
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Vijay Kumar | + 3082 word(s) | 3082 | 2022-03-08 07:39:11 | | | |
| 2 | Beatrix Zheng | + 416 word(s) | 3498 | 2022-03-09 02:54:16 | | |
Video Upload Options
The Ventx family is one of the subfamilies of the ANTP (antennapedia) superfamily and belongs to the NK-like (NKL) subclass. Ventx is a homeobox transcription factor and has a DNA-interacting domain that is evolutionarily conserved throughout vertebrates. It has been extensively studied in Xenopus, zebrafish, and humans. The Ventx family contains transcriptional repressors widely involved in embryonic development and tumorigenesis in vertebrates. Several studies have documented that the Ventx family inhibited dorsal mesodermal formation, neural induction, and head formation in Xenopus and zebrafish. Moreover, Ventx2.2 showed functional similarities to Nanog and Barx1, leading to pluripotency and neural-crest migration in vertebrates. Among them, Ventx protein is an orthologue of the Ventx family in humans. Studies have demonstrated that human Ventx was strongly associated with myeloid-cell differentiation and acute myeloid leukemia.
1. Introduction
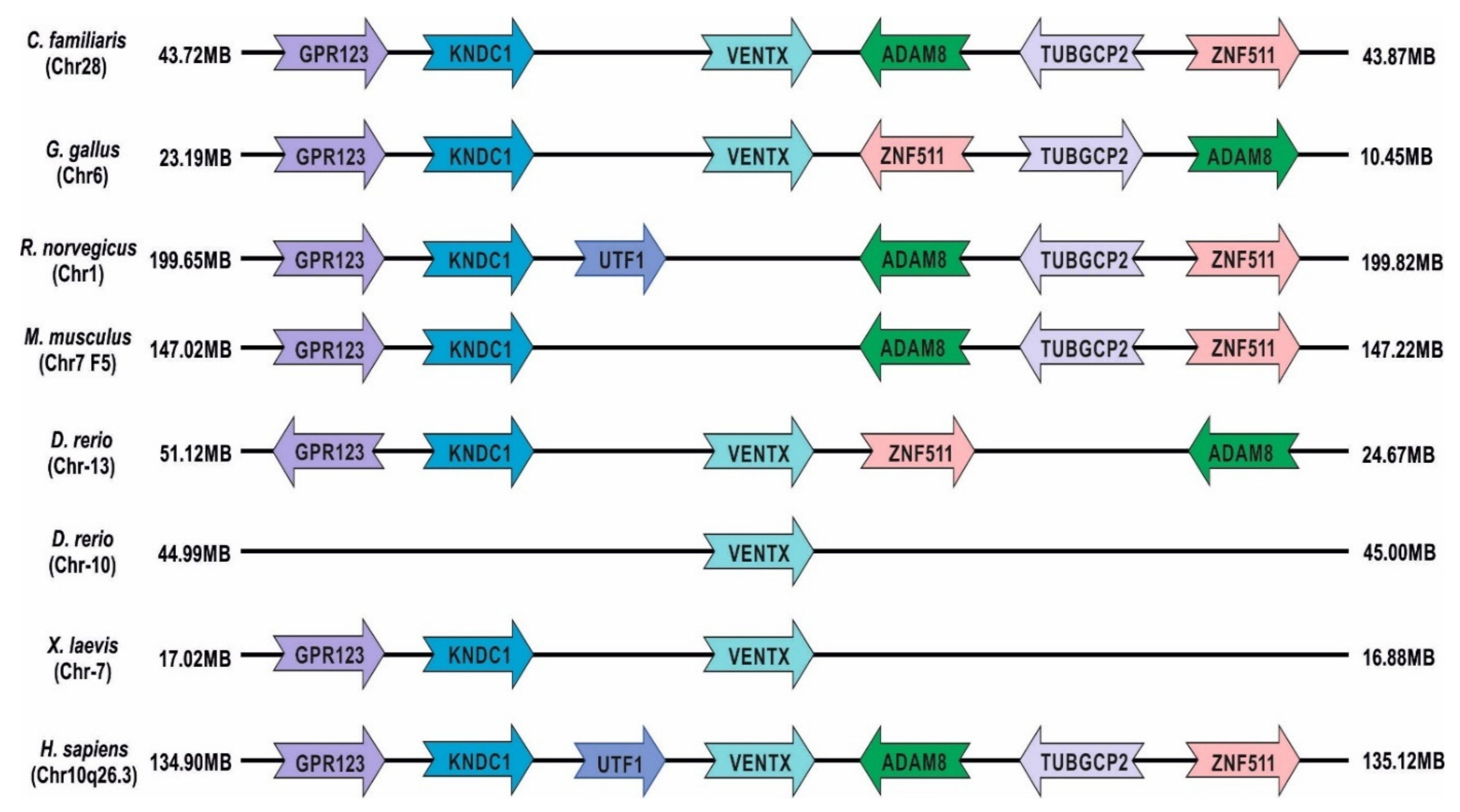
2. Human Ventx (VENT-like Homeobox Protein 2)
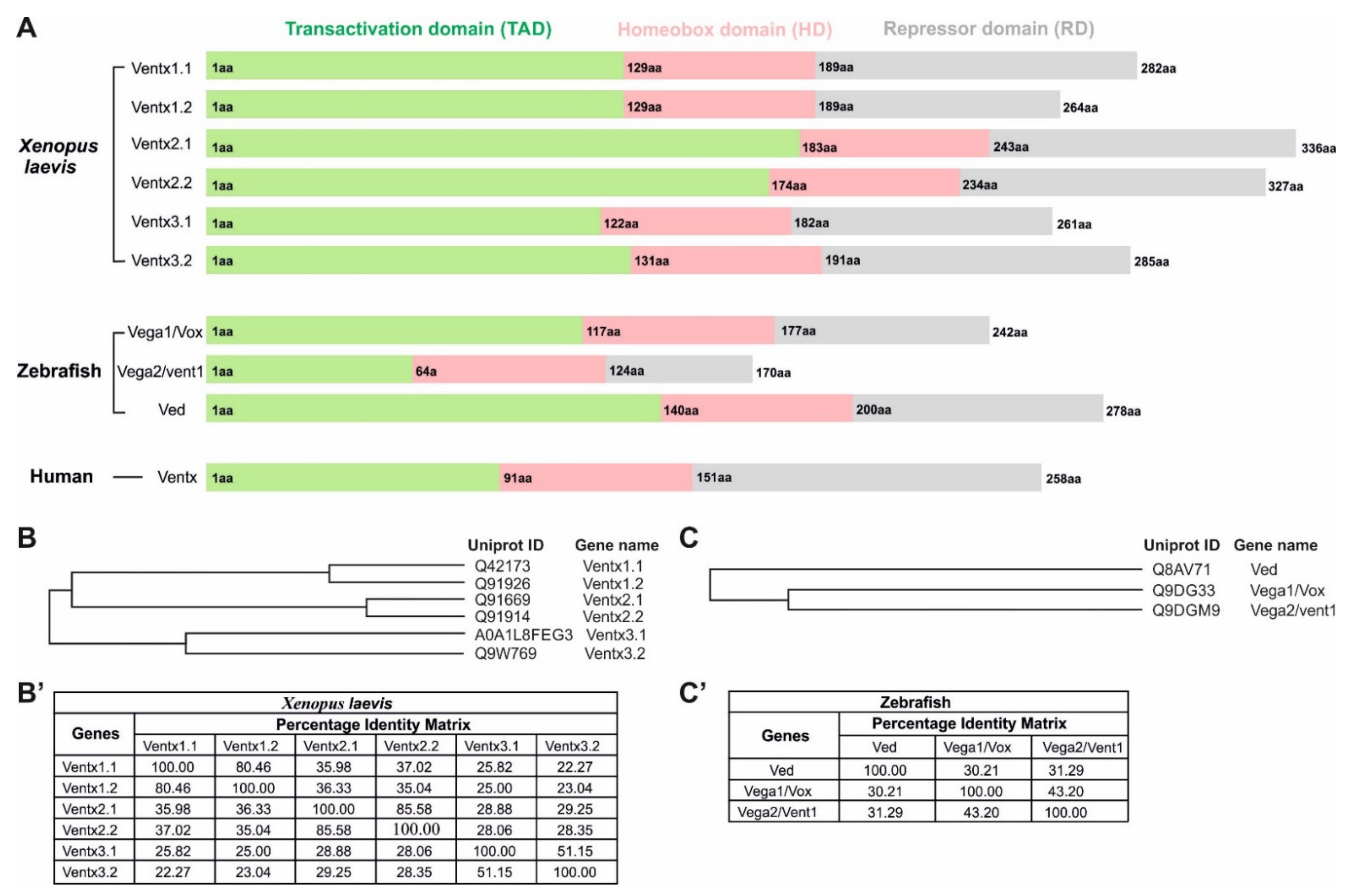
| Names | Chromosomal Localization |
