, K.; Eickmann, M.; Dolnik, O.; et al. TMPRSS2 and furin are both essential for proteolytic activation of SARS-CoV-2 in human airway cells. Life Sci. Alliance 2020, 3, e202000786.
- Johnson, B.A.; Xie, X.; Bailey, A.L.; Kalveram, B.; Lokugamage, K.G.; Muruato, A.; Zou, J.; Zhang, X.; Juelich, T.; Smith, J.K.; et al. Loss of furin cleavage site attenuates SARS-CoV-2 pathogenesis. Nature 2021, 591, 293–299.
- Xia, S.; Lan, Q.; Su, S.; Wang, X.; Xu, W.; Liu, Z.; Zhu, Y.; Wang, Q.; Lu, L.; Jiang, S. The role of furin cleavage site in SARS-CoV-2 spike protein-mediated membrane fusion in the presence or absence of trypsin. Signal Transduct. Target. Ther. 2020, 5, 92.
- Zou, X.; Chen, K.; Zou, J.; Han, P.; Hao, J.; Han, Z. Single-cell RNA-seq data analysis on the receptor ACE2 expression reveals the potential risk of different human organs vulnerable to 2019-nCoV infection. Front. Med. 2020, 14, 185–192.
- Baker, S.A.; Kwok, S.; Berry, G.J.; Montine, T.J. Angiotensin-converting enzyme 2 (ACE2) expression increases with age in patients requiring mechanical ventilation. PLoS ONE 2021, 16, e0247060.
- Sherman, E.J.; Emmer, B.T. ACE2 protein expression within isogenic cell lines is heterogeneous and associated with distinct transcriptomes. Sci. Rep. 2021, 11, 15900.
- Wrapp, D.; Wang, N.; Corbett, K.S.; Goldsmith, J.A.; Hsieh, C.-L.; Abiona, O.; Graham, B.S.; McLellan, J.S. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020, 367, 1260–1263.
- Sanda, M.; Morrison, L.; Goldman, R. N- and O-Glycosylation of the SARS-CoV-2 Spike Protein. Anal. Chem. 2021, 93, 2003–2009.
- Shajahan, A.; Supekar, N.T.; Gleinich, A.S.; Azadi, P. Deducing the N- and O-glycosylation profile of the spike protein of novel coronavirus SARS-CoV-2. Glycobiology 2020, 30, 981–988.
- Zhang, Y.; Zhao, W.; Mao, Y.; Chen, Y.; Wang, S.; Zhong, Y.; Su, T.; Gong, M.; Du, D.; Lu, X.; et al. Site-specific N-glycosylation Characterization of Recombinant SARS-CoV-2 Spike Proteins. Mol. Cell. Proteom. 2021, 20, 100058.
- Grant, O.C.; Montgomery, D.; Ito, K.; Woods, R.J. Analysis of the SARS-CoV-2 spike protein glycan shield reveals implications for immune recognition. Sci. Rep. 2020, 10, 14991.
- Ismail, A.M.; Elfiky, A.A. SARS-CoV-2 spike behavior in situ: A Cryo-EM images for a better understanding of the COVID-19 pandemic. Signal Transduct. Target. Ther. 2020, 5, 252.
- Casalino, L.; Gaieb, Z.; Goldsmith, J.A.; Hjorth, C.K.; Dommer, A.C.; Harbison, A.M.; Fogarty, C.A.; Barros, E.P.; Taylor, B.C.; McLellan, J.S.; et al. Beyond Shielding: The Roles of Glycans in the SARS-CoV-2 Spike Protein. ACS Cent. Sci. 2020, 6, 1722–1734.
- Sztain, T.; Ahn, S.-H.; Bogetti, A.T.; Casalino, L.; Goldsmith, J.A.; Seitz, E.; McCool, R.S.; Kearns, F.L.; Acosta-Reyes, F.; Maji, S.; et al. A glycan gate controls opening of the SARS-CoV-2 spike protein. Nat. Chem. 2021, 13, 963–968.
- Ali, F.; Kasry, A.; Amin, M. The new SARS-CoV-2 strain shows a stronger binding affinity to ACE2 due to N501Y mutant. Med. Drug Discov. 2021, 10, 100086.
- Laffeber, C.; de Koning, K.; Kanaar, R.; Lebbink, J.H. Experimental Evidence for Enhanced Receptor Binding by Rapidly Spreading SARS-CoV-2 Variants. J. Mol. Biol. 2021, 433, 167058.
- Barton, M.I.; MacGowan, S.A.; Kutuzov, M.A.; Dushek, O.; Barton, G.J.; van der Merwe, P.A. Effects of common mutations in the SARS-CoV-2 Spike RBD and its ligand, the human ACE2 receptor on binding affinity and kinetics. eLife 2021, 10, e70658.
