Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Laura DeOliveira-Mello | + 2268 word(s) | 2268 | 2022-02-11 10:01:12 | | | |
| 2 | Laura DeOliveira-Mello | Meta information modification | 2268 | 2022-02-11 10:21:20 | | | | |
| 3 | Laura DeOliveira-Mello | + 236 word(s) | 2504 | 2022-02-11 10:27:12 | | | | |
| 4 | Laura DeOliveira-Mello | -2 word(s) | 2266 | 2022-02-11 10:27:48 | | | | |
| 5 | Vivi Li | -34 word(s) | 2232 | 2022-02-23 05:12:41 | | | | |
| 6 | Vivi Li | -34 word(s) | 2232 | 2022-02-23 05:14:02 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Deoliveira-Mello, L. Doublecortin. Encyclopedia. Available online: https://encyclopedia.pub/entry/19360 (accessed on 26 May 2026).
Deoliveira-Mello L. Doublecortin. Encyclopedia. Available at: https://encyclopedia.pub/entry/19360. Accessed May 26, 2026.
Deoliveira-Mello, Laura. "Doublecortin" Encyclopedia, https://encyclopedia.pub/entry/19360 (accessed May 26, 2026).
Deoliveira-Mello, L. (2022, February 11). Doublecortin. In Encyclopedia. https://encyclopedia.pub/entry/19360
Deoliveira-Mello, Laura. "Doublecortin." Encyclopedia. Web. 11 February, 2022.
Copy Citation
Doublecortin (DCX) is a microtubule associated protein, essential for correct central nervous system development and lamination in the mammalian cortex. It has been demonstrated to be expressed in developing but not in mature neurons. The teleost visual system is an ideal model to study mechanisms of adult neurogenesis due to its continuous life-long growth. Immunohistochemical, in silico, and western blot analysis to detect the DCX protein in the visual system of teleost fish are described here.
doublecortin
visual system
Danio rerio
Astatotilapia burtoni
neurogenesis
1. Introduction
Doublecortin (DCX; also known as doublin or lissencephalin-X) is a microtubule-associated protein which is typically expressed in the early neuronal differentiation stage, both in precursors and immature neurons [1][2]. Due to DCX expression being nearly exclusive to developing neurons, several research groups are using it as a marker for neurogenesis in a wide range of vertebrate species, e.g., mammals [3], lampreys [4], sharks [5], and teleosts [6].
Brain formation depends on microtubules (MTs) and accompanying microtubule associated proteins (MAPs) to regulate specific migration of different neural cell types [7]. Brain development includes nuclear displacement and process formation that require the action of MTs and specific MAPs [8]. Furthermore, MTs are essential in the formation of growth cones [9]. The de-stabilization of MTs leads to the collapse of the migrating cell body and cessation of nuclear translocation [10]. Faulty of DCX expression causes critical brain defects, which implies that other MTs stabilizing proteins cannot compensate for DCX function in the central nervous system (CNS) [7][11]. In newly formed neurons, DCX are involved in the growth of neuronal processes [12][13].
The continuous life-long growth of the visual system of anamniotes (such as teleost fish) has been an intriguing phenomenon [14][15], especially since such extensive growth does not occur in mammals [16][17]. Several studies have used different fish species to understand the mechanism of adult neurogenesis in vertebrates [2][18][19]. The retina has been proposed as an ideal model to study the generation of new neurons in adults due to the presence of well-delimited neurogenic zones [20][21]: the peripheral germinal zone (PGZ), formed by stem cells; the transition zone, occupied by differentiating cells; and the layered retina, harboring completely differentiated cells, except for the generation of new rods which are added to the outer nuclear layer from rod precursors [22]. Among the many markers used to label differentiating neurons, DCX stands out both in mammals and some teleosts such as cichlid fish [6][23]. Given that commercially available anti-DCX-antibodies efficiently label processes from maturing neurons [12], researchers were interested in testing DCX as a potential marker to study differentiating neurons within the retinal transition zone of the fish retina.
