
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Harald Mangge | + 3082 word(s) | 3082 | 2021-10-11 06:08:57 | | | |
| 2 | Jessie Wu | -2 word(s) | 3080 | 2021-11-30 02:34:19 | | |
Video Upload Options
Understanding immune reactivity against SARS-CoV-2 is essential for coping with the COVID-19 pandemic. Herein, we discuss experiences and open questions about the complex immune responses to SARS-CoV-2. Some people react excellently without experiencing any clinical symptoms, they do not get sick, and they do not pass the virus on to anyone else (“sterilizing” immunity). Others produce antibodies, develop memory T cells, and do not get COVID-19 but transmit the virus to others (“protective” immunity). Some people get sick but recover. A varying percentage develops respiratory failure, systemic symptoms, clotting disorders, cytokine storms, or multi-organ failure; they subsequently decease. Some develop long COVID, a new, controversioal pathologic entity similar to fatigue syndrome or autoimmunity. In reality, COVID-19 is considered more of a systemic immune–vascular disease than a pulmonic disease, involving many tissues and the central nervous system. To fully comprehend the complex clinical manifestations, a profound understanding of the immune responses to SARS-CoV-2 is a good way to improve clinical management of COVID-19. Although neutralizing antibodies are an established approach to recognize an immune status, cellular immunity plays at least an equivalent or an even more important role. However, reliable methods to estimate the SARS-CoV-2-specific T cell capacity are not available for clinical routines. This deficit is important because an unknown percentage of people may exist with good memory T cell responsibility but a low number of or completely lacking peripheral antibodies against SARS-CoV-2. Apart from natural immune responses, vaccination against SARS-CoV-2 turned out to be very effective and much safer than naturally acquired immunity. Nevertheless, besides unwanted side effects of the currently available vector and mRNA preparations, concerns remain whether these vaccines will be strong enough to defeat the pandemic. Altogether, discussing important questions, and try to give answers based on the current knowledge and preliminary data from our laboratories.
1. Introduction
2. Questions Addressing the Human Immune Answer to Infection with SARS-CoV-2
2.1. Mild Versus Severe Disease Course: How Different Is the Immune Response?
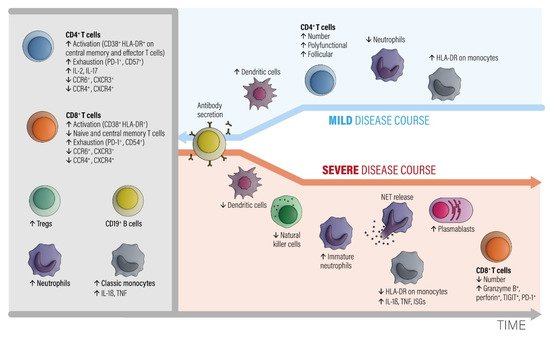
2.1.1. Grey Area
2.1.2. Blue Area
2.1.3. Red Area
2.2. How Is an Immune Response after Infection with the Virus Currently Measured?
2.3. How Do Innate and Adaptive Immune Responses Contribute to the Course of COVID-19?
- (A)
-
Innate immune response (recognition, interferon, and inflammasome activation)
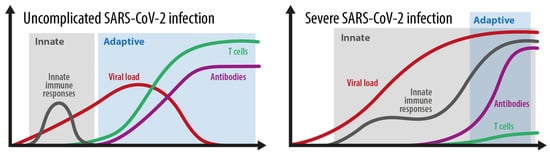
- (B)
-
Adaptive immune response (Antibody production, seroconversion, T cell memory development)
2.4. How Strongly Does the Immune Response to SARS-CoV-2 Differ between Individuals?
- (A)
-
The influence of age and lifestyle
- (B)
-
The influence of gender
- (C)
-
The influence of immune deficiency
References
- Kim, D.S.; Rowland-Jones, S.; Gea-Mallorqui, E. Will SARS-CoV-2 Infection Elicit Long-Lasting Protective or Sterilising Immunity? Implications for Vaccine Strategies (2020). Front. Immunol. 2020, 11, 571481.
- Poland, G.A.; Ovsyannikova, I.G.; Kennedy, R.B. SARS-CoV-2 immunity: Review and applications to phase 3 vaccine candidates. Lancet 2020, 396, 1595–1606.