Similarity with Human Ventx (%) | Developmental Roles | References |
|---|---|---|---|---|
| Xenopus laevis | ||||
| Venxt1.1/PV.1/Ventx1.1a/Ventx1.1b | Chr7L:21,842,291–21,845,156 Chr7S:16,920,567–16,923,476 |
59.02 | Neural repressor, Spemann-organizer inhibitor, embryonic ventralization, hematopoiesis, axial patterning | [30][33][56] |
| Ventx1.2/Xvent1/Vent1/Vent-1/Xvent-1 | Chr7L:21,802,386–21,807,247 Chr7S:16,883,662–16,887,492 |
44.95 | Neural repressor, Spemann-organizer inhibitor, embryonic ventralization, axial patterning | [46][47][57] |
| Ventx2.1a/Ventx2.1b, Vox1, Vox, Xvent2 | Chr7L:21,859,318–21,861,104 Chr7S:16,932,131–16,933,936 |
66.67 | Neural repressor, Spemann-organizer inhibitor, embryonic ventralization, Ventx1.2 and ventx1.1 activator, hematopoiesis, axial patterning | [12][15][58] |
| Ventx2.2/Xom */Xvent-2/Xbr-1 | Chr7L:21,830,246–21,832,823 Chr7S:16,908,607–16,911,052 |
66.67 | Neural repressor, VMZ activator, induce GSK3β expression, axial patterning, pluripotency marker, neural crest migration | [27][28][59][60][61] |
| Ventx3.1/vent3 | Chr7L-NA Chr7S:16,889,505–16,895,419 |
39.1 | NA | [62][63] |
| Ventx3.2/Vex1 | Chr7L:21,784,572–21,787,885 Chr7S:16,870,294–16,873,640 |
51.32 | Neural repressor, VMZ activator, axial patterning | [37][64] |
| Zebrafish | ||||
| Vega1/Vox | Chr13:50,621,929–50,624,154 | 55 | Neural repressor, VMZ, activator, axial patterning | [34] |
