
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | yan zhao | + 3942 word(s) | 3942 | 2022-01-18 04:29:27 |
Video Upload Options
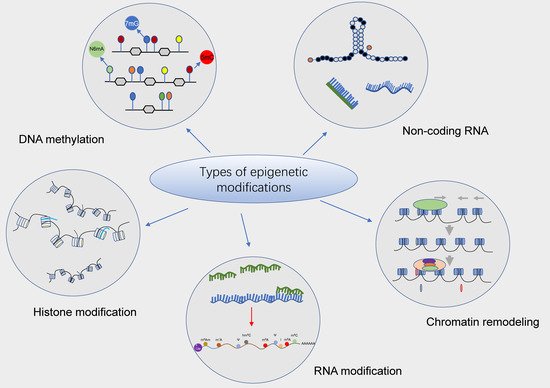
Epigenetics is identified as the study of heritable modifications in gene expression and regulation that do not involve DNA sequence alterations, such as DNA methylation, histone modifications, etc. Importantly, N6-methyladenosine (m6A) methylation modification is one of the most common epigenetic modifications of eukaryotic messenger RNA (mRNA), which plays a key role in various cellular processes. It can not only mediate various RNA metabolic processes such as RNA splicing, translation, and decay under the catalytic regulation of related enzymes but can also affect the normal development of bone marrow hematopoiesis by regulating the self-renewal, proliferation, and differentiation of pluripotent stem cells in the hematopoietic microenvironment of bone marrow. In recent years, numerous studies have demonstrated that m6A methylation modifications play an important role in the development and progression of hematologic malignancies (e.g., leukemia, lymphoma, myelodysplastic syndromes [MDS], multiple myeloma [MM], etc.). Targeting the inhibition of m6A-associated factors can contribute to increased susceptibility of patients with hematologic malignancies to therapeutic agents.
1. Introduction
2. Biological Features of m6A Methylation Modifications
2.1. Overview of m6A Methylation
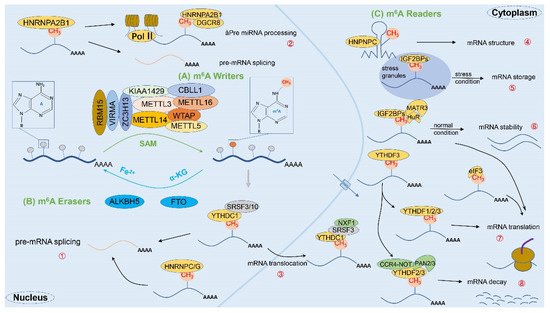
2.2. m6A-Related Enzymes
2.2.1. m6A Methyltransferase (Writers)
2.2.2. m6A Demethylases (Erasers)
3. m6A Methylation Modification and Normal Hematopoietic Regulation
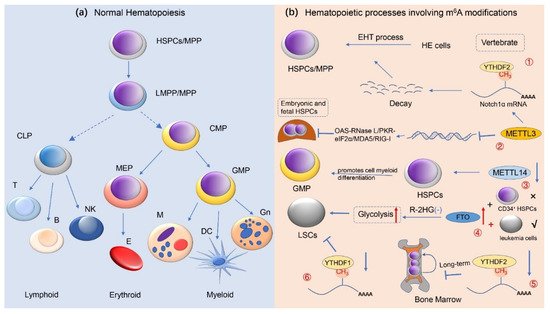
4. m6A Methylation Modifications and Hematological Malignancies
| Cancer Type | m6A Modifiers | Patients/Cell Lines | Role of m6A in Cancer | Functions | Mechanism | References |
|---|---|---|---|---|---|---|
| ALL | Writers/Erasers | In ETV6/RUNX1-positive ALL patients | oncogene | High levels of m6A “writers” (METTL3, METTL14, WTAP) and m6A “erasers” (FTO and ALKBH5) mRNA expression prior to induction therapy resulted in a high disease burden in ALL patients | Not available | [74] |
| METTL3/METTL14 | In childhood ETV6/RUNX1-positive P-ALL | oncogene | The decreased levels of METTL3 and METTL14 indicate a possible role in the pathogenesis and progression of E/R-positive P-ALL. | Not available | [75] | |
| ALKBH5 | In glucocorticoid (GC)-resistant T-ALL patients/CEM-C1 cells/mice | oncogene | Inhibition of ALKBH5-mediated m6A modification decreased USP1 expression, and downregulation of USP1 ameliorated GC resistance in T-ALL by suppressing Aurora B expression and elevating GR levels | ALKBH5/USP1/Aurora B | [76] | |
| AML | YTHDC1 | In human AML cells/LSCs in vivo in mice | oncogene | YTHDC1 is overexpressed in AML, and it contributes to the proliferation and survival of human HSPCs/AML cells, as well as the self-renewal of leukemia stem cells (LSCs) in mice | YTHDC1/ MCM4 | [77] |
| YTHDC1 | In AML cells | oncogene | nYACs maintain mRNA stability, as well as AML cell survival and undifferentiated state; moreover, nYACs protect m6A-mRNA from degradation by PAXT complex and exosome-related RNAs | YTHDC1-m6A condensates (nYACs) | [78] | |
| YTHDF2 | In leukemic cells in vitro and in mice/in AML patients | oncogene | Overexpressed in t (8; 21)-type AML patients; when YTHDF2 is knocked down, it inhibits tumor cell proliferation in vitro and in mice | AML1/ETO-HIF1α loop/YTHDF2/TNFRSF1b | [79] | |
| YTHDF2 | In mouse and human AML | oncogene | YTHDF2 contributes to the initiation of AML disease as well as proliferation and maintains the overall functional integrity of LSCs | YTHDF2/ TNFR2 | [80] | |
| YBX1/IGF2BPs | Primary human and mouse AML cells in vitro and in vivo | oncogene | Expression of YBX1 is markedly upregulated in myeloid leukemia cells, and YBX1 deficiency greatly induces apoptosis and promotes differentiation while reducing proliferation and impairing leukemic competence of primary human and mouse AML cells in vitro and in vivo | YBX1/MYC/BCL2 (mRNA) | [81] | |
| METTL3 | In MOLM-13 cell lines/recipient mice in vivo | oncogene | METTL3 is highly expressed in AML cells as well as promotes AML cell proliferation and inhibits cell differentiation; m6A modification elevates the translation levels of c-MYC, BCL2, and PTEN gene mRNAs in AML cells | METTL3/c-MYC/BCL2/PTEN (mRNA) | [82] | |