- Sahin, U.; Muik, A.; Derhovanessian, E.; Vogler, I.; Kranz, L.M.; Vormehr, M.; Baum, A.; Pascal, K.; Quandt, J.; Maurus, D.; et al. COVID-19 vaccine BNT162b1 elicits human antibody and TH1 T cell responses. Nature 2020, 586, 594–599.
- Awadasseid, A.; Wu, Y.; Tanaka, Y.; Zhang, W. Current advances in the development of SARS-CoV-2 vaccines. Int. J. Biol. Sci. 2021, 17, 8–19.
- Franchini, M.; Liumbruno, G.M.; Pezzo, M. COVID-19 vaccine-associated immune thrombosis and thrombocytopenia (VITT): Diagnostic and therapeutic recommendations for a new syndrome. Eur. J. Haematol. 2021, 107, 173–180.
- Abu Mouch, S.; Roguin, A.; Hellou, E.; Ishai, A.; Shoshan, U.; Mahamid, L.; Zoabi, M.; Aisman, M.; Goldschmid, N.; Berar Yanay, N. Myocarditis following COVID-19 mRNA vaccination. Vaccine 2021, 39, 3790–3793.
- Gargano, J.W.; Wallace, M.; Hadler, S.C.; Langley, G.; Su, J.R.; Oster, M.E.; Broder, K.R.; Gee, J.; Weintraub, E.; Shimabukuro, T.; et al. Use of mRNA COVID-19 Vaccine After Reports of Myocarditis Among Vaccine Recipients: Update from the Advisory Committee on Immunization Practices—United States, June 2021. MMWR Morb. Mortal. Wkly. Rep. 2021, 70, 977–982.
- Liu, R.; Yi, S.; Zhang, J.; Lv, Z.; Zhu, C.; Zhang, Y. Viral Load Dynamics in Sputum and Nasopharyngeal Swab in Patients with COVID-19. J. Dent. Res. 2020, 99, 1239–1244.
- Matheny, M.; Maleque, N.; Channell, N.; Eisch, A.R.; Auld, S.C.; Banerji, A.; Druey, K.M. Severe Exacerbations of Systemic Capillary Leak Syndrome After COVID-19 Vaccination: A Case Series. Ann. Intern. Med. 2021.
- Wang, E.Y.; Team, Y.I.; Mao, T.; Klein, J.; Dai, Y.; Huck, J.D.; Jaycox, J.R.; Liu, F.; Zhou, T.; Israelow, B.; et al. Diverse Functional Autoantibodies in Patients with COVID-19. Nature 2021, 595, 283–288.
- Bastard, P.; Rosen, L.B.; Zhang, Q.; Michailidis, E.; Hoffmann, H.H.; Zhang, Y.; Dorgham, K.; Philippot, Q.; Rosain, J.; Beziat, V.; et al. Autoantibodies against type I IFNs in patients with life-threatening COVID-19. Science 2020, 370.
- Zuo, Y.; Estes, S.K.; Ali, R.A.; Gandhi, A.A.; Yalavarthi, S.; Shi, H.; Sule, G.; Gockman, K.; Madison, J.A.; Zuo, M.; et al. Prothrombotic autoantibodies in serum from patients hospitalized with COVID-19. Sci. Transl. Med. 2020, 12.
- Lucas, C.; Wong, P.; Klein, J.; Castro, T.B.R.; Silva, J.; Sundaram, M.; Ellingson, M.K.; Mao, T.; Oh, J.E.; Israelow, B.; et al. Longitudinal analyses reveal immunological misfiring in severe COVID-19. Nature 2020, 584, 463–469.
- Brodin, P. Immune determinants of COVID-19 disease presentation and severity. Nat. Med. 2021, 27, 28–33.
- Farshbafnadi, M.; Zonouzi, S.K.; Sabahi, M.; Dolatshahi, M.; Aarabi, M.H. Aging & COVID-19 susceptibility, disease severity, and clinical outcomes: The role of entangled risk factors. Exp. Gerontol. 2021, 154, 111507.