2. DCX Is Present in A. burtoni Retina but Not D. rerio
Antibody stains applied to histological sections enabled the detection of DCX expression in the A. burtoni retina (Figure 1A,B). In contrast, no specific immunofluorescence was found in the zebrafish retina (Figure 1C,D). DCX positive cell bodies were located in the transition zone (Figure 1A,B), next to the PGZ, of the retina of A. burtoni and their axons passing through the nerve fiber layer to the optic nerve head (Figure 1E). No cell bodies expressing DCX were observed in the differentiated retina and/or in the optic nerve.
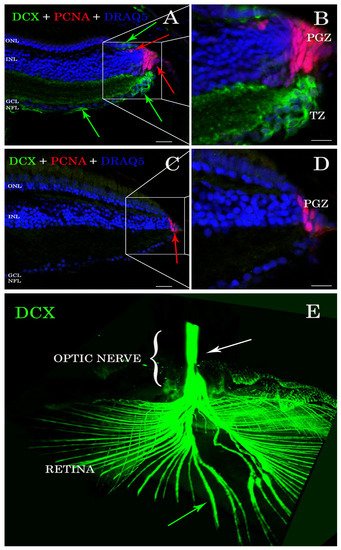
Figure 1. Sections of retina (A–D) and cleared tissue (E). DCX positive cells (green arrows) are detected in A. burtoni retina (A) but not in zebrafish (C). Magnification of (A,C) are presented in (B,D). The proliferating cell nuclear antigen (PCNA; red arrows) binding antibodies detect cells in the peripheral germinal zone (PGZ; of both species. In addition, DCX (green arrows) and PCNA (red arrows) are detected in the ONL corresponding to cones and rod precursors, respectively. Nuclei are stained with DRAQ5 (blue). No double positive cells for DCX (green arrows) and PCNA (red arrows) are found. By analyzing the whole transparent eyeball, it is possible to follow DCX positive processes in the neural fiber layer (green arrow) into the optic nerve (E; white arrow). Scale bar 20 µm (A,C); 10 µm (B,D). ONL: outer nuclear layer; INL: inner nuclear layer; GCL: ganglion cell layer; NFL: neural fiber layer.
3. DCX Proteins Are Present in All the Genomes Analyzed but Not in Zebrafish
Researchers generated a database containing 159 protein sequences belonging to 14 teleost fish (Triplophysa tibetana, Danio rerio, Betta splendens, Parambassis ranga, Oryzias latipes, Nothobranchius furzeri, Astatotilapia burtoni, Gadus morhua, Anabarilius grahami, Nothobranchius pienaari, Oryzias melastigma, Carassius auratus, Labeo rohita, and Tetraodon nigroviridis), 6 mammals (Mus musculus, Homos sapiens, Pan troglodytes, Canis lupus, Bos taurus, and Rattus norvegicus), one bird (Gallus gallus), and one shark (Callorhinchus milii), as well as D. melanogaster and C. elegans as invertebrate species (Table 1).
Table 1. Database of 159 amino acidic sequences selected to perform protein alignments. Different species are used to compare DCX and DCLK sequence identity. From this database researchers also extract DCX1 and DCX2 conserved domains for further sequence analyses.