| METTL3 | In AML cells and immunodeficient mice | oncogene | In AML cells, METTL3 promotes tumor cell proliferation and inhibits cell differentiation; downregulation of METTL3 results in the inability of immunodeficient mice to develop leukemia. | CEBPZ/ METTL3/ SP1 | [83] | |
| METTL14 | In normal HSPCs and AML cells | oncogene | METTL14 is overexpressed in AML cells and can block the differentiation of normal myeloid cells and promote malignant hematopoiesis via m6A modifications | SPI1-METTL14-MYB/MYC | [84] | |
| WTAP | In AML patients/WTAP knockout AML cells | oncogene | WTAP promotes AML cell proliferation, tumorigenesis, and inhibits cell differentiation. In addition, WTAP causes chemoresistance in AML cells | WTAP/MYC mRNA | [85] | |
| WTAP | In AML patients or in AML cells in vitro in vivo | oncogene | miR-550-1 leads to a further decrease in WWTR1 stability by downregulating the expression level of WTAP, which ultimately disrupts AML cell proliferation and tumorigenesis | miR-550-1/WTAP/ WWTR1 | [86] | |
| WTAP | In different AML cell lines, e.g., K562 cell line | oncogene | Under the regulation of functional METTL3, the expression of WTAP is upregulated and promotes the proliferation of AML cells | METTL3/WTAP | [87] | |
| FTO | In vitro, in mice, primary patient cells, and TKI-resistant cells | oncogene | SsD inhibits AML cell proliferation and promotes apoptosis and cell cycle arrest via targeting FTO/m6A signaling both in vitro and in vivo | Not available | [88] | |
| FTO | In AMLs | oncogene | FTO enhances leukemia oncogene-mediated cell transformation and leukemogenesis and suppresses all-trans retinoic acid (ATRA)-induced AML cell differentiation and apoptosis | FTO/ASB2, RARA | [89] | |
| FTO | In (R-2HG-sensitive) leukemia cells | oncogene | R-2HG abrogated FTO/m6A/YTHDF2-mediated post-transcriptional upregulation of PFKP and LDHB (two key glycolytic genes) expression, thereby attenuating aerobic glycolysis in leukemia | FTO/m6A/PFKP/LDHB axis | [90] | |
| FTO | In human AML cell lines and AML patients | oncogene | FTO inhibitors, namely FB23 and FB23-2, inhibit proliferation and promote differentiation/apoptosis in human AML cells and primary cells | Not available | [91] | |
| ALKBH5 | In human AML LSCs | oncogene | By regulating the chromatin state of the ALKBH5 locus, the expression of ALKBH5 can be elevated, thereby maintaining leukemogenesis in human AML | KDM4C, MYB, Pol II /ALKBH5/AXL Signaling Axis | [66] | |
| ALKBH5 | In human AML/in LSCs/LICs | oncogene | ALKBH5 not only facilitates the proliferation of AML cells, but also contributes to the self-renewal of leukemic stem/initiating cells (LSCs/LICs) | ALKBH5/TACC3 | [65] | |
| CML | METTL3 | In CML patients/CML cell lines | oncogene | Depletion of METTL3 strongly impairs the translation efficiency of mRNA and contributes to the proliferation of CML cells | METTL3/PES1 protein | [92] |
| METTL3 | PBMCs and CML cell lines | oncogene | Overexpression of NEAT1 inhibits cell viability and promotes apoptosis in CML cells | METTL3/NEAT1/miR-766-5p/CDKN1A axis | [93] | |
| METTL3 | In a mouse model, and in KCL22 and K562 cells | oncogene | Dysregulation of METTL3 promotes chemoresistance and inhibits autophagy in CML cells | LINC00470/METTL3/PTEN mRNA | [94] | |
| DLBCL | m6A regulators | In DLBCL patients | oncogene | In patients with DLBCL, high-risk m6A indicates worse survival when grouped according to prognostic characteristics | Not available | [95] |
| METTL3 | In DLBCL tissues and cell lines | oncogene | METTL3 promotes tumor cell proliferation | METTL3/ PEDF | [96] | |
| WTAP | In xenograft DLBCL models | oncogene | piRNA-30473 facilitates the proliferation of DLBCL cells and induces cell cycle arrest via upregulating WTAP | piRNA-30473/WTAP/HK2 m6A | [97] | |
| MM | ALKBH5 | in MM cells, xenograft models or patients | oncogene | ALKBH5 deficiency induces apoptosis and inhibits the growth of MM cells in vitro | ALKBH5/ TRAF1/NF-κB and MAPK | [98] |
| FTO | in CD138 cells from MM+ | oncogene | IDH2 promotes the growth of myeloma cells in vitro by targeting FTO to regulate the m6A RNA level of MM | IDH2/FTO/WNT7B/Wnt | [99] | |
| HNRNPA2B1 | in MM patients and in MM cells | oncogene | Overexpression of HNRNPA2B1 promotes the proliferation of MM cells in vitro and in vivo | HNRNPA2B1/ILF3 mRNA/AKT3 | [100] | |
| MDS | YTHDC1 | In MDS cells | oncogene | Causes abnormalities in hematopoietic function | YTHDC1/SRSF3 or SRSF10 | [101] |
References
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2019. CA Cancer J. Clin. 2019, 69, 7–34.