- Amin, M.T.; Fatema, K.; Arefin, S.; Hussain, F.; Bhowmik, D.R.; Hossain, M.S. Obesity, a major risk factor for immunity and severe outcomes of COVID-19. Biosci. Rep. 2021, 41, BSR20210979.
- Mehta, P.; McAuley, D.F.; Brown, M.; Sanchez, E.; Tattersall, R.S.; Manson, J.J.; Hlh Across Speciality Collaboration, U.K. COVID-19: Consider cytokine storm syndromes and immunosuppression. Lancet 2020, 395, 1033–1034.
- Cao, X. COVID-19: Immunopathology and its implications for therapy. Nat. Rev. Immunol. 2020, 20, 269–270.
- Mathew, D.; Giles, J.R.; Baxter, A.E.; Oldridge, D.A.; Greenplate, A.R.; Wu, J.E.; Alanio, C.; Kuri-Cervantes, L.; Pampena, M.B.; D’Andrea, K.; et al. Deep immune profiling of COVID-19 patients reveals distinct immunotypes with therapeutic implications. Science 2020, 369, eabc8511.
- Rodriguez, L.; Pekkarinen, P.T.; Lakshmikanth, T.; Tan, Z.; Consiglio, C.R.; Pou, C.; Chen, Y.; Mugabo, C.H.; Nguyen, N.A.; Nowlan, K.; et al. Systems-Level Immunomonitoring from Acute to Recovery Phase of Severe COVID-19. Cell Rep. Med. 2020, 1, 100078.
- Pairo-Castineira, E.; Clohisey, S.; Klaric, L.; Bretherick, A.D.; Rawlik, K.; Pasko, D.; Walker, S.; Parkinson, N.; Fourman, M.H.; Russell, C.D.; et al. Genetic mechanisms of critical illness in COVID-19. Nature 2021, 591, 92–98.
- Dennis, A.; Wamil, M.; Alberts, J.; Oben, J.; Cuthbertson, D.J.; Wootton, D.; Crooks, M.; Gabbay, M.; Brady, M.; Hishmeh, L.; et al. Multiorgan impairment in low-risk individuals with post-COVID-19 syndrome: A prospective, community-based study. BMJ Open 2021, 11, e048391.
- Ludvigsson, J.F. Case report and systematic review suggest that children may experience similar long-term effects to adults after clinical COVID-19. Acta Paediatr. 2021, 110, 914–921.
- Clark, D.V.; Kibuuka, H.; Millard, M.; Wakabi, S.; Lukwago, L.; Taylor, A.; Eller, M.A.; Eller, L.A.; Michael, N.L.; Honko, A.N.; et al. Long-term sequelae after Ebola virus disease in Bundibugyo, Uganda: A retrospective cohort study. Lancet Infect. Dis. 2015, 15, 905–912.
- Guillot, X.; Ribera, A.; Gasque, P. Chikungunya-Induced Arthritis in Reunion Island: A Long-Term Observational Follow-Up Study Showing Frequently Persistent Joint Symptoms, Some Cases of Persistent Chikungunya Immunoglobulin M Positivity, and No Anticyclic Citrullinated Peptide Seroconversion After 13 Years. J. Infect. Dis. 2020, 222, 1740–1744.
- Hickie, I.; Davenport, T.; Wakefield, D.; Vollmer-Conna, U.; Cameron, B.; Vernon, S.D.; Reeves, W.C.; Lloyd, A.; Dubbo Infection Outcomes Study, G. Post-infective and chronic fatigue syndromes precipitated by viral and non-viral pathogens: Prospective cohort study. BMJ 2006, 333, 575.
- Osuchowski, M.F.; Winkler, M.S.; Skirecki, T.; Cajander, S.; Shankar-Hari, M.; Lachmann, G.; Monneret, G.; Venet, F.; Bauer, M.; Brunkhorst, F.M.; et al. The COVID-19 puzzle: Deciphering pathophysiology and phenotypes of a new disease entity. Lancet Respir. Med. 2021, 9, 622–642.
- Lumley, S.F.; O’Donnell, D.; Stoesser, N.E.; Matthews, P.C.; Howarth, A.; Hatch, S.B.; Marsden, B.D.; Cox, S.; James, T.; Warren, F.; et al. Antibody Status and Incidence of SARS-CoV-2 Infection in Health Care Workers. N. Engl. J. Med. 2021, 384, 533–540.