| Vega2/Vent1 | Chr13:50,609,716–50,614,718 | 55 | Neural repressor, VMZ activator, axial patterning | [45] |
| Ved | Chr10:44,997,062–45,002,196 | 49.12 | Neural repressor, VMZ activator, axial patterning | [38] |
| Human | ||||
| Ventx | Chr10:133,237,855–133,241,928 | 100 | Acute myeloid leukemia, cellular senescence, myeloid and macrophage differentiation, dendritic cell differentiation, immunosuppressor, apoptotic activator | [17][18] |
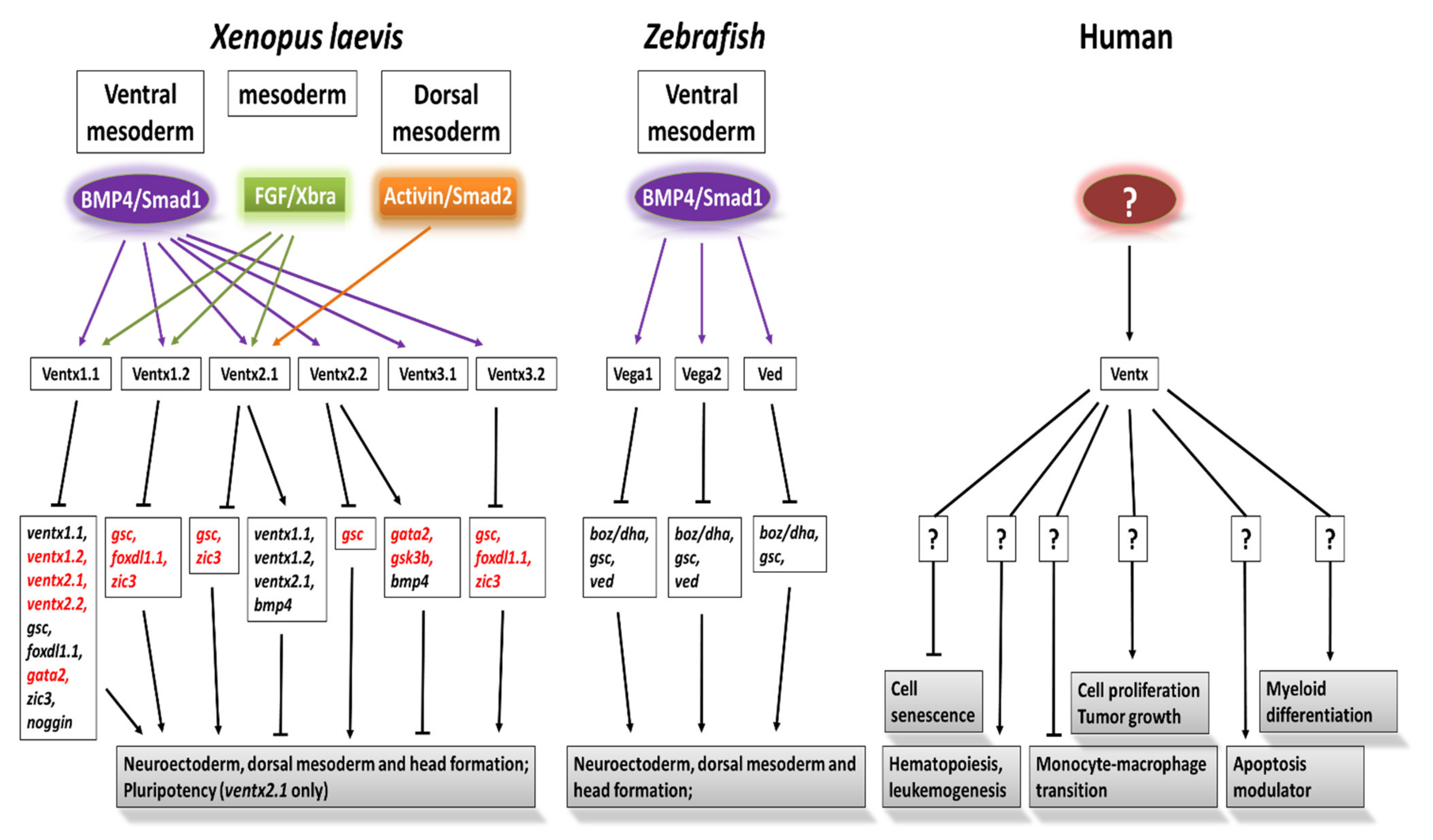
Nanog (Ventx-like) Is Considered an Oncogene in Humans
3. Current Insights
References
- Grainger, R.M. Xenopus tropicalis as a model organism for genetics and genomics: Past, present, and future. Methods Mol. Biol. 2012, 917, 3–15.
- Blum, M.; Ott, T. Xenopus: An Undervalued Model Organism to Study and Model Human Genetic Disease. Cells Tissues Organs 2018, 205, 303–313.
- Jia, Y.; Liu, X. Polyploidization and pseudogenization in allotetraploid frog Xenopus laevis promote the evolution of aquaporin family in higher vertebrates. BMC Genom. 2020, 21, 525.
- Hellsten, U.; Khokha, M.K.; Grammer, T.C.; Harland, R.M.; Richardson, P.; Rokhsar, D.S. Accelerated gene evolution and subfunctionalization in the pseudotetraploid frog Xenopus laevis. BMC Biol. 2007, 5, 31.
