
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Filipa Vinagre Marques Silva | + 1665 word(s) | 1665 | 2021-10-11 10:37:22 | | | |
| 2 | Amina Yu | -8 word(s) | 1657 | 2022-03-15 03:17:53 | | | | |
| 3 | Amina Yu | Meta information modification | 1657 | 2022-03-17 10:02:30 | | |
Video Upload Options
Brettanomyces bruxellensis is a wine spoilage concern in wineries around the world. In order to maintain wine quality during storage and ageing, it is imperative to control and monitor this yeast. Being a fastidious slow growing yeast, which requires 5 to 14 days of incubation for visible growth in agar plates, it is difficult to detect growth (colonies) by conventional agar plate count method. Yeast enumeration by impedance was investigated because previous research using other microorganisms has shown that it is potentially faster than plate counting. The relationship between plate counting and impedance detection times was investigated for Brettanomyces inoculated in red wine samples. A linear relationship between log plate count concentrations and impedance detection times was found. Incubation time was reduced from 120 h down to 0.9 and 57.7 h for samples with 6.7 × 10E7 and 1.8 × 10E2 cfu/mL, respectively, using the ‘indirect’ impedance method. The ‘indirect’ impedance method has the potential to be used by the wine industry to control and monitor the Brettanomyces numbers in wines.
1. Introduction
2. Validation, Advantages and Limitations of the ‘Direct’ and ‘Indirect’ Impedance
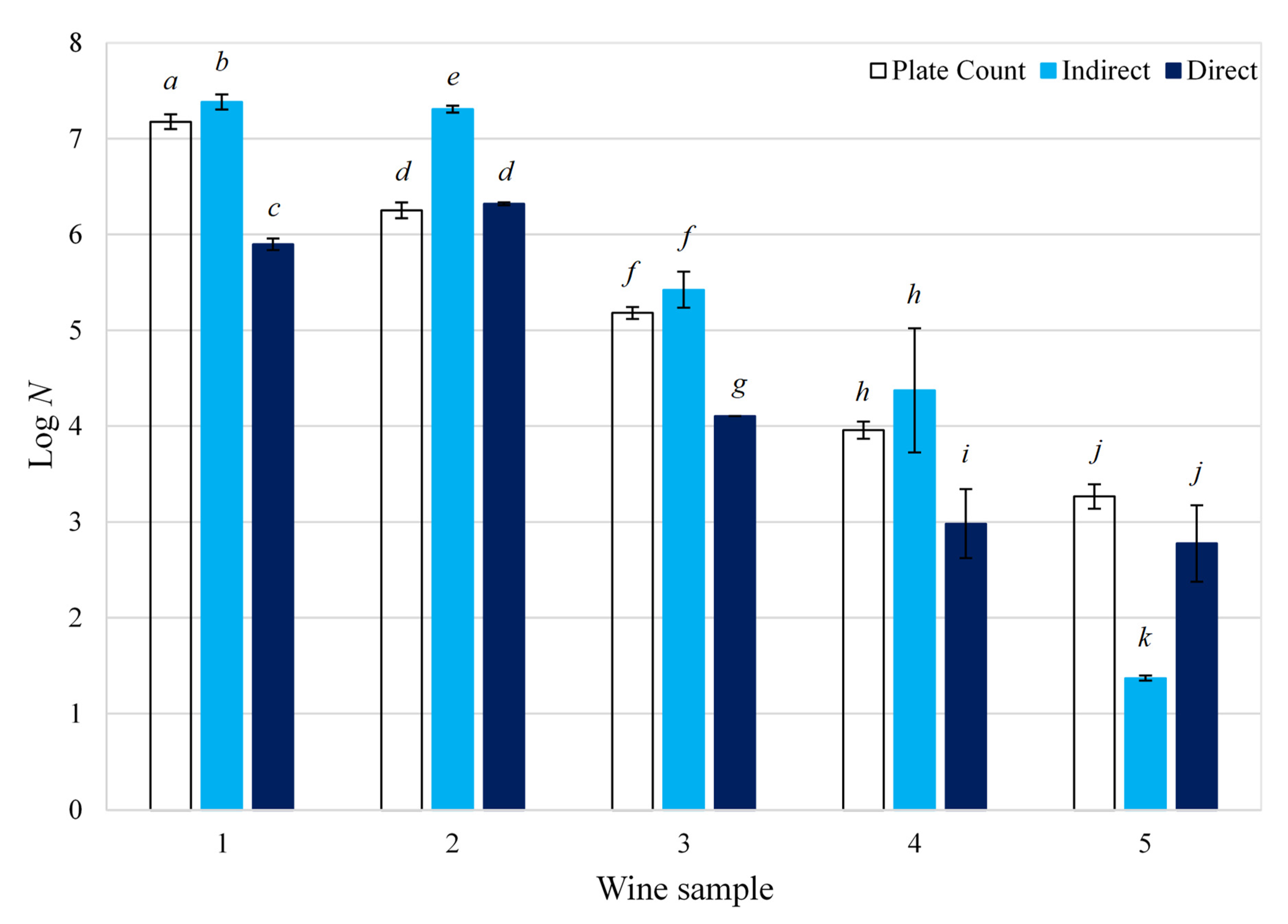
3. Conclusions
References
- Jackson, R.S. Styles and Types of Wine. In Wine Tasting, 2nd ed.; Academic Press: San Diego, CA, USA, 2009; pp. 349–386.