- Dan, J.M.; Mateus, J.; Kato, Y.; Hastie, K.M.; Yu, E.D.; Faliti, C.E.; Grifoni, A.; Ramirez, S.I.; Haupt, S.; Frazier, A.; et al. Immunological memory to SARS-CoV-2 assessed for up to 8 months after infection. Science 2021, 371.
- Cohen, K.W.; Linderman, S.L.; Moodie, Z.; Czartoski, J.; Lai, L.; Mantus, G.; Norwood, C.; Nyhoff, L.E.; Edara, V.V.; Floyd, K.; et al. Longitudinal analysis shows durable and broad immune memory after SARS-CoV-2 infection with persisting antibody responses and memory B and T cells. medRxiv 2021.
- Zhao, J.; Wang, L.; Schank, M.; Dang, X.; Lu, Z.; Cao, D.; Khanal, S.; Nguyen, L.N.; Nguyen, L.N.T.; Zhang, J.; et al. SARS-CoV-2 specific memory T cell epitopes identified in COVID-19-recovered subjects. Virus Res. 2021, 304, 198508.
- Lim, Y.X.; Ng, Y.L.; Tam, J.P.; Liu, D.X. Human Coronaviruses: A Review of Virus-Host Interactions. Diseases 2016, 4, 26.
- Fitzgerald, K.A.; Kagan, J.C. Toll-like Receptors and the Control of Immunity. Cell 2020, 180, 1044–1066.
- Nieto-Torres, J.L.; Verdia-Baguena, C.; Jimenez-Guardeno, J.M.; Regla-Nava, J.A.; Castano-Rodriguez, C.; Fernandez-Delgado, R.; Torres, J.; Aguilella, V.M.; Enjuanes, L. Severe acute respiratory syndrome coronavirus E protein transports calcium ions and activates the NLRP3 inflammasome. Virology 2015, 485, 330–339.
- Rodrigues, T.S.; de Sa, K.S.G.; Ishimoto, A.Y.; Becerra, A.; Oliveira, S.; Almeida, L.; Goncalves, A.V.; Perucello, D.B.; Andrade, W.A.; Castro, R.; et al. Inflammasomes are activated in response to SARS-CoV-2 infection and are associated with COVID-19 severity in patients. J. Exp. Med. 2021, 218.
- Wu, C.; Lu, W.; Zhang, Y.; Zhang, G.; Shi, X.; Hisada, Y.; Grover, S.P.; Zhang, X.; Li, L.; Xiang, B.; et al. Inflammasome Activation Triggers Blood Clotting and Host Death through Pyroptosis. Immunity 2019, 50, 1401–1411.
- Han, Y.; Zhang, H.; Mu, S.; Wei, W.; Jin, C.; Tong, C.; Song, Z.; Zha, Y.; Xue, Y.; Gu, G. Lactate dehydrogenase, an independent risk factor of severe COVID-19 patients: A retrospective and observational study. Aging 2020, 12, 11245–11258.
- Zust, R.; Cervantes-Barragan, L.; Habjan, M.; Maier, R.; Neuman, B.W.; Ziebuhr, J.; Szretter, K.J.; Baker, S.C.; Barchet, W.; Diamond, M.S.; et al. Ribose 2′-O-methylation provides a molecular signature for the distinction of self and non-self mRNA dependent on the RNA sensor Mda5. Nat. Immunol. 2011, 12, 137–143.
- Spiegel, M.; Pichlmair, A.; Martinez-Sobrido, L.; Cros, J.; Garcia-Sastre, A.; Haller, O.; Weber, F. Inhibition of Beta interferon induction by severe acute respiratory syndrome coronavirus suggests a two-step model for activation of interferon regulatory factor 3. J. Virol. 2005, 79, 2079–2086.
- Miorin, L.; Kehrer, T.; Sanchez-Aparicio, M.T.; Zhang, K.; Cohen, P.; Patel, R.S.; Cupic, A.; Makio, T.; Mei, M.; Moreno, E.; et al. SARS-CoV-2 Orf6 hijacks Nup98 to block STAT nuclear import and antagonize interferon signaling. Proc. Natl. Acad. Sci. USA 2020, 117, 28344–28354.