- Prakash, G.; Kaur, A.; Malhotra, P.; Khadwal, A.; Sharma, P.; Suri, V.; Varma, N.; Varma, S. Current Role of Genetics in Hematologic Malignancies. Indian J. Hematol. Blood Transfus. 2016, 32, 18–31.
- Mehrpouri, M.; Pourbagheri-Sigaroodi, A.; Bashash, D. The contributory roles of histone deacetylases (HDACs) in hematopoiesis regulation and possibilities for pharmacologic interventions in hematologic malignancies. Int. Immunopharmacol. 2021, 100, 108114.
- Guo, M.; Peng, Y.; Gao, A.; Du, C.; Herman, J.G. Epigenetic heterogeneity in cancer. Biomark. Res. 2019, 7, 23.
- Bonasio, R.; Tu, S.; Reinberg, D. Molecular signals of epigenetic states. Science 2010, 330, 612–616.
- Jones, P.A.; Issa, J.P.; Baylin, S. Targeting the cancer epigenome for therapy. Nat. Rev. Genet. 2016, 17, 630–641.
- Baylin, S.B.; Jones, P.A. Epigenetic Determinants of Cancer. Cold Spring Harb. Perspect. Biol. 2016, 8, a019505.
- Roberti, A.; Valdes, A.F.; Torrecillas, R.; Fraga, M.F.; Fernandez, A.F. Epigenetics in cancer therapy and nanomedicine. Clin. Epigenet. 2019, 11, 81.
- Liu, H.; Begik, O.; Lucas, M.C.; Ramirez, J.M.; Mason, C.E.; Wiener, D.; Schwartz, S.; Mattick, J.S.; Smith, M.A.; Novoa, E.M. Accurate detection of m(6)A RNA modifications in native RNA sequences. Nat. Commun. 2019, 10, 4079.
- Liu, N.; Pan, T. N6-methyladenosine–encoded epitranscriptomics. Nat. Struct. Mol. Biol. 2016, 23, 98–102.
- Shi, H.; Wei, J.; He, C. Where, When, and How: Context-Dependent Functions of RNA Methylation Writers, Readers, and Erasers. Mol. Cell 2019, 74, 640–650.
- Zhang, Y.; Sun, Z.; Jia, J.; Du, T.; Zhang, N.; Tang, Y.; Fang, Y.; Fang, D. Overview of Histone Modification. Adv. Exp. Med. Biol. 2021, 1283, 1–16.
- Yang, Y.; Hsu, P.J.; Chen, Y.S.; Yang, Y.G. Dynamic transcriptomic m(6)A decoration: Writers, erasers, readers and functions in RNA metabolism. Cell Res. 2018, 28, 616–624.
- Zhao, B.S.; Roundtree, I.A.; He, C. Post-transcriptional gene regulation by mRNA modifications. Nat. Rev. Mol. Cell Biol. 2017, 18, 31–42.
- Weng, H.; Huang, H.; Chen, J. RNA N (6)-Methyladenosine Modification in Normal and Malignant Hematopoiesis. Adv. Exp. Med. Biol. 2019, 1143, 75–93.
- Fong, C.Y.; Morison, J.; Dawson, M.A. Epigenetics in the hematologic malignancies. Haematologica 2014, 99, 1772–1783.
- Boccaletto, P.; Machnicka, M.A.; Purta, E.; Piatkowski, P.; Baginski, B.; Wirecki, T.K.; de Crécy-Lagard, V.; Ross, R.; Limbach, P.A.; Kotter, A.; et al. MODOMICS: A database of RNA modification pathways. 2017 update. Nucleic Acids Res. 2018, 46, D303–D307.
- Haruehanroengra, P.; Zheng, Y.Y.; Zhou, Y.; Huang, Y.; Sheng, J. RNA modifications and cancer. RNA Biol. 2020, 17, 1560–1575.