| Protein and Species | Accession Number | Protein and Species | Accession Number | Protein and Species | Accession Number |
|---|---|---|---|---|---|
| DCLK from Kanglang fish | ROJ66296 | DCLK from Human | NP_004725 | DCLK from Zebrafish | AAI63926 |
| DCLK from Kanglang fish | ROL01462 | DCLK from Human | XP_016876336 | DCLK from Zebrafish | AAI68500 |
| DCLK from Kanglang fish | ROL42582 | DCLK from Human | XP_016876337 | DCLK from Zebrafish | BAF45322 |
| DCLK from Kanglang fish | ROL54251 | DCLK from Medaka | XP_004075345 | DCLK from Zebrafish | BAF45323 |
| DCLK from Betta fish | XP_029027475 | DCLK from Medaka | XP_011474963 | DCLK from Zebrafish | BAF45324 |
| DCLK from Betta fish | XP_029027480 | DCLK from Medaka | XP_011474968 | DCLK from Zebrafish | BAF45325 |
| DCLK from Betta fish | XP_029027952 | DCLK from Medaka | XP_011474974 | DCLK from Zebrafish | BAF45326 |
| DCLK from Betta fish | XP_029027953 | DCLK from Medaka | XP_011480649 | DCLK from Zebrafish | NP_001128593 |
| DCLK from Betta fish | XP_029027954 | DCLK from Medaka | XP_011481611 | DCLK from Zebrafish | NP_001139259 |
| DCLK from Betta fish | XP_029027955 | DCLK from Medaka | XP_011485544 | DCLK from Zebrafish | NP_001139260 |
| DCLK from Betta fish | XP_029030116 | DCLK from Medaka | XP_020563782 | DCLK from Zebrafish | NP_001139261 |
| DCLK from A. burtoni fish | XP_005926513 | DCLK from Medaka | XP_020564443 | DCLK from Zebrafish | XP_005172728 |
| DCLK from A. burtoni fish | XP_005926514 | DCLK from Medaka | XP_024115177 | DCLK from Zebrafish | XP_009290868 |
| DCLK from A. burtoni fish | XP_005928154 | DCLK from Medaka | XP_024143878 | DCLK from Zebrafish | XP_009303718 |
| DCLK from A. burtoni fish | XP_005935114 | DCLK from Mouse | AAH21354 | DCLK from Zebrafish | XP_009303720 |
| DCLK from A. burtoni fish | XP_005935115 | DCLK from Mouse | AAH50903 | DCLK from Zebrafish | XP_021325742 |
| DCLK from A. burtoni fish | XP_005935116 | DCLK from Mouse | AAH64783 | DCLK from Zebrafish | XP_021334856 |
| DCLK from A. burtoni fish | XP_005951753 | DCLK from Mouse | AAI33686 | DCX from Betta fish | XP_029029871 |
| DCLK from A. burtoni fish | XP_005951754 | DCLK from Mouse | AF155819_1 | DCX from Chick | AF330009 |
| DCLK from A. burtoni fish | XP_014186335 | DCLK from Mouse | NP_001104521 | DCX from Chimp | PNI40040 |
| DCLK from A. burtoni fish | XP_014190393 | DCLK from Mouse | NP_001104522 | DCX from A. burtoni fish | XP_005922256 |
| DCLK from A. burtoni fish | XP_014190900 | DCLK from Mouse | NP_001104523 | DCX from Cow | NP_001193894 |
| DCLK from A. burtoni fish | XP_014190901 | DCLK from Mouse | NP_001182467 | DCX from Dog | XP_022271525 |
| DCLK from A. burtoni fish | XP_014190902 | DCLK from Mouse | NP_001182468 | DCX from Dog | XP_022271526 |
| DCLK from A. burtoni fish | XP_014190903 | DCLK from Mouse | NP_001344395 | DCX from Dog | XP_022271527 |
| DCLK from A. burtoni fish | XP_014190904 | DCLK from Mouse | NP_001344397 | DCX from Dog | XP_022271528 |