- Arunachalam, P.S.; Wimmers, F.; Mok, C.K.P.; Perera, R.; Scott, M.; Hagan, T.; Sigal, N.; Feng, Y.; Bristow, L.; Tak-Yin Tsang, O.; et al. Systems biological assessment of immunity to mild versus severe COVID-19 infection in humans. Science 2020, 369, 1210–1220.
- Blanco-Melo, D.; Nilsson-Payant, B.E.; Liu, W.C.; Uhl, S.; Hoagland, D.; Moller, R.; Jordan, T.X.; Oishi, K.; Panis, M.; Sachs, D.; et al. Imbalanced Host Response to SARS-CoV-2 Drives Development of COVID-19. Cell 2020, 181, 1036–1045.
- Vabret, N.; Britton, G.J.; Gruber, C.; Hegde, S.; Kim, J.; Kuksin, M.; Levantovsky, R.; Malle, L.; Moreira, A.; Park, M.D.; et al. Immunology of COVID-19: Current State of the Science. Immunity 2020, 52, 910–941.
- Zhang, Q.; Bastard, P.; Liu, Z.; Le Pen, J.; Moncada-Velez, M.; Chen, J.; Ogishi, M.; Sabli, I.K.D.; Hodeib, S.; Korol, C.; et al. Inborn errors of type I IFN immunity in patients with life-threatening COVID-19. Science 2020, 370.
- Bastard, P.; Levy, R.; Henriquez, S.; Bodemer, C.; Szwebel, T.A.; Casanova, J.L. Interferon-beta Therapy in a Patient with Incontinentia Pigmenti and Autoantibodies against Type I IFNs Infected with SARS-CoV-2. J. Clin. Immunol. 2021, 41, 931–933.
- Middeldorp, S.; Coppens, M.; van Haaps, T.F.; Foppen, M.; Vlaar, A.P.; Muller, M.C.A.; Bouman, C.C.S.; Beenen, L.F.M.; Kootte, R.S.; Heijmans, J.; et al. Incidence of venous thromboembolism in hospitalized patients with COVID-19. J. Thromb. Haemost. 2020, 18, 1995–2002.
- Sette, A.; Crotty, S. Adaptive immunity to SARS-CoV-2 and COVID-19. Cell 2021, 184, 861–880.
- Gudbjartsson, D.F.; Norddahl, G.L.; Melsted, P.; Gunnarsdottir, K.; Holm, H.; Eythorsson, E.; Arnthorsson, A.O.; Helgason, D.; Bjarnadottir, K.; Ingvarsson, R.F.; et al. Humoral Immune Response to SARS-CoV-2 in Iceland. N. Engl. J. Med. 2020, 383, 1724–1734.
- Wajnberg, A.; Amanat, F.; Firpo, A.; Altman, D.R.; Bailey, M.J.; Mansour, M.; McMahon, M.; Meade, P.; Mendu, D.R.; Muellers, K.; et al. Robust neutralizing antibodies to SARS-CoV-2 infection persist for months. Science 2020, 370, 1227–1230.
- Grifoni, A.; Weiskopf, D.; Ramirez, S.I.; Mateus, J.; Dan, J.M.; Moderbacher, C.R.; Rawlings, S.A.; Sutherland, A.; Premkumar, L.; Jadi, R.S.; et al. Targets of T Cell Responses to SARS-CoV-2 Coronavirus in Humans with COVID-19 Disease and Unexposed Individuals. Cell 2020, 181, 1489–1501.
- Tetro, J.A. Is COVID-19 receiving ADE from other coronaviruses? Microbes Infect. 2020, 22, 72–73.
- Wan, Y.; Shang, J.; Sun, S.; Tai, W.; Chen, J.; Geng, Q.; He, L.; Chen, Y.; Wu, J.; Shi, Z.; et al. Molecular Mechanism for Antibody-Dependent Enhancement of Coronavirus Entry. J. Virol. 2020, 94, e02015-19.
- Cromer, D.; Juno, J.A.; Khoury, D.; Reynaldi, A.; Wheatley, A.K.; Kent, S.J.; Davenport, M.P. Prospects for durable immune control of SARS-CoV-2 and prevention of reinfection. Nat. Rev. Immunol. 2021, 21, 395–404.