- Elurbe, D.M.; Paranjpe, S.S.; Georgiou, G.; van Kruijsbergen, I.; Bogdanovic, O.; Gibeaux, R.; Heald, R.; Lister, R.; Huynen, M.A.; van Heeringen, S.J.; et al. Regulatory remodeling in the allo-tetraploid frog Xenopus laevis. Genome Biol. 2017, 18, 198.
- Session, A.M.; Uno, Y.; Kwon, T.; Chapman, J.A.; Toyoda, A.; Takahashi, S.; Fukui, A.; Hikosaka, A.; Suzuki, A.; Kondo, M.; et al. Genome evolution in the allotetraploid frog Xenopus laevis. Nature 2016, 538, 336–343.
- Pollet, N.; Mazabraud, A. Insights from Xenopus genomes. Genome Dyn. 2006, 2, 138–153.
- Gehring, W.J.; Affolter, M.; Burglin, T. Homeodomain proteins. Annu. Rev. Biochem. 1994, 63, 487–526.
- Burglin, T.R.; Affolter, M. Homeodomain proteins: An update. Chromosoma 2016, 125, 497–521.
- Burglin, T.R. Homeodomain subtypes and functional diversity. Subcell Biochem. 2011, 52, 95–122.
- Holland, P.W.; Booth, H.A.; Bruford, E.A. Classification and nomenclature of all human homeobox genes. BMC Biol. 2007, 5, 47.
- Henningfeld, K.A.; Friedle, H.; Rastegar, S.; Knochel, W. Autoregulation of Xvent-2B; direct interaction and functional cooperation of Xvent-2 and Smad1. J. Biol. Chem. 2002, 277, 2097–2103.
- Kumar, S.; Umair, Z.; Yoon, J.; Lee, U.; Kim, S.C.; Park, J.B.; Lee, J.Y.; Kim, J. Xbra and Smad-1 cooperate to activate the transcription of neural repressor ventx1.1 in Xenopus embryos. Sci. Rep. 2018, 8, 11391.