| DCLK from A. burtoni fish | XP_014190905 | DCLK from Mouse | NP_001344398 | DCX from Dog | XP_022271529 |
| DCLK from A. burtoni fish | XP_014190906 | DCLK from Mouse | NP_001344404 | DCX from Dog | XP_022271530 |
| DCLK from A. burtoni fish | XP_014190907 | DCLK from Mouse | NP_001344405 | DCX from Dog | XP_022271531 |
| DCLK from A. burtoni fish | XP_014193189 | DCLK from Mouse | NP_064362 | DCX from Dog | XP_022271532 |
| DCLK from A. burtoni fish | XP_014193190 | DCLK from Mouse | Q9JLM8 | DCX from Fly | AAM11416 |
| DCLK from A. burtoni fish | XP_014193191 | DCLK from Mouse | XP_006501044 | DCX from Glassy fish | XP_028278320 |
| DCLK from Codfish | XP_030217849 | DCLK from Mouse | XP_006501045 | DCX from Glassy fish | XP_028278321 |
| DCLK from Glassy fish | XP_028258882 | DCLK from Mouse | XP_006501046 | DCX from Glassy fish | XP_028278322 |
| DCLK from Glassy fish | XP_028258890 | DCLK from Mouse | XP_017174936 | DCX from Human | AAC31696 |
| DCLK from Glassy fish | XP_028258898 | DCLK from Mouse | XP_030108271 | DCX from Human | AAC31797 |
| DCLK from Glassy fish | XP_028258907 | DCLK from Mouse | XP_036018776 | DCX from Human | AAC52037 |
| DCLK from Glassy fish | XP_028275059 | DCLK from Rat | NP_445795 | DCX from Human | AAH27925 |
| DCLK from Glassy fish | XP_028275060 | DCLK from Tibetan fish | KAA0702164 | DCX from Human | CAA05867 |
| DCLK from Human | AAI52457 | DCLK from Tibetan fish | KAA0704105 | DCX from Human | CAA06617 |
| DCLK from Human | NP_001317000 | DCLK from Tibetan fish | KAA0710366 | DCX from Human | NP_001182482 |
| DCLK from Human | NP_001317001 | DCLK from Tibetan fish | KAA0722200 | DCX from Human | NP_001356299 |
| DCX from Human | NP_001356300 | DCX from Mouse | AAT58219 | DCX from Rat | AAG18479 |
| DCX from Human | NP_001356301 | DCX from Mouse | AF155820_1 | DCX from Rat | NP_445831 |
| DCX from Human | NP_835364 | DCX from Mouse | BAA33387 | DCX from Rat | Q9ESI7 |
| DCX from Human | NP_835365 | DCX from Mouse | NP_001103692 | DCX from Rat | XP_006257444 |
| DCX from Human | NP_835366 | DCX from Mouse | NP_001103693 | DCX from Rat | XP_006257447 |
| DCX from Human | O43602 | DCX from Mouse | NP_001103694 | DCX from Rat | XP_006257448 |
| DCX from Killifish | AEY83972 | DCX from Mouse | NP_034155 | DCX from Rat | XP_017457656 |
| DCX from Medaka | XP_023818290 | DCX from Mouse | O88809 | DCX from Shark | AFP00992 |
| DCX from Mouse | AAC31799 | DCX from Mouse | XP_006528761 | DCX from Tibetan fish | KAA0710823 |
| DCX from Mouse | AAH56391 | DCX from Mouse | XP_030107072 | Synapsin from Zebrafish | BAH84839 |
| DCX from Mouse | AAH57010 | DCX from Mouse | XP_030107073 | ||
| DCX from Mouse | AAH62974 | DCX from Mouse | XP_030107074 |
Alignment of DCX amino acidic sequences showed a high conservation degree within and among analyzed groups (Table 2). Interestingly, shark and human DCX homologues share 96% of similarity, but no DCX coding sequence was found in the genome of zebrafish.