- Kumar, S.; Thambiraja, T.S.; Karuppanan, K.; Subramaniam, G. Omicron and Delta variant of SARS-CoV-2: A comparative computational study of spike protein. J. Med. Virol. 2021, 94, 1641–1649.
- Wu, L.; Zhou, L.; Mo, M.; Liu, T.; Wu, C.; Gong, C.; Lu, K.; Gong, L.; Zhu, W.; Xu, Z. SARS-CoV-2 Omicron RBD shows weaker binding affinity than the currently dominant Delta variant to human ACE2. Signal Transduct. Target. Ther. 2022, 7, 8.
- Iozzo, R.V. Series Introduction: Heparan sulfate proteoglycans: Intricate molecules with intriguing functions. J. Clin. Investig. 2001, 108, 165–167.
- Sarrazin, S.; Lamanna, W.C.; Esko, J.D. Heparan Sulfate Proteoglycans. Cold Spring Harb. Perspect. Biol. 2011, 3, a004952.
- Whitelock, J.; Iozzo, R. Heparan Sulfate: A Complex Polymer Charged with Biological Activity. Chem. Rev. 2005, 105, 2745–2764.
- Mulloy, B.; Linhardt, R.J. Order out of complexity—Protein structures that interact with heparin. Curr. Opin. Struct. Biol. 2001, 11, 623–628.
- Nugent, M.A. Heparin sequencing brings structure to the function of complex oligosaccharides. Proc. Natl. Acad. Sci. USA 2000, 97, 10301–10303.
- Sasisekharan, R.; Venkataraman, G. Heparin and heparan sulfate: Biosynthesis, structure and function. Curr. Opin. Chem. Biol. 2000, 4, 626–631.
- Nugent, M.A.; Zaia, J.; Spencer, J. Heparan sulfate-protein binding specificity. Biochemistry (Moscow) 2013, 78, 726–735.
- Xu, D.; Esko, J.D. Demystifying Heparan Sulfate–Protein Interactions. Annu. Rev. Biochem. 2014, 83, 129–157.
- Capila, I.; Linhardt, R.J. Heparin-protein interactions. Angew. Chem. Int. Ed. Engl. 2002, 41, 391–412.
- Liu, J.; Thorp, S.C. Cell surface heparan sulfate and its roles in assisting viral infections. Med. Res. Rev. 2002, 22, 1–25.
- Cagno, V.; Tseligka, E.D.; Jones, S.T.; Tapparel, C. Heparan Sulfate Proteoglycans and Viral Attachment: True Receptors or Adaptation Bias? Viruses 2019, 11, 596.
- Bermejo-Jambrina, M.; Eder, J.; Kaptein, T.M.; van Hamme, J.L.; Helgers, L.C.; Vlaming, K.E.; Brouwer, P.J.; van Nuenen, A.C.; Spaargaren, M.; de Bree, G.J.; et al. Infection and transmission of SARS-CoV-2 depend on heparan sulfate proteoglycans. EMBO J. 2021, 40, e106765.
- Clausen, T.M.; Sandoval, D.R.; Spliid, C.B.; Pihl, J.; Perrett, H.R.; Painter, C.D.; Narayanan, A.; Majowicz, S.A.; Kwong, E.M.; McVicar, R.N.; et al. SARS-CoV-2 Infection Depends on Cellular Heparan Sulfate and ACE2. Cell 2020, 183, 1043–1057.e15.
- Yue, J.; Jin, W.; Yang, H.; Faulkner, J.; Song, X.; Qiu, H.; Teng, M.; Azadi, P.; Zhang, F.; Linhardt, R.J.; et al. Heparan Sulfate Facilitates Spike Protein-Mediated SARS-CoV-2 Host Cell Invasion and Contributes to Increased Infection of SARS-CoV-2 G614 Mutant and in Lung Cancer. Front. Mol. Biosci. 2021, 8, 649575.
- Zhang, Q.; Chen, C.Z.; Swaroop, M.; Xu, M.; Wang, L.; Lee, J.; Wang, A.Q.; Pradhan, M.; Hagen, N.; Chen, L.; et al. Heparan sulfate assists SARS-CoV-2 in cell entry and can be targeted by approved drugs in vitro. Cell Discov. 2020, 6, 80.
- Mycroft-West, C.J.; Su, D.; Pagani, I.; Rudd, T.R.; Elli, S.; Gandhi, N.S.; Guimond, S.E.; Miller, G.J.; Meneghetti, M.C.Z.; Nader, H.B.; et al. Heparin Inhibits Cellular Invasion by SARS-CoV-2: Structural Dependence of the Interaction of the Spike S1 Receptor-Binding Domain with Heparin. Thromb. Haemost. 2020, 120, 1700–1715.