- Lee, H.S.; Lee, S.Y.; Lee, H.; Hwang, Y.S.; Cha, S.W.; Park, S.; Lee, J.Y.; Park, J.B.; Kim, S.; Park, M.J.; et al. Direct response elements of BMP within the PV.1A promoter are essential for its transcriptional regulation during early Xenopus development. PLoS ONE 2011, 6, e22621.
- Lee, H.S.; Park, M.J.; Lee, S.Y.; Hwang, Y.S.; Lee, H.; Roh, D.H.; Kim, J.I.; Park, J.B.; Lee, J.Y.; Kung, H.F.; et al. Transcriptional regulation of Xbr-1a/Xvent-2 homeobox gene: Analysis of its promoter region. Biochem. Biophys. Res. Commun. 2002, 298, 815–823.
- Faial, T.; Bernardo, A.S.; Mendjan, S.; Diamanti, E.; Ortmann, D.; Gentsch, G.E.; Mascetti, V.L.; Trotter, M.W.; Smith, J.C.; Pedersen, R.A. Brachyury and SMAD signalling collaboratively orchestrate distinct mesoderm and endoderm gene regulatory networks in differentiating human embryonic stem cells. Development 2015, 142, 2121–2135.
- Moretti, P.A.; Davidson, A.J.; Baker, E.; Lilley, B.; Zon, L.I.; D’Andrea, R.J. Molecular cloning of a human Vent-like homeobox gene. Genomics 2001, 76, 21–29.
- Rawat, V.P.; Arseni, N.; Ahmed, F.; Mulaw, M.A.; Thoene, S.; Heilmeier, B.; Sadlon, T.; D’Andrea, R.J.; Hiddemann, W.; Bohlander, S.K.; et al. The vent-like homeobox gene VENTX promotes human myeloid differentiation and is highly expressed in acute myeloid leukemia. Proc. Natl. Acad. Sci. USA 2010, 107, 16946–16951.
- Genthe, J.R.; Min, J.; Farmer, D.M.; Shelat, A.A.; Grenet, J.A.; Lin, W.; Finkelstein, D.; Vrijens, K.; Chen, T.; Guy, R.K.; et al. Ventromorphins: A New Class of Small Molecule Activators of the Canonical BMP Signaling Pathway. ACS Chem. Biol. 2017, 12, 2436–2447.
- Little, S.C.; Mullins, M.C. Extracellular modulation of BMP activity in patterning the dorsoventral axis. Birth Defects Res. C Embryo Today 2006, 78, 224–242.
- Stickney, H.L.; Imai, Y.; Draper, B.; Moens, C.; Talbot, W.S. Zebrafish bmp4 functions during late gastrulation to specify ventroposterior cell fates. Dev. Biol. 2007, 310, 71–84.
- Friedle, H.; Knochel, W. Cooperative interaction of Xvent-2 and GATA-2 in the activation of the ventral homeobox gene Xvent-1B. J. Biol. Chem. 2002, 277, 23872–23881.
- Messenger, N.J.; Kabitschke, C.; Andrews, R.; Grimmer, D.; Nunez Miguel, R.; Blundell, T.L.; Smith, J.C.; Wardle, F.C. Functional specificity of the Xenopus T-domain protein Brachyury is conferred by its ability to interact with Smad1. Dev. Cell 2005, 8, 599–610.
- Onichtchouk, D.; Gawantka, V.; Dosch, R.; Delius, H.; Hirschfeld, K.; Blumenstock, C.; Niehrs, C. The Xvent-2 homeobox gene is part of the BMP-4 signalling pathway controlling dorsoventral patterning of Xenopus mesoderm. Development 1996, 122, 3045–3053.
- Schuler-Metz, A.; Knochel, S.; Kaufmann, E.; Knochel, W. The homeodomain transcription factor Xvent-2 mediates autocatalytic regulation of BMP-4 expression in Xenopus embryos. J. Biol. Chem. 2000, 275, 34365–34374.
- Trindade, M.; Tada, M.; Smith, J.C. DNA-binding specificity and embryological function of Xom (Xvent-2). Dev. Biol. 1999, 216, 442–456.
- Ladher, R.; Mohun, T.J.; Smith, J.C.; Snape, A.M. Xom: A Xenopus homeobox gene that mediates the early effects of BMP-4. Development 1996, 122, 2385–2394.
- Papalopulu, N.; Kintner, C. A Xenopus gene, Xbr-1, defines a novel class of homeobox genes and is expressed in the dorsal ciliary margin of the eye. Dev. Biol. 1996, 174, 104–114.
- Zhong, Y.F.; Holland, P.W. The dynamics of vertebrate homeobox gene evolution: Gain and loss of genes in mouse and human lineages. BMC Evol. Biol. 2011, 11, 169.