Table 2. Sequence similarities between selected proteins containing DCX1 and DCX2 domains. Comparison of entire DCX and/or DCLK amino acid sequences to assess protein conservation between different species.
| Protein | Organism | Number of Aligned Sequences | Similarity |
|---|---|---|---|
| DCX | Human + Shark | 15 | 96.7% |
| Mammals | 47 | 89% | |
| Mammals + A. burtoni fish | 48 | 88.8% | |
| Mammals + Teleost Fish | 58 | 83.3% | |
| Teleost Fish | 11 | 82.5% | |
| Mammals + Teleost Fish + Drosophila + Shark + Chick | 61 | 78.6% | |
| DCLK | Mammals | 30 | 55.9% |
| Teleost Fish | 71 | 53.5% | |
| Mammals + Teleost Fish | 101 | 49.3% | |
| DCLK from zebrafish + DCX from D. melanogaster | 16 | 72.5% | |
| DCLKs + DCX | 159 | 50.1% | |
Since researchers could not find DCX in zebrafish, researchers also considered the DCLK proteins, which have been previously described in zebrafish, and belong to the DCX superfamily [24]. DCLK sequences showed around 50–55% pairwise identity among species (Table 2). No DCLK sequences were found in the genomes of invertebrate species (D. melanogaster and C. elegans). However, analysis of DCX proteins from D. melanogaster revealed a catalytic “protein C kinase-like” domain (IPR000719, PF00069) located between residues 477–743. Based on these results, researchers performed a multiple-sequence alignment of all the retrieved DCX and DCLK sequences. Results revealed more than 50% sequence similarity between DCX and DCLK sequences (Table 2).
Researchers then extracted the DCX1 conserved domain from DCX and DCLKs proteins, which was used to perform a multiple-sequence alignment. A similar analysis was conducted for DCX2 conserved domain. In both cases, a high conservation degree was observed between DCX and DCLKs proteins (Table 3). The same approach was applied to the kinase domain found in DCLK proteins from vertebrates, using the “protein C kinase-like” catalytic domain of the D. melanogaster DCX sequence as reference (Accession number: AAM11416). Remarkably, results revealed that the DCX2 domain is much more conserved than DCX1 and the kinase domains among species analyzed, and the latter, appears to be not much conserved in the evolutionary scale (Table 3). Comparison of DCX1 and DCX2 domains of DCLK proteins from zebrafish and of DCX proteins from A. burtoni fish revealed a high similarity of sequence and structure (Table 3).
Table 3. Sequece similarities between DCX1 and DCX2 domains after its extraction from DCX and/or DCLK. Comparison of DCX1 and DCX2 conserved domains to analyze the identity between different species.
| Domain | Organism | Number of Sequences | Similarity |
|---|---|---|---|
| DCX1 | All | 159 | 73.4% |
| DCX2 | All | 159 | 77.4% |
| Kinase | All | 159 | 47.4% |
| DCX1 | Teleost Fish | 79 | 74.5% |
| DCX2 | Teleost Fish | 79 | 79.1% |
| Kinase | Teleost Fish | 79 | 69.9% |
| DCX1 | DCLK ZF + DCLK A. burtoni + DCX from another organisms | 97 | 79.6% |
| DCX2 | DCLK ZF + DCLK A. burtoni + DCX from another organisms | 97 | 83.7% |
To verify the absence of DCX in the zebrafish genome, researchers performed a tblastn analysis using human DCX amino acidic sequence as query. Although no DCX orthologous genes were found on zebrafish, researchers found three loci putatively encoding different DCLK isoforms. Likewise, the genomes of the two invertebrate species did not show any conserved region corresponding to DCLK sequences. This supports the hypothesis that the DCLK-encoding gene is absent in invertebrates.
4. Western Blot Analysis Confirms the Presence of DCX in Burton’s Mouthbrooder Fish and Mice, but Not Zebrafish
Using a commercially available anti-DCX antibody, western blot assays revealed an intense and specific band of 40 KDa in protein samples from mice and A. burtoni fish (Figure 2), which corresponded to the weight described for DCX protein (information related to sequence similarity between mice and A. burtoni DCX can be found in Figure 3A). No specific DCX band was found in zebrafish samples (Figure 2). A positive immunoreactive band for β-actin (loading control) in all samples proved that protein lysates from zebrafish brain were correct. In contrast, DCLK expression was found in protein lysates from the three studied organisms (Figure 2); a strong immunoreactive band at 82 KDa is observed in lysates from mice brains, as well as a less intense band at 130 KDa. These two isoforms were also detected in samples from the A. burtoni brain, especially when 60 mg and 80 mg proteins were loaded, although the expression of the 82 KDa isoform is significantly lower. In the case of zebrafish, a specific immunoreactive 130 KDa band was found for the three tested protein concentrations (information related to conservation of the immunogen sequence among the studied species can be found in Figure 3B).