- Han, G.; Ugliano, M.; Currie, B.; Vidal, S.; Diéval, J.-B.; Waterhouse, A.L. Influence of closure, phenolic levels and microoxygenation on Cabernet Sauvignon wine composition after 5 years’ bottle storage. J. Sci. Food Agric. 2014, 95, 36–43.
- Oelofse, A.; Pretorius, I.; Du Toit, M. Significance of Brettanomyces and Dekkera during Winemaking: A Synoptic Review. S. Afr. J. Enol. Vitic. 2016, 29, 128–144.
- Wedral, D.; Shewfelt, R.; Frank, J. The challenge of Brettanomyces in wine. LWT Food Sci. Technol. 2010, 43, 1474–1479.
- Loureiro, V.; Malfeito-Ferreira, M. Dekkera/Brettanomyces spp. In Food Spoilage Microorganisms; Blackburn, C.W., Ed.; Woodhead Publishing: Cambridge, UK, 2006; pp. 354–398.
- Zuehlke, J.M.; Petrova, B.; Edwards, C.G. Advances in the control of wine spoilage by Zygosaccharomyces and Dekkera/Brettanomyces. Annu. Rev. Food Sci. Technol. 2013, 4, 57–78.
- Guerrero, R.F.; Cantos-Villar, E. Demonstrating the efficiency of sulphur dioxide replacements in wine: A parameter review. Trends Food Sci. Technol. 2015, 42, 27–43.
- Longin, C.; Degueurce, C.; Julliat, F.; Guilloux-Benatier, M.; Rousseaux, S.; Alexandre, H. Efficiency of population-dependent sulfite against Brettanomyces bruxellensis in red wine. Food Res. Int. 2016, 89, 620–630.
- Cocolin, L.; Rantsiou, K.; Iacumin, L.; Zironi, R.; Comi, G. Molecular Detection and Identification of Brettanomyces/Dekkera bruxellensis and Brettanomyces/Dekkera anomalus in Spoiled Wines. Appl. Environ. Microbiol. 2004, 70, 1347–1355.
- Malacrinò, P.; Zapparoli, G.; Torriani, S.; Dellaglio, F. Rapid detection of viable yeasts and bacteria in wine by flow cytometry. J. Microbiol. Methods 2001, 45, 127–134.
- Dupont, J.; Dumont, F.; Menanteau, C.; Pommepuy, M. Calibration of the impedance method for rapid quantitative estimation of Escherichia coli in live marine bivalve molluscs. J. Appl. Microbiol. 2004, 96, 894–902.
- Zhu, S.; Schnell, S.; Fischer, M. Rapid detection of Cronobacter spp. with a method combining impedance technology and rRNA based lateral flow assay. Int. J. Food Microbiol. 2012, 159, 54–58.
- van Wyk, S.; Silva, F.V.M. Impedance technology reduces the enumeration time of Brettanomyces yeast during beer fermentation. Biotechnol. J. 2016, 11, 1667–1672.
- Ruiz-Moreno, M.J.; Raposo, R.; Cayuela, J.M.; Zafrilla, P.; Piñeiro, Z.; Rojas, J.M.M.; Mulero, J.; Puertas, B.; Girón, F.; Guerrero, R.F.; et al. Valorization of grape stems. Ind. Crop. Prod. 2015, 63, 152–157.
- Ruiz-Moreno, M.J.; Raposo, R.; Moreno-Rojas, J.M.; Zafrilla, P.; Cayuela, J.M.; Mulero, J.; Puertas, B.; Guerrero, R.F.; Piñeiro, Z.; Giron, F.; et al. Efficacy of olive oil mill extract in replacing sulfur dioxide in wine model. LWT Food Sci. Technol. 2015, 61, 117–123.
- Agata, L.; Jan, P. Production of fermented goat beverage using a mixed starter culture of lactic acid bacteria and yeasts. Eng. Life Sci. 2012, 12, 486–493.
- Walker, K.; Ripandelli, N.; Flint, S. Rapid enumeration of Bifidobacterium lactis in milk powders using impedance. Int. Dairy J. 2005, 15, 183–188.
- Alkhafaji, S.R.; Farid, M. An investigation on pulsed electric fields technology using new treatment chamber design. Innov. Food Sci. Emerg. Technol. 2007, 8, 205–212.
- Steensels, J.; Daenen, L.; Malcorps, P.; Derdelinckx, G.; Verachtert, H.; Verstrepen, K.J. Brettanomyces yeasts—From spoilage organisms to valuable contributors to industrial fermentations. Int. J. Food Microbiol. 2015, 206, 24–38.