- Wei, C.M.; Gershowitz, A.; Moss, B. Methylated nucleotides block 5’ terminus of HeLa cell messenger RNA. Cell 1975, 4, 379–386.
- Desrosiers, R.; Friderici, K.; Rottman, F. Identification of methylated nucleosides in messenger RNA from Novikoff hepatoma cells. Proc. Natl. Acad. Sci. USA 1974, 71, 3971–3975.
- Geula, S.; Moshitch-Moshkovitz, S.; Dominissini, D.; Mansour, A.A.; Kol, N.; Salmon-Divon, M.; Hershkovitz, V.; Peer, E.; Mor, N.; Manor, Y.S.; et al. Stem cells. m6A mRNA methylation facilitates resolution of naïve pluripotency toward differentiation. Science 2015, 347, 1002–1006.
- Zhang, W.; Qian, Y.; Jia, G. The detection and functions of RNA modification m(6)A based on m(6)A writers and erasers. J. Biol. Chem. 2021, 297, 100973.
- Xu, H.; Dzhashiashvili, Y.; Shah, A.; Kunjamma, R.B.; Weng, Y.L.; Elbaz, B.; Fei, Q.; Jones, J.S.; Li, Y.I.; Zhuang, X.; et al. m(6)A mRNA Methylation Is Essential for Oligodendrocyte Maturation and CNS Myelination. Neuron 2020, 105, 293–309.e5.
- Meyer, K.D.; Saletore, Y.; Zumbo, P.; Elemento, O.; Mason, C.E.; Jaffrey, S.R. Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell 2012, 149, 1635–1646.
- Bodi, Z.; Bottley, A.; Archer, N.; May, S.T.; Fray, R.G. Yeast m6A Methylated mRNAs Are Enriched on Translating Ribosomes during Meiosis, and under Rapamycin Treatment. PLoS ONE 2015, 10, e0132090.
- Dominissini, D.; Moshitch-Moshkovitz, S.; Schwartz, S.; Salmon-Divon, M.; Ungar, L.; Osenberg, S.; Cesarkas, K.; Jacob-Hirsch, J.; Amariglio, N.; Kupiec, M.; et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 2012, 485, 201–206.
- Narayan, P.; Ludwiczak, R.L.; Goodwin, E.C.; Rottman, F.M. Context effects on N6-adenosine methylation sites in prolactin mRNA. Nucleic Acids Res. 1994, 22, 419–426.
- Csepany, T.; Lin, A.; Baldick, C.J., Jr.; Beemon, K. Sequence specificity of mRNA N6-adenosine methyltransferase. J. Biol. Chem. 1990, 265, 20117–20122.
- Narayan, P.; Rottman, F.M. An in vitro system for accurate methylation of internal adenosine residues in messenger RNA. Science 1988, 242, 1159–1162.
- Huang, J.; Yin, P. Structural Insights into N(6)-methyladenosine (m(6)A) Modification in the Transcriptome. Genom. Proteom. Bioinform. 2018, 16, 85–98.
- Dai, D.; Wang, H.; Zhu, L.; Jin, H.; Wang, X. N6-methyladenosine links RNA metabolism to cancer progression. Cell Death Dis. 2018, 9, 124.
- Lan, Q.; Liu, P.Y.; Haase, J.; Bell, J.L.; Hüttelmaier, S.; Liu, T. The Critical Role of RNA m(6)A Methylation in Cancer. Cancer Res. 2019, 79, 1285–1292.
- Pan, Y.; Ma, P.; Liu, Y.; Li, W.; Shu, Y. Multiple functions of m(6)A RNA methylation in cancer. J. Hematol. Oncol. 2018, 11, 48.
- Fu, Y.; Dominissini, D.; Rechavi, G.; He, C. Gene expression regulation mediated through reversible m6A RNA methylation. Nat. Rev. Genet. 2014, 15, 293–306.
- Bokar, J.A.; Shambaugh, M.E.; Polayes, D.; Matera, A.G.; Rottman, F.M. Purification and cDNA cloning of the AdoMet-binding subunit of the human mRNA (N6-adenosine)-methyltransferase. RNA 1997, 3, 1233–1247.
- Liu, J.; Yue, Y.; Han, D.; Wang, X.; Fu, Y.; Zhang, L.; Jia, G.; Yu, M.; Lu, Z.; Deng, X.; et al. A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat. Chem. Biol. 2014, 10, 93–95.
- Ping, X.L.; Sun, B.F.; Wang, L.; Xiao, W.; Yang, X.; Wang, W.J.; Adhikari, S.; Shi, Y.; Lv, Y.; Chen, Y.S.; et al. Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res. 2014, 24, 177–189.
- Bedi, R.K.; Huang, D.; Eberle, S.A.; Wiedmer, L.; Śledź, P.; Caflisch, A. Small-Molecule Inhibitors of METTL3, the Major Human Epitranscriptomic Writer. ChemMedChem 2020, 15, 744–748.
- Schöller, E.; Weichmann, F.; Treiber, T.; Ringle, S.; Treiber, N.; Flatley, A.; Feederle, R.; Bruckmann, A.; Meister, G. Interactions, localization, and phosphorylation of the m(6)A generating METTL3-METTL14-WTAP complex. RNA 2018, 24, 499–512.