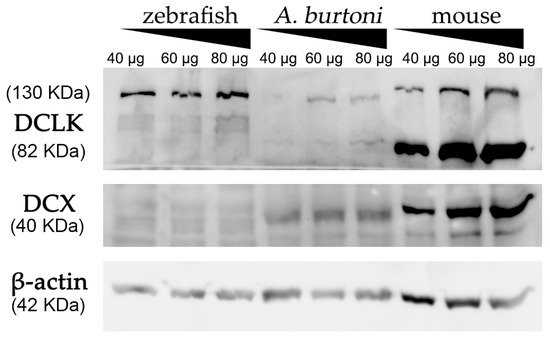
Figure 2. Western blot analysis of protein extracts derived from zebrafish, A. burtoni, and mice brains. Increasing concentrations of proteins were loaded in each well (40, 60 and 80 μg per well). Analysis shows the expression of DCX in A. burtoni and mice, but not in zebrafish extracts. β-actin was used as the loading control after the membrane stripping.
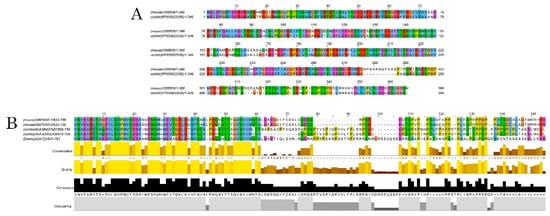
Figure 3. Alignments of DCX and DCLK amino acid sequences. Conserved residues between mice and A. burtoni DCX proteins (A); Conservation of the DCLK2 immunogen sequence use to detecting DCLK protein among the studied species (B).
References
- Gleeson, J.G.; Lin, P.T.; Flanagan, L.A.; Walsh, C. Doublecortin Is a Microtubule-Associated Protein and Is Expressed Widely by Migrating Neurons. Neuron 1999, 23, 257–271.
- Tozzini, E.T.; Baumgart, M.; Battistoni, G.; Cellerino, A. Adult neurogenesis in the short-lived teleost Nothobranchius furzeri: Localization of neurogenic niches, molecular characterization and effects of aging. Aging Cell. 2012, 11, 241–251.
- Brown, J.P.; Couillard-Despres, S.; Cooper-Kuhn, C.M.; Winkler, J.; Aigner, L.; Kuhn, H.G. Transient expression of doublecortin during adult neurogenesis. J. Comp. Neurol. 2003, 467, 1–10.
- Fernández-López, B.; Romaus-Sanjurjo, D.; Senra-Martínez, P.; Anadón, R.; Barreiro-Iglesias, A.; Rodicio, M.C. Spatiotemporal pat-tern of doublecortin expression in the retina of the sea lamprey. Front. Neuroanat. 2016, 10, 5.
- Sánchez-Farías, N.; Candal, E. Doublecortin is widely expressed in the developing and adult retina of sharks. Exp. Eye Res. 2015, 134, 90–100.
- Garcia-Pradas, L.; Gleiser, C.; Wizenmann, A.; Wolburg, H.; Mack, A.F. Glial Cells in the Fish Retinal Nerve Fiber Layer Form Tight Junctions, Separating and Surrounding Axons. Front. Mol. Neurosci. 2018, 11, 367.
- Reiner, O. LIS1 and DCX: Implications for Brain Development and Human Disease in Relation to Microtubules. Scientifica 2013, 2013, 393975.