- Paiardi, G.; Richter, S.; Oreste, P.; Urbinati, C.; Rusnati, M.; Wade, R.C. The binding of heparin to spike glycoprotein inhibits SARS-CoV-2 infection by three mechanisms. J. Biol. Chem. 2021, 298, 101507.
- Tang, N.; Bai, H.; Chen, X.; Gong, J.; Li, D.; Sun, Z. Anticoagulant treatment is associated with decreased mortality in severe coronavirus disease 2019 patients with coagulopathy. J. Thromb. Haemost. 2020, 18, 1094–1099.
- Hippensteel, J.A.; Lariviere, W.B.; Colbert, J.F.; Langouët-Astrié, C.J.; Schmidt, E.P. Heparin as a therapy for COVID-19: Current evidence and future possibilities. Am. J. Physiol. Cell. Mol. Physiol. 2020, 319, L211–L217.
- Lindahl, U.; Li, J. Heparin—An old drug with multiple potential targets in Covid-19 therapy. J. Thromb. Haemost. 2020, 18, 2422–2424.
- Gopalakrishnan, M.; Forsten-Williams, K.; Nugent, M.A.; Täuber, U.C. Effects of Receptor Clustering on Ligand Dissociation Kinetics: Theory and Simulations. Biophys. J. 2005, 89, 3686–3700.
- Chu, C.L.; Buczek-Thomas, J.A.; Nugent, M.A. Heparan sulphate proteoglycans modulate fibroblast growth factor-2 binding through a lipid raft-mediated mechanism. Biochem. J. 2004, 379 Pt 2, 331–341.
- Gudowska-Sawczuk, M.; Mroczko, B. The Role of Neuropilin-1 (NRP-1) in SARS-CoV-2 Infection: Review. J. Clin. Med. 2021, 10, 2772.
- Mayi, B.S.; Leibowitz, J.A.; Woods, A.T.; Ammon, K.A.; Liu, A.E.; Raja, A. The role of Neuropilin-1 in COVID-19. PLoS Pathog. 2021, 17, e1009153.
- Wang, K.; Chen, W.; Zhang, Z.; Deng, Y.; Lian, J.-Q.; Du, P.; Wei, D.; Zhang, Y.; Sun, X.-X.; Gong, L.; et al. CD147-spike protein is a novel route for SARS-CoV-2 infection to host cells. Signal Transduct. Target. Ther. 2020, 5, 283.
- Chaudhary, B.; Khaled, Y.; Ammori, B.J.; Elkord, E. Neuropilin 1: Function and therapeutic potential in cancer. Cancer Immunol. Immunother. 2013, 63, 81–99.
- Teran, M.; Nugent, M.A. Synergistic Binding of Vascular Endothelial Growth Factor-A and Its Receptors to Heparin Selectively Modulates Complex Affinity. J. Biol. Chem. 2015, 290, 16451–16462.
- Cantuti-Castelvetri, L.; Ojha, R.; Pedro, L.D.; Djannatian, M.; Franz, J.; Kuivanen, S.; Van Der Meer, F.; Kallio, K.; Kaya, T.; Anastasina, M.; et al. Neuropilin-1 facilitates SARS-CoV-2 cell entry and infectivity. Science 2020, 370, 856–860.
- Biswas, C.; Zhang, Y.; DeCastro, R.; Guo, H.; Nakamura, T.; Kataoka, H.; Nabeshima, K. The human tumor cell-derived collagenase stimulatory factor (renamed EMMPRIN) is a member of the immunoglobulin superfamily. Cancer Res. 1995, 55, 434–439.
- Toole, B.P. Emmprin (CD147), a cell surface regulator of matrix metalloproteinase production and function. Curr. Top. Dev. Biol. 2003, 54, 371–389.
- Fenizia, C.; Galbiati, S.; Vanetti, C.; Vago, R.; Clerici, M.; Tacchetti, C.; Daniele, T. SARS-CoV-2 Entry: At the Crossroads of CD147 and ACE2. Cells 2021, 10, 1434.
- Ragotte, R.J.; Pulido, D.; Donnellan, F.R.; Hill, M.L.; Gorini, G.; Davies, H.; Brun, J.; McHugh, K.; King, L.D.W.; Skinner, K.; et al. Human Basigin (CD147) Does Not Directly Interact with SARS-CoV-2 Spike Glycoprotein. mSphere 2021, 6, 0064721.
More
Information
Subjects:
Agriculture, Dairy & Animal Science
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.3K
Revisions:
2 times
(View History)
Update Date:
22 Mar 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