- Zhang, W.; He, L.; Liu, Z.; Ren, X.; Qi, L.; Wan, L.; Wang, W.; Tu, C.; Li, Z. Multifaceted Functions and Novel Insight Into the Regulatory Role of RNA N(6)-Methyladenosine Modification in Musculoskeletal Disorders. Front. Cell Dev. Biol. 2020, 8, 870.
- Huang, J.; Dong, X.; Gong, Z.; Qin, L.Y.; Yang, S.; Zhu, Y.L.; Wang, X.; Zhang, D.; Zou, T.; Yin, P.; et al. Solution structure of the RNA recognition domain of METTL3-METTL14 N(6)-methyladenosine methyltransferase. Protein Cell 2019, 10, 272–284.
- Wang, P.; Doxtader, K.A.; Nam, Y. Structural Basis for Cooperative Function of Mettl3 and Mettl14 Methyltransferases. Mol. Cell 2016, 63, 306–317.
- Liu, S.; Zhuo, L.; Wang, J.; Zhang, Q.; Li, Q.; Li, G.; Yan, L.; Jin, T.; Pan, T.; Sui, X.; et al. METTL3 plays multiple functions in biological processes. Am. J. Cancer Res. 2020, 10, 1631–1646.
- Zheng, W.; Dong, X.; Zhao, Y.; Wang, S.; Jiang, H.; Zhang, M.; Zheng, X.; Gu, M. Multiple Functions and Mechanisms Underlying the Role of METTL3 in Human Cancers. Front. Oncol. 2019, 9, 1403.
- Wang, Y.; Li, Y.; Toth, J.I.; Petroski, M.D.; Zhang, Z.; Zhao, J.C. N6-methyladenosine modification destabilizes developmental regulators in embryonic stem cells. Nat. Cell Biol. 2014, 16, 191–198.
- Pendleton, K.E.; Chen, B.; Liu, K.; Hunter, O.V.; Xie, Y.; Tu, B.P.; Conrad, N.K. The U6 snRNA m(6)A Methyltransferase METTL16 Regulates SAM Synthetase Intron Retention. Cell 2017, 169, 824–835.e14.
- Satterwhite, E.R.; Mansfield, K.D. RNA methyltransferase METTL16: Targets and function. Wiley Interdiscip. Rev. RNA 2021, e1681.
- Ignatova, V.V.; Stolz, P.; Kaiser, S.; Gustafsson, T.H.; Lastres, P.R.; Sanz-Moreno, A.; Cho, Y.L.; Amarie, O.V.; Aguilar-Pimentel, A.; Klein-Rodewald, T.; et al. The rRNA m(6)A methyltransferase METTL5 is involved in pluripotency and developmental programs. Genes Dev. 2020, 34, 715–729.
- Miranda-Gonçalves, V.; Lobo, J.; Guimarães-Teixeira, C.; Barros-Silva, D.; Guimarães, R.; Cantante, M.; Braga, I.; Maurício, J.; Oing, C.; Honecker, F.; et al. The component of the m(6)A writer complex VIRMA is implicated in aggressive tumor phenotype, DNA damage response and cisplatin resistance in germ cell tumors. J. Exp. Clin. Cancer Res. 2021, 40, 268.
- Jiang, X.; Liu, B.; Nie, Z.; Duan, L.; Xiong, Q.; Jin, Z.; Yang, C.; Chen, Y. The role of m6A modification in the biological functions and diseases. Signal Transduct. Target. Ther. 2021, 6, 74.
- Xu, Y.; Chen, Y.; Yao, Y.; Xie, H.; Lu, G.; Du, C.; Cheng, J.; Zhou, J. VIRMA contributes to non-small cell lung cancer progression via N(6)-methyladenosine-dependent DAPK3 post-transcriptional modification. Cancer Lett. 2021, 522, 142–154.
- Zhao, W.; Qi, X.; Liu, L.; Ma, S.; Liu, J.; Wu, J. Epigenetic Regulation of m(6)A Modifications in Human Cancer. Mol. Ther. Nucleic Acids 2020, 19, 405–412.
- Gong, P.J.; Shao, Y.C.; Yang, Y.; Song, W.J.; He, X.; Zeng, Y.F.; Huang, S.R.; Wei, L.; Zhang, J.W. Analysis of N6-Methyladenosine Methyltransferase Reveals METTL14 and ZC3H13 as Tumor Suppressor Genes in Breast Cancer. Front. Oncol. 2020, 10, 578963.
- Yue, Y.; Liu, J.; Cui, X.; Cao, J.; Luo, G.; Zhang, Z.; Cheng, T.; Gao, M.; Shu, X.; Ma, H.; et al. VIRMA mediates preferential m(6)A mRNA methylation in 3’UTR and near stop codon and associates with alternative polyadenylation. Cell Discov. 2018, 4, 10.
- Lee, Y.; Choe, J.; Park, O.H.; Kim, Y.K. Molecular Mechanisms Driving mRNA Degradation by m(6)A Modification. Trends Genet. 2020, 36, 177–188.