- Smorenberg, A.; Peters, E.J.; van Daele, P.; Nossent, E.J.; Muller, M. How does SARS-CoV-2 targets the elderly patients? A review on potential mechanisms increasing disease severity. Eur. J. Intern. Med. 2021, 83, 1–5.
- Soiza, R.L.; Scicluna, C.; Thomson, E.C. Efficacy and safety of COVID-19 vaccines in older people. Age Ageing 2021, 50, 279–283.
- Takahashi, T.; Ellingson, M.K.; Wong, P.; Israelow, B.; Lucas, C.; Klein, J.; Silva, J.; Mao, T.; Oh, J.E.; Tokuyama, M.; et al. Sex differences in immune responses that underlie COVID-19 disease outcomes. Nature 2020, 588, 315–320.
- Kowitdamrong, E.; Puthanakit, T.; Jantarabenjakul, W.; Prompetchara, E.; Suchartlikitwong, P.; Putcharoen, O.; Hirankarn, N. Antibody responses to SARS-CoV-2 in patients with differing severities of coronavirus disease 2019. PLoS ONE 2020, 15, e0240502.
- Weisberg, S.P.; Connors, T.J.; Zhu, Y.; Baldwin, M.R.; Lin, W.H.; Wontakal, S.; Szabo, P.A.; Wells, S.B.; Dogra, P.; Gray, J.; et al. Distinct antibody responses to SARS-CoV-2 in children and adults across the COVID-19 clinical spectrum. Nat. Immunol. 2021, 22, 25–31.
- Wang, X.; Guo, X.; Xin, Q.; Pan, Y.; Hu, Y.; Li, J.; Chu, Y.; Feng, Y.; Wang, Q. Neutralizing Antibody Responses to Severe Acute Respiratory Syndrome Coronavirus 2 in Coronavirus Disease 2019 Inpatients and Convalescent Patients. Clin. Infect. Dis. 2020, 71, 2688–2694.
- Pichler, D.; Baumgartner, M.; Kimpel, J.; Rossler, A.; Riepler, L.; Bates, K.; Fleischer, V.; von Laer, D.; Borena, W.; Wurzner, R. Marked increase in avidity of Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2) antibodies 7-8 months after infection is not diminished in old age. J. Infect. Dis. 2021, 224, 764–770.
- Huang, D.; Lian, X.; Song, F.; Ma, H.; Lian, Z.; Liang, Y.; Qin, T.; Chen, W.; Wang, S. Clinical features of severe patients infected with 2019 novel coronavirus: A systematic review and meta-analysis. Ann. Transl. Med. 2020, 8, 576.
- Smith, J.C.; Sausville, E.L.; Girish, V.; Yuan, M.L.; Vasudevan, A.; John, K.M.; Sheltzer, J.M. Cigarette Smoke Exposure and Inflammatory Signaling Increase the Expression of the SARS-CoV-2 Receptor ACE2 in the Respiratory Tract. Dev. Cell 2020, 53, 514–529.
- Lagunas-Rangel, F.A. Neutrophil-to-lymphocyte ratio and lymphocyte-to-C-reactive protein ratio in patients with severe coronavirus disease 2019 (COVID-19): A meta-analysis. J. Med. Virol. 2020, 92, 1733–1734.
- Yilmaz, H.; Ucan, B.; Sayki, M.; Unsal, I.; Sahin, M.; Ozbek, M.; Delibasi, T. Usefulness of the neutrophil-to-lymphocyte ratio to prediction of type 2 diabetes mellitus in morbid obesity. Diabetes Metab. Syndr. 2015, 9, 299–304.
- Molony, R.D.; Nguyen, J.T.; Kong, Y.; Montgomery, R.R.; Shaw, A.C.; Iwasaki, A. Aging impairs both primary and secondary RIG-I signaling for interferon induction in human monocytes. Sci. Signal. 2017, 10.
- McGonagle, D.; Sharif, K.; O’Regan, A.; Bridgewood, C. The Role of Cytokines including Interleukin-6 in COVID-19 induced Pneumonia and Macrophage Activation Syndrome-Like Disease. Autoimmun. Rev. 2020, 19, 102537.