- Gupta, A.; Tsai, L.-H.; Wynshaw-Boris, A. Life is a journey: A genetic look at neocortical development. Nat. Rev. Genet. 2002, 3, 342–355.
- Tint, I.; Jean, D.; Baas, P.W.; Black, M.M. Doublecortin Associates with Microtubules Preferentially in Regions of the Axon Displaying Actin-Rich Protrusive Structures. J. Neurosci. 2009, 29, 10995–11010.
- Feng, Y.; Walsh, C.A. Protein–Protein interactions, cytoskeletal regulation and neuronal migration. Nat. Rev. Neurosci. 2001, 2, 408–416.
- Fourniol, F.; Perderiset, M.; Houdusse, A.; Moores, C. Structural Studies of the Doublecortin Family of MAPs. Methods Cell Biol. 2013, 115, 27–48.
- Friocourt, G.; Koulakoff, A.; Chafey, P.; Boucher, D.; Fauchereau, F.; Chelly, J.; Francis, F. Doublecortin Functions at the Extremities of Growing Neuronal Processes. Cereb. Cortex 2003, 13, 620–626.
- Moores, C.; Perderiset, M.; Kappeler, C.; Kain, S.; Drummond, D.; Perkins, S.J.; Chelly, J.; Cross, R.; Houdusse, A.; Francis, F. Distinct roles of doublecortin modulating the microtubule cytoskeleton. EMBO J. 2006, 25, 4448–4457.
- Easter, S.; Bratton, B.; Scherer, S. Growth-related order of the retinal fiber layer in goldfish. J. Neurosci. 1984, 14, 2173–2190.
- Fernald, R.D. Teleost vision: Seeing while growing. J. Exp. Zool. 1990, 256, 167–180.
- Kitambi, S.S.; Chandrasekar, G.; Addanki, V.K. Teleost fish—Powerful models for studying development, function and diseases of the human eye. Curr. Sci. 2011, 100, 1815–1823.
- Parrilla, M.; Lillo, C.; Herrero-Turrion, M.J.; Arévalo, R.; Lara, J.M.; Aijón, J.; Velasco, A. Pax2 in the optic nerve of the goldfish, a model of continuous growth. Brain Res. 2009, 1255, 75–88.
- Ghosh, S.; Hui, S.P. Regeneration of Zebrafish CNS: Adult Neurogenesis. Neural Plast. 2016, 2016, 5815439.
- Sîrbulescu, R.F.; Ilieş, I.; Vitalo, A.G.; Trull, K.; Zhu, J.; Traniello, I.M.; Zupanc, G.K. Adult stem cells in the knifefish cerebellum. Dev. Neurobiol. 2015, 75, 39–65.
- Raymond, P.A.; Barthel, L.K.; Bernardos, R.L.; Perkowski, J.J. Molecular characterization of retinal stem cells and their niches in adult zebrafish. BMC Dev. Biol. 2006, 6, 36.
- Müller, H. Bau und wachstum der netzhaut des Guppy (Lebistes reticulates). Zool. Jahrbücher Abt. Für Allg. Zool. Und Physiol. Der Tiere. 1952, 63, 275–324.
- Johns, P.R.; Fernald, R.D. Genesis of rods in teleost fish retina. Nature 1981, 293, 141–142.
- DeOliveira-Mello, L.; Lara, J.M.; Arevalo, R.; Velasco, A.; Mack, A.F. Sox2 expression in the visual system of two teleost species. Brain Res. 2019, 1722, 146350.
- Reiner, O.; Coquelle, F.M.; Peter, B.; Levy, T.; Kaplan, A.; Sapir, T.; Orr, I.; Barkai, N.; Eichele, G.; Bergmann, S. The evolving doublecortin (DCX) superfamily-0. BMC Genom. 2006, 7, 188.
More
Information
Subjects:
Cell Biology
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.1K
Revisions:
6 times
(View History)
Update Date:
23 Feb 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