- Schwartz, S.; Mumbach, M.R.; Jovanovic, M.; Wang, T.; Maciag, K.; Bushkin, G.G.; Mertins, P.; Ter-Ovanesyan, D.; Habib, N.; Cacchiarelli, D.; et al. Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5’ sites. Cell Rep. 2014, 8, 284–296.
- Chen, T.; Hao, Y.J.; Zhang, Y.; Li, M.M.; Wang, M.; Han, W.; Wu, Y.; Lv, Y.; Hao, J.; Wang, L.; et al. m(6)A RNA methylation is regulated by microRNAs and promotes reprogramming to pluripotency. Cell Stem Cell 2015, 16, 289–301.
- Huang, H.; Weng, H.; Zhou, K.; Wu, T.; Zhao, B.S.; Sun, M.; Chen, Z.; Deng, X.; Xiao, G.; Auer, F.; et al. Histone H3 trimethylation at lysine 36 guides m(6)A RNA modification co-transcriptionally. Nature 2019, 567, 414–419.
- Jia, G.; Fu, Y.; Zhao, X.; Dai, Q.; Zheng, G.; Yang, Y.; Yi, C.; Lindahl, T.; Pan, T.; Yang, Y.G.; et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat. Chem. Biol. 2011, 7, 885–887.
- Zheng, G.; Dahl, J.A.; Niu, Y.; Fedorcsak, P.; Huang, C.M.; Li, C.J.; Vågbø, C.B.; Shi, Y.; Wang, W.L.; Song, S.H.; et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol. Cell 2013, 49, 18–29.
- Aas, A.; Isakson, P.; Bindesbøll, C.; Alemu, E.A.; Klungland, A.; Simonsen, A. Nucleocytoplasmic Shuttling of FTO Does Not Affect Starvation-Induced Autophagy. PLoS ONE 2017, 12, e0168182.
- Wu, B.; Li, L.; Huang, Y.; Ma, J.; Min, J. Readers, writers and erasers of N(6)-methylated adenosine modification. Curr. Opin. Struct. Biol. 2017, 47, 67–76.
- Zhang, X.; Wei, L.H.; Wang, Y.; Xiao, Y.; Liu, J.; Zhang, W.; Yan, N.; Amu, G.; Tang, X.; Zhang, L.; et al. Structural insights into FTO’s catalytic mechanism for the demethylation of multiple RNA substrates. Proc. Natl. Acad. Sci. USA 2019, 116, 2919–2924.
- Wang, J.; Wang, J.; Gu, Q.; Ma, Y.; Yang, Y.; Zhu, J.; Zhang, Q. The biological function of m6A demethylase ALKBH5 and its role in human disease. Cancer Cell Int. 2020, 20, 347.
- Shen, C.; Sheng, Y.; Zhu, A.C.; Robinson, S.; Jiang, X.; Dong, L.; Chen, H.; Su, R.; Yin, Z.; Li, W.; et al. RNA Demethylase ALKBH5 Selectively Promotes Tumorigenesis and Cancer Stem Cell Self-Renewal in Acute Myeloid Leukemia. Cell Stem Cell 2020, 27, 64–80.e9.
- Wang, J.; Li, Y.; Wang, P.; Han, G.; Zhang, T.; Chang, J.; Yin, R.; Shan, Y.; Wen, J.; Xie, X.; et al. Leukemogenic Chromatin Alterations Promote AML Leukemia Stem Cells via a KDM4C-ALKBH5-AXL Signaling Axis. Cell Stem Cell 2020, 27, 81–97.e8.
- Peng, Z.; Gong, Y.; Wang, X.; He, W.; Wu, L.; Zhang, L.; Xiong, L.; Huang, Y.; Su, L.; Shi, P.; et al. METTL3-m(6)A-Rubicon axis inhibits autophagy in nonalcoholic fatty liver disease. Mol. Ther. 2021.
- Qi, M.; Sun, H.; Guo, Y.; Zhou, Y.; Gu, X.; Jin, J.; Chen, X.; Wang, F.; Ma, H.; Guo, X.; et al. m(6) A reader protein YTHDF2 regulates spermatogenesis by timely clearance of phase-specific transcripts. Cell Prolif. 2021, e13164.
- Yan, G.; Yuan, Y.; He, M.; Gong, R.; Lei, H.; Zhou, H.; Wang, W.; Du, W.; Ma, T.; Liu, S.; et al. m(6)A Methylation of Precursor-miR-320/RUNX2 Controls Osteogenic Potential of Bone Marrow-Derived Mesenchymal Stem Cells. Mol. Ther. Nucleic Acids 2020, 19, 421–436.
- Liu, J.; Gao, M.; He, J.; Wu, K.; Lin, S.; Jin, L.; Chen, Y.; Liu, H.; Shi, J.; Wang, X.; et al. The RNA m(6)A reader YTHDC1 silences retrotransposons and guards ES cell identity. Nature 2021, 591, 322–326.
- Mathoux, J.; Henshall, D.C.; Brennan, G.P. Regulatory Mechanisms of the RNA Modification m(6)A and Significance in Brain Function in Health and Disease. Front. Cell Neurosci. 2021, 15, 671932.
- Choorapoikayil, S.; Kers, R.; Herbomel, P.; Kissa, K.; den Hertog, J. Pivotal role of Pten in the balance between proliferation and differentiation of hematopoietic stem cells in zebrafish. Blood 2014, 123, 184–190.