- Kang, R.; Chen, R.; Zhang, Q.; Hou, W.; Wu, S.; Cao, L.; Huang, J.; Yu, Y.; Fan, X.G.; Yan, Z.; et al. HMGB1 in health and disease. Mol. Aspects Med. 2014, 40, 1–116.
- Brodin, P. Why is COVID-19 so mild in children? Acta Paediatr. 2020, 109, 1082–1083.
- Kollmann, T.R.; Levy, O.; Montgomery, R.R.; Goriely, S. Innate immune function by Toll-like receptors: Distinct responses in newborns and the elderly. Immunity 2012, 37, 771–783.
- Saso, A.; Kampmann, B. Vaccine responses in newborns. Semin. Immunopathol. 2017, 39, 627–642.
- Torjesen, I. Covid-19: Middle aged women face greater risk of debilitating long term symptoms. BMJ 2021, 372, n829.
- Berghofer, B.; Frommer, T.; Haley, G.; Fink, L.; Bein, G.; Hackstein, H. TLR7 ligands induce higher IFN-alpha production in females. J. Immunol. 2006, 177, 2088–2096.
- Klein, S.L.; Jedlicka, A.; Pekosz, A. The Xs and Y of immune responses to viral vaccines. Lancet Infect. Dis. 2010, 10, 338–349.
- Webb, K.; Peckham, H.; Radziszewska, A.; Menon, M.; Oliveri, P.; Simpson, F.; Deakin, C.T.; Lee, S.; Ciurtin, C.; Butler, G.; et al. Sex and Pubertal Differences in the Type 1 Interferon Pathway Associate With Both X Chromosome Number and Serum Sex Hormone Concentration. Front. Immunol. 2018, 9, 3167.
- Pido-Lopez, J.; Imami, N.; Aspinall, R. Both age and gender affect thymic output: More recent thymic migrants in females than males as they age. Clin. Exp. Immunol. 2001, 125, 409–413.
- Whittaker, E.; Bamford, A.; Kenny, J.; Kaforou, M.; Jones, C.E.; Shah, P.; Ramnarayan, P.; Fraisse, A.; Miller, O.; Davies, P.; et al. Clinical Characteristics of 58 Children With a Pediatric Inflammatory Multisystem Syndrome Temporally Associated With SARS-CoV-2. JAMA 2020, 324, 259–269.
- Pulendran, B.; Li, S.; Nakaya, H.I. Systems vaccinology. Immunity 2010, 33, 516–529.
- Gao, Y.; Chen, Y.; Liu, M.; Shi, S.; Tian, J. Impacts of immunosuppression and immunodeficiency on COVID-19: A systematic review and meta-analysis. J. Infect. 2020, 81, e93–e95.
- Fung, M.; Babik, J.M. COVID-19 in Immunocompromised Hosts: What We Know So Far. Clin. Infect. Dis. 2021, 72, 340–350.
- Robilotti, E.V.; Babady, N.E.; Mead, P.A.; Rolling, T.; Perez-Johnston, R.; Bernardes, M.; Bogler, Y.; Caldararo, M.; Figueroa, C.J.; Glickman, M.S.; et al. Determinants of COVID-19 disease severity in patients with cancer. Nat. Med. 2020, 26, 1218–1223.
- Challenor, S.; Tucker, D. SARS-CoV-2-induced remission of Hodgkin lymphoma. Br. J. Haematol. 2021, 192, 415.
- Buckner, T.W.; Dunphy, C.; Fedoriw, Y.D.; van Deventer, H.W.; Foster, M.C.; Richards, K.L.; Park, S.I. Complete spontaneous remission of diffuse large B-cell lymphoma of the maxillary sinus after concurrent infections. Clin. Lymphoma Myeloma Leuk 2012, 12, 455–458.
- Kahraman, S.; Akinci, M.B.; Sendur, M.A.N.; Yalcin, B. Can the host immune response against SARS-CoV2 also cause an anticancer effect? Med. Oncol. 2021, 38, 90.
- Sonani, B.; Aslam, F.; Goyal, A.; Patel, J.; Bansal, P. COVID-19 vaccination in immunocompromised patients. Clin. Rheumatol. 2021, 40, 797–798.