- Ault, K.T.; Dirksen, M.L.; Jamrich, M. A novel homeobox gene PV.1 mediates induction of ventral mesoderm in Xenopus embryos. Proc. Natl. Acad. Sci. USA 1996, 93, 6415–6420.
- Dale, L.; Wardle, F.C. A gradient of BMP activity specifies dorsal-ventral fates in early Xenopus embryos. Semin. Cell Dev. Biol. 1999, 10, 319–326.
- Gentner, E.; Vegi, N.M.; Mulaw, M.A.; Mandal, T.; Bamezai, S.; Claus, R.; Tasdogan, A.; Quintanilla-Martinez, L.; Grunenberg, A.; Dohner, K.; et al. VENTX induces expansion of primitive erythroid cells and contributes to the development of acute myeloid leukemia in mice. Oncotarget 2016, 7, 86889–86901.
- Hwang, Y.S.; Seo, J.J.; Cha, S.W.; Lee, H.S.; Lee, S.Y.; Roh, D.H.; Kung Hf, H.F.; Kim, J.; Ja Park, M. Antimorphic PV.1 causes secondary axis by inducing ectopic organizer. Biochem. Biophys. Res. Commun. 2002, 292, 1081–1086.
- Kawahara, A.; Wilm, T.; Solnica-Krezel, L.; Dawid, I.B. Antagonistic role of vega1 and bozozok/dharma homeobox genes in organizer formation. Proc. Natl. Acad. Sci. USA 2000, 97, 12121–12126.
- Lee, S.Y.; Lim, S.K.; Cha, S.W.; Yoon, J.; Lee, S.H.; Lee, H.S.; Park, J.B.; Lee, J.Y.; Kim, S.C.; Kim, J. Inhibition of FGF signaling converts dorsal mesoderm to ventral mesoderm in early Xenopus embryos. Differentiation 2011, 82, 99–107.
- Lee, S.Y.; Yoon, J.; Lee, M.H.; Jung, S.K.; Kim, D.J.; Bode, A.M.; Kim, J.; Dong, Z. The role of heterodimeric AP-1 protein comprised of JunD and c-Fos proteins in hematopoiesis. J. Biol. Chem. 2012, 287, 31342–31348.
- Shapira, E.; Marom, K.; Yelin, R.; Levy, A.; Fainsod, A. A role for the homeobox gene Xvex-1 as part of the BMP-4 ventral signaling pathway. Mech. Dev. 1999, 86, 99–111.
- Shimizu, T.; Yamanaka, Y.; Nojima, H.; Yabe, T.; Hibi, M.; Hirano, T. A novel repressor-type homeobox gene, ved, is involved in dharma/bozozok-mediated dorsal organizer formation in zebrafish. Mech. Dev. 2002, 118, 125–138.
- De Robertis, E.M.; Kuroda, H. Dorsal-ventral patterning and neural induction in Xenopus embryos. Annu. Rev. Cell Dev. Biol. 2004, 20, 285–308.
- Hemmati-Brivanlou, A.; Thomsen, G.H. Ventral mesodermal patterning in Xenopus embryos: Expression patterns and activities of BMP-2 and BMP-4. Dev. Genet. 1995, 17, 78–89.
- Maeno, M.; Mead, P.E.; Kelley, C.; Xu, R.H.; Kung, H.F.; Suzuki, A.; Ueno, N.; Zon, L.I. The role of BMP-4 and GATA-2 in the induction and differentiation of hematopoietic mesoderm in Xenopus laevis. Blood 1996, 88, 1965–1972.
- Xu, R.H.; Ault, K.T.; Kim, J.; Park, M.J.; Hwang, Y.S.; Peng, Y.; Sredni, D.; Kung, H. Opposite effects of FGF and BMP-4 on embryonic blood formation: Roles of PV.1 and GATA-2. Dev. Biol. 1999, 208, 352–361.
- Friedle, H.; Rastegar, S.; Paul, H.; Kaufmann, E.; Knochel, W. Xvent-1 mediates BMP-4-induced suppression of the dorsal-lip-specific early response gene XFD-1’ in Xenopus embryos. EMBO J. 1998, 17, 2298–2307.
- Imai, Y.; Gates, M.A.; Melby, A.E.; Kimelman, D.; Schier, A.F.; Talbot, W.S. The homeobox genes vox and vent are redundant repressors of dorsal fates in zebrafish. Development 2001, 128, 2407–2420.