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer Statistics, 2017. CA Cancer J. Clin. 2017, 67, 7–30.
- Wang, Y.; Zeng, H.M.; Xue, Y.J.; Lu, A.D.; Jia, Y.P.; Zuo, Y.X.; Zhang, L.P. The gene expression level of m6A catalytic enzymes is increased in ETV6/RUNX1-positive acute lymphoblastic leukemia. Int. J. Lab. Hematol. 2021, 43, e89–e91.
- Sun, C.; Chang, L.; Liu, C.; Chen, X.; Zhu, X. The study of METTL3 and METTL14 expressions in childhood ETV6/RUNX1-positive acute lymphoblastic leukemia. Mol. Genet. Genom. Med. 2019, 7, e00933.
- Gong, H.T.; Liu, L.; Cui, L.N.; Ma, H.Y.; Shen, L.Y. ALKBH5-mediated m6A-demethylation of USP1 regulated T-cell acute lymphoblastic leukemia cell glucocorticoid resistance by Aurora B. Mol. Carcinog. 2021, 60, 644–657.
- Sheng, Y.; Wei, J.; Yu, F.; Xu, H.; Yu, C.; Wu, Q.; Liu, Y.; Li, L.; Cui, X.-L.; Gu, X.; et al. A Critical Role of Nuclear m6A Reader YTHDC1 in Leukemogenesis by Regulating MCM Complex-Mediated DNA Replication. Blood 2021, 138, 2838–2852.
- Cheng, Y.M.; Xie, W.; Pickering, B.F.; Chu, K.L.; Savino, A.M.; Yang, X.J.; Luo, H.Z.; Nguyen, D.T.; Mo, S.L.; Barin, E.; et al. N-6-Methyladenosine on mRNA facilitates a phase-separated nuclear body that suppresses myeloid leukemic differentiation. Cancer Cell 2021, 39, 958–972.e8.
- Chen, Z.; Shao, Y.L.; Wang, L.L.; Lin, J.; Zhang, J.B.; Ding, Y.; Gao, B.B.; Liu, D.H.; Gao, X.N. YTHDF2 is a potential target of AML1/ETO-HIF1α loop-mediated cell proliferation in t(8;21) AML. Oncogene 2021, 40, 3786–3798.
- Paris, J.; Morgan, M.; Campos, J.; Spencer, G.J.; Shmakova, A.; Ivanova, I.; Mapperley, C.; Lawson, H.; Wotherspoon, D.A.; Sepulveda, C.; et al. Targeting the RNA m(6)A Reader YTHDF2 Selectively Compromises Cancer Stem Cells in Acute Myeloid Leukemia. Cell Stem Cell 2019, 25, 137–148.e6.
- Feng, M.D.; Xie, X.Q.; Han, G.Q.; Zhang, T.T.; Li, Y.S.; Li, Y.C.; Yin, R.; Wang, Q.F.; Zhang, T.; Wang, P.P.; et al. YBX1 is required for maintaining myeloid leukemia cell survival by regulating BCL2 stability in an m(6)A-dependent manner. Blood 2021, 138, 71–85.
- Vu, L.P.; Pickering, B.F.; Cheng, Y.; Zaccara, S.; Nguyen, D.; Minuesa, G.; Chou, T.; Chow, A.; Saletore, Y.; MacKay, M.; et al. The N(6)-methyladenosine (m(6)A)-forming enzyme METTL3 controls myeloid differentiation of normal hematopoietic and leukemia cells. Nat. Med. 2017, 23, 1369–1376.
- Barbieri, I.; Tzelepis, K.; Pandolfini, L.; Shi, J.; Millán-Zambrano, G.; Robson, S.C.; Aspris, D.; Migliori, V.; Bannister, A.J.; Han, N.; et al. Promoter-bound METTL3 maintains myeloid leukaemia by m(6)A-dependent translation control. Nature 2017, 552, 126–131.
- Weng, H.; Huang, H.; Wu, H.; Qin, X.; Zhao, B.S.; Dong, L.; Shi, H.; Skibbe, J.; Shen, C.; Hu, C.; et al. METTL14 Inhibits Hematopoietic Stem/Progenitor Differentiation and Promotes Leukemogenesis via mRNA m(6)A Modification. Cell Stem Cell 2018, 22, 191–205.e9.
- Naren, D.; Yan, T.; Gong, Y.; Huang, J.; Zhang, D.; Sang, L.; Zheng, X.; Li, Y. High Wilms’ tumor 1 associating protein expression predicts poor prognosis in acute myeloid leukemia and regulates m(6)A methylation of MYC mRNA. J. Cancer Res. Clin. Oncol. 2021, 147, 33–47.
- Hu, C.; Yu, M.; Li, C.; Wang, Y.; Li, X.; Ulrich, B.; Su, R.; Dong, L.; Weng, H.; Huang, H.; et al. miR-550-1 functions as a tumor suppressor in acute myeloid leukemia via the hippo signaling pathway. Int. J. Biol. Sci. 2020, 16, 2853–2867.
- Sorci, M.; Ianniello, Z.; Cruciani, S.; Larivera, S.; Ginistrelli, L.C.; Capuano, E.; Marchioni, M.; Fazi, F.; Fatica, A. METTL3 regulates WTAP protein homeostasis. Cell Death Dis. 2018, 9, 796.