- Kawahara, A.; Wilm, T.; Solnica-Krezel, L.; Dawid, I.B. Functional interaction of vega2 and goosecoid homeobox genes in zebrafish. Genesis 2000, 28, 58–67.
- Gawantka, V.; Delius, H.; Hirschfeld, K.; Blumenstock, C.; Niehrs, C. Antagonizing the Spemann organizer: Role of the homeobox gene Xvent-1. EMBO J. 1995, 14, 6268–6279.
- Onichtchouk, D.; Glinka, A.; Niehrs, C. Requirement for Xvent-1 and Xvent-2 gene function in dorsoventral patterning of Xenopus mesoderm. Development 1998, 125, 1447–1456.
- Scerbo, P.; Girardot, F.; Vivien, C.; Markov, G.V.; Luxardi, G.; Demeneix, B.; Kodjabachian, L.; Coen, L. Ventx factors function as Nanog-like guardians of developmental potential in Xenopus. PLoS ONE 2012, 7, e36855.
- Scerbo, P.; Marchal, L.; Kodjabachian, L. Lineage commitment of embryonic cells involves MEK1-dependent clearance of pluripotency regulator Ventx2. eLife 2017, 6, e21526.
- Wu, X.; Gao, H.; Bleday, R.; Zhu, Z. Homeobox transcription factor VentX regulates differentiation and maturation of human dendritic cells. J. Biol. Chem. 2014, 289, 14633–14643.
- Wu, X.; Gao, H.; Ke, W.; Giese, R.W.; Zhu, Z. The homeobox transcription factor VentX controls human macrophage terminal differentiation and proinflammatory activation. J. Clin. Investig. 2011, 121, 2599–2613.
- Gartel, A.L.; Radhakrishnan, S.K. Lost in transcription: p21 repression, mechanisms, and consequences. Cancer Res. 2005, 65, 3980–3985.
- Deng, C.; Zhang, P.; Harper, J.W.; Elledge, S.J.; Leder, P. Mice lacking p21CIP1/WAF1 undergo normal development, but are defective in G1 checkpoint control. Cell 1995, 82, 675–684.
- Wu, X.; Gao, H.; Ke, W.; Hager, M.; Xiao, S.; Freeman, M.R.; Zhu, Z. VentX trans-activates p53 and p16ink4a to regulate cellular senescence. J. Biol. Chem. 2011, 286, 12693–12701.
- Trompouki, E.; Bowman, T.V.; Lawton, L.N.; Fan, Z.P.; Wu, D.C.; DiBiase, A.; Martin, C.S.; Cech, J.N.; Sessa, A.K.; Leblanc, J.L.; et al. Lineage regulators direct BMP and Wnt pathways to cell-specific programs during differentiation and regeneration. Cell 2011, 147, 577–589.
- Hwang, Y.S.; Lee, H.S.; Roh, D.H.; Cha, S.; Lee, S.Y.; Seo, J.J.; Kim, J.; Park, M.J. Active repression of organizer genes by C-terminal domain of PV.1. Biochem. Biophys. Res. Commun. 2003, 308, 79–86.
- Sander, V.; Reversade, B.; De Robertis, E.M. The opposing homeobox genes Goosecoid and Vent1/2 self-regulate Xenopus patterning. EMBO J. 2007, 26, 2955–2965.
- Melby, A.E.; Clements, W.K.; Kimelman, D. Regulation of dorsal gene expression in Xenopus by the ventralizing homeodomain gene Vox. Dev. Biol. 1999, 211, 293–305.
- Gao, H.; Wu, B.; Giese, R.; Zhu, Z. Xom interacts with and stimulates transcriptional activity of LEF1/TCFs: Implications for ventral cell fate determination during vertebrate embryogenesis. Cell Res. 2007, 17, 345–356.
- Wu, B.; Gao, H.; Le, Y.; Wu, X.; Zhu, Z. Xom induces proteolysis of beta-catenin through GSK3beta-mediated pathway. FEBS Lett. 2018, 592, 299–309.
- Xie, Y.; Liu, C. Xom, a ventralizing factor, regulates beta-catenin levels and cell fate. FEBS Lett. 2018, 592, 297–298.
- Leibovich, A.; Kot-Leibovich, H.; Ben-Zvi, D.; Fainsod, A. ADMP controls the size of Spemann’s organizer through a network of self-regulating expansion-restriction signals. BMC Biol. 2018, 16, 13.
- Watanabe, M.; Yasuoka, Y.; Mawaribuchi, S.; Kuretani, A.; Ito, M.; Kondo, M.; Ochi, H.; Ogino, H.; Fukui, A.; Taira, M.; et al. Conservatism and variability of gene expression profiles among homeologous transcription factors in Xenopus laevis. Dev. Biol. 2017, 426, 301–324.