- Sun, K.; Du, Y.; Hou, Y.; Zhao, M.; Li, J.; Du, Y.; Zhang, L.; Chen, C.; Yang, H.; Yan, F.; et al. Saikosaponin D exhibits anti-leukemic activity by targeting FTO/m(6)A signaling. Theranostics 2021, 11, 5831–5846.
- Li, Z.; Weng, H.; Su, R.; Weng, X.; Zuo, Z.; Li, C.; Huang, H.; Nachtergaele, S.; Dong, L.; Hu, C.; et al. FTO Plays an Oncogenic Role in Acute Myeloid Leukemia as a N(6)-Methyladenosine RNA Demethylase. Cancer Cell 2017, 31, 127–141.
- Qing, Y.; Dong, L.; Gao, L.; Li, C.; Li, Y.; Han, L.; Prince, E.; Tan, B.; Deng, X.; Wetzel, C.; et al. R-2-hydroxyglutarate attenuates aerobic glycolysis in leukemia by targeting the FTO/m(6)A/PFKP/LDHB axis. Mol. Cell 2021, 81, 922–939.e9.
- Huang, Y.; Su, R.; Sheng, Y.; Dong, L.; Dong, Z.; Xu, H.; Ni, T.; Zhang, Z.S.; Zhang, T.; Li, C.; et al. Small-Molecule Targeting of Oncogenic FTO Demethylase in Acute Myeloid Leukemia. Cancer Cell 2019, 35, 677–691.e10.
- Ianniello, Z.; Sorci, M.; Ceci Ginistrelli, L.; Iaiza, A.; Marchioni, M.; Tito, C.; Capuano, E.; Masciarelli, S.; Ottone, T.; Attrotto, C.; et al. New insight into the catalytic -dependent and -independent roles of METTL3 in sustaining aberrant translation in chronic myeloid leukemia. Cell Death Dis. 2021, 12, 870.
- Yao, F.Y.; Zhao, C.; Zhong, F.M.; Qin, T.Y.; Wen, F.; Li, M.Y.; Liu, J.; Huang, B.; Wang, X.Z. m(6)A Modification of lncRNA NEAT1 Regulates Chronic Myelocytic Leukemia Progression via miR-766-5p/CDKN1A Axis. Front. Oncol. 2021, 11, 679634.
- Lai, X.; Wei, J.; Gu, X.Z.; Yao, X.M.; Zhang, D.S.; Li, F.; Sun, Y.Y. Dysregulation of LINC00470 and METTL3 promotes chemoresistance and suppresses autophagy of chronic myelocytic leukaemia cells. J. Cell. Mol. Med. 2021, 25, 4248–4259.
- Xie, Z.; Li, M.; Hong, H.; Xu, Q.; He, Z.; Peng, Z. Expression of N(6)-methyladenosine (m(6)A) regulators correlates with immune microenvironment characteristics and predicts prognosis in diffuse large cell lymphoma (DLBCL). Bioengineered 2021, 12, 6115–6133.
- Cheng, Y.; Fu, Y.; Wang, Y.; Wang, J. The m6A Methyltransferase METTL3 Is Functionally Implicated in DLBCL Development by Regulating m6A Modification in PEDF. Front. Genet. 2020, 11, 955.
- Han, H.; Fan, G.; Song, S.; Jiang, Y.; Qian, C.; Zhang, W.; Su, Q.; Xue, X.; Zhuang, W.; Li, B. piRNA-30473 contributes to tumorigenesis and poor prognosis by regulating m6A RNA methylation in DLBCL. Blood 2021, 137, 1603–1614.
- Qu, J.; Hou, Y.; Chen, Q.; Chen, J.; Li, Y.; Zhang, E.; Gu, H.; Xu, R.; Liu, Y.; Cao, W.; et al. RNA demethylase ALKBH5 promotes tumorigenesis in multiple myeloma via TRAF1-mediated activation of NF-κB and MAPK signaling pathways. Oncogene 2021.
- Song, S.; Fan, G.; Li, Q.; Su, Q.; Zhang, X.; Xue, X.; Wang, Z.; Qian, C.; Jin, Z.; Li, B.; et al. IDH2 contributes to tumorigenesis and poor prognosis by regulating m6A RNA methylation in multiple myeloma. Oncogene 2021, 40, 5393–5402.
- Jiang, F.; Tang, X.; Tang, C.; Hua, Z.; Ke, M.; Wang, C.; Zhao, J.; Gao, S.; Jurczyszyn, A.; Janz, S.; et al. HNRNPA2B1 promotes multiple myeloma progression by increasing AKT3 expression via m6A-dependent stabilization of ILF3 mRNA. J. Hematol. Oncol. 2021, 14, 54.
- Xiao, W.; Adhikari, S.; Dahal, U.; Chen, Y.S.; Hao, Y.J.; Sun, B.F.; Sun, H.Y.; Li, A.; Ping, X.L.; Lai, W.Y.; et al. Nuclear m(6)A Reader YTHDC1 Regulates mRNA Splicing. Mol. Cell 2016, 61, 507–519.