- Shapira, E.; Marom, K.; Levy, V.; Yelin, R.; Fainsod, A. The Xvex-1 antimorph reveals the temporal competence for organizer formation and an early role for ventral homeobox genes. Mech. Dev. 2000, 90, 77–87.
- Le, Y.; Gao, H.; Bleday, R.; Zhu, Z. The homeobox protein VentX reverts immune suppression in the tumor microenvironment. Nat. Commun. 2018, 9, 2175.
- Le, Y.; Gao, H.; Richards, W.; Zhao, L.; Bleday, R.; Clancy, T.; Zhu, Z. VentX expression in tumor-associated macrophages promotes phagocytosis and immunity against pancreatic cancers. JCI Insight 2020, 5, e137088.
- Ino, Y.; Yamazaki-Itoh, R.; Shimada, K.; Iwasaki, M.; Kosuge, T.; Kanai, Y.; Hiraoka, N. Immune cell infiltration as an indicator of the immune microenvironment of pancreatic cancer. Br. J. Cancer 2013, 108, 914–923.
- Gao, H.; Wu, X.; Sun, Y.; Zhou, S.; Silberstein, L.E.; Zhu, Z. Suppression of homeobox transcription factor VentX promotes expansion of human hematopoietic stem/multipotent progenitor cells. J. Biol. Chem. 2012, 287, 29979–29987.
- Elagib, K.E.; Goldfarb, A.N. Oncogenic pathways of AML1-ETO in acute myeloid leukemia: Multifaceted manipulation of marrow maturation. Cancer Lett. 2007, 251, 179–186.
- Gao, H.; Le, Y.; Wu, X.; Silberstein, L.E.; Giese, R.W.; Zhu, Z. VentX, a novel lymphoid-enhancing factor/T-cell factor-associated transcription repressor, is a putative tumor suppressor. Cancer Res. 2010, 70, 202–211.
- Vargas, A.C.; Gray, L.A.; White, C.L.; Maclean, F.M.; Grimison, P.; Ardakani, N.M.; Bonar, F.; Algar, E.M.; Cheah, A.L.; Russell, P.; et al. Genome wide methylation profiling of selected matched soft tissue sarcomas identifies methylation changes in metastatic and recurrent disease. Sci. Rep. 2021, 11, 667.
- Gao, H.; Wu, B.; Le, Y.; Zhu, Z. Homeobox protein VentX induces p53-independent apoptosis in cancer cells. Oncotarget 2016, 7, 39719–39729.
- Chambers, I.; Silva, J.; Colby, D.; Nichols, J.; Nijmeijer, B.; Robertson, M.; Vrana, J.; Jones, K.; Grotewold, L.; Smith, A. Nanog safeguards pluripotency and mediates germline development. Nature 2007, 450, 1230–1234.
- Grubelnik, G.; Bostjancic, E.; Pavlic, A.; Kos, M.; Zidar, N. NANOG expression in human development and cancerogenesis. Exp Biol. Med. 2020, 245, 456–464.
- Nagel, S.; Scherr, M.; MacLeod, R.A.F.; Pommerenke, C.; Koeppel, M.; Meyer, C.; Kaufmann, M.; Dallmann, I.; Drexler, H.G. NKL homeobox gene activities in normal and malignant myeloid cells. PLoS ONE 2019, 14, e0226212.
- Brandner, S. Nanog, Gli, and p53: A new network of stemness in development and cancer. EMBO J. 2010, 29, 2475–2476.
- Zbinden, M.; Duquet, A.; Lorente-Trigos, A.; Ngwabyt, S.N.; Borges, I.; Ruiz i Altaba, A. NANOG regulates glioma stem cells and is essential in vivo acting in a cross-functional network with GLI1 and p53. EMBO J. 2010, 29, 2659–2674.
- Wang, S.; Fan, H.; Xu, J.; Zhao, E. Prognostic implication of NOTCH1 in early stage oral squamous cell cancer with occult metastases. Clin. Oral Investig. 2018, 22, 1131–1138.
- Schier, A.F. Axis formation and patterning in zebrafish. Curr. Opin. Genet. Dev. 2001, 11, 393–404.
- Ault, K.T.; Xu, R.H.; Kung, H.F.; Jamrich, M. The homeobox gene PV.1 mediates specification of the prospective neural ectoderm in Xenopus embryos. Dev. Biol. 1997, 192, 162–171.
- Zhu, Z.; Kirschner, M. Regulated proteolysis of Xom mediates dorsoventral pattern formation during early Xenopus development. Dev. Cell 2002, 3, 557–568.





